Abstract
Tea, derived from the leaves of Camellia sinensis, is a globally consumed beverage with considerable nutritional and economic value. Specific cultivars exhibit a striking purple leaf coloration due to anthocyanin accumulation, yet the molecular mechanisms governing this trait remain incompletely understood. In this study, we identified a sense-intronic long non-coding RNA, Cs_lncRNA.18443.6, that is co-expressed with CsUFGT (UDP-glucose: flavonoid 3-O-glucosyltransferase) and is predicted to act in cis on this gene. Together with the transcription factor CsMYB12, these components form a hypothesized three-tier regulatory module that contributes to anthocyanin accumulation in purple tea leaves. CsUFGT emerges as a potential regulatory hub in anthocyanin biosynthesis. Weighted gene co-expression network analysis (WGCNA), combined with the construction of a competing endogenous RNA network construction reveals Cs_lncRNA.18443.6 as a cis-acting lncRNA associated with CsUFGT expression. This association was supported by RNA fluorescence in situ hybridization (FISH), transient expression assays in transgenic tobacco, and RT-qPCR analysis. Dual-luciferase reporter assays provided preliminary evidence that Cs_lncRNA.18443.6 influences CsUFGT transcription by affecting CsMYB12-dependent promoter activation. These findings uncover a previously uncharacterized lncRNA association with anthocyanin biosynthesis and offer new hypotheses and provide candidate targets for the molecular breeding of anthocyanin-enriched tea cultivars.
Similar content being viewed by others
Introduction
Tea, derived from the leaves of Camellia sinensis, is one of the world’s most widely consumed non-alcoholic beverages, highly valued for its economic, cultural, and health benefits1. With over two billion cups consumed daily worldwide, tea is a significant source of income for many countries, especially developing ones, and its popularity is due in part to its bioactive compounds, such as tea polyphenols, theanine, and caffeine, which offer a variety of health benefits2. Tea secondary chemistry also enhances stress resistance in tea plants and allows them to adapt to diverse habitats3. Tea is commercially produced from the young leaves of tea plants, which are woody perennial evergreen flowering plants belonging to the genus Camellia (Theaceae). The global total tea production exceeded 6 million tons in 2024, generating an annual economic value of approximately USD 17.4 billion4.
The first, fortuitous, discovery of a natural mutant tea plant (Camellia sinensis) with purple leaves in Yunnan, China, ignited extensive research into this novel germplasm5. From this plant, the variety ‘Zijuan1’ was developed, and later other purple-leaved plants were discovered. The phenomenon of tea leaves turning purple is not uncommon and often occurs in response to environmental stressors, such as intense sunlight during summer or low temperatures in autumn6. While conventional tea cultivars revert to green leaves when favorable conditions return, purple tea cultivars maintain their distinct purple hue throughout their growth cycle7. This striking pigmentation is primarily attributed to the accumulation of anthocyanins, which can reach levels as high as 0.5–1%, surpassing typical green tea leaves by 50 to 100 times8. Owing to their high anthocyanin content, purple tea cultivars are considered high-value functional teas with enhanced nutritional benefits and growing market demand. The abundance of anthocyanins in purple tea holds significant health benefits and substantial economic value. Therefore, deciphering the mechanisms behind how tea plants produce and regulate these metabolites is an important area of research.
The alteration in leaf color observed in purple tea primarily stems from the accumulation of anthocyanins9,10. At the biochemical level, anthocyanins are water-soluble pigments that impart distinctive red, purple or blue colors when stored in the vacuoles of plant cells11. Similar to other plants, the biosynthesis of anthocyanins in tea plants (Fig. 1) commences with the conversion of phenylalanine into p-coumaryl-CoA, catalyzed by enzymes such as phenylalanine ammonialyase (PAL), cinnamate-4-hydroxylase (C4H), and 4-coumaryl-CoA ligase (4CL), thereby channeling metabolic flux into flavonoid metabolism12,13. Critical early steps in the flavonoid pathway are regulated by enzymes such as chalcone synthase (CHS), chalcone isomerase (CHI), flavanone 3-hydroxylase (F3H), flavonoid 3’-hydroxylase (F3′H), and flavonoid 3’,5’-hydroxylase (F3′5′H)14,15. These enzymes guide the synthesis of various types of proanthocyanidins by introducing different groups at the R1 and R3 positions of the B ring. Further downstream, genes encoding late biosynthesis enzymes (LBGs), including dihydroflavonol 4-reductase (DFR), anthocyanidin synthase (ANS), and UDP-glycosyltransferase (UFGT), control the key branches and metabolic flux redirection16. Despite extensive research into the genes regulating this process and their functional identification, the role of non-coding RNAs (ncRNAs) in this intricate pathway remains underexplored.
The schematic diagram illustrates the major enzymatic steps involved in flavonoid and anthocyanin biosynthesis, including phenylpropanoid metabolism, flavonoid backbone formation, and anthocyanin glycosylation. Key enzymes such as PAL, C4H, 4CL, CHS, CHI, F3H, F3′H, F3′5′H, DFR, ANS, and UFGT are indicated. The pathway was adapted and modified from the KEGG pathway database (map00941: Flavonoid biosynthesis)12,13.
Non-coding RNAs (ncRNAs) constitute a class of functional transcripts that lack or possess minimal protein-coding capacity and are abundantly dispersed throughout eukaryotic genomes17 (Amaral et al., 2011), encompassing microRNAs (miRNAs), long non-coding RNAs (lncRNAs), and circular RNAs (circRNAs). In plant biology, lncRNAs have garnered substantial attention for their pivotal roles in diverse biological processes, notably in plant growth and development18,19. Their significance has been well-established as essential regulators of plant life, and they modulate processes ranging from organogenesis to the response to biotic and abiotic stresses. The canonical definition of lncRNAs stipulates that they must comprise at least 200 nucleotides and cannot code for proteins. Furthermore, they are typically distinguished from other abundant RNAs such as ribosomal RNA (rRNA), transfer RNA (tRNA), small nuclear RNA (snRNA), and small nucleolar RNA (snoRNA). In the context of plants, most lncRNAs are transcribed by RNA polymerase II, while a minority are generated by RNA polymerase III or IV/V19. Among these, only those transcribed by RNA polymerase II and exhibiting stability are classified as typical lncRNAs. According to their genomic proximity to protein-coding genes, typical lncRNAs can be categorized as long intergenic non-coding RNAs (lincRNAs), long non-coding nature antisense transcripts, or long intronic non-coding RNAs20,21. Understanding the interplay of lncRNAs with anthocyanin biosynthesis genes and their roles in this intricate pathway is an emerging area of research facilitated by advancements in high-throughput sequencing, transcriptomics, and genome annotation, particularly in the context of the tea plant genome. For instance, lncRNAs expression promotes overexpression of endogenous target mimic and SPLs, thereby promoting anthocyanin accumulation in apple fruit3. So, does such an ncRNA regulatory module exist in purple tea? If it exists, how does it carry out its regulatory function?
In the present study, to explore the putative function of transcripts in gene regulation in the formation of anthocyanin glycoside in three kinds of purple tea leaves, we conducted a comprehensive network analysis of differential expression (DE) genes to identify the lncRNAs potentially associated with the period of color transition of purple tea leaves, using two cultivars and one wild variety of tea trees with purple leaves. Compared to the two purple tea cultivars (which start purple when young and then turn green), C. sinensis var. assamica ‘Zijuan’ (ZJ) and C. sinensis var. sinensis ‘Zikui’ (ZK), which have been marketed, a purple tea variety, C. sinensis var. pubilimba ‘WangmoZC’ (WZC), discovered and cultivated in the wild by our research team from the southwestern region of Guizhou, China, has a unique pattern of pigmented leaf transition (starting green then turn purple and finally revert to green). If such a regulatory module exists, it should be reflected in the transcriptomic association patterns of these three purple tea varieties during their color transition.
Methods
Plant materials and sampling
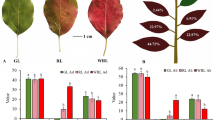
Two cultivars, ZJ from Yunan and ZK from Guizhou, and one wild variety, WZC from Guizhou, of tea tree with purple leaves were grown in the breeding nursery of Tea Tree Resources Team at the College of Tea Science, Guizhou University. These three types of tea plants were all grown in adjacent rows within the same nursery, receiving identical conditions of light, water, and other environmental factors. During the plant development, variety WZC has leaves that start green then turn purple and finally revert to green. The cultivar ZJ leaves start purple when young and then turn green. The cultivar ZK similarly starts purple and turns green.
The samples WZC were collected for three periods: WZC_1 (green leaves), WZC_2 (purple leaves), and WZC_3 (green leaves). The samples ZJ and ZK were collected for two periods, ZJ_1 (purple leaves), ZJ_2 (green leaves), ZK_1 (purple leaves), and ZK_2 (green leaves), respectively (Fig. 2). Each sample was collected by mixing leaves of the same color from different individuals (two or three for each tea tree variety) of that variety of tea tree during the same period of growth, and three biological replicates. The leaves of each tea trees were marked after field collection and immediately stored in liquid nitrogen until subsequent RNA-seq experiments.
The phenotype of two purple tea cultivars (C. sinensis var. assamica ‘Zijuan’, C. sinensis var. sinensis ‘Zikui’) and one wild variety (C. sinensis var. pubilimba ‘WangmoZC’) sampled in the study. The red circles refer to materials used for RNA-seq and hematoxylin and eosin staining experiments.
RNA extraction, library preparation, and sequencing
Total RNA of all samples was extracted from the leaves using the Total RNA Isolation Kit (Vazyme Biotech, Nanjing, Jiangsu, China), following the manufacturer’s instructions. The quality and purity of RNA were assessed using an Agilent 2100 Bioanalyzer (Agilent Technologies, California, USA) and RNA 6000 Nano LabChip Kit (Agilent Technologies, California, USA). Ribosome RNA (rRNA) was removed from total RNA using Epicentre Ribo-Zero™ Kit (Epicentre, Madison, Wisconsin, USA). The cDNA library was constructed using the NEB Next Ultra Directional RNA Library Prep Kit for Illumina (NEB, Ipswich, Massachusetts, USA). Sequencing of the libraries was performed with the Illumina NovaSeq 6000 System (Illumina, San Diego, California, USA). The construction of the cDNA library and sequencing were performed by Biomarker Technologies Co. Ltd (Beijing, China).
Genome alignment and lncRNAs identification
Raw reads obtained from the Illumina NovaSeq 6000 System were filtered and trimmed to remove the low-quality, polyN-containing and adapter reads, using fastp software22 to obtain clean reads. These clean reads were firstly aligned against a chromosome-scale genome for C. sinensis var. sinensis cv. ‘Tieguanyin’ (TGY), a Chinese Oolong tea variety23 using Hisat2 program24 after construction of the reference genome index. The mRNAs of transcripts were assembled according to the annotation of the reference genome using StringTie25 with params (–rf –m 200 –a 10 –conservative –g 50 –u), and then were merged. Each transcript quantified its expression as the read counts and transcripts per million (TPM) matrixes using the R package featureCounts.
Identifying lncRNAs in the transcripts was performed with stringent criteria. First, the transcripts of more than 200 nt in length that did not intersect with the mRNA were selected as candidates for lncRNA. Then, the programs CNCI, PLEK, CPC2, and FEELnc (params –m pl) were applied together to select the lncRNAs, using default parameters26,27,28,29. Moreover, the sequence-similarity-based programs of diamond (Swiss-Prot release 2023_01) and HMMER (Pfam v35.0, params –E 1e-5) were applied to the lncRNAs30. Finally, the intersection of the above results was determined.
Analysis of differential expression (DE) transcripts and annotation
The analysis of DE mRNAs and lncRNAs was performed with the above read counts matrix using a perl script adapted from DESeq231, where the values of log2FoldChange (lg2FC), log-fold change standard error (lfcSE), and adjusted p-value (padj) parameters were counted. The final results were determined with the criterion of lg2FC ≥ 1 and padj < 0.01. Functional annotation of coding genes was performed using the TGY’s protein sequence submitted to the online tool eggNOG-mapper v2.1.9 with eggNOG 5.0 (http://eggnog-mapper.embl.de/)32. The clusters of orthologous genes (COG) categories, gene ontology (GO), and pathway annotation were determined by using Zicha’s Organism Database (OrgDB) with an R script.
Target prediction and screening of miRNA-mRNA, miRNA-lncRNA, and lncRNA-mRNA
The mature miRNA of Camellia sinensis were downloaded from PmiREN (https://www.pmiren.com, accessed May 18, 2024). The potential targeted mRNA and lncRNA prediction of these miRNAs was performed by using a perl script from TarHunter33. Then, the Pearson’s correlation coefficient and p-value between target RNAs and miRNA were calculated with the above read counts matrix using an R script. The screening of competing endogenous RNAs (ceRNAs) for miRNAs that regulate other target RNAs was performed using the correlation value > 0.4 and p-value < 0.001. The screening of lncRNA-regulated target mRNAs in samples from four critical pair periods, including WZC_2 vs. WZC_1, WZC_3 vs. WZC_2, ZJ_2 vs. ZJ_1, and ZK_2 vs. ZK_1, was performed by using the correlation value > 0.4, p-value < 0.001, and isBest = 1 (“isBest=1” denotes the top-ranked interaction returned by TarHunter for a given regulator–target pair, according to its integrated scoring scheme). The functional annotation of the above target mRNAs was performed as per “Analysis of differential expression (DE) transcripts and annotation”.
Determination of the mRNAs and lncRNAs in flavonoid and anthocyanin biosynthetic pathways
To explore the key genes for leaf color changes of these purple tea cultivars, the mRNAs of DESs were enriched to the flavonoid (KEGG map00941) and anthocyanin (KEGG map00942) pathways by using the R packages magrittr and pathview. Then, miRNAs and lncRNAs paired with these mRNAs were filtered out from the above miRNA-mRNA and lncRNA-mRNA datasets, respectively, according to the correlation value > 0.5.
Construction of interaction network
Weighted gene co‑expression network construction (WGCNA) of DE mRNAs and lncRNAs was performed using the R package WGCNA34. WGCNA was performed on a VST-normalized expression matrix (DESeq2) derived from raw read counts, including all expressed transcripts (mRNAs and lncRNAs; TPM > 1 in ≥50% of samples). A signed network was constructed with β = 10, deepSplit = 2, minModuleSize = 30, and mergeCutHeight = 0.25. We combined all the lncRNAs and the corresponding target mRNAs in the module enriched with the transcripts involving flavonoid and anthocyanin biosynthesis and then constructed the putative interaction network mediated by these lncRNAs using Cytoscape 3.10.0 software35 was then utilized to draw the putative interaction network.
Verification of the lncRNAs and regulated mRNAs
Real‑time quantitative reverse transcription PCR (RT-qPCR) was employed to confirm the above results. Total RNAs from all samples were reverse transcribed into cDNA using oligo(dT) as a primer using a PrimeScript™ RT reagent Kit (Takara, Shiga, Japan) in a 15 μl volume. Specific experimental procedures for this section are detailed as follows.
Total RNAs from all samples were reverse transcribed into cDNA using oligo(dT) as a primer using a PrimeScript™ RT reagent Kit (Takara, Shiga, Japan) in a 15 μl volume. The cDNA was diluted to 100 μl with ddH2O serving as a PCR template. The RT-qRT-PCR experiments were performed with a reaction system (20 μl) included 0.5 μl (250 nM) of each forward or reverse primer, 2 μl of cDNA template, 10 μl of 2× Tag PCR MasterMix Il (TIANGEN), and ddH2O, using a qTower 2.2 (Analytik Jena AG, Jena, Germany). The thermal profiles were: 94 °C for 10 min, and 40 cycles of 94 °C for 15 s, 60 °C for 40 s, and 72 °C for 50 s. The gene GAPDH in C. sinensis36 was selected as the internal control. The relative expression was calculated according to the 2−∆∆Ct method with three biological replicates37.
All the primer sequences studied were included in Table S10.
Visualization of target lncRNA transcripts in leaves
Three purple tea varieties were sampled with FAA (50% ethanol: 37–40% formaldehyde: glacial acetic acid = 90:5:5 (v/v/v)) fixation in the field for 24 h at room temperature with three biological replicates during the purple leaf period (WZC_2, ZJ_1, and ZK_1). Hematoxylin and eosin (H&E) staining was employed to observe the leaf structure of these purple tea cultivars and differentiate nuclear and cytoplasmic components. RNA-FISH was employed to confirm the key lncRNAs in the flavonoid or anthocyanin pathways. The leaf tissue sections were subjected to deparaffinization and hydration by washing with 1X phosphate-buffered saline (PBS), followed by incubation in a pre-chilled permeabilization solution containing 0.5% Triton X-100 in PBS. The detection of RNA transcripts was accomplished using a probe. Specific experimental procedures for this section were detailed as follows.
Three purple tea varieties were sampled with FAA (formaldehyde, acetic acid and ethanol) fixation in the field for 24 hours (h) at room temperature with three biological replicates during the purple leaf period (WZC_2, ZJ_1, and ZK_1). The paraffin blocks were cut at a thickness of 5–10 mm, trimming the block to obtain a flat surface for sectioning, and then sectioning the embedded tissue using the microtome, with subsequent collection of sections on glass slides. Hematoxylin and eosin (H&E) staining was employed to observe the leaf structure of these purple tea cultivars and differentiate nuclear and cytoplasmic components. The hematoxylin was made up as hematoxylin 2.5 g, anhydrous ethanol 25 ml, aluminum potassium sulfate 50 g, distilled water 500 ml, mercuric oxide 1.25 g, and glacial acetic acid 20 ml. The eosin staining solution is eosin Y 2 g, distilled water 500 ml, and glacial acetic acid 1 ml. The experimental procedure was as follows: paraffin removal was performed by heating the slides at 70 °C for 5 min, followed by two rounds of xylene treatment (10 min each). Subsequently, the slides were washed with 100% ethanol to remove xylene and then hydrated using decreasing ethanol concentrations (95%, 85%, and 75%) for 5 min each, followed by distilled water. Staining was conducted with hematoxylin for 10 min, followed by rinsing and differentiation in 1% hydrochloric acid-alcohol solution. The slides were rinsed, stained with eosin for 10 min, and differentiated using 80% ethanol. Finally, the slides were dehydrated, made transparent using xylene, and sealed with neutral gum for examination using a DM2000 microscope (LEICA, Germany) with automatic pathology scanner panoramic MIDI II (3DHISTECH, Hungary).
RNA-FISH was employed to confirm the key lncRNAs in the flavonoid or anthocyanin pathways. The leaf tissue sections were subjected to deparaffinization and hydration by washing with 1X phosphate-buffered saline (PBS), followed by incubation in a pre-chilled permeabilization solution containing 0.5% Triton X-100 in PBS. The detection of RNA transcripts was accomplished using a probe. Pre-hybridization was performed by incubating the sections with the pre-hybridization solution at 37 °C for 30 min. Simultaneously, the hybridization solution was preheated at 37 °C. The probe was added to the hybridization solution, and the sections were incubated with the probe hybridization mixture overnight at 37 °C under dark conditions. Subsequently, the sections underwent a series of washes with hybridization wash buffers I, II, and III at 42 °C. Finally, the sections were washed with 1X PBS and DNA staining using 1X DAPI staining solution. The sections were sealed with a cover slip, and fluorescence detection was carried out under dark conditions using inverted fluorescence microscope TS2 (NIKON, Japan), ulnsclean bench BSC-1300-II-A2 (HUAYU, China), and automatic pathology scanner panoramic MIDI II (3DHISTECH, Hungary).
Basic local alignment search tool (BLAST) analysis of target lncRNA sequence in related species genomes
The whole genome sequences of seven tea plants (five cultivated C. sinensis and two wild tea trees) (Table 1), two model plants (Nicotiana tabacum and Arabidopsis thaliana), and several species closely related to the Camelliaceae were downloaded from the National Genomics Data Center (NGDC) and National Center for Biotechnology Information (NCBI) public database website. Then, a database was constructed based on the genome sequences of each species. The target lncRNA sequences selected based on the previous experiments were subjected to sequence alignment using the NCBI blastn program.
Construction, purification, and expression of tobacco plants with transformation Cs_lncRNA.18443.6
The Gateway recombinant cloning system38 was used for constructing the transformation vectors of Cs_lncRNA.18443.6. Considering the characteristics of Cs_lncRNA.18443.6 sequence (the first part of the entire sequence with less than 10% GC content and large number of repetitive sequences), its desired DNA sequence (Table S9) was directly synthesized by Beijing Ruibiotech Limited (CN).
In this study, to verify the expression of the specific lncRNA and avoid the potential interference that may be caused by reporter gene expression, we excised the GUS sequence of the pBI121 vector to simplify the experiment. The targeted lncRNA was subcloned into the expression vector pBI121 (Takara, JA). Then, the cloned gene sequences in recombinant colonies pBI121-Cs_lncRNA.18443.6 were double-verified by PCR and Sanger sequencing. The Agrobacterium suspension was slowly injected into the lower epidermis of tobacco leaves using a 1 mL syringe. Subsequently, the plants were placed in darkness within a growth chamber for 48 h before observing their phenotypic traits. The RT-qPCR experiments in transgenic tobacco with target traits were performed using the same methods as in “Verification of the lncRNAs and regulated mRNAs”. Specific experimental procedures for this section are detailed as follows.
The Gateway recombinant cloning system37 was developed for constructing the transformation vectors of Cs_lncRNA.18443.6. Considering the characteristics of Cs_lncRNA.18443.6 sequence (the first part of the entire sequence with less than 10% GC content and large number of repetitive sequences), its desired DNA sequence (Table S9) was directly synthesized by Beijing Ruibiotech Limited (CN). Then, it was cloned into the pCR2.1-TOPO TA cloning vector following the manufacturer’s protocol (Takara, JA). The pCR2.1-TOPO_TA-Cs_lncRNA.18443.6 entry vector was then transferred into the plant transformation destination. Each single colony containing each target construct was confirmed by PCR. The targeted lncRNA was subcloned into the expression vector pBI121 (Takara, JA). The cloned gene sequences in recombinant colonies pBI121-Cs_lncRNA.18443.6 were double-verified by PCR and Sanger sequencing. Subsequently, the pBI121-Cs_lncRNA.18443.6 expression vectors and empty vectors were transformed into E. coli Novablue (DE3) competent cells (Novagen, Schwalbach, Germany).
Using an adapted version of the Cui et al. protocol39, 1 μl of plasmid was transferred into freshly thawed Agrobacterium tumefaciens cells in a state of competence and maintained on ice for 10 min before being plunged into liquid nitrogen for 1–2 min. Subsequently, the cells were thawed in a 37 °C water bath until completely melted, then returned to ice for 2 min. In a sterile environment, 200 μl of YEB medium without antibiotics was added, and the mixture was incubated at 28 °C with shaking at 220 rpm for 1 h. Afterward, 30–50 μl of the bacterial suspension was streaked onto a solid YEB medium supplemented with kanamycin (plasmid resistance) and rifampicin (Agrobacterium resistance) and incubated at 28 °C in an incubator for 48 h. With a 10 μl pipette tip within a sterile workspace, single colonies were selected from the YEB agar plates and transferred into 1.5 ml centrifuge tubes containing 400 μl of liquid YEB medium with kanamycin and rifampicin. The tubes were then incubated overnight at 28 °C with shaking at 220 rpm. Following overnight growth of single colonies in the 1.5 ml centrifuge tubes, 200 μl of the colonies was streaked onto YEB agar plates supplemented with kanamycin and rifampicin, followed by overnight incubation for further expansion of the colonies, readying it for subsequent applications. In parallel, healthy tobacco plants were selected and cultured under 16 h of light and 8 h of darkness, with a temperature of 22 °C and a relative humidity of 70%, for approximately 3–4 weeks.
The Agrobacterium suspension was slowly injected into the lower epidermis of tobacco leaves using a 1 mL syringe. Subsequently, the plants were placed in darkness within a growth chamber for 48 h before observing their phenotypic traits. The RT-qRT-PCR experiments in transgenic tobacco with target traits were performed using the same methods as in “Verification of the lncRNAs and regulated mRNAs”.
Dual-luciferase reporter assay of CsMYB12, Cs_lncRNA.18443.6, and CsUFGT promoter
To investigate the regulatory relationships among the transcription factor CsMYB12, the long non-coding RNA Cs_lncRNA.18443.6, and CsUFGT promoter, we performed a dual-luciferase (Dual-LUC) assay in Nicotiana benthamiana leaves. The full-length coding sequence of CsMYB12 was cloned into the pGreenII62-SK effector vector under the CaMV 35S promoter (35S:CsMYB12). The 1.0-kb promoter region of CsUFGT and the promoter (~1226 bp) of Cs_lncRNA.18443.6 were separately inserted into the pGreenII 0800-LUC reporter vector to drive firefly luciferase expression (pGreenII0800-LUC-pCsUFGT and pGreenII0800-LUC-lncRNA). The Renilla luciferase cassette in the same vector served as an internal reference.
Agrobacterium tumefaciens strains (GV3101) harboring effector and reporter constructs were cultured, induced with acetosyringone, and co-infiltrated into fully expanded N. benthamiana leaves. After 18 h of dark incubation followed by 24 h under normal light, leaf discs were collected for luminescence detection. Firefly and Renilla luciferase activities were quantified using the Dual-Luciferase® Reporter Assay System (Promega, USA). The LUC/REN ratio was used to evaluate promoter activation and the regulatory effects of CsMYB12 and Cs_lncRNA.18443.6.
Yeast one-hybrid (Y1H) assay for CsMYB12 binding to CsUFGT promoter
A yeast one-hybrid (Y1H) assay was conducted to verify the physical interaction between CsMYB12 and CsUFGT promoter. The promoter fragment of CsUFGT (WT) and a version with mutated MYB-binding sites (mut-MBS) were cloned into the pLacZi2µ reporter vector to generate promoter–LacZ fusion constructs (pLacZi2µ-pCsUFGT and pLacZi2µ-pCsUFGT(mut-MBS)). The coding sequence of CsMYB12 was inserted into the pJG4-5 activation-domain vector (pJG4-5-CsMYB12), while the empty pJG4-5 vector served as the negative control.
Reporter and effector plasmids were co-transformed into the yeast strain EGY48 following the manufacturer’s protocol (Clontech). Transformants were selected on SD/-Ura/-Trp medium, and serial dilutions (10⁰, 10⁻¹, 10⁻², 10⁻³) of yeast cultures were spotted onto SD/-Leu/X-α-Gal plates. Yeast growth and blue coloration were recorded after 48–72 h to assess promoter activation. Activation of CsUFGT promoter by CsMYB12 was indicated by robust growth and X-α-Gal hydrolysis, whereas mut-MBS constructs abolished promoter activation, confirming binding specificity.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Results
Assembly, characterization, and reconstruction of transcripts
Seven kinds, and a total of 21 libraries were constructed, including the three sequential stages (green to purple to green) of WZC leaves and two sequential stages (purple to green) of ZJ and ZK leaves. Over 436 Gb of transcriptomic clean data was generated, and each sequencing sample averaged 20.79 Gb, with more than 92.27% of bases scoring Q30 (Table S1). A total of 2904 Mb reads with more than 130 Mb/each were successfully mapped to the TGY genome23, with matching rates in the range of 86.12–93.42%, except for 75.58% of the b2ZJ sample (Table S2). (Although b2ZJ exhibited a lower mapping rate, PCA confirmed its correct clustering, and sensitivity tests showed that excluding this sample did not alter downstream DE or WGCNA results.)
To understand the mRNA-lncRNA regulatory mechanism during the color change of tea plant leaves, a strand-specific transcripts analysis for all RNA-seq data was conducted using StringTie, based on the known annotated genes (mRNA) of the TGY genome. As a result, 63,249 potential lncRNA transcripts were generated after primary screening using the FEELnc based on the criteria: minimum lncRNA size of 200 bases and number of exons ≥1.
Identification and characterization of lncRNAs
The prediction of ORF length, length of nucleotides, and coding capacity were essential to identify lncRNAs. Moreover, sequence comparison and similarity search (BLAST) comparison and detection of conserved domains and patterns (hmm method) of proteins are also required. Based on the preliminary 63,249 transcripts, the FEELnc, CPC2, PLEK, and CNCI methods predicted 55,016, 47,659, 44,285, and 42,681 transcripts to have non-coding potential transcripts, respectively (Fig. 3a). The BLAST and hmm methods generated 44,955 and 63,242 transcripts, respectively. In total, 26,470 lncRNAs were obtained by taking the intersection of the above results. Most lncRNAs were between 300 and 500 bp in length, with lncRNAs smaller than 1000 bp accounting for more than 80% of the total (Fig. 3b). Compared with the number of types of mRNAs with one exon, the predicted potential lncRNAs all have at least two exons and are the most numerous (Fig. 3c), but their overall expression is significantly lower than that of mRNAs (Fig. 3d).
a Prediction of protein-coding mRNA with different programs. b Transcript length distribution of lncRNAs and mRNAs. c Exon number distribution of lncRNAs and mRNAs. d Expression distribution in different samples. e Classification statistics and correlation coefficients of lncRNAs. f Principal component analysis of 21 purple tea samples based on expression (specifically labeled hub genes with the highest contribution of mRNAs and lncRNAs).
The positional classification of the predicted novel lncRNAs showed that 194 were bi-directional (transcribed from a protein-coding gene promoter, but in the opposite direction), 14,371 were intergenic (>5 kb from nearest gene), 2192 were sense intergenic upstream (<1 kb), 4230 were sense intronic, 3250 were antisense intronic, 499 were antisense exonic, and 1734 were other types of lncRNAs (Fig. 3e). The differences in frequency between the intergenic and sense intergenic upstream, as well as the sense intronic and antisense intronic were highly statistically significant. The results of the principal component analysis (PCA) of the 21 samples (Fig. 3f) showed that three replicates of the same variety and the same period were more obviously clustered together by the PCA based on the mRNAs expression, with several genes, such as CsTGY04G0002441.t1, contributing the most to the analysis. The PCA based on the lncRNAs expression showed that the three replicate samples of the same variety and the same period were together. Still, the overall clustering was not apparent, among which was the contribution of several genes, such as Cs_lncRNA.111879.116, which was the highest.
DE, GO, and KEGG analysis of transcripts
To investigate biosynthesis-related genes before and after color change in purple tea leaves, we conducted a statistical analysis on DE transcripts in the period of WZC_2 vs. WZC_1, WZC_3 vs. WZC_2, ZJ_2 vs. ZJ_1, and ZK_2 vs. ZK_1 based on the raw counts of all transcripts (Figs. S1–S4). Among them, 8589, 6544, 10,334, and 9581 DE mRNAs were significantly enriched (|log2FoldChange| ≥ 1 & adjusted P-value < 0.01), respectively; and 1855, 1544, 2931, and 1801 DE lncRNAs were significantly enriched (|log2FoldChange | ≥1 & adjusted P-value < 0.01), respectively. For DE lncRNAs, there were 500 down-regulated and 835 up-regulated in the period of WZC_2 vs. WZC_1, 692 down-regulated and 383 up-regulated in the period of WZC_3 vs. WZC_2, 1381 down-regulated and 1365 up-regulated in the period of ZJ_2 vs. ZJ_1, 623 down-regulated and 821 up-regulated in the period of ZK_2 vs. ZK_1.
The biological processes enriched by these DE mRNAs in the transition period of purple-green leaves (WZC_3 vs. WZC_2, ZJ_2 vs. ZJ_1, and ZK_2 vs. ZK_1) were mostly annotated to ‘cell wall organization or biogenesis’, ‘secondary metabolic process’, and ‘mitotic cell cycle’ using GO term analysis (Fig. S5). And the transition period of green-purple leaves was mostly annotated to ‘secondary metabolic process’, ‘generation of precursor metabolites and energy’, and ‘photosynthesis’. The KEGG enrichment results for DE mRNAs in the three periods of purple-green leaves were nearly the same, and ‘flavonoid biosynthesis’ was one of the pathways to which DE mRNAs were significant for all periods of color change (Fig. S6).
ceRNA network and predicted interaction between miRNAs, lncRNAs, and mRNAs
This study annotated all target genes of DE lncRNAs in the period of color change and found 9764 potential regulated pairs (Table S3). The transcript hierarchical clustering tree included 62 different modules (Fig. 4a) based on all 21 samples using WGCNA. The result of module gene annotation showed that the DE lncRNAs in the flavonoid and anthocyanin biosynthesis pathway indicated only one, and it was clustered in the turquoise module. Taking the intersection of the two results above, the potential ceRNA networks of miRNAs-lncRNAs-mRNAs in the turquoise module (top 10) were mapped using the Cytoscape (Fig. 4b), including five Csi-miR156 (e, h, i, j, and k), four Csi-miR828 (a, b, c, and d). However, the results of the screened miRNA-mediated regulatory networks did not reveal the presence of existing mature miRNAs with potential regulatory relationships for these DE mRNAs and lncRNAs when taken in intersection with DE transcripts annotated to the flavonoid and anthocyanin biosynthesis pathway. Therefore, only seven pairs of lncRNAs-mRNAs in the flavonoid biosynthesis pathway were identified based on RNA-seq data with the correlation value > 0.4 and p-value < 0.001, using a perl script from TarHunter33. They were CsTGY04G0001430.t1/Cs_lncRNA.32020.1, CsTGY06G0000959.t1/Cs_lncRNA.48423.3, CsTGY08G0001425.t1/Cs_lncRNA.67822.2, CsTGY09G0001135.t1/ s_lncRNA.373762.1, CsTGY09G0001136.t1/Cs_lncRNA.73765.2, CsTGY02G0002971.t1/Cs_lncRNA.18443.6, and CsTGY04G0001430.t1/Cs_lncRNA.32020.1 (Fig. 4c).
a mRNA hierarchical clustering tree of different modules. Each main branch of the tree represents a module, each leaf of the tree represents a transcript, and different modules are marked with different colors. b Detailed network dynamics of nodes (top 10 miRNA hub) in turquoise module (no. 1) based on DE transcripts. Yellow ellipses represent miRNAs, pink triangles represent lncRNAs, and light blue rhombuses represent mRNAs. c DE transcripts enriched in flavonoid and anthocyanin biosynthesis pathways. Error bars represent mean ± SD from n = 3 biologically independent experiments.
Analysis of the mRNAs and lncRNAs in the flavonoid and anthocyanin biosynthesis pathway
Specific to the flavonoid biosynthesis pathway, there were 44, 62, and 62 key DE mRNAs, respectively, in the three purple-to-green leaf contrasts (WZC_3 vs. WZC_2, ZJ_2 vs. ZJ_1, and ZK_2 vs. ZK_1; Figs. S7–S9) and 27 key DE mRNAs in the green to purple leaf contrast (WZC_2 vs. WZC_1; Fig. S10). Moreover, there was evidence that 7, 12, and 9 lncRNAs were key in regulating some of the mRNAs in the three purple-to-green contrasts and 4 in the green-to-purple contrast (Tables S4–S7). However, of these, only a single pair of DE mRNAs/lncRNAs are referrable to the anthocyanin biosynthesis pathway, namely CsTGY02G0002971.t1/Cs_lncRNA.18443.6. The intersection of the above results gives seven regulatory lncRNA/mRNA pairs consistently showing in all periods of color change (Fig. 4c), including the anthocyanin biosynthetic pathway pair. Based on the eggNOG-mapper and KEGG pathway annotations, CsTGY02G0002971.t1 was identified as a UDP-glucose: anthocyanidin/flavonol 3-O-glucosyltransferase (UFGT; i.e., the Bronze gene in maize, BZ1), a critical regulatory gene for anthocyanin glycoside synthesis (Fig. 5). The sequences of CsTGY02G0002971.t1 and Cs_lncRNA.18443.6 are shown in Tables S8 and S9. The lncRNA Cs_lncRNA.18443.6 is sense intronic and is transcribed from within its target gene, UFGT (Fig. 5). There was no evidence of miRNA involvement in a ceRNA network in any flavonoid and anthocyanin biosynthesis pathway in this study.
Enzymes in red font indicate that their genes and related lncRNAs were detected with significant up- or down-regulated changes in the DE transcripts.
Determination of the lncRNAs and regulated genes responding to leaves color change
To further validate the results based on RNA-seq data, the seven regulatory lncRNA/mRNA pairs identified above, as related to flavonoid or anthocyanin biosynthesis, were selected to test their expression levels using RT-qPCR (Fig. 6; primer information is provided in Table S10). The RT-qPCR results showed that only two of the seven lncRNA/mRNA pairs, UFGT (CsTGY02G0002971.t1/Cs_lncRNA.18443.6) and FLS (CsTGY04G0001430.t1/Cs_lncRNA.32020.1), exhibited statistically significant expression changes (P < 0.05), whereas the remaining five pairs, including caffeoyl-CoA O-methyltransferase (CCoAOMT) (CsTGY06G0000959.t1/s_lncRNA.48423.3), and three shikimate O-hydroxycinnamoyl transferases (HCT) (CsTGY08G0001425.t1/Cs_lncRNA.67822.2, CsTGY09G0001135.t1/Cs_lncRNA.373762.1, and CsTGY09G0001136.t1/Cs_lncRNA.73765.2) (The location the pathway shown in Fig. 1), displayed similar but statistically non-significant expression trends across developmental stages and varieties.
a Genomic organization and relative expression of Cs_lncRNA.18443.6 and its cis-associated CsTGY02G0002971.t1 (UFGT) on chromosome 2. b Genomic organization and relative expression of Cs_lncRNA.32019.3 and CsTGY04G0001430.t1 (FLS) on chromosome 4. c Genomic organization and relative expression of Cs_lncRNA.32020.1 and CsTGY04G0001430.t1 on chromosome 4. d Genomic organization and relative expression of Cs_lncRNA.48423.3 and CsTGY06G0000959.t1 on chromosome 6. e Genomic organization and relative expression of Cs_lncRNA.67822.2 and CsTGY08G0001425.t1 on chromosome 8. f Genomic organization and relative expression of Cs_lncRNA.73762.1 and CsTGY09G0001135.t1 on chromosome 9. g Genomic organization and relative expression of Cs_lncRNA.73765.2 and CsTGY09G0001136.t1 on chromosome 9. Relative expression levels were determined by RT–qPCR during purple-to-green leaf transition in three tea cultivars. * indicates statistical significance at P < 0.01. Error bars represent mean ± SD from n = 5 biologically independent experiments. The numerical source data for the graphs can be found in Supplementary Data 2.
The lack of statistical significance for these five pairs may reflect relatively low transcript abundance, tissue heterogeneity, or pathway-specific regulatory dynamics, rather than a complete absence of regulatory association. Overall, these RT-qPCR results are largely consistent with the differential expression patterns observed in the RNA-seq analysis (Fig. 5). Combining the annotation of anthocyanin biosynthesis in KEGG database and the results of previous work40 that implicated UFGT but not FLS in the purple leaf transition, we considered UFGT a highly promising candidate for proceeding to further validation.
Identification of the genes regulated by Cs_lncRNA.18443.6
Considering the critical position of UFGT/BZ1 in anthocyanin glycoside synthesis, we employed RNA-FISH experiments to verify the co-expression in leaf tissue of the postulated regulator, Cs_lncRNA.18443.6 with the level of anthocyanin occurrence. Each leaf selected from nine samples (three biological replicates of each of the three purple tea varieties, represented by A, B, and C) during the purple leaf period (WZC_2, ZJ_1, and ZK_1) showed a standard leaf structure after staining the nucleus and cytoplasm by H&E staining (Fig. 7a). Samples WZC_2C, ZJ_1A, and ZK_1B were selected to reveal the RNA expression signals after using electron microscopic observation of leaf sections and comparison among these samples (Fig. 7b). The primer sequences of molecular probes for the detection of Cs_lncRNA.18443.6 are shown in Table S10. Two control experiments were autofluorescence assays and negative controls. Autofluorescence validation experiments showed that the two fluorescents used in this study, green (probe binding to target sequences) and blue (DAPI-stained nuclei were blue under ultraviolet excitation), emitted weak fluorescence without using a quencher and no fluorescence after using a quencher. Negative control experiments using the same probe (green fluorescent agent) in RNA-FISH experiments on green leaves of the same tea tree did not observe significant fluorescent signals, suggesting that the specificity of the probes used in this study was good (Fig. 3c). The RNA-FISH results (Fig. 7d) showed that the three samples were positive expression with green fluorescence. Overall, the intensity of green fluorescence in the three samples is the highest for ZJ_1A, followed by ZK_1B, and the weakest for WZC_2C, and this is consistent with the trend of color depth in the corresponding samples (Fig. 7b). In particular, the RNA expression signals were detected in tissues across the entire leaf cross-section but were mainly concentrated in the cells of the upper epidermis and phloem.
a Sampled leaves and their cross-sectional scans after H&E staining under the microscope. b Sampled leaves for FISH detection and their cross-sectional scans. c Both autofluorescence assays and negative controls were used for the control experiments. The autofluorescence of the two green and blue fluorescents used in this study is without a quenching agent. No significant green fluorescence signal was observed in the negative control group using green leaves of the same tea tree as negative control for RNA-FISH experiments. d Expression of Cs_lncRNA.18443.6 in purple tea leaf tissue structures labeled with a sequence-specific probe. DAPI-stained nuclei are blue under UV excitation, and positive expression is green fluorescence labeled with the corresponding fluorescein.
Sequence analysis of Cs_lncRNA.18443.6 in genus Camellia plants
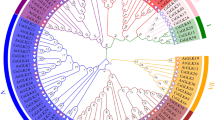
CsUFGT is a critical regulatory gene for anthocyanin and flavonoid biosynthesis in tea plants41. Moreover, Cs_lncRNA.18443.6 can cis-regulate CsUFGT (CsTGY02G0002971.t1), which in turn affects anthocyanin glycoside synthesis and thus contributes significantly to the variation of leaf color in tea plants. To explore the evolutionary relationship of this novel lncRNA in closely related species as well as its variation in tea group plants, its complete sequence was spliced from the strand-specific transcriptome sequencing data of the present study and then compared with the full genome sequences of each species for BLAST. The alignment results showed that the lncRNA is moderately conserved, being present in all Camellia species for which appropriate genomic resources exist (Table 1, Fig. S11). This novel lncRNA had BLAST results only in the family Theaceae, and the alignment was close, especially in cultivated tea plants, with >95% concordance of the corresponding coding sequence and alignable over the entire length (Table 1).
To further characterize structural features of Cs_lncRNA.18443.6, potential open reading frames (ORFs) predicted by NCBI’s ORFfinder. The analysis revealed the presence of multiple short ORFs (13–17 ORFs per transcript) with comparable length distributions and positional patterns across different Camellia species (Fig. S11; Tables S11–S18). Importantly, these ORFs predicted by ORFfinder represent incidental open reading frames commonly found in long non-coding transcripts and were used solely as part of coding-potential assessment rather than as evidence of conserved protein-coding capacity.
Heterologous expression and verification of Cs_lncRNA.18443.6 and Dual-LUC reporter assays
The expression vector pBI121-Cs_lncRNA.18443.6 was successfully constructed and verified through PCR amplification and Sanger sequencing (Fig. 8a, b; Figs. S12 and S13). Phenotypic observation of transiently transformed Nicotiana benthamiana leaves revealed no visible pigmentation changes compared with WT and empty-vector controls (Fig. 8a). RT–qPCR analysis demonstrated that heterologous expression of Cs_lncRNA.18443.6 resulted in a marked increase in both the lncRNA itself and its putative cis-regulated target CsUFGT in infiltrated tobacco tissue (Fig. 8d). In contrast, upstream regulatory enzymes in the flavonoid pathway (FLS, DFR, and ANS) showed no significant change, consistent with Cs_lncRNA.18443.6 acting at the terminal anthocyanin glycosylation step (Fig. 8e). These results confirm that the recombinant lncRNA is transcriptionally active in planta and is capable of enhancing UFGT expression without inducing visible coloration, likely due to species-specific differences in upstream pathway activation.
a Phenotypic comparison of Nicotiana benthamiana leaves transiently expressing Cs_lncRNA.18443.6. Whole plants and enlarged leaves are shown for WT, pBI121 empty vector, and pBI121–Cs_lncRNA.18443.6, indicating no visible anthocyanin phenotype. b PCR verification of Cs_lncRNA.18443.6 expression in infiltrated tobacco leaves. c Schematic map of the T-DNA region in the pBI121–Cs_lncRNA.18443.6 construct used for transient expression. d RT–qPCR analysis of Cs_lncRNA.18443.6, CsUFGT, NtFLS, NtDFR, and NtANS in infiltrated tobacco leaves (mean ± SD, n = 3 biological replicates; one-way ANOVA with Tukey’s post hoc test; * indicates statistical significance at P < 0.05). e Proposed position of Cs_lncRNA.18443.6 within the anthocyanin biosynthetic pathway, acting at the terminal glycosylation step catalyzed by UFGT. f Yeast one-hybrid (Y1H) assay showing that CsMYB12 binds the CsUFGT promoter. Ten-fold serial dilutions (10⁰–10⁻³) on SD/-Leu + X-α-Gal plates indicate strong promoter activation by CsMYB12, whereas mutation of MYB-binding sites (mut-MBS) abolishes activation. g Effector and reporter constructs used in Dual-LUC assays. Effectors include 35S:CsMYB12, 35S:CsMYB12-DBDmut, and 35S:Cs_lncRNA.18443.6. Reporter constructs contain pCsUFGT(WT)::LUC or pCsUFGT(mut-MBS)::LUC, with REN as internal control. h Dual-LUC assays testing the specificity of CsMYB12-mediated activation of CsUFGT promoter. Luminescence images (left) and LUC/REN quantification (right) show strong activation by CsMYB12, while mut-MBS and DBDmut abolish promoter activity. i CsMYB12 activates the CsUFGT promoter. Luminescence and LUC/REN values are shown for (1) control and (2) CsMYB12 + pCsUFGT(WT). j Co-expression of Cs_lncRNA.18443.6 enhances CsMYB12-mediated activation of the CsUFGT promoter. k Dose-dependent activation of the CsUFGT promoter by increasing concentrations of CsMYB12 (20%, 30%, 40%, 50%). h–k Statistical significance is indicated as P < 0.05 (*), P < 0.01 (**), and P < 0.001 (***); Error bars represent mean ± SD from n = 4 biologically independent experiments. And the numerical source data for the graphs can be found in Supplementary Data 2.
To evaluate whether CsMYB12 directly binds CsUFGT promoter, Y1H assay was performed using WT and MYB-binding-site-mutated (mut-MBS) promoter fragments. CsMYB12 strongly activated CsUFGT promoter, producing robust growth and X-α-Gal hydrolysis across dilution series, whereas mut-MBS constructs abolished activation (Fig. 8f), demonstrating specific recognition of CsUFGT promoter by CsMYB12.
To further assess transcriptional regulation in planta, dual-luciferase (Dual-LUC) assays were conducted using effector constructs expressing CsMYB12, CsMYB12-DBDmut, or Cs_lncRNA.18443.6, and reporter constructs containing WT or mut-MBS CsUFGT promoter (Fig. 8g). Co-expression of CsMYB12 with the WT CsUFGT promoter produced strong promoter activation, whereas DBD-mutated CsMYB12 or mut-MBS promoters lost activation capacity (Fig. 8i, j). Notably, co-expression of Cs_lncRNA.18443.6 significantly enhanced CsMYB12-mediated CsUFGT promoter activation (Fig. 8h), supporting a functional interaction between the lncRNA and MYB12-dependent transcriptional regulation. Moreover, incremental delivery of the CsMYB12 effector resulted in dose-dependent increases in promoter activation (Fig. 8k), further supporting MYB12-dependent transcriptional control.
Discussion
Purple tea, distinguished by its high flavonoid and anthocyanin content, is an excellent source for specialized tea products. Varieties like ‘Zijuan’ and ‘Zikui’ have established themselves in the Chinese market41,42, while ‘WangmoZC,’ a wild-type cultivar, is currently being developed as a novel tea product16 (Li et al., 2023). These three purple tea varieties constitute promising materials for investigating the regulatory mechanisms governing the accumulation of flavonoids, specifically anthocyanins, in tea leaves, including the involvement of ncRNA.
Numerous studies have demonstrated that lncRNAs play pivotal roles in secondary metabolite biosynthesis by modulating the expression of relevant genes18,19. However, research on lncRNAs in tea, especially their involvement in responses to pathogenic invasion, salt stress, and secondary metabolite regulation, has been limited18 (Zhou et al., 2022). Previous research identifying lncRNAs of Camellia sinensis relied on bioinformatic prediction methods19,43, which, while informative, often requires more comprehensive validation. For example, Varshney et al. (2019) employed bioinformatics tools such as CPC and CPAT to predict 33,400 lncRNAs based on publicly available RNA-seq data from 11 harvestable tissues44. Another study45 (Zhu et al., 2019) systematically analyzed lncRNAs in tea leaf production. However, these approaches were limited by their dependence on the quality of the reference genome and suffered from accuracy-related issues.
Identification of lncRNAs from RNA sequencing data typically involves the application of specialized algorithms or pipelines. These methods are often supplemented with similarity searches against established gene databases, such as Rfam, a repository of RNA families46, and Pfam, a comprehensive protein family database47. Moreover, an additional filter based on open reading frame (ORF) length, typically exceeding 100 amino acids, is commonly employed. These methodologies have played a pivotal role in discovering functionally important lncRNAs in eukaryotes17,48.
In this study, we combined foundational aspects of previous methods while customizing our approach to the color changes of purple tea leaves. Our workflow began with meticulously constructing strand-specific libraries, preceded by removing rRNA to ensure data integrity. Subsequently, we conducted high-depth RNA-seq experiments for each sample (Table S1). For the precise identification of tea tree lncRNAs, we adopted a comprehensive strategy. Initially, we employed two kinds of algorithm tools: one that utilizes coding potential prediction algorithms, including CPC2, CNCI, and PLEK, and the other is FEElnc, which is a machine learning-based software trained on known lncRNA sequences. Furthermore, we implemented a sequence similarity-based strategy involving BLAST algorithms and HMMER comparisons. The final compilation of identified lncRNAs was achieved by taking the intersection of results from these diverse methodologies. We argue that this systematic approach enhances the reliability and robustness of our analytical outcomes.
LncRNAs have an impact on molecular mechanisms at different levels, such as epigenetics, transcription factor regulation, and RNA post-transcriptional regulation, and play a role in complex molecular networks through multilevel regulation, which is closely related to numerous cellular activities49. Building upon our bioinformatics analyses, we propose that Cs_lncRNA.18443.6 is a potential regulatory factor in the regulation of anthocyanin biosynthesis in purple tea leaves, supported by annotations from public databases such as KEGG Pathway. As Cs_lncRNA.18443.6 is transcribed from within its target gene (it is sense intronic to UFGT) it must be considered to act in cis, potentially as an enhancer. Cis-acting lncRNAs are defined as those lncRNAs for which regulation of gene expression is influenced by their location in close proximity to their target50. Subsequent RT-qPCR results in a variety of tea leaves (Fig. 6) confirmed relative expression matching between CsTGY02G0002971.t1 (CsUFGT) and Cs_lncRNA.18443.6, adding evidence that Cs_lncRNA.18443.6 could be involved in cis-regulatory control over CsUFGT, ultimately contributing to the enhanced accumulation of anthocyanins in purple tea leaves.
Furthermore, RNA-FISH results confirm the widespread expression of Cs_lncRNA.18443.6 in the leaf, notably in the upper epidermal parenchyma cells and phloem cells, areas associated with anthocyanin accumulation, providing supportive evidence for a regulatory role of Cs_lncRNA.18443.6 (Fig. 7). Additionally, the variation in green fluorescence intensity between samples (ZJ_1A, ZK_1B, and WZC_2C) correlates with the depth of purple coloration in their leaves. Given that Cs_lncRNA.18443.6 lacks coding potential and acts at the RNA level, its regulatory influence on CsUFGT is likely mediated through transcriptional or chromatin-associated mechanisms rather than peptide-dependent processes. This further supports the association between Cs_lncRNA.18443.6 and CsUFGT expression levels (and, subsequently, anthocyanin content). However, these experiments strictly provide indirect evidence. Further nucleic acid-nucleic acid interaction experiments, such as dual-luciferase assays, need to be designed to validate this conclusion.
Genetic manipulation of the lncRNA in tea plants would provide additional functional information. Some studies have found that knocking down lncRNA expression can specifically affect gene expression and function in particular cell types42. Because of the absence of an efficient and stable genetic transformation system in tea plants51, we used heterologous expression of Cs_lncRNA.18443.6 in tobacco to test whether the lncRNA could be induced to act heterologously in trans. If successful, this would strongly validate our study. Even if the regulation works natively in cis, it has been pointed out that when overexpressed in trans, it is possible that the lncRNA is able to “flood the system”, enabling sufficient lncRNA transcript to reach the site of transcription of the target gene50. Although the transgenic tobacco plants did not exhibit the target phenotypic trait (altered leaf color), remarkably, the RT-qPCR validation showed that this lncRNA may promote an increase in the expression of its predicted target mRNA (UFGT).
The absence of a discernible phenotype may stem from species-specific factors because there are no similar sequences to this lncRNA found in tobacco (Table 1). Alternatively, the expression of this mRNA may require the co-participation of other enzyme genes to show a color phenotype in the plant. However, the heterologous functional effect of the lncRNA suggests that although the target UFGT is evolutionarily divergent in tobacco, the relevant regulatory elements of tobacco UFGT are sufficiently well conserved for the lncRNA to act heterologously, although the trend in its regulated expression is much lower than that of the homologous genes (Figs. 6 and 8e).
The Dual-LUC reporter assay enabled preliminary exploration of the TF–lncRNA–target gene regulatory module and demonstrated that Cs_lncRNA.18443.6 may enhance CsUFGT transcription, potentially by facilitating CsMYB12-mediated promoter activation. Co-expression of CsMYB12 with the CsUFGT promoter construct significantly increased luciferase activity, and this effect was further amplified by the addition of Cs_lncRNA.18443.6, suggesting a possible synergistic interaction (Fig. 8i). The enhanced promoter activation in the presence of both the lncRNA and TF implies that Cs_lncRNA.18443.6 could stabilize CsMYB12 binding or modulate chromatin accessibility at the CsUFGT promoter. The observed dose-dependent increase in promoter activity with escalating CsMYB12 levels further supports a hypothetical cooperative model.
To further validate the module, we conducted a Y1H assay, which confirmed the direct binding of CsMYB12 to the CsUFGT promoter, while the lncRNA displayed no DNA-binding activity, consistent with its predicted non-coding nature. Although these assays were performed in a heterologous tobacco system, the functional enhancement of CsMYB12-mediated promoter activation in the presence of Cs_lncRNA.18443.6 suggests that key components of this regulatory mechanism may be conserved. Integrating these results, we propose a working model in which Cs_lncRNA.18443.6 acts as a co-regulatory factor that enhances CsMYB12-driven activation of CsUFGT transcription. While the updated Dual-LUC and Y1H experiments strengthen the evidence for an lncRNA–TF–target gene regulatory module, further biochemical and genetic validation will be required to fully elucidate the underlying mechanism and confirm its in planta relevance.
Comparative analyses can help elucidate the origin and function of lncRNAs52. At present, we believe this lncRNA is specific to the Theaceae. All sequenced species of Camellia and all varieties of C. sinensis have sequences with high blast similarity (Table 1; Fig. S11). No genomic resources currently exist for other genera in the Theaceae, but it would be interesting to survey other Theaceae genera, such as Franklinia, Gordonia, Polyspora, Schima and Stewartia. Related families in the Ericales for which whole genome data exist, such as the Actinidiaceae (kiwi-fruit family), contain no detectably similar sequences. This accords with what is known about the fast evolutionary turnover of lncRNA sequences52. However, the apparent ubiquity of these sequences throughout the genus Camellia means that these sequences could find wide application in tea breeding.
Data availability
All the data used in this research have been deposited to the National Center for Biotechnology Information (NCBI) under the BioProject accession number PRJNA1019822 and BioSample: SAMN37497780. All Supplementary Tables are found in Supplementary Data 1. The numerical source data (Figs. 6, 8d, h–k) for the graphs can be found in Supplementary Data 2. Supplementary Fig. 1, Data 1 and 2 are available online.
Code availability
The main code sources for the prediction of lncRNA used in this study are available at Zenodo: https://zenodo.org/records/10976644.
References
Xia, E. H. et al. The tea tree genome provides insights into tea flavor and independent evolution of caffeine biosynthesis. Mol. Plant 10, 866–877 (2017).
Liao, Y., Smyth, G. & Shi, W. FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923 (2014).
Zeng, L. T., Watanabe, N. & Yang, Z. Y. Understanding the biosyntheses and stress response mechanisms of aroma compounds in tea (Camellia sinensis) to safely and effectively improve tea aroma. Crit. Rev. Food Sci. Nutr. 59, 2321–2334 (2019).
Grand View Research. Tea Market Size and Outlook: Horizon Databooks. https://www.grandviewresearch.com/horizon/outlook/tea-market-size/global (Grand View Research, Inc., 2024).
Li, G. T. et al. Research progress of Yunnan endemic tea variety-zijuan. China Tea 35, 10–12 (2013).
Li, X. X. et al. Anthocyanin metabolism and its differential regulation in purple tea (Camellia sinensis). Plant Physiol. Biochem. 201, 107875 (2023).
Wang, K. R., Liang, Y. R., Li, M. & Zhang, L. J. Progress in the development of germplasm resources for albino and purple tea cultivars. China Tea Processing 3, 5–8 (2015).
Ji, Q. Y., Liu, Y., Zhou, H. J., Shao, J. N. & He, W. Z. Biochemical component analysis and germplasm selection of tea tree germplasm resources with reddish violet leaves. Jiangsu Agric. Sci. 50, 110–116 (2022).
Mozos, I. et al. Effects of anthocyanins on vascular health. Biomolecules 11, 811 (2021).
Nistor, M. et al. Anthocyanins as key phytochemicals acting for the prevention of metabolic diseases: an overview. Molecules 27, 4254 (2022).
Zhang, W. et al. Genome assembly of wild tea tree DASZ reveals pedigree and selection history of tea varieties. Nat. Commun. 11, 3719 (2020).
Ferrer, J. L., Austin, M. B., Stewart, C. Jr & Noel, J. P. Structure and function of enzymes involved in the biosynthesis of phenylpropanoids. Plant Physiol. Biochem. 46, 356–370 (2008).
Kanehisa, M., Furumichi, M., Sato, Y., Ishiguro-Watanabe, M. & Tanabe, M. KEGG: integrating viruses and cellular organisms. Nucleic Acids Res. 49, D545–D551 (2021).
Lai, Y. S. et al. The dark-purple tea cultivar ‘ziyan’ accumulates a large amount of delphinidin-related anthocyanins. J. Agric. Food Chem. 64, 2719–2726 (2016).
Kumari, M. et al. Regulation of color transition in purple tea (Camellia sinensis). Planta 251, 35 (2020).
Li, F. et al. Integrated transcriptome and metabolome provide insights into flavonoid biosynthesis in ‘P113’, a new purple tea of Camellia tachangensis. Beverage Plant Res. 3, 3 (2023). 14.
Amaral, P. P., Clark, M. B., Gascoigne, D. K., Dinger, M. E. & Mattick, J. S. lncRNAdb: a reference database for long noncoding RNAs. Nucleic Acids Res. 39, D146–D151 (2011).
Zhou, C. Z. et al. Hidden players in the regulation of secondary metabolism in tea plant: focus on non-coding RNAs. Beverage Plant Res. 2, 19 (2022).
Palos, K., Yu, L., Railey, C. E., Nelson Dittrich, A. C. & Nelson, A. D. L. Linking discoveries, mechanisms, and technologies to develop a clearer perspective on plant long noncoding RNAs. Plant Cell 35, 1762–1786 (2023).
Rinn, J. L. & Chang, H. Y. Genome regulation by long noncoding RNAs. Annu. Rev. Biochem. 81, 145–166 (2012).
Ma, L., Bajic, V. B. & Zhang, Z. On the classification of long non-coding RNAs. RNA Biol. 10, 925–933 (2013).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Zhang, X. et al. Haplotype-resolved genome assembly provides insights into evolutionary history of the tea plant Camellia sinensis. Nat. Genet. 53, 1250–1259 (2021).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Sun, L. et al. Utilizing sequence intrinsic composition to classify protein-coding and long non-coding transcripts. Nucleic Acids Res. 41, e166 (2013).
Li, A., Zhang, J. & Zhou, Z. PLEK: a tool for predicting long non-coding RNAs and messenger RNAs based on an improved k-mer scheme. BMC Bioinforma. 15, 311 (2014).
Kang, Y. J. et al. CPC2: a fast and accurate coding potential calculator based on sequence intrinsic features. Nucleic Acids Res. 45, W12–W16 (2017).
Wucher, V. et al. FEELnc: a tool for long non-coding RNA annotation and its application to the dog transcriptome. Nucleic Acids Res. 45, e57 (2017).
Buchfink, B., Reuter, K. & Drost, H. G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat. Methods 18, 366–368 (2021).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
Ma, X. et al. TarHunter, a tool for predicting conserved microRNA targets and target mimics in plants. Bioinformatics 34, 1574–1576 (2018).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinforma. 9, 559 (2008).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Sun et al. Reference genes for real-time fluorescence quantitative PCR in Camellia sinensis. Chin. Bull. Bot. 45, 579–587 (2010).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408 (2001).
Hartley, J. L., Temple, G. F. & Brasch, M. A. DNA cloning using in vitro site-specific recombination. Genome Res. 10, 1788–1795 (2000).
Cui, L. et al. Identification of UDP-glycosyltransferases involved in the biosynthesis of astringent taste compounds in tea (Camellia sinensis). J. Exp. Bot. 67, 2285–2297 (2016).
He, X. J. et al. Isolation and characterization of key genes that promote flavonoid accumulation in purple-leaf tea (Camellia sinensis L. Sci. Rep. 8, 130 (2018).
Shen, J. et al. Metabolic analyses reveal different mechanisms of leaf color change in two purple-leaf tea plant (Camellia sinensis L.) cultivars. Horticulture Res. 5, 7 (2018).
Cai, J. et al. Integrative analysis of metabolomics and transcriptomics reveals molecular mechanisms of anthocyanin metabolism in the Zikui tea plant (Camellia sinensis cv. Zikui). Int. J. Mol. Sci. 23, 4780 (2022).
Liu, S. J. et al. CRISPRi-based genome-scale identification of functional long noncoding RNA loci in human cells. Science 355, aah7111 (2017).
Varshney, D. et al. Tissue specific long non-coding RNAs are involved in aroma formation of black tea. Ind. Crops Products 133, 79–89 (2019).
Zhu, C. et al. Transcriptome and phytochemical analyses provide new insights into long non-coding RNAs modulating characteristic secondary metabolites of oolong tea (Camellia sinensis) in solar-withering. Front. Plant Sci. 10, 1638 (2019).
Griffiths-Jones, S., Bateman, A., Marshall, M., Khanna, A. & Eddy, S. R. Rfam: an RNA family database. Nucleic Acids Res. 31, 439–441 (2003).
Finn, R. D. et al. Pfam: the protein families database. Nucleic Acids Res. 42, D222–D230 (2014).
Liu, J. et al. Genome-wide analysis uncovers regulation of long intergenic noncoding RNAs in Arabidopsis. Plant Cell 24, 4333–4345 (2012).
Mattick, J. S. et al. Long non-coding RNAs: definitions, functions, challenges and recommendations. Nat. Rev. Mol. Cell Biol. 24, 430–447 (2023).
Gil, N. & Ulitsky, I. Regulation of gene expression by cis-acting long non-coding RNAs. Nat. Rev. Genet. 21, 102–117 (2020).
Zhou, C. Z. et al. Establishment of an efficient in planta transformation method for Camellia sinensis. Biotechnol. Bull. 38, 263–265 (2022).
Ulitsky, I. Evolution to the rescue: using comparative genomics to understand long non-coding RNAs. Nat. Rev. Genet. 17, 601–614 (2016).
Acknowledgements
This work was supported by Guizhou Province Major Science and Technology Project (2025) 044, the National Natural Science Foundation of China (32260086), China Scholarship Council. Grant (No. 202108525029) and the Cultivation Project of Guizhou University (Gzu.2020 No.65).
Author information
Authors and Affiliations
Contributions
S.N. and B.X. conceived the project and designed the study; Q.S. and L.Z. collected and raised the plants; Q.L. and Y.Y. sampled the materials; B.X., Q.L., and Y.Y. performed the formal experiments; B.W., Q.L., and Y.Y. performed the transgenic tobacco experiments; B.X. and L.Z. performed the bioinformatic data analysis; B.X., Q.L., and Y.Y. performed the formal analysis and data curation; B.X. and Q.L. designed and visualized the tables and figures; B.X. and L.Z. wrote the first manuscript; Q.C. provided suggestions and facilities and edited and revised the manuscript; S.N., Q.C., and B.X. supervised the entire project; All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Zhimin Qiu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Kaliya Georgieva.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xiong, B., Zhang, L., Li, Q. et al. A long noncoding RNA modulates anthocyanin biosynthesis in Camellia sinensis. Commun Biol 9, 675 (2026). https://doi.org/10.1038/s42003-026-09785-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-026-09785-7