Abstract
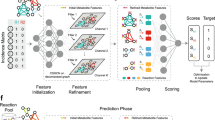
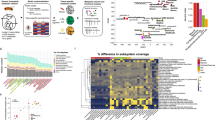
Genome-scale metabolic models (GEMs) are indispensable tools for probing cellular metabolism, enabling predictions of metabolic fluxes, guiding strain optimization, and advancing biomedical research. However, their predictive capacity is often compromised by incomplete reaction networks, stemming from gaps in biochemical knowledge, annotation inaccuracies, and insufficient experimental validations. Here we present MuSHIN (Multi-way SMILES-based Hypergraph Interface Network), a deep hypergraph learning method that integrates network topology with biochemical domain knowledge to predict missing reactions in GEMs. Evaluated on 926 high- and intermediate-quality GEMs with artificially removed reactions, MuSHIN achieves up to a 17% improvement over the current state-of-the-art method across multiple evaluation metrics. Furthermore, MuSHIN substantially enhances phenotypic predictions in 24 draft GEMs associated with fermentation by resolving critical metabolic gaps, as validated against experimental measurements. Together, these findings highlight MuSHIN’s potential to advance GEM reconstruction and accelerate discoveries in systems biology, metabolic engineering, and precision medicine.
Similar content being viewed by others
Data availability
The datasets used and analyzed during the current study are included within this article and its Supplementary Information file. The raw data were collected from publicly available databases: ChEBI (https://www.ebi.ac.uk/chebi/), BiGG Models (http://bigg.ucsd.edu/), AGORA Models (https://www.vmh.life). The source data for the figures are provided in the Supplementary Data file. More details can be found in Supplementary Note 5.
Code availability
The source code for our framework is available at Github51 [https://github.com/cyixiao/MuSHIN].
References
Thiele, I., Price, N. D., Vo, T. D. & Palsson, B. Ø Candidate metabolic network states in human mitochondria: Impact of diabetes, ischemia, and diet. J. Biol. Chem. 280, 11683–11695 (2005).
Thiele, I., Jamshidi, N., Fleming, R. M. & Palsson, B. Ø Genome-scale reconstruction of Escherichia coli’s transcriptional and translational machinery: a knowledge base, its mathematical formulation, and its functional characterization. PLoS Comput. Biol. 5, e1000312 (2009).
Lee, S. Y. & Kim, H. U. Systems strategies for developing industrial microbial strains. Nat. Biotechnol. 33, 1061–1072 (2015).
Gu, C., Kim, G. B., Kim, W. J., Kim, H. U. & Lee, S. Y. Current status and applications of genome-scale metabolic models. Genome Biol. 20, 121 (2019).
Lieven, C. et al. Memote for standardized genome-scale metabolic model testing. Nat. Biotechnol. 38, 272–276 (2020).
Simeonidis, E. & Price, N. D. Genome-scale modeling for metabolic engineering. J. Ind. Microbiol. Biotechnol. 42, 327–338 (2015).
Kim, B., Kim, W. J., Kim, D. I. & Lee, S. Y. Applications of genome-scale metabolic network model in metabolic engineering. J. Ind. Microbiol. Biotechnol. 42, 339–348 (2015).
Raškevičius, V. et al. Genome scale metabolic models as tools for drug design and personalized medicine. PloS One 13, e0190636 (2018).
Robinson, J. L. & Nielsen, J. Anticancer drug discovery through genome-scale metabolic modeling. Curr. Opin. Syst. Biol. 4, 1–8 (2017).
King, Z. A. et al. Bigg models: a platform for integrating, standardizing and sharing genome-scale models. Nucleic Acids Res. 44, D515–D522 (2016).
Nielsen, J. & Keasling, J. D. Engineering cellular metabolism. Cell 164, 1185–1197 (2016).
Chen, C., Liao, C. & Liu, Y.-Y. Teasing out missing reactions in genome-scale metabolic networks through hypergraph learning. Nat. Commun. 14, 2375 (2023).
Liu, X. et al. A generalizable framework for unlocking missing reactions in genome-scale metabolic networks using deep learning. arXiv preprint arXiv:2409.13259 (2024).
Pan, S. & Reed, J. L. Advances in gap-filling genome-scale metabolic models and model-driven experiments lead to novel metabolic discoveries. Curr. Opin. Biotechnol. 51, 103–108 (2018).
Benedict, M. N., Mundy, M. B., Henry, C. S., Chia, N. & Price, N. D. Likelihood-based gene annotations for gap filling and quality assessment in genome-scale metabolic models. PLoS Comput. Biol. 10, e1003882 (2014).
Karp, P. D., Weaver, D. & Latendresse, M. How accurate is automated gap filling of metabolic models? BMC Syst. Biol. 12, 73 (2018).
Orth, J. D. & Palsson, B. Ø Systematizing the generation of missing metabolic knowledge. Biotechnol. Bioeng. 107, 403–412 (2010).
Schroeder, W. L. & Saha, R. Optfill: a tool for infeasible cycle-free gapfilling of stoichiometric metabolic models. IScience 23, 100783 (2020).
Prigent, S. et al. Meneco, a topology-based gap-filling tool applicable to degraded genome-wide metabolic networks. PLoS Comput. Biol. 13, e1005276 (2017).
Satish Kumar, V., Dasika, M. S. & Maranas, C. D. Optimization based automated curation of metabolic reconstructions. BMC Bioinforma. 8, 212 (2007).
Henry, C. S. et al. High-throughput generation, optimization and analysis of genome-scale metabolic models. Nat. Biotechnol. 28, 977–982 (2010).
Almeida, A. et al. A new genomic blueprint of the human gut microbiota. Nature 568, 499–504 (2019).
Machado, D., Andrejev, S., Tramontano, M. & Patil, K. R. Fast automated reconstruction of genome-scale metabolic models for microbial species and communities. Nucleic Acids Res. 46, 7542–7553 (2018).
Chen, C. & Liu, Y.-Y. A survey on hyperlink prediction. IEEE Trans. Neural Netw. Learn. Syst. 35, 15034–15050 (2023).
Feng, Y., You, H., Zhang, Z., Ji, R. & Gao, Y. Hypergraph neural networks. Proc. AAAI Conf. Artif. Intell. 33, 3558–3565 (2019).
Bai, S., Zhang, F. & Torr, P. H. Hypergraph convolution and hypergraph attention. Pattern Recognit. 110, 107637 (2021).
Chen, C., Surana, A., Bloch, A. M. & Rajapakse, I. Controllability of hypergraphs. IEEE Trans. Netw. Sci. Eng. 8, 1646–1657 (2021).
Chen, C. & Rajapakse, I. Tensor entropy for uniform hypergraphs. IEEE Trans. Netw. Sci. Eng. 7, 2889–2900 (2020).
Berge, C. Hypergraphs: Combinatorics of Finite Sets Vol. 45 (Elsevier, 1984).
Zhou, D., Huang, J. & Schölkopf, B. Learning with hypergraphs: clustering, classification, and embedding. Adv. Neural Inform. Process. Syst. 19, 1601–1608 (2006).
Gao, Y. et al. Hypergraph learning: methods and practices. IEEE Trans. Pattern Anal. Mach. Intell. 44, 2548–2566 (2020).
Zhang, M., Cui, Z., Jiang, S. & Chen, Y. Beyond link prediction: Predicting hyperlinks in adjacency space. In Proceedings of the AAAI Conference on Artificial Intelligence Vol. 32, (2018).
Sharma, G., Patil, P. & Murty, M. N. C3mm: clique-closure based hyperlink prediction. IJCAI 20, 3364–3370 (2020).
Yadati, N. et al. Nhp: Neural hypergraph link prediction. In Proceedings of the 29th ACM International Conference on Information & Knowledge Management, 1705–1714 (2020).
Chithrananda, S., Grand, G. & Ramsundar, B. Chemberta: large-scale self-supervised pretraining for molecular property prediction. arXiv preprint arXiv:2010.09885 (2020).
Schwaller, P. et al. Mapping the space of chemical reactions using attention-based neural networks. Nat. Mach. Intell. 3, 144–152 (2021).
Vaswani, A. et al. Attention is all you need. Adv. Neural Inform. Process. Syst. 30, 6000–6010 (2017).
Magnúsdóttir, S. et al. Generation of genome-scale metabolic reconstructions for 773 members of the human gut microbiota. Nat. Biotechnol. 35, 81–89 (2017).
Oyetunde, T., Zhang, M., Chen, Y., Tang, Y. & Lo, C. Boostgapfill: improving the fidelity of metabolic network reconstructions through integrated constraint and pattern-based methods. Bioinformatics 33, 608–611 (2017).
Bernstein, D. B., Sulheim, S., Almaas, E. & Segrè, D. Addressing uncertainty in genome-scale metabolic model reconstruction and analysis. Genome Biol. 22, 64 (2021).
Lü, W. et al. The formate channel foca exports the products of mixed-acid fermentation. Proc. Natl. Acad. Sci. USA 109, 13254–13259 (2012).
van ’t Hof, M. et al. High-quality genome-scale metabolic network reconstruction of probiotic bacterium Escherichia coli nissle 1917. BMC Bioinforma. 23, 566 (2022).
Bu, X. et al. Engineering endogenous ABC transporter with improving ATP supply and membrane flexibility enhances the secretion of β-carotene in Saccharomyces cerevisiae. Biotechnol. Biofuels 13, 168 (2020).
Danchin, A. Zinc, an unexpected integrator of metabolism? Microb. Biotechnol. 13, 895–898 (2020).
Hastings, J. et al. The CHEBI reference database and ontology for biologically relevant chemistry: enhancements for 2013. Nucleic Acids Res. 41, D456–D463 (2012).
Mikolov, T., Chen, K., Corrado, G. & Dean, J. Efficient estimation of word representations in vector space. https://arxiv.org/abs/1301.3781 (2013).
Gao, Y., Feng, Y., Ji, S. & Ji, R. Hgnn+: general hypergraph neural networks. IEEE Trans. Pattern Anal. Mach. Intell. 45, 3181–3199 (2022).
Jiang, J., Wei, Y., Feng, Y., Cao, J. & Gao, Y. Dynamic hypergraph neural networks. In IJCAI 2635–2641 (2019).
Kim, S. et al. A survey on hypergraph neural networks: an in-depth and step-by-step guide. In Proceedings of the 30th ACM SIGKDD Conference on Knowledge Discovery and Data Mining, 6534–6544 (2024).
Yi, J. & Park, J. Hypergraph convolutional recurrent neural network. In Proceedings of the 26th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining, 3366–3376 (2020).
Chen, Y. & Zhao, Y. cyixiao/mushin: mushin (2026). Zenodo. Version v1.0. https://doi.org/10.5281/zenodo.18427362 (2026).
Acknowledgements
The authors would like to thank Dr. Chen Liao from Dartmouth College for his contributions to understanding MuSHIN’s metabolic gap-filling processes during the phenotypic prediction experiment.
Author information
Authors and Affiliations
Contributions
C.C. conceived and designed the project. Y.Z., Y.C., and X.L. developed the MuSHIN algorithm. Y.C. performed the internal validation. Y.Z. performed the external validation. Y.Z. and Y.C. interpreted the results. Y.C., Y.Z., Y.Y., and J.D. prepared the manuscript. J.W., Q.S., R.W., and C.C. edited and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ove Øyås and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Silvio Waschina and Laura Rodríguez. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhao, Y., Chen, Y., Yu, Y. et al. A multi-way SMILES-based hypergraph inference network for metabolic model reconstruction. Commun Biol (2026). https://doi.org/10.1038/s42003-026-09761-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-026-09761-1