Abstract
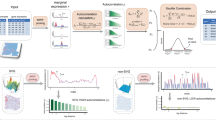
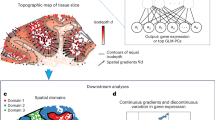
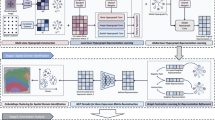
Spatial transcriptomics enables in situ gene expression profiling, yet precise spatial domain identification and marker gene detection remain challenging. We present HarveST, a heterogeneous graph-based framework that integrates spatial, transcriptomic, and gene-gene interaction data through a unified computational model. HarveST employs dual learning strategies: self-supervised learning for feature extraction and partially supervised refinement for domain delineation. Additionally, it implements a Random Walk with Restart algorithm for identifying spatial domain-marker spatially variable genes (SVGs). Applied to human cortical tissue, mouse olfactory bulb, and tumor microenvironments across multiple platforms, HarveST demonstrates superior performance in detecting biologically meaningful spatial domains and associated marker genes. HarveST further supports joint analysis across consecutive spatial transcriptomics sections, enabling consistent reconstruction of functional domains across slices. By capturing both spatial topology and molecular relationships in a single graph-theoretical framework, HarveST advances spatial transcriptomics analysis beyond conventional clustering approaches, offering deeper insights into tissue architecture and cellular interactions in normal and pathological contexts.
Similar content being viewed by others
Data availability
The spatial transcriptomics datasets analyzed in this study are publicly available from the following repositories. The Human Dorsolateral Prefrontal Cortex (DLPFC) dataset29,65 is available at http://spatial.libd.org/spatialLIBD/. The 10x Visium Human Breast Cancer dataset is available from the 10x Genomics website at https://www.10xgenomics.com/resources/datasets. The Mouse Olfactory Bulb (Stereo-seq) dataset31,66 is available from CNGBdb (MOSTA) at https://db.cngb.org/data_resources/project/CNP0001543. The Mouse Olfactory Bulb (Slide-seqV2) dataset21,67 is available via the Broad Institute Single Cell Portal at https://singlecell.broadinstitute.org/single_cell/study/SCP815. The PDAC dataset is available from the Gene Expression Omnibus (GEO) under accession number GSE11167250,68 at https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE111672. Processed data supporting the findings of this study are available in the GitHub repository (https://github.com/Seven595/HarveST) and are archived on Zenodo (https://doi.org/10.5281/zenodo.18532348). Source data underlying the plots in the main figures are provided with this paper. Any remaining data are available from the corresponding author upon reasonable request.
Code availability
The source code for HarveST, including the implementation of the heterogeneous graph learning framework and scripts for reproducing the analysis, is openly available on GitHub at https://github.com/Seven595/HarveST. A persistent version of the software code and processed data used to generate the results in this manuscript is archived in the Zenodo repository (https://doi.org/10.5281/zenodo.18532348)69.
References
Asp, M. et al. A spatiotemporal organ-wide gene expression and cell atlas of the developing human heart. Cell 179, 1647–1660 (2019).
Armingol, E., Officer, A., Harismendy, O. & Lewis, N. E. Deciphering cell–cell interactions and communication from gene expression. Nat. Rev. Genet. 22, 71–88 (2021).
Dries, R. et al. Advances in spatial transcriptomic data analysis. Genome Res. 31, 1706–1718 (2021).
Ferreira, R. M. et al. Integration of spatial and single-cell transcriptomics localizes epithelial cell–immune cross-talk in kidney injury. JCI Insight 6, e147703 (2021).
Cheung, M. D. et al. Resident macrophage subpopulations occupy distinct microenvironments in the kidney. JCI Insight 7, e161078 (2022).
Zhou, R., Yang, G., Zhang, Y. & Wang, Y. Spatial transcriptomics in development and disease. Mol. Biomed. 4, 32 (2023).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. Theory Exp. 2008, P10008 (2008).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Zhao, E. et al. Spatial transcriptomics at subspot resolution with bayesspace. Nat. Biotechnol. 39, 1375–1384 (2021).
Xu, H. et al. Unsupervised spatially embedded deep representation of spatial transcriptomics. Genome Med. 16, 12 (2024).
Dong, K. & Zhang, S. Deciphering spatial domains from spatially resolved transcriptomics with an adaptive graph attention auto-encoder. Nat. Commun. 13, 1739 (2022).
Long, Y. et al. Spatially informed clustering, integration, and deconvolution of spatial transcriptomics with graphst. Nat. Commun. 14, 1155 (2023).
Ren, H., Walker, B. L., Cang, Z. & Nie, Q. Identifying multicellular spatiotemporal organization of cells with spaceflow. Nat. Commun. 13, 4076 (2022).
Xu, K., Xu, Y., Wang, Z., Zhou, X. M. & Zhang, L. stdyer enables spatial domain clustering with dynamic graph embedding. Genome Biol. 26, 1–25 (2025).
Wang, Y., Liu, Z. & Ma, X. Mnmst: topology of cell networks leverages identification of spatial domains from spatial transcriptomics data. Genome Biol. 25, 133 (2024).
Wang, H., Zhao, J., Nie, Q., Zheng, C. & Sun, X. Dissecting spatiotemporal structures in spatial transcriptomics via diffusion-based adversarial learning. Research 7, 0390 (2024).
Zuo, C., Xia, J. & Chen, L. Dissecting tumor microenvironment from spatially resolved transcriptomics data by heterogeneous graph learning. Nat. Commun. 15, 5057 (2024).
Duan, Z. et al. Impeller: a path-based heterogeneous graph learning method for spatial transcriptomic data imputation. Bioinformatics 40, btae339 (2024).
Varrone, M., Tavernari, D., Santamaria-Martínez, A., Walsh, L. A. & Ciriello, G. Cellcharter reveals spatial cell niches associated with tissue remodeling and cell plasticity. Nat. Genet. 56, 74–84 (2024).
Ji, A. L. et al. Multimodal analysis of composition and spatial architecture in human squamous cell carcinoma. Cell 182, 497–514 (2020).
Stickels, R. R. et al. Highly sensitive spatial transcriptomics at near-cellular resolution with slide-seqv2. Nat. Biotechnol. 39, 313–319 (2021).
Yan, G., Hua, S. H. & Li, J. J. Categorization of 34 computational methods to detect spatially variable genes from spatially resolved transcriptomics data. Nat. Commun. 16, 1141 (2025).
Edsgärd, D., Johnsson, P. & Sandberg, R. Identification of spatial expression trends in single-cell gene expression data. Nat. Methods 15, 339–342 (2018).
Svensson, V., Teichmann, S. A. & Stegle, O. Spatialde: identification of spatially variable genes. Nat. Methods 15, 343–346 (2018).
Sun, S., Zhu, J. & Zhou, X. Statistical analysis of spatial expression patterns for spatially resolved transcriptomic studies. Nat. Methods 17, 193–200 (2020).
Wolf, F. A., Angerer, P. & Theis, F. J. Scanpy: large-scale single-cell gene expression data analysis. Genome Biol. 19, 1–5 (2018).
Hu, J. et al. Spagcn: Integrating gene expression, spatial location and histology to identify spatial domains and spatially variable genes by graph convolutional network. Nat. Methods 18, 1342–1351 (2021).
Cai, P., Robinson, M. D. & Tiberi, S. Despace: spatially variable gene detection via differential expression testing of spatial clusters. Bioinformatics 40, btae027 (2024).
Maynard, K. R. et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat. Neurosci. 24, 425–436 (2021).
Shi, X., Zhu, J., Long, Y. & Liang, C. Identifying spatial domains of spatially resolved transcriptomics via multi-view graph convolutional networks. Brief. Bioinform. 24, bbad278 (2023).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 (2022).
Charych, E. I., Liu, F., Moss, S. J. & Brandon, N. J. Gabaa receptors and their associated proteins: implications in the etiology and treatment of schizophrenia and related disorders. Neuropharmacology 57, 481–495 (2009).
Kadowaki, K. et al. Phosphohippolin expression in the rat central nervous system. Mol. Brain Res. 125, 105–112 (2004).
Sunkin, S. M. et al. Allen brain atlas: an integrated spatio-temporal portal for exploring the central nervous system. Nucleic Acids Res. 41, D996–D1008 (2012).
Zacharias, D. A. & Kappen, C. Developmental expression of the four plasma membrane calcium atpase (pmca) genes in the mouse. Biochim. Biophys. Acta Gen. Subj. 1428, 397–405 (1999).
Castelli, L. M. et al. Srsf1-dependent inhibition of c9orf72-repeat rna nuclear export: genome-wide mechanisms for neuroprotection in amyotrophic lateral sclerosis. Mol. Neurodegener. 16, 53 (2021).
Neftel, C. et al. An integrative model of cellular states, plasticity, and genetics for glioblastoma. Cell 178, 835–849 (2019).
Wei, Y., Yang, X., Gao, L., Xu, Y. & Yi, C. Differences in potential key genes and pathways between primary and radiation-associated angiosarcoma of the breast. Transl. Oncol. 19, 101385 (2022).
Plasschaert, L. W. et al. A single-cell atlas of the airway epithelium reveals the CFTR-rich pulmonary ionocyte. Nature 560, 377–381 (2018).
Young, M. D. et al. Single-cell transcriptomes from human kidneys reveal the cellular identity of renal tumors. science 361, 594–599 (2018).
Yamada, A. et al. High expression of atp-binding cassette transporter abcc11 in breast tumors is associated with aggressive subtypes and low disease-free survival. Breast Cancer Res. Treat. 137, 773–782 (2013).
O’Flanagan, C. H. et al. Dissociation of solid tumor tissues with cold active protease for single-cell rna-seq minimizes conserved collagenase-associated stress responses. Genome Biol. 20, 1–13 (2019).
Shikang, Z., Xin, J. & Song, X. Expression of rasgrp2 in lung adenocarcinoma and its effect on immune microenvironment. Zhongguo Fei Ai Za Zhi 24, 404–411 (2021).
Wu, S. Z. et al. Stromal cell diversity associated with immune evasion in human triple-negative breast cancer. EMBO J. 39, e104063 (2020).
Xydia, M. et al. Common clonal origin of conventional t cells and induced regulatory t cells in breast cancer patients. Nat. Commun. 12, 1119 (2021).
Kester, L. et al. Differential survival and therapy benefit of patients with breast cancer are characterized by distinct epithelial and immune cell microenvironments. Clin. Cancer Res. 28, 960–971 (2022).
Li, H., Calder, C. A. & Cressie, N. Beyond moran’s i: testing for spatial dependence based on the spatial autoregressive model. Geogr. Anal. 39, 357–375 (2007).
Abdelaal, T., Mourragui, S., Mahfouz, A. & Reinders, M. J. Spage: spatial gene enhancement using scrna-seq. Nucleic Acids Res. 48, e107 (2020).
Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat. Biotechnol. 40, 517–526 (2022).
Moncada, R. et al. Integrating microarray-based spatial transcriptomics and single-cell rna-seq reveals tissue architecture in pancreatic ductal adenocarcinomas. Nat. Biotechnol. 38, 333–342 (2020).
De Kanter, J. K., Lijnzaad, P., Candelli, T., Margaritis, T. & Holstege, F. C. Chetah: a selective, hierarchical cell type identification method for single-cell RNA sequencing. Nucleic Acids Res. 47, e95 (2019).
Qadir, M. M. F. et al. Single-cell resolution analysis of the human pancreatic ductal progenitor cell niche. Proc. Natl. Acad. Sci. USA 117, 10876–10887 (2020).
Gonçalves, C. A. et al. A 3d system to model human pancreas development and its reference single-cell transcriptome atlas identify signaling pathways required for progenitor expansion. Nat. Commun. 12, 3144 (2021).
Tosti, L. et al. Single-nucleus and in situ rna–sequencing reveal cell topographies in the human pancreas. Gastroenterology 160, 1330–1344 (2021).
Sun, H. et al. Dissecting the heterogeneity and tumorigenesis of BRCA1 deficient mammary tumors via single cell RNA sequencing. Theranostics 11, 9967 (2021).
Li, C., Guo, L., Li, S. & Hua, K. Single-cell transcriptomics reveals the landscape of intra-tumoral heterogeneity and transcriptional activities of ECS in cc. Mol. Ther. Nucleic Acids 24, 682–694 (2021).
Kim, J. Y. et al. Stratifin (sfn) regulates lung cancer progression via nucleating the Vps34-BeCN1-Traf6 complex for autophagy induction. Clin. Transl. Med. 12, e896 (2022).
Egelston, C. A. et al. Tumor-infiltrating exhausted CD8+ T cells dictate reduced survival in premenopausal estrogen receptor–positive breast cancer. JCI Insight 7, e153963 (2022).
Merz, M. et al. Deciphering spatial genomic heterogeneity at a single cell resolution in multiple myeloma. Nat. Commun. 13, 807 (2022).
Ge, P. et al. Identifying drug candidates for pancreatic ductal adenocarcinoma based on integrative multiomics analysis. J. Gastrointest. Oncol. 15, 1265 (2024).
Zhang, M. et al. Development and validation of cancer-associated fibroblasts-related gene landscape in prognosis and immune microenvironment of bladder cancer. Front. Oncol. 13, 1174252 (2023).
Han, J., DePinho, R. A. & Maitra, A. Single-cell rna sequencing in pancreatic cancer. Nat. Rev. Gastroenterol. Hepatol. 18, 451–452 (2021).
Majdalawieh, A. F., Massri, M. & Ro, H.-S. Aebp1 is a novel oncogene: mechanisms of action and signaling pathways. J. Oncol. 2020, 8097872 (2020).
Cortes, C. & Vapnik, V. Support-vector networks. Mach. learn. 20, 273–297 (1995).
Maynard, K. et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex [data set]. http://spatial.libd.org/spatialLIBD/ (2021).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using dna nanoball-patterned arrays [data set]. https://db.cngb.org/stomics/mosta/ (2022).
Stickels, R. et al. Highly sensitive spatial transcriptomics at near-cellular resolution with slide-seqv2 [data set]. https://singlecell.broadinstitute.org/single_cell/study/SCP815 (2021).
Moncada, R. et al. Integrating microarray-based spatial transcriptomics and single-cell rna-seq reveals tissue architecture in pancreatic ductal adenocarcinomas [data set]. https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE111672 (2020).
Feng, J., Yu, T. & Zhang, Y. Harvest: heterogeneous graph learning framework for spatial transcriptomics [data set/software]. Preprint at https://doi.org/10.5281/zenodo.18532348 (2025).
Acknowledgements
This work was partially supported by Shenzhen Science and Technology Program (Grant Nos. ZDSYS20230626091302006 and JCYJ20240813113536047). Y.Z. is partially supported by a Guangdong Provincial Project (2024QN11N085). During the preparation of this work, the author(s) utilized OpenAI’s ChatGPT-4 to enhance the readability and clarity of the text, given that the primary authors are non-native English speakers. The author(s) subsequently reviewed and edited the AI-generated content as necessary and take full responsibility for the final content of the publication.
Author information
Authors and Affiliations
Contributions
Conceptualization: J.F., T.Y., and Y.Z.; Methodology: J.F. and Y.Z.; Software: J.F.; Validation: J.F. and T.Y.; Formal analysis: J.F.; Investigation: J.F. and Y.Z.; Resources: T.Y. and Y.Z.; Data curation: J.F.; Writing—original draft: J.F.; Writing—review & editing: J.F., T.Y., and Y.Z.; Visualization: J.F.; Supervision: T.Y. and Y.Z.; Project administration: Y.Z.; Funding acquisition: T.Y. and Y.Z.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Bin Li and Jun Ding for their contribution to the peer review of this work. Primary handling editors: Leelavati Narlikar, Aylin Bircan and George Inglis. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Feng, J., Yu, T. & Zhang, Y. HarveST uses a heterogeneous graph learning framework to reveal spatial transcriptomics patterns. Commun Biol (2026). https://doi.org/10.1038/s42003-026-09841-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-026-09841-2