Abstract
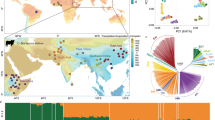
Domestic cattle (Bos taurus and Bos indicus) underpin food security and livelihoods worldwide but face intensifying pressures from climate change, infectious disease, and inconsistent feed supplies. African and European indigenous cattle provide a natural comparative framework spanning gradients of climate, pathogen burden, and husbandry, and possess genomic mosaics comprising African taurine, European taurine, and indicine ancestry. We analyzed whole-genome sequences from 519 cattle across 24 African and European indigenous populations and 117 publicly available genomes from Africa, Asia, Europe, and the Americas. This dataset reveals admixture mosaics among major lineages and identifies 36 candidate genes exhibiting adaptive retention of ancestral alleles associated with response to heat stress (e.g., HSPA12B, DDIT3), immunity (IRAK3), productivity (ACSF3), and reproductivity (SSMEM1, SPEF1). Our study suggests that historical admixture introduced variation shaped by local ecological selection, clarifying how environmental heterogeneity drives the retention of advantageous alleles and informing sustainable breeding and diversity conservation.
Similar content being viewed by others
Data availability
Population-level variant calls (VCF files) for the 636 indigenous and reference cattle analyzed in this study have been deposited in the European Variation Archive under accession PRJEB102975 (ERP184337)141. Whole-genome sequence data for the 519 African and European indigenous cattle analyzed in this study are available from the European Nucleotide Archive under accessions PRJEB90914142 and PRJEB90816143. Publicly available reference variant datasets were obtained from the 1000 Bull Genomes Project (PRJNA391427; ERZ14211345)144 and the African Genomic Reference Resource (PRJEB74565; Muturu)145. Sample IDs for the ancestry reference panels (AFT, n = 34; EUT, n = 29; AFI, n = 25; AAI, n = 29) are provided in Supplementary Data 1. Supplementary Data 2 contains genome-wide haplotype-mosaic files for locus-level inspection, provided for two AFT reference configurations (N’Dama + Muturu and Muturu only). Supplementary Data 3 lists putative ancestry-retention genes together with retention rates. Supplementary Data 4 provides window-based nucleotide diversity (π) and genetic differentiation (weighted Fst) for Mertolenga and EUT populations. All supplementary datasets are available via the project repository on GitHub: https://github.com/junxingao888/ancestral_retention_signals_cattle/. Source data underlying the figures are provided as follows: Supplementary Table 3 (Fig. 2a,b), Supplementary Table 4 (Fig. 2c,d), Supplementary Table 5 (Fig. 4c), Supplementary Table 6 (Fig. 4e), and Supplementary Data 4 (Fig. 5d,e).
Code availability
All custom code and workflows (Linux shell, R, and Python) for read processing, variant calling, local-ancestry inference, selection scans, and figure generation are available at GitHub (https://github.com/junxingao888/ancestral_retention_signals_cattle/tree/main/Code_availability) and Zenodo140.
References
Godde, C. M., Mason-D’Croz, D., Mayberry, D. E., Thornton, P. K. & Herrero, M. Impacts of climate change on the livestock food supply chain; a review of the evidence. Glob. Food Secur. 28, 100488 (2021).
Kim, K. et al. The mosaic genome of indigenous African cattle as a unique genetic resource for African pastoralism. Nat. Genet. 52, 1099–1110 (2020).
Felius, M. Cattle Breeds of the World. (BRILL, 2024).
Malek, Ž et al. Improving the representation of cattle grazing patterns in the European Union. Environ. Res. Lett. 19, 114077 (2024).
Friedrich, J. et al. Mapping restricted introgression across the genomes of admixed indigenous African cattle breeds. Genet. Sel. Evol. 55, 91 (2023).
Ward, J. A. et al. Genome-wide local ancestry and evidence for mitonuclear coadaptation in African hybrid cattle populations. iScience 25, 104672 (2022).
Chen, N. et al. Whole-genome resequencing reveals world-wide ancestry and adaptive introgression events of domesticated cattle in East Asia. Nat. Commun. 9, 2337 (2018).
Achilli, A. et al. Mitochondrial genomes of extinct aurochs survive in domestic cattle. Curr. Biol. 18, R157–R158 (2008).
Chen, N. et al. Global genetic diversity, introgression, and evolutionary adaptation of indicine cattle revealed by whole genome sequencing. Nat. Commun. 14, 7803 (2023).
Bollongino, R. et al. Modern taurine cattle descended from a small number of Near Eastern founders. Mol. Biol. Evol. 29, 2101–2104 (2012).
Chen, S. et al. Zebu cattle are an exclusive legacy of the South Asia neolithic. Mol. Biol. Evol. 27, 1–6 (2010).
Loftus, R. T., MacHugh, D. E., Bradley, D. G., Sharp, P. M. & Cunningham, P. Evidence for two independent domestications of cattle. Proc. Natl. Acad. Sci. 91, 2757–2761 (1994).
Scheu, A. et al. The genetic prehistory of domesticated cattle from their origin to the spread across Europe. BMC Genet. 16, 54 (2015).
Brito, L. F. et al. Review: Genetic selection of high-yielding dairy cattle toward sustainable farming systems in a rapidly changing world. Animal 15, 100292 (2021).
Flori, L. et al. The genome response to artificial selection: a case study in dairy cattle. PLoS One 4, e6595 (2009).
Park, S. D. et al. Genome sequencing of the extinct Eurasian wild aurochs, Bos primigenius, illuminates the phylogeography and evolution of cattle. Genome Biol. 16, 234 (2015).
Verdugo, M. P. et al. Ancient cattle genomics, origins, and rapid turnover in the Fertile Crescent. Science 365, 173–176 (2019).
Wragg, D. et al. A locus conferring tolerance to Theileria infection in African cattle. PLoS Genet. 18, e1010099 (2022).
Kambal, S. et al. Candidate signatures of positive selection for environmental adaptation in indigenous African cattle: A review. Anim. Genet. 54, 689–708 (2023).
Naessens, J. Bovine trypanotolerance: A natural ability to prevent severe anaemia and haemophagocytic syndrome?. Int J. Parasitol. 36, 521–528 (2006).
Nicolotti, M. & Guérin, C. Le zébu (Bos indicus) dans l’Egypte ancienne. Archaeozoologia 5, 87–108 (1992).
Hanotte, O. et al. African pastoralism: genetic imprints of origins and migrations. Science 296, 336–339 (2002).
Epstein, H. The origin of the domestic animals of Africa. Vol. II. 2 (1971).
Gifford-Gonzalez, D. & Hanotte, O. Domesticating animals in Africa: implications of genetic and archaeological findings. J. World Prehist. 24, 1–23 (2011).
Fuller, D. Q. & Boivin, N. Crops, cattle and commensals across the Indian Ocean. Current and potential archaeobiological evidence. Études Océ Indien 13, 46 (2009).
Felius, M. Cattle breeds: An encyclopedia. (1995).
Mwai, O., Hanotte, O., Kwon, Y.-J. & Cho, S. African indigenous cattle: unique genetic resources in a rapidly changing world. Asian-Australas. J. Anim. Sci. 28, 911 (2015).
Beja-Pereira, A. et al. The origin of European cattle: evidence from modern and ancient DNA. Proc. Natl. Acad. Sci. 103, 8113–8118 (2006).
Scholtz, M. & Theunissen, A. The use of indigenous cattle in terminal cross-breeding to improve beef cattle production in Sub-Saharan Africa. Anim. Genet. Resour. 46, 33–39 (2010).
da Fonseca, R. R. et al. Consequences of breed formation on patterns of genomic diversity and differentiation: the case of highly diverse peripheral Iberian cattle. BMC Genomics 20, 1–13 (2019).
Cymbron, T., Loftus, R. T., Malheiro, M. I. & Bradley, D. G. Mitochondrial sequence variation suggests an African influence in Portuguese cattle. Proc. R. Soc. Lond. Ser. B: Biol. Sci. 266, 597–603 (1999).
Decker, J. E. et al. Worldwide patterns of ancestry, divergence, and admixture in domesticated cattle. PLoS Genet. 10, e1004254 (2014).
Upadhyay, M. et al. Deciphering the patterns of genetic admixture and diversity in southern European cattle using genome-wide SNPs. Evolut. Appl. 12, 951–963 (2019).
Kim, J. et al. The genome landscape of indigenous African cattle. Genome Biol. 18, 34 (2017).
Nielsen, R., Hellmann, I., Hubisz, M., Bustamante, C. & Clark, A. G. Recent and ongoing selection in the human genome. Nat. Rev. Genet. 8, 857–868 (2007).
Huerta-Sánchez, E. et al. Altitude adaptation in Tibetans caused by introgression of Denisovan-like DNA. Nature 512, 194–197 (2014).
Qiu, Q. et al. The yak genome and adaptation to life at high altitude. Nat. Genet. 44, 946–949 (2012).
Scholtz, M., Bester, J., Mamabolo, J. & Ramsay, K. Results of the national cattle survey undertaken in South Africa, with emphasis on beef. Appl. Anim. Husb. Rural. Develop. 1, 1–9 (2008).
Kim, E.-S. & Rothschild, M. F. Genomic adaptation of admixed dairy cattle in East Africa. Front. Genet. 5, 443 (2014).
Upadhyay, M. Genomic variation across European cattle: contribution of gene flow. (Wageningen University and Research, 2019).
Daetwyler, H. D. et al. Whole-genome sequencing of 234 bulls facilitates mapping of monogenic and complex traits in cattle. Nat. Genet. 46, 858–865 (2014).
Hayes, B. J. & Daetwyler, H. D. 1000 bull genomes project to map simple and complex genetic traits in cattle: applications and outcomes. Annu. Rev. Anim. Biosci. 7, 89–102 (2019).
Moorjani, P. et al. The history of African gene flow into Southern Europeans, Levantines, and Jews. PLoS Genet. 7, e1001373 (2011).
Liang, M., Shishkin, M., Mikhailova, A., Shchur, V. & Nielsen, R. Estimating the timing of multiple admixture events using 3-locus linkage disequilibrium. PLoS Genet. 18, e1010281 (2022).
Dias-Alves, T., Mairal, J. & Blum, M. G. Loter: a software package to infer local ancestry for a wide range of species. Mol. Biol. Evol. 35, 2318–2326 (2018).
Racimo, F., Sankararaman, S., Nielsen, R. & Huerta-Sánchez, E. Evidence for archaic adaptive introgression in humans. Nat. Rev. Genet. 16, 359–371 (2015).
Garattini, E., Mendel, R., Romão, M. J., Wright, R. & Terao, M. Mammalian molybdo-flavoenzymes, an expanding family of proteins: structure, genetics, regulation, function and pathophysiology. Biochem. J. 372, 15–32 (2003).
Huntley, S. et al. A comprehensive catalog of human KRAB-associated zinc finger genes: insights into the evolutionary history of a large family of transcriptional repressors. Genome Res. 16, 669–677 (2006).
Kobayashi, K. et al. IRAK-M is a negative regulator of Toll-like receptor signaling. Cell 110, 191–202 (2002).
Harding, H. P. et al. Regulated translation initiation controls stress-induced gene expression in mammalian cells. Mol. Cell 6, 1099–1108 (2000).
Schroer, T. A. Dynactin. Annu. Rev. Cell Dev. Biol. 20, 759–779 (2004).
Utsunomiya, Y. et al. Genomic clues of the evolutionary history of Bos indicus cattle. Anim. Genet. 50, 557–568 (2019).
Li, M. et al. DDIT3 directs a dual mechanism to balance glycolysis and oxidative phosphorylation during glutamine deprivation. Adv. Sci. 8, 2003732 (2021).
Pereira, A. M., Baccari, F., Titto, E. A. & Almeida, J. A. Effect of thermal stress on physiological parameters, feed intake and plasma thyroid hormones concentration in Alentejana, Mertolenga, Frisian and Limousine cattle breeds. Int. J. Biometeorol. 52, 199–208 (2008).
Ginja, C. et al. Iron age genomic data from Althiburos-Tunisia renew the debate on the origins of African taurine cattle. Iscience 26 (2023).
Mengistie, D. Origin of cattle breeds in East Africa and introduction to general breeding science: A–review. World N. Nat. Sci. 49, 88–110 (2023).
Murray, M., Trail, J., Davis, C. & Black, S. J. Genetic resistance to African trypanosomiasis. J. Infect. Dis. 149, 311–319 (1984).
Zegeye, T., Belay, G., Vallejo-Trujillo, A., Han, J. & Hanotte, O. Genome-wide diversity and admixture of five indigenous cattle populations from the Tigray region of northern Ethiopia. Front. Genet. 14, 1050365 (2023).
Kim, K. et al. Inference of Admixture Origins in Indigenous African Cattle. Mol. Biol. Evol. 40, https://doi.org/10.1093/molbev/msad257 (2023).
Hanotte, O. et al. Geographic distribution and frequency of a taurine Bos taurus and an indicine Bos indicus Y specific allele amongst sub-saharan African cattle breeds. Mol. Ecol. 9, 387–396 (2000).
Dai, S. et al. Global pangenome analysis highlights the critical role of structural variants in cattle improvement and identifies a unique event as a novel enhancer in IGFBP7+ cells. Mol. Biol. Evol. 42, msaf205 (2025).
Coffey, E., Horan, B., Evans, R. & Berry, D. Milk production and fertility performance of Holstein, Friesian, and Jersey purebred cows and their respective crosses in seasonal-calving commercial farms. J. Dairy Sci. 99, 5681–5689 (2016).
Xia, X. et al. Global dispersal and adaptive evolution of domestic cattle: a genomic perspective. Stress Biol. 3, 8 (2023).
Taye, M. et al. Exploring the genomes of East African Indicine cattle breeds reveals signature of selection for tropical environmental adaptation traits. Cogent Food Agric. 4, 1552552 (2018).
Jonsson, N., Piper, E. & Constantinoiu, C. Host resistance in cattle to infestation with the cattle tick R hipicephalus microplus. Parasite Immunol. 36, 553–559 (2014).
Magee, D. A., MacHugh, D. E. & Edwards, C. J. Interrogation of modern and ancient genomes reveals the complex domestic history of cattle. Anim. Front. 4, 7–22 (2014).
Martín-Burriel, I. et al. Genetic diversity, structure, and breed relationships in Iberian cattle. J. Anim. Sci. 89, 893–906 (2011).
Groh, J. S. & Coop, G. The temporal and genomic scale of selection following hybridization. Proc. Natl. Acad. Sci. 121, e2309168121 (2024).
Sankararaman, S. et al. The genomic landscape of Neanderthal ancestry in present-day humans. Nature 507, 354–357 (2014).
Smetko, A. et al. Trypanosomosis: potential driver of selection in African cattle. Front. Genet. 6, 137 (2015).
Hunter, C. A. & Kastelein, R. Interleukin-27: balancing protective and pathological immunity. Immunity 37, 960–969 (2012).
Lamp, O. et al. Metabolic Heat Stress Adaption in Transition Cows: Differences in Macronutrient Oxidation between Late-Gestating and Early-Lactating German Holstein Dairy Cows. PLoS One 10, e0125264 (2015).
Ma, Z., Tanis, J. E., Taruno, A. & Foskett, J. K. Calcium homeostasis modulator (CALHM) ion channels. Pflug. Arch. 468, 395–403 (2016).
Liu, D. et al. Genome-wide selection signatures detection in Shanghai Holstein cattle population identified genes related to adaption, health and reproduction traits. BMC Genomics 22, 747 (2021).
Kusano, T., Nishino, T., Okamoto, K., Hille, R. & Nishino, T. The mechanism and significance of the conversion of xanthine dehydrogenase to xanthine oxidase in mammalian secretory gland cells. Redox Biol. 59, 102573 (2023).
Chen, C. et al. Dermatan sulfate: structure, biosynthesis, and biological roles. Biomolecules 15, https://doi.org/10.3390/biom15081158 (2025).
Larkin, D. M. et al. Whole-genome resequencing of two elite sires for the detection of haplotypes under selection in dairy cattle. Proc. Natl. Acad. Sci. 109, 7693–7698 (2012).
Nayeri, S. et al. Genome-wide association analysis for β-hydroxybutyrate concentration in Milk in Holstein dairy cattle. BMC Genet. 20, 58 (2019).
Nozawa, K. et al. Knockout of serine-rich single-pass membrane protein 1 (Ssmem1) causes globozoospermia and sterility in male mice†. Biol. Reprod. 103, 244–253 (2020).
Mulim, H. A. et al. Detection and evaluation of parameters influencing the identification of heterozygous-enriched regions in Holstein cattle based on SNP chip or whole-genome sequence data. BMC Genomics 25, 726 (2024).
He, W. et al. Function identification of bovine ACSF3 gene and its Association with lipid metabolism traits in beef cattle. Front. Vet. Sci. 8, 766765 (2022).
Dirandeh, E., Ansari-Pirsaraei, Z. & Thatcher, W. Melatonin as a smart protector of pregnancy in dairy cows. Antioxidants. 11, https://doi.org/10.3390/antiox11020292 (2022).
de Camargo, G. M. et al. Polymorphisms in TOX and NCOA2 genes and their associations with reproductive traits in cattle. Reprod. Fertil. Dev. 27, 523–528 (2015).
Wang, T. et al. The biological properties of the FAS and TACR3 genes and the association of single-nucleotide polymorphisms with milk quality traits in Gannan Yak. Foods 14, https://doi.org/10.3390/foods14091575 (2025).
Jiang, L.-Y. et al. Ring finger protein 145 (RNF145) is a ubiquitin ligase for sterol-induced degradation of HMG-CoA reductase. J. Biol. Chem. 293, 4047–4055 (2018).
Rothi, M. H. et al. The 18S rRNA methyltransferase DIMT-1 regulates lifespan in the germline later in life. Nat. Commun. 16, 6944 (2025).
Benfica, L. F. et al. Genome-wide association study between copy number variation and feeding behavior, feed efficiency, and growth traits in Nellore cattle. BMC Genomics 25, 54 (2024).
Singh, A. et al. Genomewide expression analysis of the heat stress response in dermal fibroblasts of Tharparkar (zebu) and Karan-Fries (zebu× taurine) cattle. Cell Stress Chaperones 25, 327–344 (2020).
Tijjani, A. et al. Genomic signatures for drylands adaptation at gene-rich regions in African zebu cattle. Genomics 114, 110423 (2022).
Gujar, G. et al. Characterization of thermo-physiological, hematological, and molecular changes in response to seasonal variations in two tropically adapted native cattle breeds of Bos indicus lineage in hot arid ambience of Thar Desert. Int. J. Biometeorol. 66, 1515–1529 (2022).
Mengistie Yirsaw, D. Genome-Wide Signature of Positive Selection, Breed-Specific SNPs and Linkage Disequilibrium in Ethiopian Indigenous and European Beef Cattle Breeds, Addis Ababa University, (2021).
Liu, Y. et al. Hypoxia-induced GPCPD1 depalmitoylation triggers mitophagy via regulating PRKN-mediated ubiquitination of VDAC1. Autophagy 19, 2443–2463 (2023).
Freihat, L. A. et al. IRAK3 modulates downstream innate immune signalling through its guanylate cyclase activity. Sci. Rep. 9, 15468 (2019).
Flori, L. et al. A genomic map of climate adaptation in Mediterranean cattle breeds. Mol. Ecol. 28, 1009–1029 (2019).
Shinoda, K. et al. The dystonia gene THAP1 controls DNA double-strand break repair choice. Mol. Cell 81, 2611–2624.e2610 (2021).
Bressler, K. R. et al. Depletion of eukaryotic initiation factor 5B (eIF5B) reprograms the cellular transcriptome and leads to activation of endoplasmic reticulum (ER) stress and c-Jun N-terminal kinase (JNK). Cell Stress Chaperones 26, 253–264 (2021).
Ruthig, V. A. et al. The RNA-binding protein DND1 acts sequentially as a negative regulator of pluripotency and a positive regulator of epigenetic modifiers required for germ cell reprogramming. Development 146, dev175950 (2019).
Purfield, D. C., Bradley, D. G., Evans, R. D., Kearney, F. J. & Berry, D. P. Genome-wide association study for calving performance using high-density genotypes in dairy and beef cattle. Genet. Sel. Evol. 47, 47 (2015).
Guzmán, L. F. et al. Expression of heat shock protein genes in Simmental cattle exposed to heat stress. Anim. Biosci. 36, 704 (2023).
Kim, J. et al. Expansion of the HSP70 gene family in Tegillarca granosa and expression profiles in response to zinc toxicity. Cell Stress Chaperones 29, 97–112 (2024).
Chen, Q. et al. A brown fat-enriched adipokine Adissp controls adipose thermogenesis and glucose homeostasis. Nat. Commun. 13, 7633 (2022).
Bo, D. et al. Whole-genome resequencing reveals genetic diversity and growth trait-related genes in Pinan cattle. Animals 14, 2163 (2024).
Asai, Y. et al. Transgenic Tmc2 expression preserves inner ear hair cells and vestibular function in mice lacking Tmc1. Sci. Rep. 8, 12124 (2018).
Steiner, P. & Zierler, S. Inter-Organellar Ca(2+) Homeostasis in plant and animal systems. Cells 14, https://doi.org/10.3390/cells14151204 (2025).
Almhanna, H. et al. Comparison of Siglec-1 protein networks and expression patterns in sperm and male reproductive tracts of mice, rats, and humans. Vet. World 17, 645–657 (2024).
Spetter, M. J. et al. Genetic diversity, Admixture, and selection signatures in a Rarámuri Criollo cattle population introduced to the Southwestern United States. Int. J. Mol. Sci. 26, 4649 (2025).
Almhanna, H. et al. Comparison of Siglec-1 protein networks and expression patterns in sperm and male reproductive tracts of mice, rats, and humans. Vet. World 17, 645 (2024).
Jones, J. M. & First, N. L. Expression of the cell cycle control protein cdc25 in cleavage stage bovine embryos. Zygote 3, 133–139 (1995).
Tijjani, A. et al. Genomic reference resource for African cattle: genome sequences and high-density array variants. Sci. Data 11, 801 (2024).
Chen, S. Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp. Imeta 2, e107 (2023).
Vasimuddin, M., Misra, S., Li, H. & Aluru, S. in 2019 IEEE International Parallel And Distributed Processing Symposium (IPDPS). 314–324 (IEEE).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Okonechnikov, K., Conesa, A. & García-Alcalde, F. Qualimap 2: advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics 32, 292–294 (2016).
Garrison, E. & Marth, G. Haplotype-based variant detection from short-read sequencing. arXiv preprint arXiv:1207.3907 (2012).
Browning, B. L., Zhou, Y. & Browning, S. R. A one-penny imputed genome from next-generation reference panels. Am. J. Hum. Genet. 103, 338–348 (2018).
Browning, B. L., Tian, X., Zhou, Y. & Browning, S. R. Fast two-stage phasing of large-scale sequence data. Am. J. Hum. Genet. 108, 1880–1890 (2021).
Lindenbaum, P. JVarkit: Java-based utilities for Bioinformatics. figshare 10, m9 (2015).
Chen, C. et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 13, 1194–1202 (2020).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Ayalew, W. et al. Whole genome scan uncovers candidate genes related to milk production traits in Barka Cattle. Int. J. Mol. Sci. 25, 6142 (2024).
Gómez-Rubio, V. ggplot2-elegant graphics for data analysis. J. stat. softw. 77, 1–3 (2017).
Paradis, E. & Schliep, K. ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucl. Acids Res., gkae268 (2024).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Qanbari, S. & Wittenburg, D. Male recombination map of the autosomal genome in German Holstein. Genet. Sel. Evol. 52, 73 (2020).
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012).
Peter, B. M. Admixture, population structure, and F-statistics. Genetics 202, 1485–1501 (2016).
Chintalapati, M., Patterson, N. & Moorjani, P. The spatiotemporal patterns of major human admixture events during the European Holocene. Elife 11, https://doi.org/10.7554/eLife.77625 (2022).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. fly 6, 80–92 (2012).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Holsinger, K. E. & Weir, B. S. Genetics in geographically structured populations: defining, estimating and interpreting F ST. Nat. Rev. Genet. 10, 639–650 (2009).
Durrett, R. & Durrett, R. Probability models for DNA sequence evolution. 2 (Springer, 2008).
Nei, M. & Li, W.-H. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl. Acad. Sci. 76, 5269–5273 (1979).
Pan, B. et al. TMC1 and TMC2 are components of the mechanotransduction channel in hair cells of the mammalian inner ear. Neuron 79, 504–515 (2013).
Weir, B. S. & Cockerham, C. C. Estimating F-statistics for the analysis of population structure. evolution, 1358–1370 (1984).
Borowsky, R. L. Estimating nucleotide diversity from random amplified polymorphic DNA and amplified fragment length polymorphism data. Mol. Phylogenet. Evol. 18, 143–148 (2001).
McLaren, W. et al. The Ensembl Variant Effect Predictor. Genome Biol. 17, 1–14 (2016).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Meng, E. C. et al. UCSF ChimeraX: Tools for structure building and analysis. Protein Sci. 32, e4792 (2023).
Gao, J. Ancestral retention signals cattle [Workflow]. Zenodo https://doi.org/10.5281/zenodo.18378574 (2026).
EVA European Variation Archive. https://identifiers.org/ena.embl:PRJEB102975 (2025).
ENA European Nucleotide Archive. https://identifiers.org/ena.embl:PRJEB90914 (2025).
ENA European Nucleotide Archive. https://identifiers.org/ena.embl:PRJEB90816 (2026).
ENA European Nucleotide Archive. https://identifiers.org/ena.embl:PRJNA391427 (2017).
ENA European Nucleotide Archive. https://identifiers.org/ena.embl:PRJEB74565 (2024).
Hamada, M. et al. Prognostic association of starvation-induced genes in head and neck cancer. (2021).
Kumar, A. et al. 2-Deoxyglucose drives plasticity via an adaptive ER stress-ATF4 pathway and elicits stroke recovery and Alzheimer’s resilience. Neuron 111, 2831–2846. e2810 (2023).
Oyadomari, S. & Mori, M. Roles of CHOP/GADD153 in endoplasmic reticulum stress. Cell Death Differ. 11, 381–389 (2004).
Baral, K. & Rotwein, P. ZMAT2 in Humans and Other Primates: A Highly Conserved and Understudied Gene. Evol. Bioinform. Online 16, 1176934320941500 (2020).
Warmack, R. A. et al. Human Protein-l-isoaspartate O-Methyltransferase Domain-Containing Protein 1 (PCMTD1) Associates with Cullin-RING Ligase Proteins. Biochemistry 61, 879–894 (2022).
Sdiri, C., Ben Souf, I., Ben Salem, I., M’Hamdi, N. & Ben Hamouda, M. Assessment of Genetic and Health Management of Tunisian Holstein Dairy Herds with a Focus on Longevity. Genes. 14, https://doi.org/10.3390/genes14030670 (2023).
Acknowledgements
This study was supported by the Long-term EU-Africa Research and Innovation Partnership on Food and Nutrition Security and Sustainable Agriculture (LEAP-Agri) as part of the OPTIBOV project (LEAP-Agri-326), and by the European Union’s Horizon 2020 Research and Innovation Program (grant agreement No. 727715). Additional funding was provided by Fundação Nacional para a Ciência e a Tecnologia (FCT), Portugal (Leap Agri-326/LEAPAgri/0003/2017 and 2020.02754.CEECIND, C.G.); the Research Council of Finland (319987); the Netherlands Organization for Scientific Research (NWO-WOTRO, 2018/WOTRO/00488849); the Science, Technology & Innovation Funding Authority of Egypt (STDF, LEAP-Agri 326); the Ministry of Science, Technology and Innovations of Uganda (MoSTI/LEAP-11); the National Research Foundation of South Africa (NRF, 115577); and the China Scholarship Council (CSC, 202208610017). The funding bodies had no role in study design, data collection, analysis, interpretation, or manuscript writing.
We thank all collaborators for their assistance with sample collection, laboratory work, and technical support, including Bert Dibbits, Kimberley Laport, Rania Agamy, Mohamed Hamada Elsawy, Filipe Ribeiro, Ricardo Loureiro, Daniel Gaspar, Ludmilla Blaschikoff, Ana Elisabete Pires, Carolina Bruno-de-Sousa, Heli Lindberg, Tiina Reilas, Dr. Avhashoni Zwane, Khanyisani Nxumalo, Maano Malima, and all breeders and breed associations involved. We also acknowledge the 1000 Bull Genomes consortium and the African Genomic Reference Resource for providing sequence data.
Author information
Authors and Affiliations
Contributions
J.G. and R.C. conceived the study. J.G. drafted the manuscript and interpreted the results. C.G. defined reference data sets. J.G. and Y.L. participated in data analysis. C.G., J.K., N.G., D.K., M.M., R.O., and R.C. collected the samples. H.B., M.G., and R.C. supervised the study. All the authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Alana Alexander and the other anonymous reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: George Inglis. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gao, J., Ginja, C., Liu, Y. et al. Distinct adaptation and ancestral retention signals in African and European indigenous cattle genomes. Commun Biol (2026). https://doi.org/10.1038/s42003-026-09856-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-026-09856-9