Abstract
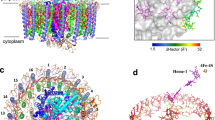
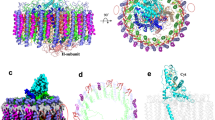
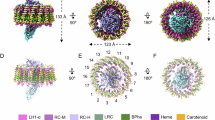
The marine purple nonsulfur phototrophic bacterium Rhodovulum (Rdv.) sulfidophilum (Alphaproteobacteria) has been a model organism for bacterial photosynthesis research because of its unusual ability to grow phototrophically (anoxic/light) using high concentrations of inorganic or organic sulfur compounds as electron donors or by respiration under fully oxic conditions. Here we present a 1.81 Å-resolution cryo-EM structure of the light-harvesting 1–reaction center (LH1–RC) photocomplex from the Rdv. sulfidophilum type strain W4 with a focus on RC structure and function. The Rdv. sulfidophilum RC is characterized by its cytochrome (Cyt) subunit that contains three heme groups and is anchored by its intact N-terminal domain in the membrane. In contrast to a methionine as the 6th axial ligand to the heme-2 in other bacterial RC-bound triheme and tetraheme Cyt subunits, the outmost heme-2 in the Rdv. sulfidophilum Cyt subunit is ligated by a cysteine residue, resulting in a significant downshift of reduction potential of 470 mV compared to that of a methionine-ligated heme-2. A nonheme Fe ligated by a histidine of the Cyt subunit and five water molecules was identified in close proximity to heme-2, implying a potential role in electron transport from soluble electron donors to heme-2. The Rdv. sulfidophilum LH1 complex forms an open ring structure consisting of 16 αβ-subunits with a gap formed where the N-terminal transmembrane domain of the RC Cyt subunit and a newly identified protein with three helical domains (designated as protein-3h) are located. Protein-3h corresponds to the truncated N-terminal fragment of a gene product encoded by the pseudo-gene urf1 in the NADH:ubiquinone oxidoreductase (complex I) nuo operon in the genome of Rdv. sulfidophilum W4. Genes urf1 are also found in other purple nonsulfur bacteria and in aerobic anoxygenic phototrophic bacteria, and their putative products all share a common structural motif of N-terminal transmembrane U-shaped tandem helices. Based on structural and spectroscopic data, possible electron transfer pathways between the Rdv. sulfidophilum RC Cyt subunit and soluble electron donors and potential roles of protein-3h in the structural integrity of LH1–RC are discussed.
Similar content being viewed by others
Data availability
Maps and models have been deposited in the EMDB and PDB with the accession codes: EMD-66181 and extended PDB ID pdb_00009WQV (PDB-9WQV) for the Rdv. sulfidophilum LH1–RC. The numerical source values underlying Fig. 1d, Fig. 3, and Supplementary Fig. 3 can be found in Supplementary data. Uncropped and unedited gel images underlying Fig. 1d and Supplementary Fig. 12a can be found in Supplementary Fig. 15. All other data are available from the authors upon reasonable request.
References
Hansen, T. A. & Veldkamp, H. Rhodopseudomonas sulfidophila, nov. spec., a new species of the purple nonsulfur bacteria. Arch. Mikrobiol. 92, 45–58 (1973).
Hagemann, G. E., Katsiou, E., Forkl, H., Steindorf, A. C. J. & Tadros, M. H. Gene cloning and regulation of gene expression of the puc operon from Rhodovulum sulfidophilum. Biochim. Biophys. Acta 1351, 341–358 (1997).
Masuda, S. et al. Structural and functional analyses of photosynthetic regulatory genes regA and regB from Rhodovulum sulfidophilum, Roseobacter denitrificans, and Rhodobacter capsulatus. J. Bacteriol. 181, 4205–4215 (1999).
Masuda, S., Nagashima, K. V. P., Shimada, K. & Matsuura, K. Transcriptional control of expression of genes for photosynthetic reaction center and light-harvesting proteins in the purple bacterium Rhodovulum sulfidophilum. J. Bacteriol. 182, 2778–2786 (2000).
Masuda, S., Yoshida, M., Nagashima, K. V. P., Shimada, K. & Matsuura, K. A new cytochrome subunit bound to the photosynthetic reaction center in the purple bacterium, Rhodovulum sulfidophilum. J. Biol. Chem. 274, 10795–10801 (1999).
Alric, J. et al. Structural and functional characterization of the unusual triheme cytochrome bound to the reaction center of Rhodovulum sulfidophilum. J. Biol. Chem. 279, 26090–26097 (2004).
Dracheva, S. M. et al. Electrogenic steps in the redox reactions catalyzed by photosynthetic reaction-center complex from Rhodopseudomonas viridis. Eur. J. Biochem. 171, 253–264 (1988).
Osyczka, A. et al. Interaction site for soluble cytochromes on the tetraheme cytochrome subunit bound to the bacterial photosynthetic reaction center mapped by site-directed mutagenesis. Biochemistry 37, 11732–11744 (1998).
Nitschke, W., Jubault-Bregler, M. & Rutherford, A. W. The reaction center associated tetraheme cytochrome subunit from Chromatium vinosum revisited: A reexamination of its EPS properties. Biochemistry 32, 8871–8879 (1993).
Masuda, S. et al. Mutational analyses of the photosynthetic reaction center-bound triheme cytochrome subunit and cytochrome c2 in the purple bacterium Rhodovulum sulfidophilum. Biochemistry 41, 11211–11217 (2002).
Kimura, Y. et al. A new membrane-bound cytochrome c works as an electron donor to the photosynthetic reaction center complex in the purple bacterium, Rhodovulum sulfidophilum. J. Biol. Chem. 282, 6463–6472 (2007).
Wang, W. et al. Cryo-EM analysis of a tri-heme cytochrome-associated RC-LH1 complex from the marine photoheterotrophic bacterium Dinoroseobacter shibae. Adv. Sci. 12, 2413456 (2025).
Liu, Z.-K. et al. Structural basis for aerobic anoxygenic photosynthesis in the reaction center–light-harvesting 1 (RC–LH1) supercomplex of Dinoroseobacter shibae. Comms. Biol. 8, 1565 (2025).
Gucwa, M., Bijak, V., Zheng, H., Murzyn, K. & Minor, W. CheckMyMetal (CMM): validating metal-binding sites in X-ray and cryo-EM data. IUCrJ 11, 871–877 (2024).
Kimura, Y. et al. Salt- and pH-dependent thermal stability of photocomplexes from extremophilic bacteriochlorophyll b-containing Halorhodospira species. Microorganisms 10, 959 (2022).
Qi, C.-H. et al. Structural insights into the unusual core photocomplex from a triply extremophilic purple bacterium, Halorhodospira halochloris. J. Integr. Plant Biol. 66, 2262–2272 (2024).
Tani, K. et al. A native LH1–RC–HiPIP supercomplex from an extremophilic phototroph. Commun. Biol. 8, 42 (2025).
Tani, K. et al. Structure and biochemistry of the LH1–RC photocomplex from the halophilic purple bacterium, Rhodothalassium salexigens. Biochemistry 64, 3070–3080 (2025).
Qian, P., Siebert, C. A., Wang, P., Canniffe, D. P. & Hunter, C. N. Cryo-EM structure of the Blastochloris viridis LH1-RC complex at 2.9 Å. Nature 556, 203–208 (2018).
Aasa, R. Powder line shapes in the electron paramagnetic resonance spectra of high-spin ferric complexes. J. Chem. Phys. 52, 3919–3930 (1970).
Gaffney, B. EPR of mononuclear non-heme iron proteins. in High Resolution EPR: Applications to Metalloenzymes and Metals in Medicine (eds. Hanson, G. & Berliner, L.) 233-268 (Springer, 2009).
Huck, O., Schiltz, E., Drews, G. & Labahn, A. Sequence analysis reveals new membrane anchor of reaction centre-bound cytochromes possibly related to PufX. FEBS Lett. 535, 166–170 (2003).
Tani, K. et al. An LH1–RC photocomplex from an extremophilic phototroph provides insight into origins of two photosynthesis proteins. Commun. Biol. 5, 1197 (2022).
Wang, X.-P. et al. Insights into the divergence of the photosynthetic LH1 complex obtained from structural analysis of the unusual photocomplexes of Roseospirillum parvum. Commun. Biol. 7, 1658 (2024).
Xin, Y. et al. Cryo-EM structure of the RC-LH core complex from an early branching photosynthetic prokaryote. Nat. Commun. 9, 1568 (2018).
Huang, G. et al. Cryo-EM structure of a minimal reaction center–light-harvesting complex from the phototrophic bacterium Chloroflexus aurantiacus. J. Integr. Plant Biol. 67, 967–978 (2025).
Dupuis, A. et al. The complex I from Rhodobacter capsulatus. Biochim. Biophys. Acta 1364, 147–165 (1998).
Tichi, M. A., Meijer, W. G. & Tabita, F. R. Complex I and its involvement in redox homeostasis and carbon and nitrogen metabolism in Rhodobacter capsulatus. J. Bacteriol. 183, 7285–7294 (2001).
Swainsbury, D. J. K., Qian, P., Hitchcock, A. & Hunter, C. N. The structure and assembly of reaction centre-light-harvesting 1 complexes in photosynthetic bacteria. Biosci. Rep. 43, BRS20220089 (2023).
Liu, L.-N., Bracun, L. & Li, M. Structural diversity and modularity of photosynthetic RC–LH1 complexes. Trends Microbiol 32, 38–52 (2024).
Kimura, Y. et al. The thermo-stable LH1–RC complex of a hot spring purple bacterium powers photosynthesis with extremely low-energy near-infrared light. Biochemistry 64, 170–179 (2025).
Tani, K. et al. A distinct double-ring LH1–LH2 photocomplex from an extremophilic phototroph. Nat. Commun. 16, 1410 (2025).
Yu, L.-J., Suga, M., Wang-Otomo, Z.-Y. & Shen, J.-R. Structure of photosynthetic LH1-RC supercomplex at 1.9 Å resolution. Nature 556, 209–213 (2018).
Qian, P. et al. Cryo-EM structure of the monomeric Rhodobacter sphaeroides RC-LH1 core complex at 2.5 Å. Biochem. J. 478, 3775–3790 (2021).
Qian, P. et al. Cryo-EM structure of the dimeric Rhodobacter sphaeroides RC-LH1 core complex at 2.9 Å: the structural basis for dimerisation. Biochem. J. 478, 3923–3937 (2021).
Tani, K. et al. Asymmetric structure of the native Rhodobacter sphaeroides dimeric LH1-RC complex. Nat. Commun. 13, 1904 (2022).
Cao, P. et al. Structural basis for the assembly and quinone transport mechanisms of the dimeric photosynthetic RC–LH1 supercomplex. Nat. Commun. 13, 1977 (2022).
Tani, K. et al. Rhodobacter capsulatus forms an unusually compact crescent-shaped LH1–RC photocomplex. Nat. Commun. 14, 846 (2023).
Bracun, L., Yamagata, A., Christianson, B. M., Shirouzu, M. & Liu, L.-N. Cryo-EM structure of a monomeric RC-LH1-PufX supercomplex with high-carotenoid content from Rhodobacter capsulatus. Structure 31, 318–328 (2023).
Wang, P. et al. Architectures of photosynthetic RC-LH1 supercomplexes from Rhodobacter blasticus. Sci. Adv. 10, eadp6678 (2024).
Kimura, Y., Tani, K., Madigan, M. T. & Wang-Otomo, Z.-Y. Advances in the spectroscopic and structural characterization of core light-harvesting complexes from purple phototrophic bacteria. J. Phys. Chem. B 127, 6–17 (2023).
Tsukatani, Y. et al. Phylogenetic distribution of unusual triheme to tetraheme cytochrome subunit in the reaction center complex of purple bacteria. Photosynth. Res. 79, 83–91 (2004).
Knaff, D. B. et al. Reaction of cytochrome c2 with photosynthetic reaction centers from Rhodopseudomonas viridis. Biochemistry 30, 1303–1310 (1991).
Ortega, J. M., Drepper, F. & Mathis, P. Electron transfer between cytochrome c2 and the tetraheme cytochrome c in Rhodopseudomonas viridis. Photosynth. Res. 59, 147–157 (1999).
Osyczka, A., Nagashima, K. V. P., Shimada, K. & Matsuura, K. Interaction site for high-potential iron-sulfur protein on the tetraheme cytochrome subunit bound to the photosynthetic reaction center of Rubrivivax gelatinosus. Biochemistry 38, 2861–2865 (1999).
Osyczka, A. et al. Comparison of the binding sites for high-potential iron-sulfur protein and cytochrome c on the tetraheme cytochrome subunit bound to the bacterial photosynthetic reaction center. Biochemistry 38, 15779–15790 (1999).
Venturoli, G. et al. Electron transfer from HiPIP to the photooxidized tetraheme cytochrome subunit of Allochromatium vinosum reaction center: new insights from site-directed mutagenesis and computational studies. Biochemistry 43, 437–445 (2004).
Kawakami, T. et al. Crystal structure of a photosynthetic LH1-RC in complex with its electron donor HiPIP. Nat. Commun. 12, 1104 (2021).
Evans, R. et al. Protein complex prediction with AlphaFold-Multimer. bioRxiv, https://doi.org/10.1101/2021.10.04.463034 (2022).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Khanppnavar, B., Leka, O., Pal, S. K., Korkhov, V. M. & Kammerer, R. A. Cryo-EM structure of the botulinum neurotoxin A/SV2B complex and its implications for translocation. Nat. Commun. 16, 1224 (2025).
Heering, H. A., Bulsink, Y. B. M., Hagen, W. R. & Meyer, T. E. Influence of charge and polarity on the redox potentials of high-potential Iron–sulfur proteins: evidence for the existence of two groups. Biochemistry 34, 14675–14686 (1995).
Lieutaud, C., Alric, J., Bauzan, M., Nitschke, W. & Schoepp-Cothenet, B. Study of the high-potential iron sulfur protein in Halorhodospira halophila confirms that it is distinct from cytochrome c as electron carrier. Proc. Natl. Acad. Sci. 102, 3260–3265 (2005).
Hanlon, S. P., Holt, R. A., Moore, G. R. & McEwan, A. G. Isolation and characterization of a strain of Rhodobacter sulfidophilus: a bacterium which grows autotrophically with dimethylsulfide as electron donor. Microbiology 140, 1953–1958 (1994).
McDevitt, C. A., Hugenholtz, P., Hanson, G. & McEwan, A. G. Molecular analysis of dimethyl sulphide dehydrogenase from Rhodovulum sulfidophilum: its place in the dimethyl sulphoxide reductase family of microbial molybdopterin-containing enzymes. Mol. Microbiol. 44, 1575–1587 (2002).
Herter, S. M., Kortlüke, C. M. & Drews, G. Complex I of Rhodobacter capsulatus and its role in reverted electron transport. Arch. Microbiol. 169, 98–105 (1998).
Tani, K. et al. A previously unrecognized membrane protein in the Rhodobacter sphaeroides LH1-RC photocomplex. Nat. Commun. 12, 6300 (2021).
Sekine, F. et al. Gene sequencing and characterization of the light-harvesting complex 2 from thermophilic purple sulfur bacterium Thermochromatium tepidum. Photosynth. Res. 111, 9–18 (2012).
Takaichi, S. et al. Direct injection of pigment–protein complexes and membrane fragments suspended in water from phototrophs to C18 HPLC. Photosynth. Res. 144, 101–107 (2020).
Kimura, Y. et al. Characterization of the quinones in purple sulfur bacterium Thermochromatium tepidum. FEBS Lett. 589, 1761–1765 (2015).
Nagatsuma, S. et al. Phospholipid distributions in purple phototrophic bacteria and LH1-RC core complexes. Biochim. Biophys. Acta – Bioenerg. 1860, 461–468 (2019).
Stoll, S. & Schweiger, A. EasySpin, a comprehensive software package for spectral simulation and analysis in EPR. J. Magn. Reson. 178, 42–55 (2006).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for a rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Wagner, T. & Raunser, S. The evolution of SPHIRE-crYOLO particle picking and its application in automated cryo-EM processing workflows. Commun. Biol. 3, 61 (2020).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Cryst. D66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Cryst. D66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 66, 12–21 (2010).
DeLano, W.L. The PyMOL molecular graphics system, DeLano Scientific, LCC, San Carlos, CA, USA. (2004).
Pettersen, E. F. et al. UCSF ChimeraX: Structure visualization for researchers, editors, and developers. Protein Sci. 30, 70–82 (2021).
Acknowledgements
The high-resolution cryo-EM data used in this study were acquired at the Core-Facility Portal of Okayama University (CFPOU RIIS-n01). We thank Prof. Jian-Ren Shen and Dr. Nobutaka Numoto for their assistance in data collection. This research was partially supported by the National Key R&D Program of China (No. 2022YFC3401800), Platform Project for Supporting Drug Discovery and Life Science Research (Basis for Supporting Innovative Drug Discovery and Life Science Research (BINDS)) from AMED under Grant Numbers JP21am0101118 and JP21am0101116, and JP23ama121004. R.K., E.R.P., T. M., and B.M.H. acknowledge the generous support of Dr. Malgorzata Hall, the Okinawa Institute of Science and Technology (OIST), Scientific Computing & Data Analysis Section, and Scientific Imaging Section at OIST, and the Japanese Cabinet Office. R.K. acknowledges the support from Prof. Tsumoru Shintake. M.T.M. was supported in part by NASA Cooperative Agreement 80NSSC21M0355. This work was supported in part by JSPS KAKENHI (Grant Numbers 22K06111, 23K05822, 24H02084, 24K01620 and 24H02078), Center for Quantum and Information Life Sciences, University of Tsukuba, JST-Mirai Program (Grant Number JPMJMI22I3) and MEXT Joint Usage/Research Promotion Project: CURE JPMXP1323015488 (Spin-L program No spin25XN018).
Author information
Authors and Affiliations
Contributions
L.-J.Y., Z.-Y.W.-O., Y.Ki., and K.T. designed the work, K.V.P.N. provided materials, X.-Y.Y., G.-L.W., S.K., and Y.-L. W., Y.Ko., T.S., K.T., R.K., E.R.P., S.T., T.M., and K.V.P.N. performed the experiments, K.T., L.-J.Y., H.M., M.T.M., A.M., B.M.H., Y.Ki., and Z.-Y.W.-O. analyzed data, L.-J.Y., Z.-Y.W.-O., H.M., K.T., Y.Ki. and M.T.M. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Xiaoling Xu and Laura Rodríguez Pérez. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yue, XY., Wang, GL., Kosaki, S. et al. Structural insights into the photochemistry of the LH1–RC complex from the marine purple phototrophic bacterium Rhodovulum sulfidophilum. Commun Biol (2026). https://doi.org/10.1038/s42003-026-09755-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-026-09755-z