Abstract
The Tibetan sheep is an ideal model animal for investigating adaptations to hypoxia and an important livestock species; however, the reproductive mechanisms that characterize its adaptive responses in extreme environments remain poorly understood. In this study, we employ single-nucleus RNA sequencing to characterize the transcriptomic landscape of the Tibetan sheep testis at four key developmental stages: newborn, pre-sexual maturity, and post-sexual maturity to adulthood. We constructed a single-nucleus transcriptomic atlas and identified two distinct subtypes of spermatogonial stem cells (SSCs): quiescent and active. Notably, we observed that pre-sexually mature Sertoli cells exhibit three distinct transcriptional states. Furthermore, we also identified a transitional state Sertoli cell that links immature and mature Sertoli cells. Analysis of testicular intercellular communication networks provides evidence for understanding somatic-germ cell interactions during spermatogenesis. Our study provides a comprehensive atlas of Tibetan sheep testicular development, revealing key insights into the dynamic changes and regulatory mechanisms of spermatogenesis and somatic cell maturation from birth to adulthood. These findings contribute to a detailed understanding of the genetic mechanisms underlying testicular development in Tibetan sheep and offer valuable insights and references for the study and comprehension of reproductive adaptations in other highland male domestic animals.
Similar content being viewed by others
Introduction
Tibetan sheep (Ovis aries) are a main livestock species raised on the Qinghai-Tibet Plateau and its adjacent areas at altitudes of around 3000 m above sea level. This sheep breed, developed through extensive natural selection and artificial breeding, is highly regarded for its significant contribution to local economic development and the maintenance of ecological balance in alpine grassland regions1. To adapt to extreme cold and low-oxygen environments in the plateau region, Tibetan sheep have developed a distinctive physiological mechanism characterized by an extended development cycle, delayed sexual maturity, and low fecundity2,3, which poses a serious challenge to the efficient production and sustainable development of the Tibetan sheep industry. These reproductive traits, shaped by long-term husbandry in alpine environments, make them an ideal model for studying developmental heterochrony in ungulates under ecological constraints.
Mammalian testicular tissue is a highly heterogeneous organ composed of germ cells at various developmental stages, along with different types of somatic cells. The continuity, robustness, and regeneration of the male germline depend on spermatogonial stem cell (SSC) populations4,5, which are established through a multifaceted developmental process initiated during embryogenesis and progressing through fetal, juvenile, pubertal, and adult stages6. Cell division and progressive differentiation constitute the foundational biological processes of spermatogenesis. Typically, SSCs maintain the SSC pool through a precise equilibrium between self-renewal and differentiation, and continuously produce progenitor spermatogonia available for differentiation, which then undergo niche-guided transitions during a variety of cellular states and processes (including mitotic divisions of spermatogonia, meiotic progression of spermatocytes, and spermiogenesis) to maintain continuous sperm production7,8. Gaining a comprehensive understanding of the developmental pathways of various germ cell types is crucial for elucidating the regulatory mechanisms governing spermatogenesis. Although the core machinery of spermatogenesis is conserved, distinct mechanistic differences in regulating stage-specific transitions within this process, both within and among species, have evolved, including differences in SSC state and the proliferative capacity of progenitor spermatogonia8,9,10. Thus, understanding testicular cell fate and transformation remains challenging, and many questions remain to be addressed.
The successful execution of the germ cell developmental program depends on the specialized testicular microenvironment, which is composed mainly of somatic cells. Physical and biochemical interactions, together with signaling events between somatic and germ cells at various stages of germ cell development, are essential for sustaining continuous spermatogenesis. Sertoli cells are the principal somatic cell type within the testicular microenvironment, not only providing physical support for germ cells and establishing the immune-privileged niche essential for spermatogenic processes but also regulating germ cell development and differentiation through complex molecular mechanisms and cell-cell interactions. Although most studies have categorized Sertoli cell development postnatally into two stages—immature and mature—whether any intermediate or transitional states exist remains largely unknown11,12,13. In addition, as research has progressed, interactions between Sertoli cells and germ cells have been increasingly elucidated, yet species-specific differences in the underlying molecular mechanisms remain to be fully defined. For example, a study exploring intercellular interactions in porcine testes found that the ligands in the glial cell line-derived neurotrophic factor (GDNF) signaling pathway associated with SSC maintenance differ from those in humans and mice (i.e., Sertoli cells express NRTN but not GDNF)14. This finding highlights species-specific differences in SSC maintenance by Sertoli cells and suggests that species-specific regulatory factors must be considered to obtain a more accurate understanding of reproductive regulatory mechanisms.
Although single-cell transcriptomic analyses of ovine testes have advanced our understanding of spermatogenesis, limitations remain in developmental coverage, cell type resolution, and breed representation. Current studies primarily focus on single or narrow developmental windows. For instance, scRNA-seq of 8-month-old Mongolian sheep testes revealed conserved and divergent transcriptomic features across mammalian spermatogenesis, yet it captured only a single post-pubertal stage15. Similarly, research on 1.5-year-old Hu sheep failed to identify spermatogonial populations (including SSCs and progenitors), highlighting the critical influence of developmental stage at sampling on cell type detection16. A recent multi-stage scRNA-seq atlas of Hu sheep testes (from birth to 540 days) represents a significant advancement, as it dynamically maps transcriptional changes across postnatal development17. However, despite the expanded developmental coverage achieved in Hu sheep (spanning birth to adulthood), critical knowledge gaps persist regarding Tibetan sheep, particularly in the molecular dynamics governing sexual maturation. This oversight is compounded by the predominant reliance on low-altitude breeds (e.g., Hu sheep) in existing studies, which fails to capture the unique adaptations of high-altitude species. The exclusion of Tibetan sheep represents a significant scientific gap, as this breed exhibits specialized transcriptional networks (e.g., hypoxia-response pathways) and epigenetic regulators (e.g., miRNAs) that are essential for testicular function in plateau environments. Analogous research on yak hybrids further underscores the role of cell-autonomous gene dysregulation in high-altitude spermatogenic failure18,19, highlighting the importance of studying plateau-adapted species. Given that sexual maturation represents a period of profound cellular reorganization and functional transition within the testis, targeted sampling before, during, and after this key developmental window is crucial for understanding the mechanisms governing the establishment and maintenance of spermatogenesis. Our current understanding of these processes in the unique context of Tibetan sheep – particularly the molecular underpinnings of SSC self-renewal, Sertoli cell maturation, and Sertoli-germ cell interactions during this critical phase – remains limited.
Single-nucleus RNA sequencing (snRNA-seq) now offers a powerful approach to dissect this cellular and molecular heterogeneity, reconstruct differentiation trajectories, and elucidate cellular functions and interactions within the native testicular microenvironment20,21. Here, we established a testicular cell atlas of Tibetan sheep at four postnatal developmental stages (newborn, pre-sexual maturity, post-sexual maturity, and adult) using snRNA-seq to systematically probe the molecular signatures, developmental status, and intercellular communication networks that define the transition to and establishment of full reproductive capacity in this breed. We identified 21 distinct cell clusters encompassing all major cell types within the testis, revealed highly heterogeneous cell populations in the testes, and generated individual spatial maps of cluster-specific marker expression through immunofluorescence analysis. In addition, we mapped the global intercellular signaling networks, providing insights into cellular interactions during spermatogenesis. Together, these data provide a theoretical basis for optimizing reproductive performance in rams and constitute an important resource for subsequent gene function studies and molecular genetic improvement. In conclusion, these findings not only dissect the molecular mechanisms underlying the reproductive adaptations of male Tibetan sheep but also provide valuable data resources and insights for studying reproductive adaptations in other highland male domestic animals.
Results
Single-nucleus transcriptomic profiles of the developing Tibetan sheep testes
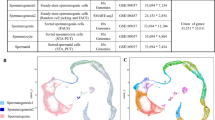
We collected whole testes from Tibetan sheep at various postnatal ages, including T0M, T3M, T1Y, and T3Y, encompassing the entire period from newborn to adult stages. Histological examination of the testis revealed dynamic changes in cell morphology and composition during this developmental timeline: (i) 1-day-old (newborn: seminiferous cords contained gonocytes and spermatogonia), (ii) 3-month-old (pre-sexual maturity: very few spermatocytes were found in addition to gonocytes and spermatogonia), a period in which the testis undergoes structurally and functionally important transformations, (iii) 1-year-old (post-sexual maturity: three to five layers of germ cells at various stages of division (from spermatogonia to spermatids) were observed in neatly arranged seminiferous tubules), corresponding to the stage at which the testis is functionally mature and capable of continuously producing and releasing spermatids, and (iv) 3-year-old (adult: a marked increase in the number of spermatogenic cell layers (7–9 layers), with spermatogonia, spermatocytes, round spermatids, and elongating spermatids distributed sequentially from the basement membrane to the lumen), a stage characterized by a significant degree of functional stability and maturity in the testes (Fig. 1A). In addition, immunofluorescence staining further supported the histological observations by confirming the expected localization of germ cells, as indicated by DDX4 staining (Fig. 1B). To characterize the molecular features associated with this developmental process and to parse testicular cellular heterogeneity, we isolated single nuclei from these testicular tissues and conducted snRNA-seq using the 10× Genomics platform (Fig. 1C). For each developmental stage, two independent biological replicates were conducted, each consisting of pooled testicular tissues from two randomly selected individuals within that stage, yielding a total of eight datasets. Out of a total of 36,199 nuclei, 33,773 passed standard quality control filters and were subsequently retained for downstream analysis (Supplementary Fig. S1A, B). Two biological replicates in the same group were highly similar (Supplementary Fig. S1C). After removing low-quality nuclei, batch effects across datasets were mitigated using the Harmony algorithm to facilitate downstream analysis22. The t-distributed stochastic neighbor embedding (t-SNE) embedding before Harmony integration is provided in Supplementary Fig. S1D. We then performed a graph-based Louvain clustering algorithm in Seurat23 and visualized the resulting clusters using t-SNE24. Further clustering analyses yielded a global cell/molecular atlas of Tibetan sheep testis (Fig. 1D). Notably, we observed a trend wherein cells from T0M and T3M groups exhibited close proximity, as did cells from T1Y and T3Y groups (Fig. 1D). Additionally, the four developmental stages demonstrated temporal variations in the composition of specific cell types (Supplementary Fig. S1E), consistent with the histological characteristics of testes at different developmental stages.
A Hematoxylin and Eosin (H&E) staining of sections of Tibetan sheep testes demonstrating histomorphologic features. Age: one-day-old, T0M; three-month-old, T3M; one-year-old, T1Y; three-year-old, T3Y. Scale bars = 20 μm. B Representative images show DDX4-positive germ cells in seminiferous tubules. Scale bars = 10 μm. C A brief schematic diagram of single-nucleus RNA sequencing workflow. By Figdraw. D t-SNE and clustering analysis of single-nucleus transcriptome data from Tibetan sheep testes at different developmental stages (two biological replicates per stage; 8 libraries total; n = 33,773 nuclei). Each dot represents a single nucleus and is colored by developmental stage or by cluster identity, as indicated. E Bubble plots showing expression of cell-type-specific marker genes for all identified cell clusters. For each cell type, representative markers are shown in the main figure, accompanied by a gallery of additional markers in Supplementary Fig. S1F, G. F The identification of cell clusters expressing the specified marker genes enabled the alignment of these clusters with distinct spermatogenic cell types.
Subsequently, based on established cell type-specific markers and the dynamic changes in cell number, we annotated each cluster along a putative continuum of cell proliferation and differentiation during spermatogenesis, identifying the represented cell types, including the major germ cell populations—spermatogonia, spermatocytes, and spermatids—as well as their respective subtypes (Fig. 1D). These assignments are qualitatively supported by protein immunolocalization patterns that are broadly consistent with our annotations (as detailed in the corresponding sections), and the key marker genes together with their supporting references are summarized in Supplementary Data 1. Based on their high expression of spermatogonia-associated genes (NCL15,25, HMGB225, and CNOT326), clusters 14, 10, 16, 3, and 2 were collectively assigned to the spermatogonial compartment (Fig. 1E, F and Supplementary Fig. S1F). In accordance with the classical subdivision of postnatal spermatogonia into undifferentiated and differentiating fractions, with the undifferentiated pool comprising SSCs and progenitors27,28, we further resolved these clusters into sequential states along this continuum. Clusters 14 and 10 (designated Undiff-SPG1 and Undiff-SPG2) represented the most primitive undifferentiated populations, showing prominent expression of SSC-associated genes such as TKTL113, LIN28B18, PODXL229, MYBL230,31,32,33, and UTF129,32,33,34,35. Cluster 16 (Undiff-SPG3) likely corresponded to a progenitor spermatogonial population transitioning toward differentiation, characterized by expression of progenitor-associated genes such as SOX236,37,38, SALL339, and PRDM1437. In contrast, clusters 3 and 2 displayed elevated expression of RPLP132,33,34, DMRTC240, PTMA17,41, TPT133, HMGN241, MIF17, and other spermatogonia-associated genes, consistent with previous reports describing these genes as characteristic of differentiating spermatogonia in other mammalian species. Accordingly, these five cell clusters were identified as representing three developmental stages of spermatogonial programmed proliferation and differentiation (clusters 14/10: SSCs; cluster 16: progenitors; clusters 3/2: differentiating spermatogonia, designated Diff-SPG1 and Diff-SPG2). Similarly, SPATA16 and SPATA1732, FSIP232,42, PIWIL133,43, SPAG611,12,32,44, and other meiosis-associated genes were highly expressed in clusters 4, 11, and 5. Based on these expression patterns, clusters 4, 11, and 5 were therefore identified as meiotic germ cells during spermatogenesis. Combined with the expression patterns of the spermatid markers CHD533, TNP1 and TNP212,33,35, SPEM132,33,35, and PRM332,33,44, clusters 7 and 0 were identified as cell populations representing the stage of sperm morphogenesis (spermiogenesis). Additionally, we identified separate clusters corresponding to testicular somatic cells (Fig. 1D), particularly Sertoli cells (FATE114 and SOX913, etc.), which are the principal support cells for spermatogenesis, as well as other somatic cell types, including peritubular myoid cells, Leydig cells, endothelial cells, lymphocytes, and macrophages (Fig. 1E and Supplementary Fig. S1G for additional markers). Based on the dynamic expression of these marker genes, we identified a total of 21 cell clusters in the developing Tibetan sheep testis, encompassing germ cells from spermatogonia to spermatids, as well as six types of somatic cells (Fig. 1D, F). It is worth noting that, as shown in Supplementary Fig. S2, germ cell marker genes were highly conserved among sheep populations (Tibetan sheep, Hu sheep, and Mongolia sheep), whereas orthologous genes exhibited limited functional conservation across mammalian species.
Coordinated gene expression patterns and cell type-specific characteristics of spermatogenesis
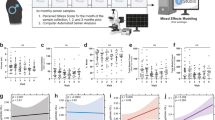
We utilized Monocle2 pseudotime trajectory analyses to construct a developmental trajectory of germ cells. Unsupervised pseudotime analysis from SSCs to spermatids recapitulated the developmental trajectory during Tibetan sheep spermatogenesis (Fig. 2A and Supplementary Fig. S3A). Meanwhile, five different gene cohorts were generated along the germ cell differentiation trajectory (Supplementary Data 1). Gene ontology (GO) analysis of these clusters revealed a dynamic progression of cellular, developmental, and metabolic processes that align with well-organized germline development (Fig. 2B and Supplementary Data 1). We then examined the expression of key marker genes during spermatogenesis by providing scatter plots for individual genes in each cell (Fig. 2C and Supplementary Fig. S3B, C). It is worth noting that candidate spermatogonia markers (ITGA645, PHGDH34, MSL346, ADGR335, PIWIL434, NCL15,25, etc.) were highly expressed only in the early pseudotime, while candidate spermatocytes markers (TEX1133 and MYBL133) were highly expressed in the middle pseudotime. In contrast, candidate spermatids markers (DDX4347, MTF146, HDAC1135, DNAH548, etc.) were highly expressed from mid to late pseudotime. The consistent alignment of these genes in pseudotime with the process of spermatogenesis in both rodents and primates strongly indicates that pseudotime accurately reflects the biological characteristics of testicular development. Germ cells were subsequently parsed according to developmental stage to assess their relative composition across various stages of development. Interestingly, germ cells from T0M samples consisted of undifferentiated and differentiating spermatogonia, consistent with previous observations in Hu sheep49. In the T3M sample, meiotic cells began to emerge, but the number was very small, whereas in the T1Y sample, the relative proportion of spermatocytes considerably increased, consistent with the proliferative expansion of spermatocytes during meiosis. The T1Y sample largely resembled the T3Y sample, though post-meiotic cells increased in proportion, suggesting a more robust commitment to meiosis and an almost complete progression of spermatogenesis (Supplementary Figs. S1C, S3D). In addition, differential expression analysis (P < 0.01; log2FC ≥ 0.36 or log2FC < 0) identified all differentially up- and down-regulated genes in the 10 germ cell clusters (Fig. 2D). Notably, more than 4000 genes (2184 up and 2481 down) were differentially expressed during meiotic initiation, exhibiting the most dramatic transcriptomic changes. To delineate cluster-specific transcriptional programs, we generated a heatmap of the top 10 up-regulated genes for each germ cell cluster (Supplementary Fig. S2E); consistent with these signatures, GO enrichment of the up-regulated genes corroborated the assigned cell-type identities (Fig. 1D and Supplementary Data 1).
A Pseudotime developmental trajectory of germ cell subclusters. Cells are colored by the predicted pseudotime. B Heatmap showing the dynamic changes of gene expression in germ cells along pseudotime. The most representative GO terms in each gene cohort are noted on the right. C Expression levels of representative dynamic genes among germ cells ordered in pseudotime. Each dot represents a single nucleus ordered along pseudotime; the y-axis shows the expression level of the indicated gene, and dot colors denote cluster identity (color scheme as in Fig. 1D). D Differentially expressed genes (up- and down-regulated) and associated GO terms in germ cell types. The represented GO terms from the up-regulated genes of each cell cluster are shown.
Dynamic gene expression patterns of spermatogonia and signaling pathways and metabolic drivers regulating their differentiation fate
Based on t-SNE analysis, we identified five clusters that specifically express spermatogonia marker genes (Fig. 1D, E), and these expression patterns were further validated by immunofluorescence staining (Fig. 3A and S4A). To further resolve the heterogeneity within undifferentiated spermatogonia, we next performed a detailed comparison of the Undiff-SPG1, Undiff-SPG2, and Undiff-SPG3 clusters (Fig. 3 and Supplementary Fig. S4). We found that cells within the Undiff-SPG1 and Undiff-SPG2 clusters have similar transcriptional profiles (Supplementary Fig. S4B) and, at first glance, co-express many key stem cell transcription factors such as TCF1250,51, ATM52, PIWIL232,53,54, ICA1L34, PODXL229, MYBL230,31,32,33, ZBTB4332, and POU5F155,56,57,58,59 (Fig. 3B). Notably, although POU5F1 is classically defined as a marker of primordial germ cells and pluripotency, developmental studies in mammals indicate that, after birth, its expression shifts from centrally located neonatal gonocytes to basement membrane-associated undifferentiated spermatogonia in the adult testis55,56,57,58,59. Consistent with this, our immunofluorescence analyses detected POU5F1 in centrally located gonocytes at neonatal stages and in undifferentiated spermatogonia along the basement membrane at later postnatal stages (Supplementary Fig. S4C), likely reflecting the maintenance of POU5F1 expression into postnatal life57. Despite these shared stem cell factors, Undiff-SPG1 and Undiff-SPG2 also enriched for different subsets of canonical and noncanonical genes (Undiff-SPG1: DNAJB632, PTRHD129,34, etc.; Undiff-SPG2: DMRT140,60, ZBTB1632,33,59,60, CPEB129, PLPPR329,34, UTF129,32,33,34,35, SOHLH132,33,61, FOXO132,33, TSPAN3332,34, etc.) that clearly distinguish Undiff-SPG1 from Undiff-SPG2 (Fig. 3B). We next explored whether Undiff-SPG1 and Undiff-SPG2 clusters represent cells in different phases of the mitotic cycle. We used known cell cycle genes (Supplementary Data 2) to determine the stage that each cell may be in and found that the Undiff-SPG1 and Undiff-SPG2 clusters are similarly distributed across the cell cycle (Supplementary Fig. S4D). This suggests that the cell cycle state does not dictate the clusters; rather, the clusters reflect two distinct transcriptional states of undifferentiated spermatogonia. Given these findings, we conclude that the Undiff-SPG1 and Undiff-SPG2 clusters define transcriptionally discrete states of SSCs and that they may represent metastable (heterogeneous) cellular phenotypes that allow SSCs to adapt to dynamic testicular niche environments. Interestingly, we found that cells in Undiff-SPG2 expressed DMRT1, ZBTB16, MKI67, CDK14, and numerous cyclins (Fig. 3B, C), indicating that the population is more actively proliferative. It is unknown whether an active mechanism is required to maintain differentiation and proliferation potential in SSCs, and whether this active state is necessary for expanding the transit-amplifying progenitors requires further exploration (The results are detailed in subsequent sections). Unlike Undiff-SPG1 and Undiff-SPG2, the Undiff-SPG3 cluster downregulated some SSC-associated genes (ATM, PIWIL2, MYBL2, TCF12, ZBTB16, etc.) while maintaining progenitor-associated genes (Fig. 3B), including SOX236,37,38,62, FABP762,63, PIWIL433, CDH164, SALL339, PRDM1437, UPP133,37, and FOXA265. We speculate that this cluster may represent a transitional state (progenitors) between undifferentiated and differentiating spermatogonia. Consistent with findings in high-altitude species such as the yak18, we also detected expression of the conserved pluripotency-associated gene LIN28B across all undifferentiated spermatogonia clusters (Undiff-SPG1/2/3; Figs. 1D and 3D), suggesting its potential role in maintaining stemness under diverse physiological conditions. Although we did not perform functional assays to determine the function of cells in the Undiff-SPG1/2 and Undiff-SPG3 clusters, the enriched gene expression patterns indicate that Undiff-SPG1/2 is rich in SSCs, while Undiff-SPG3 is rich in progenitors. Notably, several of these genes (Fig. 3B), which have been used in other species to infer SSC- or progenitor-enriched states, were examined only post hoc in our dataset, i.e., after the initial annotation based on canonical genes (Fig. 1E, F). As cluster-enriched transcripts, they therefore serve as supportive rather than primary defining markers and provide additional molecular features that help distinguish SSC-enriched (Undiff-SPG1/2) from progenitor-enriched (Undiff-SPG3) states.
A Representative images of immunofluorescence staining. Scale bars = 10 μm. B Bubble plots showing SSC- and progenitor-associated genes that are shared and unique across undifferentiated spermatogonia. C Violin plots showing the dynamic expression of cell cycle-related genes. Plots are shown at single-nucleus resolution (each dot represents one nucleus); nuclei were obtained from n = 2 biologically independent animals per developmental stage. Source data are provided in Supplementary Data 7. D Representative images of immunofluorescence staining. Scale bars = 10 μm. E Summary of metabolic changes from undifferentiated to differentiating spermatogonia. By Figdraw. F Heatmap showing the dynamic expression of genes associated with glycolysis and oxidative phosphorylation.
Alterations in cellular state and function, such as the differentiation of stem and progenitor cells into diverse cell types, are frequently associated with substantial modifications in metabolic demands and characteristics, often driven by changes in the extracellular environment66. However, our understanding of the metabolic properties associated with SSCs—including self-renewal and differentiation—and the specific molecular features of their transformation during differentiation remains limited across species. To explore the unique metabolic mechanisms and differentiation characteristics of Tibetan sheep spermatogonia subpopulations during this process, we performed differential expression analysis across the five spermatogonia clusters and observed 1783, 2735, 219, 249, and 240 up-regulated genes in Undiff-SPG1, Undiff-SPG2, Undiff-SPG3, Diff-SPG1, and Diff-SPG2 (Supplementary Fig. S4E), respectively. The results indicate that, within heterogeneous spermatogonia subpopulations, substantial differences in transcript abundance and characteristics are readily apparent even among closely related cell types, which may underlie the maintenance of SSCs, the transition of SSCs to progenitors, and the programmed proliferative differentiation of SSCs into differentiating spermatogonia. GO enrichment analysis showed significant enrichment of terms associated with metabolic processes (Supplementary Data 1), such as “cellular macromolecule metabolic process, DNA metabolic process, and regulation of cellular metabolic process, etc.” This suggests that altered gene expression influencing distinct metabolic pathways may be interconnected with variations in cell fate decisions among spermatogonial subpopulations. In general, a well-documented correlation exists between glycolytic metabolism and stem cell properties, whereas a transition to oxidative phosphorylation usually occurs upon the initiation of the differentiation process. Consistent with this notion, GO enrichment analysis showed that the up-regulated genes in Undiff-SPG1 and Undiff-SPG2 are predominantly involved in glycolysis-related signaling pathways, whereas the up-regulated genes in Undiff-SPG3, Diff-SPG1, and Diff-SPG2 are mainly involved in mitochondrial function and oxidative phosphorylation-related signaling pathways (Fig. 3E, F and Supplementary Data 1). In addition, KEGG enrichment analysis showed that genes up-regulated in Undiff-SPG1 and Undiff-SPG2 are significantly enriched in signaling pathways regulating pluripotency of stem cells, while genes in Diff-SPG1/2 are significantly enriched in the oxidative phosphorylation pathway (Supplementary Data 2). Together, these results suggest that the metabolic shift from glycolysis to oxidative metabolism during stem cell differentiation is at least partly governed by changes in the expression of genes related to glycolysis and mitochondrial activity. This glycolysis-to-OXPHOS transition reflects the established hypoxia-response paradigm, whereby glycolytic metabolism maintains stem cell quiescence and survival in hypoxic niches, while OXPHOS fuels differentiation upon oxygen availability, consistent with metabolic reprogramming described in stem cells, including SSCs11,66. Some studies have demonstrated that the MAPK, PI3K-Akt, and mTOR signaling pathways in mouse testis are usually related to both the interconversion between quiescent and activated SSCs and the differentiation of active SSCs into progenitors67. In our dataset, activated SSCs show a significant increase in MAPK and mTOR signaling activity compared with quiescent SSCs, and enhanced mTOR activation was exclusively observed in activated SSCs but not in progenitors (Supplementary Fig. S4G). This observation suggests that the quiescent SSC populations generate partially activated SSCs primarily via the MAPK/mTOR signaling pathway, leading to the formation of early progenitors by the activated SSCs. Although Tibetan sheep SSCs and progenitors exhibit signaling characteristics that are not identical to those reported in mice, we also found that, consistent with findings in mice67, RPS6—a downstream target critical for mTOR signaling—is highly expressed in progenitors (Supplementary Fig. S4G, H). Collectively, these findings link active MAPK, PI3K-Akt, and mTOR signaling with the progressive transition from quiescent SSCs to active SSCs to progenitors.
Molecular characterization of the self-renewal or differentiation fate of spermatogonial stem cells
Spermatogonial differentiation is a critical step in spermatogenesis and marks the initiation of an irreversible commitment to germ-cell differentiation. To reveal the transcriptome dynamics of this process in Tibetan sheep, we performed a pseudotime trajectory analysis of all spermatogonia subclusters using Monocle2 to align individual cells along developmental trajectories and elucidate the molecular features from stem cells to progenitor cells and finally to differentiated cells. The results showed that these five spermatogonia subclusters have the following developmental trajectory: SSCs (Undiff-SPG1 and Undiff-SPG2 clusters, which were automatically merged during trajectory inference) → progenitors (Undiff-SPG3) → differentiating spermatogonia (Diff-SPG1 and Diff-SPG2) (Fig. 4A). Interestingly, pseudotime trajectory analysis revealed a branching point along the inferred differentiation path, indicating two distinct cellular trajectories: one branched group of cells corresponds to terminal differentiation from undifferentiated to differentiating spermatogonia, whereas the other represents a transitional state that bridges the undifferentiated and differentiating spermatogonia stages. This branch is likely enriched for the SSC pool and may simultaneously generate progenitors to initiate spermatogenesis (Fig. 4A). To elucidate the molecular dynamics that distinguish these two branches, we subsequently examined gene-expression changes along the spermatogonia differentiation trajectory to explore cell-type characteristics and identify potential candidate genes associated with cell-fate transitions. By mapping gene expression onto pseudotime, we identified five gene cohorts (a total of ~3510 genes; Fig. 4B and Supplementary Data 2). Previous work has shown that SSCs can be classified into two categories based on JAK1 mRNA abundance, with JAK1- SSCs associated with self-renewal and JAK1+ SSCs biased toward producing differentiating spermatogonia68. To test whether a similar dichotomy exists in Tibetan sheep, we analyzed the dynamic expression of genes associated with stem cell self-renewal or differentiation along pseudotime (Fig. 4C). Stem cell-related genes grouped into cohorts 4, 3, and 5 began to increase at the starting point of pseudotime, whereas those in cohorts 1 increased around the midpoint of pseudotime. These gene sets continued to increase along the branch leading to differentiating spermatogonia and decreased along the trajectory toward the basal SSC pool. Based on these patterns, we hypothesize that higher expression of genes such as JAK1, ITGB1, ATM, PHGDH, PTRHD1, UTF1, DNAJB6, SOHLH2, ICA1L, PPP1R36, EPCAM, MKI67, and others is preferentially initiated in SSCs that are poised to produce differentiating spermatogonia, rather than in SSCs undergoing self-renewal, and that these transcripts may subsequently persist in differentiating spermatogonia (Fig. 4B–D). We next evaluated the protein-level distribution of JAK1 and EPCAM by immunofluorescence analysis. Positive signals for JAK1 and EPCAM were detected in both SSCs and differentiating spermatogonia (Fig. 4E), supporting this hypothesis. Furthermore, DMRT1 and ZBTB16 were included among the gene cohort responsible for sustaining the SSC pool (Fig. 4C). Previous studies in mice have shown that DMRT1 promotes both maintenance and replenishment of the SSC pool, at least in part by activating ZBTB16 transcription to regulate self-renewal60. Consistent with these findings, our data indicate that expression of DMRT1 and ZBTB16 is initiated in the SSCs in a self-renewing mode (Fig. 4D). Although it is unclear what mechanism or signal directly triggers stem cells to express these markers, identifying these triggers will significantly enhance our understanding of spermatogonial fate decisions. Taken together, our datasets and comparative analyses elucidate the developmental trajectories and gene-expression profiles associated with fate transitions in undifferentiated spermatogonia in Tibetan sheep. The undifferentiated spermatogonial populations contain both a core SSC pool and transit-amplifying progenitors, which are governed by a corresponding set of dominant genes that guide subsequent development toward either self-renewal or differentiation fates.
A Pseudotime trajectory of the spermatogonia subclusters. Cells are colored by (i) cell state and (ii) cell subclusters. B Heatmap showing pseudotime-dependent differentially expressed genes in each cluster at the branch points of differentiating spermatogonia and SSC pools. C Expression levels (vertical axis) of representative dynamic genes among spermatogonia ordered in pseudotime. Each dot represents a single nucleus that is color-coded by the cell state of the trajectory analysis. D A hypothetical model for the expression and asymmetric distribution of genes related to self-renewal or differentiation in stem cells. By Figdraw. E Representative images of immunofluorescence staining. Scale bars = 10 μm.
Dynamic transcriptional program during meiotic developmental process
We singled out cells from clusters 4, 11, and 5 (characterized by the expression of PIWIL1; Fig. 5A) for re-clustering analysis, which revealed five subclusters (Fig. 5B). The numbers of nuclei in each subcluster after reclustering are summarized in Supplementary Fig. S5A. Next, we identified pre-leptotene, leptotene, zygotene/early pachytene, late pachytene, and diplotene spermatocyte subtypes using established markers from meiosis databases (https://mcg.ustc.edu.cn/bsc/meiosis/) and literature sources34,35,69,70,71,72,73 (Fig. 5C and Supplementary Fig. S5B). During meiosis, global transcriptional activity decreased during early prophase (leptotene, zygotene, and early pachytene) and recovered at the late pachytene stage, and key genes required for spermatogenesis were significantly up-regulated from early pachytene onward (Supplementary Fig. S5C and Supplementary Data 3), in general agreement with previous studies in mice and humans69. Sertoli cells form the blood-testis barrier (BTB) through diverse cellular junctions, creating a specialized immune and nutritional microenvironment for germ cell differentiation74. Primary spermatocytes successfully traverse this barrier to enter the adluminal compartment, a process that depends on BTB reorganization involving dynamic changes and interactions among multiple proteins. Our results show that at the onset of BTB reorganization (pre-leptotene), GO terms such as cell morphogenesis, cell junction, protein binding, immune response-regulating cell surface receptor signaling pathway, and cell migration are significantly enriched. When leptotene spermatocytes traverse the BTB, tight junctions and adherens junctions undergo further dynamic changes that increase BTB permeability and facilitate spermatocyte migration toward the adluminal compartment. The subsequent zygotene, pachytene, and diplotene phases are mainly enriched for GO terms and genes related to chromosome and cytoarchitectural stability, such as microtubule bundle, centrosome duplication, telomere maintenance, and sister chromatid segregation (Supplementary Fig. S5D, E and Supplementary Data 3). This transition reflects alterations in the prioritized biological events that spermatocytes engage in at successive stages of meiosis.
A Representative images of immunofluorescence staining. Scale bars = 10 μm. B Focused analysis of the cells from clusters 4, 11, and 5 showing developmental progression throughout meiosis. C Expression patterns of known meiotic markers projected onto the t-SNE plot. Red indicates high expression and gray indicates low or no expression. D Pseudotime trajectory of spermatocyte subclusters. Cells are colored according to (upper panel) pseudotime and (lower panel) cell subclusters. E Heatmap showing the dynamic changes of gene expression in spermatocytes along pseudotime. The most representative GO terms in each gene cohort are noted on the right. F, G Expression levels of representative dynamic genes among spermatocytes ordered in pseudotime. Each dot corresponds to a single nucleus, color-coded according to t-SNE clusters.
Next, pseudotime trajectory analysis revealed that these spermatocyte subclusters appeared sequentially along pseudotime, corresponding to distinct meiotic stages (Fig. 5D). The underrepresentation of secondary spermatocytes may be due to their rapid progression into round spermatids75, which is consistent with the chronological characterization of mammalian spermatogenesis. Pseudotime-based differential gene-clustering analysis identified significant transcriptional changes at each stage of meiosis with five distinct molecular signatures (a total of ~6045 genes; Supplementary Data 3), particularly during the transition from the pre-leptotene/leptotene stages—marked by expression of USP50, MND1, and MAEL—to the zygotene/pachytene/diplotene stages—characterized by expression of TEX11, SCML2, SYCP1, FSIP2, TFDP1, RHCG, MLH3, and HMBOX1 (Fig. 5E, F and Supplementary Fig. S5F). Consistent with these dynamics, pseudotime scatter plots further revealed significant up-regulation of key spermatogenesis genes along meiotic progression (Fig. 5G and Supplementary Fig. S5G). Meanwhile, we identified several meiotic recombination-associated genes, including TEX11/14, RAD51AP1, MSH2/3/6, BRCA1, and TOPBP1, that were up-regulated from the leptotene stage to the pachytene stage (Fig. 5E). These results suggest that the accumulation of these mRNAs may prime cells for the subsequent formation of double-stranded DNA breaks as well as homologous chromosome pairing and synapsis. In addition, histone variants play a key role in chromatin recombination and gene expression regulation during spermatogenesis76. Notably, we observed a gradual accumulation of histone genes coincident with the onset/completion of meiotic recombination (Fig. 5E), supporting the idea that meiotic recombination occurs during the transition from leptotene to pachytene69. This transcriptional trajectory reflects dynamic gene expression reprogramming during meiotic progression from leptotene to diplotene, characterized by the sequential activation of meiotic recombination machinery (e.g., TEX11, RAD51AP1) and histone variant accumulation, which are critical for homologous chromosome pairing and chromatin remodeling. Functional analyses were performed to better understand stage-specific changes in various biological processes (Fig. 5E). Taken together, these results reveal stage-specific transcriptional changes and functional diversity of genes in specific biological processes as well as meiotic progression, reflecting the functional hierarchy of molecular events governing meiotic fidelity.
Gene dynamic expression during spermatid morphogenesis
From the Tibetan sheep steady-state spermatogenic cell dataset (Fig. 1D), we identified spermatid-containing clusters—cluster 7 (round spermatids) and cluster 0 (elongated spermatids)—based on the expression of established marker genes (Fig. 1E and Supplementary Fig. S6A). A total of 1469 and 520 up-regulated genes were observed in clusters 7 and 0, respectively (Supplementary Fig. S6B), and were further validated by immunofluorescence analysis (Supplementary Fig. S6C). Transcriptional activity persists predominantly in round spermatids before subsequent genome condensation during spermatid morphogenesis, and almost all transcriptional activity is repressed in mature sperm77. The observed partial transcriptional repression in elongated spermatids suggests that this spermatid cluster is not yet fully mature, which is consistent with the physiological and positional characteristics of sperm maturation in the epididymis. Enrichment analysis showed that “sexual reproduction” and “gamete generation” were highly enriched GO terms in all spermatid clusters, and several of these genes have been shown to be associated with spermatogenesis, such as DDX25, PRM3, and TEX15 (Supplementary Fig. S6D and Supplementary Data 1).
Pseudotime analysis showed that spermatid clusters overlapped minimally in the developmental trajectory (Fig. 6A and Supplementary Fig. S6E), indicating that they represent distinct stages during spermiogenesis. Interestingly, pseudotime trajectory analysis partitioned the round spermatids into two distinct states (State 1 and State 2), which converged into a single path that eventually gave rise to elongated spermatids (Fig. 6A and Supplementary Fig. S6E). We observed different functional features between State 1 and State 2 round spermatids (Fig. 6B, C and Supplementary Data 4). Specifically, GO analysis highlighted an enrichment of terms such as DNA repair, signaling, and gamete generation in State 1, supporting genomic stability. In contrast, State 2 round spermatids lacked the State 1 features described above and were enriched in GO terms related to protein synthesis, translation, and intracellular localization, suggesting preparatory programs for the subsequent development of highly specialized spermatozoa. Analysis of spermatid morphogenesis via pseudotime revealed dynamic expression of ~5018 genes (Fig. 6D) that differed between the round spermatids and elongated spermatid stages. Meanwhile, we identified distinct gene cohorts that exhibited six-wave patterns, reaching peak expression during narrow windows relatively early in round spermatid pseudotime (DNAH3, ARG2, CHD5, and TTN; gene cohorts 2 and 1), slightly later during spermatid development (STPG2, ZNF683, and DEFB2; gene cohorts 5 and 3), or late in elongated spermatid pseudotime (TNP2, TNP1, LDHA, PRM3, ODF1, and TEX54; gene cohorts 4 and 6). In addition, residual expression of meiotic genes (RAD51 and FSIP2) in initial round spermatids coincided with meiotic exit and progression into spermiogenesis. Afterwards, activation of genes necessary for sperm motility (DOP1B and ACTG1), as well as post-meiotic activation of transition proteins required for sperm DNA packaging (TNP1 and TNP2) and protamine (PRM3), becaome noticeable at the midpoint of the pseudotime developmental trajectory (Fig. 6D, E). Consistent with the transcriptional results, immunofluorescence signals for TNP2 began to appear in the post-meiotic spermatid nuclei and increased progressively thereafter (Fig. 6F).
A Pseudotime trajectory of the spermatid subclusters. Cells are colored according to cell state. B Transcriptional differences in round spermatids from different differentiation states. C Box plots showing expression of the top differentially expressed genes (state 1 vs. state 2 round spermatids) across pseudotime-defined differentiation states. Plots are shown at single-nucleus resolution (each dot represents one nucleus); nuclei were obtained from n = 2 biologically independent animals per developmental stage. Source data are provided in Supplementary Data 7. D Heatmap showing the dynamic changes in gene expression in spermatids along pseudotime. The most representative GO terms in each gene cohort are noted on the right. E Expression levels of representative dynamic genes among spermatids ordered in pseudotime. Each dot corresponds to a single nucleus and is color-coded by the cell state of the trajectory analysis. F Representative images of immunofluorescence staining. Scale bars = 10 μm.
Sperm head shaping is a central event in sperm morphogenesis, orchestrated by nuclear condensation. Notably, several components of this core event related to histone-protamine substitution were abundantly present during Tibetan sheep spermiogenesis. Histone expression peaked and protamine expression was elevated at the stage when round spermatids undergo morphological and biochemical changes to form elongated spermatids (Fig. 6D and Supplementary Fig. S6F), which is consistent with the histone-protamine substitution occurring in early elongated spermatids but not in round spermatids. In addition, genes responsible for encoding second messenger enzymes, kinases, and ion channels associated with sperm motility were expressed throughout the sperm morphogenesis stages (Fig. 6D), ensuring the normal physiological function of spermatids at different stages. Taken together, these data from Tibetan sheep testis both depict the temporal trajectory of sperm morphogenesis and reveal the ordered activation of transcriptional waves throughout the process.
Gene dynamic expression during Sertoli cell maturation
Next, we focused on the developmental trajectory and molecular characterization of Sertoli cells in Tibetan sheep. Research indicates that Sertoli cells display distinct variations in size and marker gene expression patterns at various stages of the spermatogenic epithelial cycle12. To characterize the morphological reorganization of Sertoli cell populations, as well as the fundamental differences in transcriptome-wide functional properties among different developmental stages, the Sertoli cells were re-clustered (characterized by the expression of SOX9; Fig. 7A), revealing five subclusters with distinct functional attributes (Fig. 7B and Supplementary Data 5, see below). Our analysis identified immature Sertoli cell clusters (States 1, 2, and 3, characterized by the expression of GATA4 and AMH), transition state Sertoli cell clusters (State 4), and mature Sertoli cell clusters (State 5, characterized by the expression of ABAT and AR) (Fig. 7C). Sertoli cells were subsequently analyzed to assess their composition across various stages of development (Supplementary Fig. S7A). Immature Sertoli cells primarily were derived from sexually immature samples (T0M and T3M) and display a G1/S phase bias (Supplementary Fig. S7A, B). Notably, transition-state Sertoli cells were present across all developmental stages, and mature Sertoli cells primarily comprised cells that develop after sexual maturation (T1Y). This suggests that Sertoli cell populations mature progressively from early life and reach a relatively stable state after sexual maturation.
A Representative images of immunofluorescence staining. Scale bars = 10 μm. B Focused analysis (t-SNE and re-clustering) of Sertoli cell subclusters (subclusters 6, 13, and 9 from Fig. 1D). C Expression patterns of known markers projected onto the t-SNE plot. Red indicates high expression and gray indicates low or no expression. D Distinct states of Sertoli cells showed differential expression of selected key genes. Plots are shown at single-nucleus resolution (each dot represents one nucleus); nuclei were obtained from n = 2 biologically independent animals per developmental stage. Source data are provided in Supplementary Data 7. E Distribution of Sertoli cells along the pseudotime developmental trajectory. F Focused analysis (t-SNE and re-clustering) of Sertoli cells, with cells colored by age (top left) and cell-cycle phase (top right). Monocle pseudotime trajectories of Sertoli cell subclusters, colored by age (lower panel). G Heatmap showing the dynamic changes in gene expression in Sertoli cells along pseudotime. The most representative GO terms in each gene cohort are shown on the right. H Expression levels (vertical axis) of immune-related genes during Sertoli cell maturation. Each dot corresponds to a single nucleus, color-coded according to t-SNE clusters.
Based on subcluster-specific up-regulated gene analysis (Supplementary Fig. S7C), we identified various characteristics of immature, transition-state, and mature Sertoli cells, such as the expression of AR target genes (determined by a set of 26 known AR-dependent genes, of which 7 were repressed and 19 were induced) and ribosomal protein genes (Fig. 7D). To better understand the differences in biological processes reflected in stage-specific changes, further GO functional annotation revealed that, in addition to terms typically associated with Sertoli cell function (such as extracellular matrix component, anchoring junction, cell-cell junction, etc.), unique terms related to functional maintenance and metabolic characterization in three distinct cell states were also identified. For example, GO terms including “regulation of gene expression, epigenetic,” “demethylase activity,” “methyltransferase complex,” and “glycerophospholipid biosynthetic process” were markedly enriched in immature State 1. In contrast, GO terms such as “germ cell development,” “exocrine system development,” “steroid hormone receptor binding,” and “DNA metabolic process” were significantly enriched in immature State 2. Furthermore, GO terms, such as “cell migration,” “regulation of cell communication,” “morphogenesis of an epithelium,” “stem cell proliferation,” “glycolytic process,” “second-messenger-mediated signaling,” and “regulation of cytokine-mediated signaling pathway”, were notably enriched in immature State 3. Collectively, these immature Sertoli cells exhibit distinct variations in metabolism, post-translational modifications, signaling and intercellular communication (as indicated by transcriptional signatures). Transition-state cells lacked the immature-state features described above and were enriched for GO terms related to the electron transport chain, mitochondrial membrane part, lamellipodium organization and ribosome (Supplementary Fig. S7D and Supplementary Data 5). They exhibit higher transcription of mitochondrial membrane proteins (e.g., NDUFB2 and TIMM8B), mitochondrial ATPases genes (e.g., ATP5IF1 and ATP5MPL), and ribosomal proteins (e.g., RRP1B and MRPS25) and contain a higher fraction of cells in the G2M phase (Fig. 7D and Supplementary Fig. S7B, and Supplementary Data 5). In mature Sertoli cells, in addition to the various processes described above, ligase activity, microtubule-based process, vacuolar transport, autophagosome assembly, and activation of innate immune response were significantly enriched (Supplementary Fig. S7D and Supplementary Data 5). This suggests that the primary roles of mature Sertoli cells are to protect germ cells from immunological destruction, coordinate intercellular information exchange, phagocytose senescent or dead germ cells and cellular debris, and maintain a stable microenvironment for spermatogenesis to proceed smoothly.
An unexpected finding is the presence of marker genes associated with sperm-specific nucleus protein (e.g., TNP1/2, PRM3) as well as a large number of genes/terms related to cilia in both transition-state and mature Sertoli cells (Supplementary Fig. S7E, F and Supplementary Data 5). One possible explanation is that these mRNAs in the cytoplasm of Sertoli cells originate from phagocytosed germ cells or residual bodies12. The persistence of transition nucleus protein- and protamine-related genes in the Sertoli cell nuclei is surprising because, according to the conventional view, mRNA derived from phagocytosed germ cells or residual bodies should be degraded rapidly by lysosomes. Immunofluorescence validation showed varying degrees of positive signals for TNP1/2 protein in spermatids at almost all stages of the seminiferous tubule cycle, whereas no positive signal was detected in Sertoli cells (Fig. 6F and Supplementary Fig. S6C). This observation aligns with the hypothesis that TNP1/2 transcripts are not actively transcribed in Sertoli cells. These genes in the Sertoli cell nuclei suggest that there may be protective retention mechanism that is not fully understood, or that these genes perform important previously unrecognized functions in specific physiological processes, thus avoiding the expected fate of degradation. While the predominant explanation involves phagocytic uptake, the functional implications of transiently retained germ cell transcripts in Sertoli cells warrant further investigation. For instance, residual mRNAs may influence Sertoli cell signaling during niche remodeling78, though direct evidence in this context remains limited. Future studies integrating spatial transcriptomics and lysosomal activity profiling could clarify whether these transcripts serve as transient signaling molecules or are merely degradation intermediates.
We next examined Sertoli cell transcriptomic dynamics along pseudotime and visualized the distribution of cells from each developmental stage along the inferred trajectory (Fig. 7E, F). Analysis of Sertoli cell maturation via pseudotime identified dynamic expression in about ~5790 genes that vary across the immature, transitional, and mature stages (Fig. 7G). As anticipated, genes related to the cytoskeleton, extracellular matrix, microtubules, and cellular morphogenesis are increasingly expressed during Sertoli cell maturation (Fig. 7G). Notably, many innate immunity genes, including the β-defensin genes, are upregulated during Sertoli cell maturation (Fig. 7G, H), which supports the important function of Sertoli cells in defending the testes from external stimuli, especially after sexual maturation. Overall, our data characterize the diverse states of Sertoli cells in Tibetan sheep postnatal testes and reveal differences in gene regulation underlying these cell states.
Construction of cellular communication networks in the testicular microenvironment
Intercellular communication is usually characterized by the binding of ligands to receptors and the activation of downstream signaling pathways. To investigate the complex signaling network and its dynamics in the testicular microenvironment, we integrated the ligand-receptor pairs from the CellPhoneDB and CellChat databases, followed by performing cell-cell interaction analysis using the CellChat algorithm. In total, 10,829 significant ligand-receptor pairs were detected, which were further categorized into 57 signaling pathways (Supplementary Data 6). Also, we constructed and quantified global signaling crosstalk atlases (Fig. 8A), which provide both the number and the weight/strength of interactions, uncovering complex somatic-germline interactions that support an open niche model for the testis. SSCs and spermatocytes among germ cells appear to engage in more potential interactions than other germ cells, aligning with the function of the BTB, which blocks meiotic and post-meiotic germ cells from direct communication with the interstitial compartment. Among somatic cells, Sertoli cells usually account for most of the interactions with germ cells, suggesting that Sertoli cells may act as hubs in the intercellular interaction network during spermatogenesis (Fig. 8A and Supplementary Data 6). Furthermore, considering that some interactions were detected at multiple developmental stages, we also quantified the developmental stage at which each ligand-receptor pair was detected. We found that the number of interactions varied across different developmental stages, with unique ligand-receptor pairs present at each stage (Supplementary Fig. S8A and Supplementary Data 6).
A Global testicular intercellular communication is visualized using circle plots showing both the quantity (left) and the weight/strength (right) of significant ligand-receptor interactions among all testicular cell types. B The communication networks inferred among all cell types are presented through hierarchical plots. C The expression of ligands (red) and receptors (blue) associated with the specified signaling pathways is illustrated on the t-SNE plot from Fig. 1D. D Significant ligand-receptor interactions contribute to Sertoli-germ cell communication. The number of detected significant interactions in both panels is indicated in the bottom left of each plot. E Bubble plot showing ligand-receptor interactions between Sertoli cells (source) and germ cells (target). Bubble color represents communication probability. Ligand-receptor pairs are classified into two major groups based on mode of action: physical contact signals (cell-cell adhesion and ECM-receptor interactions) and secreted signals. F A schematic representation of specific signaling pathways involved in the interactions between somatic cells and germ cells in the testes of Tibetan sheep. By Figdraw. G Representative images of immunofluorescence staining. Scale bars = 10 μm.
Considering the considerable signaling differences among different cell types, we investigated signaling pathways with high communication strength in the testicular microenvironment in more detail to provide insights into somatic-germline interactions. We observed that several pathways with high inferred communication strength between somatic and germline compartments (e.g., COLLAGEN, LAMININ, CADM) appear qualitatively similar to those reported in other mammals (Fig. 8B and Supplementary Data 6); however, in Tibetan sheep, the predominant senders/receivers often shift (e.g., altered somatic sources or germline targets), suggesting species- or lineage-specific rewiring of otherwise familiar modules. Interestingly, these pathways have not been highlighted as central in previous studies on testicular cell communication. In addition to physical cell-to-cell contact, intercellular communication through secreted factors also plays a key role in the testicular microenvironment69. In the present study, we found that pleiotrophin (PTN) signaling was the main pathway contributing to communication in somatic-germline interactions, with Leydig cells occupying the center of the cell-cell communication (Fig. 8C). Within the ACTIVIN pathway that enhances spermatogenesis, State 1 Sertoli cells are central to cell-cell communication, acting as the primary signaling sender and receiver (Fig. 8C). We interpret these patterns as hypothesis-generating features of Tibetan sheep rather than universal organizational principles, pending cross-species integration and orthogonal validation. Furthermore, proper progression of spermatogenesis also requires an appropriate immune microenvironment created by somatic cells. We also identified immune signaling pathways (CD46, CD30, and MIF; Supplementary Fig. S8B).
Given that the Sertoli-germ cell communication is a key event in guaranteeing an orderly spermatogenesis. We next focused on the interactions between Sertoli cells (senders) and germ cells (receivers). In State 1, State 2, State 3, State 4, and State 5, we found 438, 733, 10, 301, and 364 interactions between Sertoli cell ligands and receptors from other cells, respectively (Supplementary Fig. S8C). This suggests that the way Sertoli cells interact with other testicular cells changes across developmental states and cell types. Focusing on the interactions between Sertoli cells (senders) and germ cells (receivers), we found that: 1) most of the Sertoli-germ cell interactions involve physical contact signals; 2) while State 3 Sertoli cells interact most frequently with germ cells, State 1 and State 5 Sertoli cells exhibit the highest communication probability with germ cells; 3) most of the ligand-receptor interactions are common, but their interaction strengths vary, and only a few are unique to specific stages (Fig. 8D, E and Supplementary Data 6). These patterns of intercellular communication in Tibetan sheep suggest that while some ligand-receptor pairs may be used in broadly similar way to regulate comparable stages of germ cell development, the timing of their activation and the source of the signaling may differ. The CADM1-NECTIN3, CADM1-CADM1, and CDH2-CDH2 ligand-receptor pairs show the highest communication probability in Sertoli-germ cell communication (Fig. 8E). Cell adhesion molecule 1 (CADM1), an immunoglobulin superfamily member, plays an important role in cell-cell interactions, primarily through hemophilic binding to CADM1 in neighboring cells or heterophilic binding to other Nectin family members79. CDH2 is a calcium-dependent adhesion glycoprotein and a component of adherens junctions that mediate cell-cell adhesion and contribute to BTB80. Thus, these ligand-receptor pairs in Sertoli-germ cell communication likely support the testicular immune microenvironment and maintain BTB integrity in Tibetan sheep81. Immunofluorescence staining for CDH2 supported this interpretation by illustrating the specific localization of CDH2 at the Sertoli cell-germ cell junction (Fig. 8F). Furthermore, intercellular communication analyses in Tibetan sheep testis also identified some ligand-receptor pairs not previously linked to Sertoli-germ cell interaction (Fig. 8G). In summary, we delineate the signaling pathways and key Sertoli–germ cell ligand–receptor pairs that organize somatic–germline communication in Tibetan sheep, offering insight into spermatogenesis while treating cross-mammalian parallels as qualitative context and interpreting differences within a species/ruminant-specific—not universal—framework.
Discussion
The mammalian testis is a dynamic and heterogeneous organ that undergoes significant developmental and structural transformations during puberty, involving complex molecular regulation to achieve somatic cell proliferation and maturation, as well as the initiation of spermatogenesis. To date, molecular and genetic analyses of human and mouse testes at different postnatal stages have provided valuable insights into spermatogenesis, often serving as a reference for studies in other mammals11,44. Nevertheless, many studies rely on a limited number of markers to identify hypothetical equivalent cellular states across different species, potentially overlooking crucial functional differences. To better understand this dynamic developmental process in the Tibetan sheep testis at the molecular level, we generated a comprehensive single-nucleus transcriptional atlas of testes across four postnatal stages, revealing germ and Sertoli cell states, their transcriptional regulation, and the ligand-receptor networks mediating somatic-germline communication under long-term cold, low-oxygen conditions. These findings not only help to dissect the molecular mechanisms underlying the reproductive adaptations of male Tibetan sheep but also provide a valuable reference and conceptual framework for studying and understanding reproductive adaptations in other highland male domestic animals.
At a comprehensive level, we obtained all primary germ cell types in postnatal Tibetan sheep and reconstructed cell fate decisions and differentiation trajectories with unparalleled resolution. However, interspecific and intraspecific variations cause many marker genes to display broad expression patterns, limiting their specificity as molecular markers for particular cell types in the Tibetan sheep testis (Supplementary Fig. S2). Instead, a list of tightly regulated genes along the germ cell differentiation trajectory is needed to accurately classify cell types in Tibetan sheep. Our data depict a dynamic progression of cellular, developmental, and metabolic processes that aligns with well-organized germline development and is comparable to previous primate and rodent milestone datasets.
Undifferentiated spermatogonia subpopulations demonstrate transcriptional plasticity in mammalian testicular niches12. In this study, we systematically characterized five heterogeneous spermatogonial subpopulations (Undiff-SPG1/2/3 and Diff-SPG1/2) by single-nucleus transcriptomic profiling, with particular emphasis on delineating their regulatory signaling pathways and metabolic determinants governing differentiation commitment. A key finding is that MKI67 serves as a robust biomarker for distinguishing activated SSC populations. Although undifferentiated spermatogonia within the seminiferous tubules usually remain quiescent and function as a primitive SSC reservoir35, our data indicate that they retain the potential to transition into an active state. This state transition is closely associated with activation of the MAPK, PI3K-Akt, and mTOR signaling pathways67. Comparative analysis suggests evolutionary divergence in SSC maintenance strategies: human SSCs predominantly rely on self-renewing quiescence, whereas in mice, interconvertible SSC subpopulations have been reported. Our data extend this paradigm to Tibetan sheep, identifying coexisting quiescent (MKI67-) and activated (MKI67+) SSC populations, corresponding to the Undiff-SPG1 and Undiff-SPG2 populations, respectively. Although it remains unclear whether transitions between these two states are fully reversible, our data support a predominantly directional transition from Undiff-SPG1 to Undiff-SPG2. This process may be driven by several mechanisms: 1) quiescent Undiff-SPG1 cells exhibit low proliferative activity, and their transition to an activated state facilitates the establishment and expansion of both the SSC and progenitor pools; 2) Undiff-SPG2 shows markedly increased MAPK-mTOR pathway activity, consistent with its proliferative status; 3) this population begins to express progenitor-associated genes such as FABP7, indicating that it has progressively entered a differentiation trajectory.
Studies have shown that two populations of undifferentiated spermatogonia populations in humans and macaques cannot be strictly mapped onto classical Adark and Apale spermatogonia32. This suggests that these nuclear morphological descriptions are not strictly separable at the transcriptional level. Consistent with this, the bifurcated structure revealed by pseudotime analysis suggests that the SSC pool is not functionally homogeneous; rather, its constituents (including Undiff-SPG1 and Undiff-SPG2) possess dual potential for generating differentiating progeny and maintaining self-renewal. Therefore, we believe that Undiff-SPG1 and Undiff-SPG2 do not correspond to a simple dichotomy of self-renewal versus destined-to-differentiate cells, but rather they define transcriptionally discrete states of SSCs, allowing SSCs to adjust to a dynamic niche environment and maintain homeostatic balance in the testis. Interestingly, we found heterogeneity in the spatial distribution of stem cell-related genes within SSCs; that is, the self-renewal and differentiation of SSCs are temporally regulated through precise transcriptional programming. This finding corroborates the established paradigm of predominantly quiescent SSC pools under homeostatic conditions, where stochastic activation of a small proportion drives progenitor production. Through integrative subpopulation analysis, we propose a revised model of SSC fate regulation, wherein transcriptional plasticity rather than rigid lineage commitment governs the self-renewal-differentiation equilibrium.
During the preliminary review of all cells designated as spermatocytes (pre-leptotene, leptotene, zygotene, pachytene, and diplotene), we were unable to clearly distinguish spermatocyte subtypes. Only by performing reclustering could we successfully distinguish these subpopulations. This refined analysis identified five distinct spermatocyte subtypes that corresponded to spermatocyte populations previously described histologically. However, our analyses did not distinguish between zygotene and pachytene, nor did they capture features of secondary spermatocytes, suggesting that neither cell number nor sequencing depth was sufficient to discern very subtle transcriptomic variations. Nevertheless, we propose that meiotic recombination is most active during the transition from leptotene to pachytene and that meiotic progression is characterized by chromatin-associated signatures and dynamic metabolic changes. In addition, we characterized specific gene sets that show wave-like expression during post-meiotic spermiogenesis.
Subsequently, we resolved finer-level heterogeneity within Sertoli cells, delineated subtype-specific morphological remodeling and fundamental differences in transcriptomic functions and developmental states, thereby elucidating the molecular and cellular events underlying Sertoli cell maturation in Tibetan sheep. Most studies suggest that Sertoli cells undergo two developmental stages, transitioning from immature to mature states82,83. Another study focused on pubertal Sertoli cell development and discovered that immature Sertoli cells are partitioned into two distinct states, which eventually merge into one mature state11. However, the current research identified three major subtypes of Sertoli cells. Interestingly, we identified a transitional state (State 4) that persists from birth into adulthood. At sexual maturity, Sertoli cells in State 4 proliferate vigorously, contributing to an increase in the volume of seminiferous tubules and preparing for structural maturation. This developmental plasticity may be closely related to their functional adaptation in microenvironment regulation. The dynamic expression of cilia-related genes during Sertoli cell maturation, together with the timing of BTB remodeling, further supports this view. The transient upregulation of cilia-associated components coincides with the establishment of a “new” BTB behind migrating pre-leptotene/leptotene spermatocytes, and the controlled degeneration of the “old” BTB84, implying a role for cilia-mediated sensing and communication in coordinating niche formation. Once the testicular microenvironment is fully established and cellular associations are stabilized, the reduced requirement for cilia is reflected in their declining expression. Collectively, these transcriptional changes indicate that Sertoli cells undergo at least three distinct sequential developmental stages, and that all three states coexist in sexually mature testes, forming a stable cellular mosaic that preserves seminiferous tubule architecture and supports long-term homeostasis of the spermatogenic microenvironment.
Having clarified the various cell types in the testicular microenvironment, we next constructed an unbiased ligand-receptor interaction network among different cell types, enabling quantitative inference of cell-to-cell signaling and providing a more comprehensive view of intercellular communication mechanisms. Notably, we also identified several ligand-receptor pairs that have not been previously characterized in Sertoli-germ cell interactions. These findings highlight potential disparities across species and between datasets within the same species, establishing a foundation for future functional investigations. For example, in PTN signaling, SDC2, ALK, and PTPRZ1 were identified as receptors in Tibetan sheep, whereas SDC2 and NCL were identified in Mongolia sheep15. In human single-cell RNA sequencing data, the COL1A2 ligand of the COLLAGEN signaling pathway is primarily observed in Sertoli cells, whereas the receptor integrins α1β1, α10β1, and α2β1 are only expressed in endothelial cells13. However, our study also identified additional ligand-receptor pairs within this pathway that may mediate Sertoli-germ cell interactions. Additionally, a study of cellular communication in porcine testes showed that the ligand LAMB1 does not regulate Sertoli-germ cell interactions14, again pointing to possible species-specific differences. However, the specific roles and regulatory mechanisms of these ligand-receptor pairs in Sertoli-germ cell interactions remain unclear and warrant further investigation. Revisiting both the known and newly identified cell-cell interactions in the testis may help to decipher the functional heterogeneity of testicular cell populations and to ensure the fine-tuned regulation of the testicular microenvironment throughout spermatogenesis.
We acknowledge several limitations in the current study. First, we primarily focus on transcriptomic profiling, supplemented by protein-level validation of key markers. It is important to note, however, that RNA expression does not consistently exhibit a linear correlation with protein abundance. Additionally, while our immunofluorescence images broadly support the compartmental distribution of key germ cell and somatic markers, their resolution and signal quality do not permit definitive assignment of marker expression to individual clusters defined by snRNA-seq, especially within the undifferentiated spermatogonial compartment. Accordingly, the immunofluorescence data should be interpreted as qualitative, supportive evidence that complements the transcriptome-based cell type annotations. Second, given that sexual maturation is a gradual process, subsequent studies with larger sample sizes may provide additional details regarding developmental processes and cell-state transitions. Third, while the high-altitude environment provides crucial ecological context for this study, we did not perform direct comparative analyses with low-altitude breeds to isolate specific mechanisms underlying hypoxia or cold adaptation—primarily due to the lack of suitable control data. Consequently, future comparative studies based on this atlas are needed to directly elucidate such adaptive mechanisms. Overall, this study offers significant insights into the molecular regulatory mechanisms governing spermatogenesis in Tibetan sheep. Our data further strengthen the concept that spermatogenesis requires complex interactions among multiple signaling pathways, and these results provide the foundation for developing in vitro sheep SSC culture systems. The downstream utilization of these data will offer a theoretical basis for optimizing reproductive performance in rams and provide targets for subsequent gene function studies and molecular genetic improvement.
Materials and methods
Tibetan sheep testis sample collection
All animal research was performed in compliance with the Guide for the Care and Use of Laboratory Animals and received approval from the Ethics Committee of the Laboratory Animal Centre at Gansu Agricultural University (Ethics No. GSAU-Eth-ASF2022-008). We also complied with all relevant ethical regulations for animal use. Healthy Tibetan sheep at four developmental stages, one-day-old (T0M, newborn), three-months-old (T3M, pre-sexual maturity), one-year-old (T1Y, post-sexual maturity), and three-years-old (T3Y, adult), were selected from the Ganjia Tibetan Sheep Breeding Base (Xiahe, China). A total of 16 animals were included, with four animals per age group. To robustly capture population-level transcriptomic profiles while accounting for inter-individual variation inherent in this high-altitude-adapted species, we incorporated biological replicates in the snRNA-seq design. For each developmental stage, two independent pooled samples were prepared as biological replicates, with each pool consisting of equal amounts of testicular tissue from two randomly selected individuals (four individuals per stage in total), and each pool was processed for nuclei isolation, library construction, and sequencing. The experimental sheep were humanely euthanized according to standard procedures (electrical stunning followed by exsanguination). The left testis was rapidly collected, and morphologically intact tissue with a uniform texture was selected for subsequent paraffin section preparation and snRNA-seq, respectively.
H&E and immunofluorescence staining of testicular tissue
Fresh testis tissue blocks (~1 cm³) were fixed in 15 mL of testis tissue fixative (Servicebio, G1121) at 4 °C for 24 h. The samples were then rinsed three times with 1× PBS (10 min each) and processed for paraffin embedding using standard procedures. Paraffin sections (4–5 μm thick) were cut using a microtome (Servicebio, PM-24) and mounted onto glass slides. Before H&E and immunofluorescence staining, tissue sections were deparaffinized twice in xylene (15 min each) and rehydrated through a graded ethanol series (100%, 100%, 95%, 95%, 85%, 85%, 75%, 75%, and 50%), followed by rinsing in distilled water.
For H&E staining, sections were stained with hematoxylin followed by eosin counterstaining (Servicebio, G1005), mounted with neutral resin (Servicebio, WG10004160), and imaged using an upright bright-field microscope (Nikon Eclipse E100, Nikon, Japan) equipped with a Nikon DS-U3 imaging system (Nikon, Japan).
For immunofluorescence staining, antigen retrieval was performed by microwave heating in EDTA buffer (pH 8.0; Servicebio, G1207). After cooling to room temperature, sections were blocked with 5% BSA (Solarbio, SW3015) for 30 min at room temperature and then incubated overnight at 4 °C in a humidified chamber with primary antibodies. Sections were washed three times with 1× PBS (10 min each) and incubated with corresponding secondary antibodies for 1 h at room temperature in the dark; all antibodies were listed in Supplementary Table 1. After secondary antibody incubation, sections were washed three times with 1× PBS (10 min each). DAPI (Servicebio, G1012) working solution was then applied to the sections for nuclear counterstaining for 15 min in the dark, followed by three additional washes with 1× PBS (10 min each). Finally, sections were mounted with an antifade mounting medium (Servicebio, G1401-5ML) and protected from light before imaging. Images were acquired using a Nikon C1 confocal microscope (Nikon, Japan).
Testis nuclei isolation for snRNA-seq
After sampling, the testes of Tibetan sheep were placed in a sterile 10 cm disposable culture dish and rinsed twice with sterile PBS under aseptic conditions. The epididymis, visible connective tissue, and tunica albuginea were removed, and the tissue surface was gently blotted dry with sterile absorbent paper. The tissue was then kept on ice and minced into ~1–2 mm³ pieces, transferred to cryovials, snap-frozen in liquid nitrogen for 5 min, and stored at −80 °C for short-term storage. For quality control (QC), an additional piece of tissue from the same batch and of similar size (without mincing) was processed in parallel, snap-frozen in liquid nitrogen, and stored at −80 °C. Before the formal experiment, the QC sample was thawed, and total RNA was extracted for quality assessment; subsequent nuclei isolation and library construction were performed only after confirming that the RNA showed no obvious signs of degradation.
For nuclei isolation, frozen tissue was transferred to a pre-chilled Dounce homogenizer and homogenized in ice-cold lysis buffer (0.25 M sucrose, 5 mM CaCl₂, 3 mM MgAc₂, 10 mM Tris-HCl (pH 8.0), 1 mM DTT, 0.1 mM EDTA, 1× protease inhibitor, and 1 U/μL RiboLock RNase inhibitor) until fully dissociated. Nuclei wash buffer (PBS supplemented with 0.04% BSA, 0.2 U/μL RiboLock RNase inhibitor, 500 mM mannitol, and 0.1 mM PMSF protease inhibitor) was then added, and the homogenate was filtered through a 70 μm strainer to remove tissue debris, yielding ~1 mL of filtrate. The filtrate was gently mixed with an equal volume of ice-cold 50% iodixanol solution (0.16 M sucrose, 10 mM NaCl, 3 mM MgCl₂, 10 mM Tris-HCl (pH 7.4), 1 U/μL RiboLock RNase inhibitor, 1 mM DTT, and 0.1 mM PMSF protease inhibitor). A density gradient was prepared in a 15-mL tube by sequentially layering 1 mL of 33% and 2 mL of 30% ice-cold iodixanol, and the nuclei-containing sample was carefully layered on top of the 30% fraction. Samples were centrifuged at 4 °C and 10,000 × g for 20 min, and the white nuclei band at the 30–33% interface was collected. The nuclei fraction was washed with nuclei wash buffer and pelleted at 4 °C and 500 × g for 8 min; after repeated washing, nuclei were resuspended and filtered through a 40 μm strainer to remove residual debris and aggregates. Nuclei count and integrity were assessed by trypan blue staining using a hemocytometer, and the nuclei concentration was adjusted to 700–1200 nuclei/μL in PBS for downstream loading.
snRNA-seq library preparation and sequencing
Libraries were prepared using the Chromium Single Cell 3′ workflow (v3 chemistry) with the Chromium Single Cell 3′ GEM Kit v3, Chromium Single Cell 3′ Library Kit v3, Chromium Single Cell 3′ GEM Gel Bead Kit v3, Chromium Chip B Single Cell Kit, and Chromium i7 Multiplex Kit (10x Genomics, Pleasanton, CA, USA), according to the manufacturer’s instructions. Briefly, nuclei input was optimized based on counts and the recommended loading strategy. Nuclei were partitioned into Gel Beads-in-Emulsions (GEMs) for cell barcodes and UMI labeling, followed by in-GEM reverse transcription (53 °C for 45 min; 85 °C for 5 min). After breaking emulsions with the Recovery Agent, barcoded cDNA was purified using Dynabeads MyOne™ SILANE (10x Genomics). cDNA was amplified (98 °C for 3 min; 11 cycles of 98 °C for 15 s, 63 °C for 20 s, and 72 °C for 1 min; 72 °C for 1 min) and purified using SPRIselect beads (Beckman Coulter, Brea, CA, USA). Sequencing libraries were constructed by fragmentation/end repair and A-tailing (32 °C for 5 min; 65 °C for 30 min), size selection, adaptor ligation (20 °C for 15 min), and index PCR (98 °C for 45 s; 12 cycles of 98 °C for 20 s, 54 °C for 30 s, and 72 °C for 20 s; 72 °C for 1 min), each followed by bead purification. Final libraries were quantified using the Qubit Fluorometer (Thermo Fisher Scientific, Waltham, MA, USA) and assessed on an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). A total of eight libraries were generated, with two biological replicates per developmental stage. Paired-end sequencing (PE150) was performed on an Illumina NovaSeq 6000 (Illumina, San Diego, CA, USA) platform, generating ≥120 Gb raw data per sample.
Quantification and statistical analysis of snRNA-seq data
Raw sequencing data were processed using Cell Ranger (v3.1.0; 10x Genomics) for demultiplexing, initial quality control, read alignment, and gene expression quantification. Cell Ranger first filtered low-quality barcodes and UMIs, and then invoked STAR to align Read 2 to a Cell Ranger reference built from the CAU_O.aries_1.0 genome assembly (NCBI accession GCA_017524585.1)85 and the corresponding GTF annotation file for this assembly. Cell Ranger subsequently performed barcode/UMI correction and deduplication, conducted UMI counting based on uniquely mapped reads, and generated the feature–barcode matrix. Downstream quality control and filtering were performed in Seurat (v3.1.1) in R22. Doublets were identified and removed using DoubletFinder86 based on the proportion of artificial nearest neighbors (pANN). Nuclei were further filtered by excluding those with <200 or >5900 detected genes (nFeature_RNA), >10% mitochondrial transcripts (percent.mt), or >26,000 total UMI counts (nCount_RNA).
Nucleus clustering and cell-type annotation
After retaining high-quality nuclei and performing log-normalization, highly variable genes were identified in Seurat (v3.1.1), and the datasets were integrated prior to dimensionality reduction. We integrated the samples, conducted PCA for dimensionality reduction, and applied Harmony22 for batch effect correction using a strong correction mode (theta = 2, lambda = 1) with convergence ensured (max.iter.harmony = 10). Correction was applied over the first 50 principal components targeting the sample batch variable (orig.ident) to effectively remove technical variation while preserving biological heterogeneity. Subsequent clustering and visualization were performed based on the Harmony-corrected first 50 principal components. Specifically, a shared nearest neighbor (SNN) graph was constructed, and unsupervised clustering was performed using the Louvain algorithm to identify nuclear subpopulations. For two-dimensional visualization, t-distributed stochastic neighbor embedding (t-SNE) was applied. Cluster-specific marker genes were identified for all cell clusters using the FindAllMarkers function. Cell types were annotated by integrating information from the CellMarker 2.0 database and published literature, together with the gene expression characteristics of each cluster. Canonical markers were visualized using FeaturePlot on the t-SNE embedding and DotPlot (bubble plot) in Seurat to support cell-type annotation.
Identification and functional enrichment of differentially expressed genes
Differentially expressed genes among certain clusters or samples were identified by the Wilcoxon rank sum test in Seurat87, according to the following screening criteria: a fold change (FC) of gene expression with a cut-off of log2FC ≥ 0.36 for up-regulated genes or log2FC < 0 for down-regulated genes; a P-value ≤ 0.01; and the gene must be present in more than 25% of the cells within the targeted subpopulation. Functional enrichment analysis of differentially expressed genes was conducted using Gene Ontology (GO) and Kyoto Encyclopaedia of Genes and Genomes (KEGG) enrichment analyses. Heatmaps and Venn diagrams were generated using the online visualization platform OmicShare (https://www.omicshare.com/).
Cell cycle analysis
Gene sets for cell cycle evaluation were defined in a previous study88, resulting in four distinct gene sets: G1/S, S, G2/M, and M, corresponding to different phases of the cell cycle. Based on this, we employed the AddModuleScore function in Seurat to perform cell cycle scoring analysis on single-cell transcriptomic data, using characteristic gene sets for each phase. The specific workflow was as follows: first, all genes were sorted by expression level and divided into modules; for each phase-specific gene set, an equal number of genes were randomly selected from the corresponding module to serve as the background set; the difference in average expression between the target set and the background set was calculated for each cell as the score for that phase. The phase with the highest score was assigned as the predicted cell cycle stage for each cell. If all phase scores were below 0, the cell was considered non-cycling.
Cell trajectory analysis
Cell differentiation trajectories were generated using the Monocle2 (Version 2.10.1)89 software tool based on cell and gene expression matrices. Differentiation trajectories were produced along the inferred temporal order and colored according to pseudotime, cell state, and cell clusters. Key genes associated with developmental and differentiation processes were identified using an FDR threshold of 1e−5, and genes exhibiting similar expression patterns were grouped. Differentially expressed genes were visualized using the “plot_pseudotime_heatmap” function, based on the pseudotime of cell clusters, to create a pseudotime heatmap and a differentiation fate heatmap.
Cell-cell communication analysis
In the absence of a database for ligand-receptor pairs specific to sheep, the integration of CellPhoneDB90 and CellChat91 was employed as a repository of ligand-receptor pairs, utilizing the default configuration to evaluate cell-cell communication among various cell types, using the UMI count matrix produced by Seurat to delineate ligand-receptor pairs.
Statistics and reproducibility
Testis samples were collected from Tibetan sheep at four developmental stages. For each stage, two independent pooled samples were prepared as biological replicates (each pool contained equal amounts of testicular tissue from two individuals). In the computational analysis, the statistical analysis outlined in the specific packages was utilized; detailed procedures are provided in the Methods section.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Single-nucleus transcriptomic datasets generated in this study are available in the Genome Sequence Archive (GSA) under accession number PRJCA035469. Source data underlying all graphs are provided in Supplementary Data 7. All additional data supporting the findings of this study are provided in Supplementary Data 1–6.
Code availability
Details of publicly available software used in the study are given in the Methods section. No custom code was used in the study.
References
Yang, J. et al. Whole-genome sequencing of native sheep provides insights into rapid adaptations toextreme environments. Mol. Biol. Evol. 33, 2576–2592 (2016).
Yan, Z. et al. A time-resolved multi-omics atlas of transcriptional regulation in response to high-altitudehypoxia across whole-body tissues. Nat. Commun. 15, 3970 (2024).
Han, B. et al. Multiomics analyses provide new insight into genetic variation of reproductiveadaptability in Tibetan sheep. Mol. Biol. Evol. 41, https://doi.org/10.1093/molbev/msae058 (2024).
Morimoto, H. et al. ROS are required for mouse spermatogonial stem cell self-renewal. Cell Stem Cell 12, 774–786 (2013).
Nishimura, H. & L’Hernault, S. W. Spermatogenesis. Curr. Biol. 27, R988–R994 (2017).
Guo, J. et al. Single-cell analysis of the developing human testis reveals somatic niche cell specificationand fetal germline stem cell establishment. Cell Stem Cell 28, 764–778 (2021).
Lord, T., Oatley, M. J. & Oatley, J. M. Testicular architecture is critical for mediation of retinoic acidresponsiveness by undifferentiated spermatogonial subtypes in the mouse. Stem Cell Rep 10, 538–552 (2018).
Tan, J. H. L. et al. Infertility-causing haploinsufficiency reveals TRIM28/KAP1 requirement inspermatogonia. Stem. Cell Rep. 14, 818–827 (2020).
De Rooij, D. G. Proliferation and differentiation of spermatogonial stem cells. Reproduction 121, 347–354 (2001).
Gassei, K., Valli, H. & Orwig, K. E. Whole-mount immunohistochemistry to study spermatogonial stemcells and spermatogenic lineage development in mice, monkeys, and humans. Methods Mol. Biol. 1210, 193–202 (2014).
Guo, J. et al. The dynamic transcriptional cell atlas of testis development during human puberty. CellStem Cell 26, 262–276 (2020).
Green, C. D. et al. A comprehensive roadmap of murine spermatogenesis defined by single-cell RNA-seq. Dev. Cell 46, 651–667 (2018).
Zhao, L. et al. Single-cell analysis of developing and azoospermia human testicles reveals central roleof Sertoli cells. Nat. Commun. 11, 5683 (2020).
Wang, X. et al. Single-cell transcriptomic and cross-species comparison analyses reveal distinctmolecular changes of porcine testes during puberty. Commun. Biol. 7, 1478 (2024).
Tian, Y. et al. Single-cell RNA sequencing of the Mongolia sheep testis reveals a conserved anddivergent transcriptome landscape of mammalian spermatogenesis. FASEB J. 36, e22348 (2022).
Yang, H. et al. Characterization of sheep spermatogenesis through single-cell RNA sequencing. FASEB J. 35, e21187 (2021).
Su, J. et al. A dynamic transcriptional cell atlas of testes development after birth in Hu sheep. BMC Biol. 23, 78 (2025).
Mipam, T. et al. Single-cell transcriptome analysis and in vitro differentiation of testicular cells revealnovel insights into male sterility of the interspecific hybrid cattle-yak. BMC Genomics 24, 149 (2023).
Zhang, Y. W. et al. Single-cell analysis identifies critical regulators of spermatogonial development anddifferentiation in cattle-yak bulls. J. Dairy Sci. 107, 7317–7336 (2024).
Huang, D. et al. A single-nucleus transcriptomic atlas of primate testicular aging reveals exhaustion ofthe spermatogonial stem cell reservoir and loss of Sertoli cell homeostasis. Protein Cell 14, 888–907 (2023).
Murat, F. et al. The molecular evolution of spermatogenesis across mammals. Nature 613, 308–316 (2023).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Rotta, R., Noack, A. Multilevel local search algorithms for modularity clustering. ACM J. Exp.Algorithmics 16, https://doi.org/10.1145/1963190.1970376 (2011).
van der Maaten, L. J. P. & Hinton, G. E. Visualizing data using t-SNE. J. Mach. Learn. Res 9, 2579–2605 (2008).
Zhang, G. et al. Single-cell and spatial transcriptomic investigation reveals the spatiotemporalspecificity of the beta-defensin gene family during mouse sperm maturation. Cell Commun. Signal 22, 267 (2024).
Chen, Q. et al. CNOT3 is required for male germ cell development and spermatogonial stem cellmaintenance. Development 152, https://doi.org/10.1242/dev.204557 (2025).
Griswold, M. D. Spermatogenesis: the commitment to meiosis. Physiol. Rev. 96, 1–17 (2016).
Ribeiro, J. & Crossan, G. P. GCNA is a histone binding protein required for spermatogonial stem cellmaintenance. Nucleic Acids Res. 51, 4791–4813 (2023).
Sohni, A. et al. The neonatal and adult human testis defined at the single-cell level. Cell Rep. 26, 1501–1517 (2019).
Papetti, M. & Augenlicht, L. H. MYBL2, a link between proliferation and differentiation in maturingcolon epithelial cells. J. Cell Physiol. 226, 785–791 (2011).
Boheler, K. R. Stem cell pluripotency: a cellular trait that depends on transcription factors, chromatinstate and a checkpoint deficient cell cycle. J. Cell Physiol. 221, 10–17 (2009).
Shami, A. N. et al. Single-cell RNA sequencing of human, macaque, and mouse testes uncoversconserved and divergent features of mammalian spermatogenesis. Dev. Cell 54, 529–547 (2020).
Hermann, B. P. et al. The Mammalian spermatogenesis single-cell transcriptome, from spermatogonialstem cells to spermatids. Cell Rep. 25, 1650–1667 (2018).
Guo, J. et al. The adult human testis transcriptional cell atlas. Cell Res. 28, 1141–1157 (2018).
Wang, M. et al. Single-cell RNA sequencing analysis reveals sequential cell fate transition duringhuman spermatogenesis. Cell Stem Cell 23, 599–614 (2018).
Arnold, K. et al. Sox2+ adult stem and progenitor cells are important for tissue regeneration andsurvival of mice. Cell Stem Cell 9, 317–329 (2011).
La, H. M. et al. Identification of dynamic undifferentiated cell states within the male germline. Nat.Commun. 9, 2819 (2018).
Adikusuma, F., Pederick, D., McAninch, D., Hughes, J. & Thomas, P. Functional equivalence of theSOX2 and SOX3 transcription factors in the developing mouse brain and testes. Genetics 206, 1495–1503 (2017).
Givelet, M. et al. Transcriptional profiling of β-2M(-)SPα-6(+)THY1(+) spermatogonial stem cells inhuman spermatogenesis. Stem. Cell Rep 17, 936–952 (2022).
Wang, S. et al. scRNA-seq and scATAC-seq reveal that Sertoli cell mediates spermatogenesis disordersthrough stage-specific communications in non-obstructive azoospermia. Elife 13, https://doi.org/10.7554/eLife.97958 (2025).
Lin, W. et al. Isoform-resolution single-cell RNA sequencing reveals the transcriptional panorama ofadult Baoshan pig testis cells. BMC Genomics 26, 459 (2025).
Litchfield, K. et al. Whole-exome sequencing reveals the mutational spectrum of testicular germ celltumours. Nat. Commun. 6, 5973 (2015).
Ernst, C., Odom, D. T. & Kutter, C. The emergence of piRNAs against transposon invasion to preservemammalian genome integrity. Nat. Commun. 8, 1411 (2017).
Nie, X. et al. Single-cell analysis of human testis aging and correlation with elevated body mass index. Dev. Cell 57, 1160–1176 (2022).
Kubota, H. et al. Glial cell line-derived neurotrophic factor and endothelial cells promote self-renewalof rabbit germ cells with spermatogonial stem cell properties. FASEB J. 25, 2604–2614 (2011).
Lau, X., Munusamy, P., Ng, M. J. & Sangrithi, M. Single-cell RNA sequencing of the cynomolgusmacaque testis reveals conserved transcriptional profiles during mammalian spermatogenesis. Dev.Cell 54, 548–566 (2020).
Tan, H. et al. Single-cell RNA-seq uncovers dynamic processes orchestrated by RNA-binding proteinDDX43 in chromatin remodeling during spermiogenesis. Nat. Commun. 14, 2499 (2023).
Fliegauf, M. et al. Mislocalization of DNAH5 and DNAH9 in respiratory cells from patients with primaryciliary dyskinesia. Am. J. Respir. Crit. Care Med. 171, 1343–1349 (2005).
Su, J. et al. Dosage compensation of the X chromosome during sheep testis development revealed bysingle-cell RNA sequencing. Animals 12, https://doi.org/10.3390/ani12172169 (2022).
Whelan, E. C. et al. Single-cell multiomic comparison of mouse and rat spermatogenesis reveals generegulatory networks conserved for over 20 million years. Stem Cell Rep. 20, 102449 (2025).
Yi, S. et al. Tcf12, A member of basic helix-loop-helix transcription factors, mediates bone marrowmesenchymal stem cell osteogenic differentiation in vitro and in vivo. Stem Cells 35, 386–397 (2017).
Takubo, K. et al. Stem cell defects in ATM-deficient undifferentiated spermatogonia through DNAdamage-induced cell-cycle arrest. Cell Stem Cell 2, 170–182 (2008).
Unhavaithaya, Y. et al. MILI, a PIWI-interacting RNA-binding protein, is required for germ line stem cellself-renewal and appears to positively regulate translation. J. Biol. Chem. 284, 6507–6519 (2009).
Wang, X. et al. A homozygous PIWIL2 frameshift variant affects the formation and maintenance ofhuman-induced pluripotent stem cell-derived spermatogonial stem cells and causes Sertoli cell-onlysyndrome. Stem Cell Res. Ther. 13, 480 (2022).
Lei, N. et al. Sex-specific differences in mouse DMRT1 expression are both cell type- and stage-dependent during gonad development. Biol. Reprod. 77, 466–475 (2007).
Li, C. H. et al. Long-term propagation of tree shrew spermatogonial stem cells in culture andsuccessful generation of transgenic offspring. Cell Res. 27, 241–252 (2017).
Law, N. C., Oatley, M. J. & Oatley, J. M. Developmental kinetics and transcriptome dynamics of stemcell specification in the spermatogenic lineage. Nat. Commun. 10, 2787 (2019).
Dann, C. T. et al. Spermatogonial stem cell self-renewal requires OCT4, a factor downregulated duringretinoic acid-induced differentiation. Stem Cells 26, 2928–2937 (2008).
Buaas, F. W. et al. Plzf is required in adult male germ cells for stem cell self-renewal. Nat. Genet. 36, 647–652 (2004).
Zhang, T., Oatley, J., Bardwell, V. J. & Zarkower, D. DMRT1 is required for mouse spermatogonial stemcell maintenance and replenishment. PLoS Genet 12, e1006293 (2016).
Suzuki, H. et al. SOHLH1 and SOHLH2 coordinate spermatogonial differentiation. Dev. Biol. 361, 301–312 (2012).
Li, Z. et al. Transcriptional priming as a conserved mechanism of lineage diversification in thedeveloping mouse and human neocortex. Sci. Adv. 6, eabd2068 (2020).
Delile, J. et al. Single cell transcriptomics reveals spatial and temporal dynamics of gene expression inthe developing mouse spinal cord. Development 146, https://doi.org/10.1242/dev.173807 (2019).
Valli, H. et al. Fluorescence- and magnetic-activated cell sorting strategies to isolate and enrichhuman spermatogonial stem cells. Fertil. Steril. 102, 566–580 (2014).
Bardot, E. et al. Foxa2 identifies a cardiac progenitor population with ventricular differentiationpotential. Nat. Commun. 8, 14428 (2017).
Lord, T. & Nixon, B. Metabolic changes accompanying spermatogonial stem cell differentiation. Dev.Cell 52, 399–411 (2020).
Suzuki, S., McCarrey, J. R. & Hermann, B. P. An mTORC1-dependent switch orchestrates the transitionbetween mouse spermatogonial stem cells and clones of progenitor spermatogonia. Cell Rep 34, 108752 (2021).
Hyakutake, K. et al. Asymmetrical allocation of JAK1 mRNA during spermatogonial stem cell division inXenopus laevis. Dev. Growth Differ. 57, 389–399 (2015).
Wang, H. Q. et al. Meiotic transcriptional reprogramming mediated by cell-cell communications inhumans and mice revealed by scATAC-seq and scRNA-seq. Zool. Res. 45, 601–616 (2024).
Di Persio, S. & Neuhaus, N. Human spermatogonial stem cells and their niche in male (in) fertility:novel concepts from single-cell RNA-sequencing. Hum. Reprod. 38, 1–13 (2023).
Handel, M. A. & Schimenti, J. C. Genetics of mammalian meiosis: regulation, dynamics and impact onfertility. Nat. Rev. Genet. 11, 124–136 (2010).
Suzuki, S., Diaz, V. D. & Hermann, B. P. What has single-cell RNA-seq taught us about mammalianspermatogenesis? Biol. Reprod. 101, 617–634 (2019).
Li, L. et al. A novel frameshift mutation in ubiquitin-specific protease 26 gene in a patient with severeoligozoospermia. Biosci. Rep. 40, https://doi.org/10.1042/bsr20191902 (2020).
Zhao, Y. et al. Connexin-43 is a promising target for lycopene preventing phthalate-inducedspermatogenic disorders. J. Adv. Res. 49, 115–126 (2023).
Mays-Hoopes, L. L., Bolen, J., Riggs, A. D. & Singer-Sam, J. Preparation of spermatogonia,spermatocytes, and round spermatids for analysis of gene expression using fluorescence-activatedcell sorting. Biol. Reprod. 53, 1003–1011 (1995).
Sabari, B. R., Zhang, D., Allis, C. D. & Zhao, Y. Metabolic regulation of gene expression through histoneacylations. Nat. Rev. Mol. Cell Biol. 18, 90–101 (2017).
Ward, W. S. Function of sperm chromatin structural elements in fertilization and development. Mol.Hum. Reprod. 16, 30–36 (2010).
Liu, C. et al. Autophagy is required for ectoplasmic specialization assembly in sertoli cells. Autophagy 12, 814–832 (2016).
Gewaily, M. S. et al. Comparative expression of cell adhesion molecule1 (CADM1) in the testes ofexperimental mice and some farm animals. Acta Histochem 122, 151456 (2020).
Cai, Z. et al. ALKBH5 in mouse testicular Sertoli cells regulates CDH2 mRNA translation to maintainblood-testis barrier integrity. Cell Mol. Biol. Lett. 27, 101 (2022).
Mruk, D. D. & Cheng, C. Y. The mammalian blood-testis barrier: its biology and regulation. Endocr. Rev. 36, 564–591 (2015).
Simorangkir, D. R. et al. Sertoli cell differentiation in rhesus monkey (Macaca mulatta) is an earlyevent in puberty and precedes attainment of the adult complement of undifferentiatedspermatogonia. Reproduction 143, 513–522 (2012).
Meroni, S. B. et al. Molecular mechanisms and signaling pathways involved in Sertoli cell proliferation. Front Endocrinol 10, 224 (2019).
Ou, Y. et al. Primary cilia in the developing pig testis. Cell Tissue Res 358, 597–605 (2014).
Li, X. et al. Genomic analyses of wild argali, domestic sheep, and their hybrids provide insights intochromosome evolution, phenotypic variation, and germplasm innovation. Genome Res. 32, 1669–1684 (2022).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNAsequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337 (2019).
Camp, J. G. et al. Multilineage communication regulates human liver bud development frompluripotency. Nature 546, 533–538 (2017).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporalordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell-cellcommunication from combined expression of multi-subunit ligand-receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 32260833 & No. 32202675), and Discipline Team Project of Gansu Agricultural University (NO. GAU-XKTD-2022-20). We thank the Guangzhou Genedenovo Biotechnology Co., Ltd., for assisting in single-nucleus RNA sequencing. Schematic illustration elements in Figs. 1C, F, 3E, 4D, and 8F were created using Figdraw (www.figdraw.com), an online platform for scientific illustration. The use of these illustration elements complies with the platform’s Terms of Use for academic publishing. License information (Agreement No.) is as follows: Fig. 1C, SSUYUdd744; Fig. 1F, PRRSOc2e8d; Fig. 3E, OIPTOb8ffc; Fig. 4D, YUPUPae06c; Fig. 8F, YSRAO2ad8d.
Author information
Authors and Affiliations
Contributions
Conceptualization: H.H.W., Y.J.M. Methodology: H.H.W., D.P.L., Z.L.L., C.H.W. Investigation: X.X.Z., Y.J.Z., T.T.L., T.Z.S. Formal analysis: H.H.W., T.T.L. Visualization: H.H.W. Supervision: T.Z.S., Y.J.M. Writing—original draft: H.H.W. Writing—review & editing: H.H.W., T.T.L., Y.J.M. Funding acquisition: T.T.L., Y.J.M. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: David Favero.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Hh., Li, Tt., Li, Dp. et al. Single-nucleus RNA sequencing provides insights into the genetic mechanisms underlying reproductive adaptability in Tibetan sheep (Ovis aries). Commun Biol 9, 452 (2026). https://doi.org/10.1038/s42003-026-09729-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-026-09729-1