Abstract
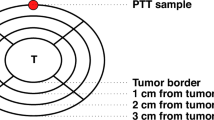
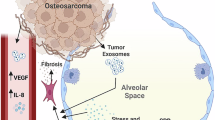
The non-tumor tissue adjacent to metastases can appear morphologically unremarkable under a microscope; however, it is exposed to a milieu of secretory factors and proteins derived from tumor cells, stromal cells, and immune cells within the surrounding tumor microenvironment. Studies investigating the peritumoral tissue (PTT) or so-called Normal tissue Adjacent to Tumor tissue (NAT) have identified distinct differences between the genomic and transcriptomic profiles of healthy and tumor-adjacent non-tumor tissues. These alterations are hypothesized to have significant implications in local tumor progression, metastasis, and patient outcome. Most NAT/PTT studies focus on the primary tumor microenvironment (TME) with comparisons between patients with and without cancer. The study described herein expands upon this work by investigating the metastatic TME with comparisons between met-recipient and met-free tissues, both derived from a canine osteosarcoma clinical trial. Our study identifies shared and tissue-specific changes in met-recipient non-tumor lung, liver, and kidney which overlap with transcriptional alterations described in human cancers. These findings improve our understanding of the landscape of the peritumoral TME of metastatic osteosarcoma and further underscore the translational relevance of the canine patient as a model of human disease.
Similar content being viewed by others
Data availability
The canine clinical trial datasets generated and/or analyzed during the current study are available in the GEO database (RRID:SCR_005012; accession ID: GSE309676). Source data is available in Supplementary Data 2.
Code availability
Codes are deposited at GitHub (https://github.com/Anjaligarg006/NAT.git).
References
Beird, H. C. et al. Osteosarcoma. Nat. Rev. Dis. Prim. 8, 77 (2022).
Aljubran, A. H., Griffin, A., Pintilie, M. & Blackstein, M. Osteosarcoma in adolescents and adults: survival analysis with and without lung metastases. Ann. Oncol. 20, 1136–1141 (2009).
Liu, Z. L., Chen, H. H., Zheng, L. L., Sun, L. P. & Shi, L. Angiogenic signaling pathways and anti-angiogenic therapy for cancer. Signal Transduct. Target Ther. 8, 198 (2023).
Yang, D., Liu, J., Qian, H. & Zhuang, Q. Cancer-associated fibroblasts: from basic science to anticancer therapy. Exp. Mol. Med. 55, 1322–1332 (2023).
Gaikwad, S., Agrawal, M. Y., Kaushik, I., Ramachandran, S. & Srivastava, S. K. Immune checkpoint proteins: signaling mechanisms and molecular interactions in cancer immunotherapy. Semin Cancer Biol. 86, 137–150 (2022).
Cappell, K. M. & Kochenderfer, J. N. Long-term outcomes following CAR T cell therapy: what we know so far. Nat. Rev. Clin. Oncol. 20, 359–371 (2023).
Beck, J. et al. Canine and Murine models of osteosarcoma. Vet. Pathol. 59, 399–414 (2022).
LeBlanc, A. K. & Mazcko, C. N. Improving human cancer therapy through the evaluation of pet dogs. Nat. Rev. Cancer 20, 727–742 (2020).
Silver, K. I. et al. Patterns of metastatic progression and association with clinical outcomes in canine osteosarcoma: A necropsy study of 83 dogs. Vet. Comp. Oncol. 21, 646–655 (2023).
Chi, S. N. et al. The patterns of relapse in osteosarcoma: the Memorial Sloan-Kettering experience. Pediatr. Blood Cancer 42, 46–51 (2004).
Jeffree, G. M., Price, C. H. & Sissons, H. A. The metastatic patterns of osteosarcoma. Br. J. Cancer 32, 87–107 (1975).
Koca, D., Abedi-Ardekani, B., LeMaoult, J. & Guyon, L. Peritumoral tissue (PTT): increasing need for naming convention. Br. J. Cancer 131, 1111–1115 (2024).
Aran, D. et al. Comprehensive analysis of normal adjacent to tumor transcriptomes. Nat. Commun. 8, 1077 (2017).
Heaphy, C. M., Griffith, J. K. & Bisoffi, M. Mammary field cancerization: molecular evidence and clinical importance. Breast Cancer Res Treat. 118, 229–239 (2009).
Morla-Barcelo, P. M. et al. Unraveling malignant phenotype of peritumoral tissue: transcriptomic insights into early-stage breast cancer. Breast Cancer Res. 26, 89 (2024).
Troester, M. A. et al. Activation of host wound responses in breast cancer microenvironment. Clin. Cancer Res. 15, 7020–7028 (2009).
Kim, J. et al. Transcriptomes of the tumor-adjacent normal tissues are more informative than tumors in predicting recurrence in colorectal cancer patients. J. Transl. Med. 21, 209 (2023).
Hou, Y. et al. Integrated analysis of tumor and adjacent non-tumor proteomic data reveals SERPINH1 as a recurrence biomarker and drug target in hepatocellular carcinoma. Int. J. Biol. Sci. 20, 5191–5207 (2024).
Michot, A. et al. Analysis of the peritumoral tissue unveils cellular changes associated with a high risk of recurrence. Cancers (Basel) 15, 3450 (2023).
Huang, X., Stern, D. F. & Zhao, H. Transcriptional profiles from paired normal samples offer complementary information on cancer patient survival-evidence from TCGA pan-cancer data. Sci. Rep. 6, 20567 (2016).
LeBlanc, A. K. et al. Adjuvant sirolimus does not improve outcome in pet dogs receiving standard-of-care therapy for appendicular osteosarcoma: a prospective, randomized trial of 324 dogs. Clin. Cancer Res. 27, 3005–3016 (2021).
Hassan, S., Syed, S. & Kehar, S. I. Glial fibrillary acidic protein (GFAP) as a mesenchymal marker of early hepatic stellate cells activation in liver fibrosis in chronic hepatitis C infection. Pak. J. Med Sci. 30, 1027–1032 (2014).
Mulugeta, S. & Beers, M. F. Surfactant protein C: its unique properties and emerging immunomodulatory role in the lung. Microbes Infect. 8, 2317–2323 (2006).
Patkar, S. et al. Large-scale comparative analysis of canine and human osteosarcomas uncovers conserved clinically relevant tumor microenvironment subtypes. Clin. Cancer Res. 30, 5630–5642 (2024).
Eigenbrood, J. et al. Spatial profiling identifies regionally distinct microenvironments and targetable immunosuppressive mechanisms in pediatric osteosarcoma pulmonary metastases. Cancer Res. 85, 2320–2337 (2025).
Ammons, D. T. et al. Single-cell RNA sequencing reveals the cellular and molecular heterogeneity of treatment-naive primary osteosarcoma in dogs. Commun. Biol. 7, 496 (2024).
Kang, M.-J. et al. Inhibition of hepatic stellate cell activation suppresses tumorigenicity of hepatocellular carcinoma in mice. Am. J. Pathol. 191, 2219–2230 (2021).
Kang, N., Gores, G. J. & Shah, V. H. Hepatic stellate cells: partners in crime for liver metastases? Hepatology 54, 707–713 (2011).
Cogliati, B., Yashaswini, C. N., Wang, S., Sia, D. & Friedman, S. L. Friend or foe? The elusive role of hepatic stellate cells in liver cancer. Nat. Rev. Gastroenterol. Hepatol. 20, 647–661 (2023).
Kiefer, A. et al. DMBT1 is upregulated in cystic fibrosis, affects ciliary motility, and is reduced by acetylcysteine. Mol. Cell. Pediatr. 9, 4 (2022).
Ruaro, B. et al. The history and mystery of alveolar epithelial type II cells: focus on their physiologic and pathologic role in lung. Int. J. Mol. Sci. 22, 2566 (2021).
McGee, L. E. et al. The tumor microenvironment of metastatic osteosarcoma in the human and canine lung. Commun. Biol. 8, 756 (2025).
Reinecke, J. B. et al. Aberrant activation of wound-healing programs within the metastatic niche facilitates lung colonization by osteosarcoma cells. Clin. Cancer Res. 31, 414–429 (2025).
Regan, D. P. et al. The angiotensin receptor blocker losartan suppresses growth of pulmonary metastases via AT1R-independent inhibition of CCR2 signaling and monocyte recruitment. J. Immunol. 202, 3087–3102 (2019).
Regan, D. P. et al. Losartan blocks osteosarcoma-elicited monocyte recruitment, and combined with the kinase inhibitor toceranib, exerts significant clinical benefit in canine metastatic osteosarcoma. Clin. Cancer Res. 28, 662–676 (2022).
Klosowski, M. et al. Naturally occurring canine sarcomas: bridging the gap from mouse models to human patients through cross-disciplinary research partnerships. Front Oncol. 13, 1130215 (2023).
Wightman, S. C. et al. Oncogenic CXCL10 signalling drives metastasis development and poor clinical outcome. Br. J. Cancer 113, 327–335 (2015).
Liu, M., Guo, S. & Stiles, J. K. The emerging role of CXCL10 in cancer (Review). Oncol. Lett. 2, 583–589 (2011).
Flores, R. J. et al. A novel prognostic model for osteosarcoma using circulating CXCL10 and FLT3LG. Cancer 123, 144–154 (2017).
Tokunaga, R. et al. CXCL9, CXCL10, CXCL11/CXCR3 axis for immune activation - A target for novel cancer therapy. Cancer Treat. Rev. 63, 40–47 (2018).
Ohmori, Y. et al. Tumor necrosis factor-alpha induces cell type and tissue-specific expression of chemoattractant cytokines in vivo. Am. J. Pathol. 142, 861–870 (1993).
Ohmori, Y., Schreiber, R. D. & Hamilton, T. A. Synergy between interferon-gamma and tumor necrosis factor-alpha in transcriptional activation is mediated by cooperation between signal transducer and activator of transcription 1 and nuclear factor kappaB. J. Biol. Chem. 272, 14899–14907 (1997).
Gordon, I., Paoloni, M., Mazcko, C. & Khanna, C. The comparative oncology trials consortium: using spontaneously occurring cancers in dogs to inform the cancer drug development pathway. PLoS Med. 6, e1000161 (2009).
Paoloni, M., Lana, S., Thamm, D., Mazcko, C. & Withrow, S. The creation of the comparative oncology trials consortium pharmacodynamic core: infrastructure for a virtual laboratory. Vet. J. 185, 88–89 (2010).
Talhouk, A. et al. Single-patient molecular testing with nanostring ncounter data using a reference-based strategy for batch effect correction. PLoS One 11, e0153844 (2016).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 26, 139–140 (2010).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innov. (Camb.) 2, 100141 (2021).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 16, 284–287 (2012).
Becht, E. et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 17, 218 (2016).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Acknowledgements
We would like to thank the NCI-COTC member institutions, canine clinical trials teams, and the families who enrolled their dogs into COTC-021/022. We would also like to acknowledge the Molecular Histopathology Laboratory for their assistance with FFPE RNA isolation, and the Genomics Technology Laboratory for their assistance with RNA isolation and the NanoString IO Panel. This research was supported by the Intramural Research Program (Z01-BC006161) of the National Institutes of Health (NIH). The contributions of the NIH author(s) were made as part of their official duties as NIH federal employees, are in compliance with agency policy requirements, and are considered Works of the United States Government. However, the findings and conclusions presented in this paper are those of the author(s) and do not necessarily reflect the views of the NIH or the U.S. Department of Health and Human Services. Figure 1 (https://biorender.com/w5r5qeg), Fig. 2D (https://biorender.com/t9y5khn), and Fig. 5 (https://biorender.com/r7ef8fg) were created in BioRender.
Author information
Authors and Affiliations
Contributions
J.A.B.: conceptualization, methodology, project administration, data curation, investigation, visualization, writing—original draft; A.G.: formal analysis, visualization, writing—review & editing; P.C.: resources, supervision, writing—review & editing; C.M.: data curation, writing—review & editing; A.L.: supervision, project administration, writing—review & editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Bruce Smith, Berkley Gryder and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Christina Karlsson Rosenthal.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Beck, J.A., Garg, A., Choyke, P.L. et al. Comprehensive analysis of non-tumor lung, liver, and kidney transcriptomes in canine metastatic osteosarcoma. Commun Biol (2026). https://doi.org/10.1038/s42003-026-09870-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-026-09870-x