Abstract
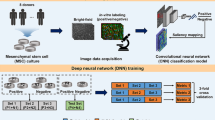
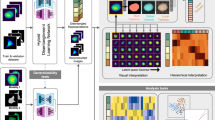
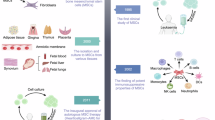
Cellular morphology, a critical manifestation of biological characteristics, is linked to functions. In traditional cell detection, invasive labeling and detection methods not only compromise cellular viability but also entail labor-intensive workflows. Here we presented a non-invasive artificial intelligence framework that integrated deep learning (DL) and machine learning (ML) to predict the immunomodulatory capacity of mesenchymal stem cells (MSCs) through morphological profiling. The improved PreAct-ResNet50 encoder-decoder architecture was used to achieve high-accuracy instance segmentation of cells and nuclei, enabling quantification of morphological features. A LightGBM-based predictive model was subsequently employed to predict MSCs immunomodulatory biomarkers through morphological features. This dual-model system demonstrated satisfactory cell segmentation and biological characteristics prediction capabilities through performance testing. Our method provided an efficient, non- invasive tool for real-time MSCs potency assessment, which could enhance quality controls in cell therapy manufacturing.
Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and Supplementary data 5. A subset of the training and test data is publicly accessible on Figshare (https://doi.org/10.6084/m9.figshare.31175839)41. Complete training and test data of this study are available on request from the corresponding author.
Code availability
The code for the fully trained model is available on GitHub at https://github.com/Rye-052D/A-dual-model-system-for-MSC-segmentation-and-function-prediction/tree/main. Additionally, the code used for model training is publicly and permanently available on Figshare (https://doi.org/10.6084/m9.figshare.31175839), under the Apache 2.0 license.
References
Yuqian Z., Jianxin L., Huifen L., Guiyuan L. Improved watershed algorithm for dowels image segmentation. In: 2008 7th World Congress on Intelligent Control and Automation) (2008).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. ImageNet classification with deep convolutional neural networks. Commun ACM 60, 84–90 (2017).
Blanc, K. L. et al. ISCT MSC committee statement on the US FDA approval of allogenic bone-marrow mesenchymal stromal cells. Cytotherapy, (2025).
Liang, J. et al. Allogenic mesenchymal stem cells transplantation in refractory systemic lupus erythematosus: a pilot clinical study. Ann Rheum Dis 69, 1423–1429 (2010).
Uccelli, A. et al. Safety, tolerability, and activity of mesenchymal stem cells versus placebo in multiple sclerosis (MESEMS): a phase 2, randomised, double-blind crossover trial. Lancet Neurol 20, 917–929 (2021).
Dominici, M. et al. Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy 8, 315–317 (2006).
Zha, K. et al. Heterogeneity of mesenchymal stem cells in cartilage regeneration: from characterization to application. npj Regenerative Medicine 6, 14 (2021).
Zhou, Y. et al. Assessing Biomaterial-Induced Stem Cell Lineage Fate by Machine Learning-Based Artificial Intelligence. Adv Mater 35, e2210637 (2023).
Ramezani, M. et al. A genome-wide atlas of human cell morphology. Nat Methods, (2025).
Harrison, P. J. et al. Evaluating the utility of brightfield image data for mechanism of action prediction. PLOS Computational Biology 19, e1011323 (2023).
Klinker, M. W., Marklein, R. A., Lo Surdo, J. L., Wei, C.-H. & Bauer, S. R. Morphological features of IFN-γ–stimulated mesenchymal stromal cells predict overall immunosuppressive capacity. Proceedings of the National Academy of Sciences 114, E2598–E2607 (2017).
Marklein, R. A. et al. Morphological profiling using machine learning reveals emergent subpopulations of interferon-γ–stimulated mesenchymal stromal cells that predict immunosuppression. Cytotherapy 21, 17–31 (2019).
Rui, K. et al. Differential Phase Contrast Imaging to Predict MSC Immune Function. Adv Healthc Mater 14, e2501553 (2025).
Kong, Y. et al. Evaluating Differentiation Status of Mesenchymal Stem Cells by Label-Free Microscopy System and Machine Learning. Cells 12, (2023).
He, L. et al. Morphology-based deep learning enables accurate detection of senescence in mesenchymal stem cell cultures. BMC Biology 22, (2024).
Christiansen, E. M. et al. In Silico Labeling: Predicting Fluorescent Labels in Unlabeled Images. Cell 173, 792–803.e719 (2018).
Solopov, M. et al. Comparative Study of Deep Transfer Learning Models for Semantic Segmentation of Human Mesenchymal Stem Cell Micrographs. Int. J. Mol. Sci. 26, (2025).
Park, H. Y. et al. Priming Mesenchymal Stem/Stromal Cells with a Combination of a Low Dose of IFN-γ and Bortezomib Results in Potent Suppression of Pathogenic Th17 Immunity Through the IDO1-AHR Axis. Stem Cells 41, 64–76 (2023).
Kuçi, Z. et al. Expression of HLA-DR by mesenchymal stromal cells in the platelet lysate era: an obsolete release criterion for MSCs? J Transl Med 22, 39 (2024).
Deuse, T. et al. Immunogenicity and immunomodulatory properties of umbilical cord lining mesenchymal stem cells. Cell Transplant 20, 655–667 (2011).
Kurawaki, S. et al. Mesenchymal stem cells pretreated with interferon-gamma attenuate renal fibrosis by enhancing regulatory T cell induction. Sci Rep 14, 10251 (2024).
López-García L., Castro-Manrreza M. E. TNF-α and IFN-γ Participate in Improving the Immunoregulatory Capacity of Mesenchymal Stem/Stromal Cells: Importance of Cell-Cell Contact and Extracellular Vesicles. Int. J. Mol. Sci. 22, (2021).
Ghannam, S., Pène, J., Moquet-Torcy, G., Jorgensen, C. & Yssel, H. Mesenchymal stem cells inhibit human Th17 cell differentiation and function and induce a T regulatory cell phenotype. J Immunol 185, 302–312 (2010).
Rozenberg, A. et al. Human Mesenchymal Stem Cells Impact Th17 and Th1 Responses Through a Prostaglandin E2 and Myeloid-Dependent Mechanism. Stem Cells Transl Med 5, 1506–1514 (2016).
Bai, L. et al. Human bone marrow-derived mesenchymal stem cells induce Th2-polarized immune response and promote endogenous repair in animal models of multiple sclerosis. Glia 57, 1192–1203 (2009).
Azevedo, R. I. et al. Mesenchymal stromal cells induce regulatory T cells via epigenetic conversion of human conventional CD4 T cells in vitro. Stem Cells 38, 1007–1019 (2020).
Naserian, S., Shamdani, S., Arouche, N. & Uzan, G. Regulatory T cell induction by mesenchymal stem cells depends on the expression of TNFR2 by T cells. Stem Cell Research & Therapy 11, 534 (2020).
Krampera, M. et al. Role for interferon-gamma in the immunomodulatory activity of human bone marrow mesenchymal stem cells. Stem Cells 24, 386–398 (2006).
Hoogduijn, M. J. Indoleamine 2,3-Dioxygenase Does It. Transplantation 99, 1751–1752 (2015).
Ren, G. et al. Inflammatory cytokine-induced intercellular adhesion molecule-1 and vascular cell adhesion molecule-1 in mesenchymal stem cells are critical for immunosuppression. J Immunol 184, 2321–2328 (2010).
Ren, G., Roberts, A. I. & Shi, Y. Adhesion molecules: key players in Mesenchymal stem cell-mediated immunosuppression. Cell Adh Migr 5, 20–22 (2011).
Yu, Y. et al. Preconditioning with interleukin-1 beta and interferon-gamma enhances the efficacy of human umbilical cord blood-derived mesenchymal stem cells-based therapy via enhancing prostaglandin E2 secretion and indoleamine 2,3-dioxygenase activity in dextran sulfate sodium-induced colitis. J Tissue Eng Regen Med 13, 1792–1804 (2019).
Kim, D. S. et al. Enhanced Immunosuppressive Properties of Human Mesenchymal Stem Cells Primed by Interferon-γ. EBioMedicine 28, 261–273 (2018).
Montesinos, J. J. et al. Human Bone Marrow Mesenchymal Stem/Stromal Cells Exposed to an Inflammatory Environment Increase the Expression of ICAM-1 and Release Microvesicles Enriched in This Adhesive Molecule: Analysis of the Participation of TNF-α and IFN-γ. J Immunol Res 2020, 8839625 (2020).
van Zutphen, T. & van der Klei, I. J. Quantitative analysis of organelle abundance, morphology and dynamics. Curr Opin Biotechnol 22, 127–132 (2011).
Heinrich, L. et al. Whole-cell organelle segmentation in volume electron microscopy. Nature 599, 141–146 (2021).
Miko, M., Danišovič, L., Majidi, A. & Varga, I. Ultrastructural analysis of different human mesenchymal stem cells after in vitro expansion: a technical review. Eur J Histochem 59, 2528 (2015).
He K., Zhang X., Ren S., Sun J. Identity Mappings in Deep Residual Networks. In: Computer Vision – ECCV 2016 (eds Leibe B., Matas J., Sebe N., Welling M.). Springer International Publishing (2016).
Stirling, D. R. et al. CellProfiler 4: improvements in speed, utility and usability. BMC Bioinformatics 22, 433 (2021).
Cai Z., Huang H., Sun G., Li Z., Ouyang C. Advancing Predictive Models: Unveiling LightGBM Machine Learning for Data Analysis. In: 2023 4th International Conference on Computer, Big Data and Artificial Intelligence (ICCBD+AI)) (2023).
Liu, Z.-y Deep learning-based in silico labeling for analyzing morphological features of MSCs to predict immunomodulatory capacity. figshare (2026).
Acknowledgements
The authors would like to thank Xiao Liang at Harbin Beike Health Technology Co., Ltd for their deployment with algorithmic results of this study in the established automated stem cell manufacturing platform for more MSCs data. This work was supported by grants from Shenzhen Non-invasive Cell Quality Online Monitoring and Analysis Platform (F-2022-Z99-502233 to M.L.), National Engineering Research Center of Foundational Technologies for CGT Industry (NDRC-High-Technology [2023] No. 447 to M.L.), Special Project on the Integrated Application of Biotechnology and Information Technology (Harbin Songbei Development and Reform Commission Letter [2023] No. 79), Joint Funds for the innovation of science and Technology, Fujian province (Grant number: 2024J011042).
Author information
Authors and Affiliations
Contributions
Zhiyu Liu: Writing—original draft, Methodology, Data curation, Project administration. Gang An: Methodology, Investigation, Validation. Xiao Liang: Conceptualization, Validation. Xumin Wu: Methodology, Validation, Formal analysis. Junyuan Hu: Resources, Conceptualization, Funding acquisition. Haijun Wang: Software, Visualization. Jingfeng Ou: Software, Visualization. Xiuping Zeng: Methodology, Validation. Zhiliang Xia: Software, Visualization. Kaixiang Hou: Methodology. Wanglong Chu: Methodology. Jianbin Ye: Supervision. Cui Liao: Supervision. Zhengmian Zhang: Funding acquisition. Muyun Liu: Writing—review and editing, Conceptualization, Project administration, Funding acquisition.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Shan E Ahmed Raza and Ophelia Bu. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, Z., An, G., Liang, X. et al. Deep learning-based in silico labeling for analyzing morphological features of MSCs to predict immunomodulatory capacity. Commun Biol (2026). https://doi.org/10.1038/s42003-026-09833-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-026-09833-2