Abstract
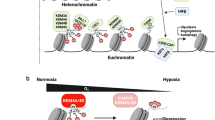
Adaptation to reduced levels of oxygen (hypoxia) is an essential feature of eukaryotic life. Within the animal kingdom, cellular responses are orchestrated by the transcription factor HIF (Hypoxia Inducible Factor) which is regulated by specific 2-oxoglutarate-dependent oxygenases. This family of enzymes also includes histone demethylases, and histone methylation has also been observed to increase in hypoxia. Since histone methylation is associated with gene expression, this has raised questions about whether this also contributes to transcriptional regulation in hypoxia. However, to date pangenomic studies have not been normalised in a way that preserves these bulk changes. Using drosophila chromatin spike-in normalisation, we have shown widespread increases in histone H3K4/9/27/36me3 in hypoxia at almost all gene loci that occur irrespective of whether gene expression is increased or reduced. However, methylation of H3K4me3 and H3K36me3 increases most at direct transcriptional targets of HIF and this is abrogated by inactivation of HIF. Taken together this suggests that global H3 trimethylation increases in hypoxia are widespread and not sufficient to predict transcriptional direction, whereas enhanced H3K4me3/H3K36me3 at direct HIF targets appears consequent to HIF binding and transcriptional engagement.
Similar content being viewed by others
Data availability
ChIP-seq and RNA-seq data are available at Gene Expression Omnibus (GEO): GSE296192, GSE313247, GSE200203 (HIF-1α and HIF-2α ChIP-seq), and GSE130989 (HIF-1β ChIP-seq). Original Western blot images are available in Supplementary Fig. 9 and source data are available in Supplementary Data. This paper does not report original code. Any additional information required to reanalyse the data reported in this paper is available from the lead contact upon request.
References
Li, L. et al. Searching for molecular hypoxia sensors among oxygen-dependent enzymes. eLife 12, https://doi.org/10.7554/eLife.87705 (2023).
Kindrick, J. D. & Mole, D. R. Hypoxic Regulation of Gene Transcription and Chromatin: Cause and Effect. International journal of molecular sciences 21, https://doi.org/10.3390/ijms21218320 (2020).
Hong, S. et al. Identification of JmjC domain-containing UTX and JMJD3 as histone H3 lysine 27 demethylases. In Proc. National Academy of Sciences of the United States of America, Vol. 104, 18439–18444 (2007).
Hillringhaus, L. et al. Structural and evolutionary basis for the dual substrate selectivity of human KDM4 histone demethylase family. The Journal of biological chemistry 286, 41616–41625 (2011).
Xia, X. et al. Integrative analysis of HIF binding and transactivation reveals its role in maintaining histone methylation homeostasis. Proceedings of the National Academy of Sciences of the United States of America 106, 4260–4265 (2009).
Batie, M. et al. Hypoxia induces rapid changes to histone methylation and reprograms chromatin. Science 363, 1222–1226 (2019).
Johnson, A. B., Denko, N. & Barton, M. C. Hypoxia induces a novel signature of chromatin modifications and global repression of transcription. Mutat. Res. 640, 174–179 (2008).
Lee, S. et al. Multi-dimensional histone methylations for coordinated regulation of gene expression under hypoxia. Nucleic acids research 45, 11643–11657 (2017).
Prickaerts, P. et al. Hypoxia increases genome-wide bivalent epigenetic marking by specific gain of H3K27me3. Epigenetics & chromatin 9, 46 (2016).
Wu, M. Z. et al. Interplay between HDAC3 and WDR5 is essential for hypoxia-induced epithelial-mesenchymal transition. Molecular cell 43, 811–822 (2011).
Zhou, X. et al. Hypoxia induces trimethylated H3 lysine 4 by inhibition of JARID1A demethylase. Cancer research 70, 4214–4221 (2010).
Hancock, R. L., Masson, N., Dunne, K., Flashman, E. & Kawamura, A. The Activity of JmjC Histone Lysine Demethylase KDM4A is Highly Sensitive to Oxygen Concentrations. ACS Chem Biol 12, 1011–1019 (2017).
Dobrynin, G. et al. KDM4A regulates HIF-1 levels through H3K9me3. Scientific reports 7, 11094 (2017).
Tausendschon, M., Dehne, N. & Brune, B. Hypoxia causes epigenetic gene regulation in macrophages by attenuating Jumonji histone demethylase activity. Cytokine 53, 256–262 (2011).
Costa, M. et al. Nickel carcinogenesis: epigenetics and hypoxia signaling. Mutat. Res. 592, 79–88 (2005).
Chen, H., Yan, Y., Davidson, T. L., Shinkai, Y. & Costa, M. Hypoxic stress induces dimethylated histone H3 lysine 9 through histone methyltransferase G9a in mammalian cells. Cancer research 66, 9009–9016 (2006).
Olcina, M. M. et al. Replication stress and chromatin context link ATM activation to a role in DNA replication. Molecular cell 52, 758–766 (2013).
Osumek, J. E., Revesz, A., Morton, J. S., Davidge, S. T. & Hardy, D. B. Enhanced trimethylation of histone h3 mediates impaired expression of hepatic glucose 6-phosphatase expression in offspring from rat dams exposed to hypoxia during pregnancy. Reprod Sci 21, 112–121 (2014).
Olcina, M. M. et al. H3K9me3 facilitates hypoxia-induced p53-dependent apoptosis through repression of APAK. Oncogene 35, 793–799 (2016).
Chakraborty, A. A. et al. Histone demethylase KDM6A directly senses oxygen to control chromatin and cell fate. Science 363, 1217–1222 (2019).
Li, Y. et al. Acetate supplementation restores chromatin accessibility and promotes tumor cell differentiation under hypoxia. Cell Death Dis 11, 102 (2020).
Platt, J. L. et al. Capture-C reveals preformed chromatin interactions between HIF-binding sites and distant promoters. EMBO Rep 17, 1410–1421 (2016).
Choudhry, H. et al. Extensive regulation of the non-coding transcriptome by hypoxia: role of HIF in releasing paused RNApol2. EMBO Rep 15, 70–76 (2014).
Adriaens, M. E. et al. Quantitative analysis of ChIP-seq data uncovers dynamic and sustained H3K4me3 and H3K27me3 modulation in cancer cells under hypoxia. Epigenetics & chromatin 9, 48 (2016).
Orlando, D. A. et al. Quantitative ChIP-Seq normalization reveals global modulation of the epigenome. Cell reports 9, 1163–1170 (2014).
Egan, B. et al. An Alternative Approach to ChIP-Seq Normalization Enables Detection of Genome-Wide Changes in Histone H3 Lysine 27 Trimethylation upon EZH2 Inhibition. PLoS ONE 11, e0166438 (2016).
Bonhoure, N. et al. Quantifying ChIP-seq data: a spiking method providing an internal reference for sample-to-sample normalization. Genome research 24, 1157–1168 (2014).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Kirmes, I. et al. A transient ischemic environment induces reversible compaction of chromatin. Genome biology 16, 246 (2015).
Lombardi, O. et al. Pan-cancer analysis of tissue and single-cell HIF-pathway activation using a conserved gene signature. Cell reports 41, 111652 (2022).
Schmid, V. et al. Co-incidence of RCC-susceptibility polymorphisms with HIF cis-acting sequences supports a pathway tuning model of cancer. Sci Rep 9, 18768 (2019).
Perez, M. F. & Sarkies, P. Histone methyltransferase activity affects metabolism in human cells independently of transcriptional regulation. PLoS biology 21, e3002354 (2023).
Clouaire, T. et al. Cfp1 integrates both CpG content and gene activity for accurate H3K4me3 deposition in embryonic stem cells. Genes & development 26, 1714–1728 (2012).
Taylor, B. C. & Young, N. L. Combinations of histone post-translational modifications. The Biochemical journal 478, 511–532 (2021).
Su, Z. & Denu, J. M. Reading the Combinatorial Histone Language. ACS Chem Biol 11, 564–574 (2016).
Ye, C. & Tu, B. P. Sink into the Epigenome: Histones as Repositories That Influence Cellular Metabolism. Trends in endocrinology and metabolism: TEM 29, 626–637 (2018).
Luco, R. F. et al. Regulation of alternative splicing by histone modifications. Science 327, 996–1000 (2010).
Mushtaq, A. et al. Role of Histone Methylation in Maintenance of Genome Integrity. Genes (Basel) 12, https://doi.org/10.3390/genes12071000 (2021).
Greer, E. L. & Shi, Y. Histone methylation: a dynamic mark in health, disease and inheritance. Nature reviews. Genetics 13, 343–357 (2012).
Farina, A. R. et al. Hypoxia-induced alternative splicing: the 11th Hallmark of Cancer. Journal of experimental & clinical cancer research: CR 39, 110 (2020).
Bristow, R. G. & Hill, R. P. Hypoxia and metabolism. Hypoxia, DNA repair and genetic instability. Nature reviews 8, 180–192 (2008).
Ward, M. C., Banovich, N. E., Sarkar, A., Stephens, M. & Gilad, Y. Dynamic effects of genetic variation on gene expression revealed following hypoxic stress in cardiomyocytes. eLife 10, https://doi.org/10.7554/eLife.57345 (2021).
Xin, J. et al. Chromatin accessibility landscape and regulatory network of high-altitude hypoxia adaptation. Nature communications 11, 4928 (2020).
Wang, J., Wang, Y., Duan, Z. & Hu, W. Hypoxia-induced alterations of transcriptome and chromatin accessibility in HL-1 cells. IUBMB life 72, 1737–1746 (2020).
Batie, M., Frost, J., Shakir, D. & Rocha, S. Regulation of chromatin accessibility by hypoxia and HIF. The Biochemical journal 479, 767–786 (2022).
Ortmann, B. M. et al. The HIF complex recruits the histone methyltransferase SET1B to activate specific hypoxia-inducible genes. Nature genetics 53, 1022–1035 (2021).
Wysocka, J. et al. A PHD finger of NURF couples histone H3 lysine 4 trimethylation with chromatin remodelling. Nature 442, 86–90 (2006).
Edmunds, J. W., Mahadevan, L. C. & Clayton, A. L. Dynamic histone H3 methylation during gene induction: HYPB/Setd2 mediates all H3K36 trimethylation. The EMBO journal 27, 406–420 (2008).
Le Martelot, G. et al. Genome-wide RNA polymerase II profiles and RNA accumulation reveal kinetics of transcription and associated epigenetic changes during diurnal cycles. PLoS biology 10, e1001442 (2012).
Consortium, T. E. ENCODE Guidelines for Experiments Generating ChIP-seq Data January 2017. https://www.encodeproject.org/documents/ceb172ef-7474-4cd6-bfd2-5e8e6e38592e/@@download/attachment/ChIP-seq_ENCODE3_v3.0.pdf (2017).
Consortium, T. E. ENCODE Guidelines and Best Practices for RNA-Seq: Revised December 2016. https://www.encodeproject.org/documents/cede0cbe-d324-4ce7-ace4-f0c3eddf5972/@@download/attachment/ENCODE%20Best%20Practices%20for%20RNA_v2.pdf (2016).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Dale, R. K., Pedersen, B. S. & Quinlan, A. R. Pybedtools: a flexible Python library for manipulating genomic datasets and annotations. Bioinformatics 27, 3423–3424 (2011).
Sivakumar, S. et al. Pan cancer patterns of allelic imbalance from chromosomal alterations in 33 tumor types. Genetics 217, 1–12 (2021).
Shen, L., Shao, N., Liu, X. & Nestler, E. ngs.plot: Quick mining and visualization of next-generation sequencing data by integrating genomic databases. BMC Genomics 15, 284 (2014).
Hower, V., Evans, S. N. & Pachter, L. Shape-based peak identification for ChIP-Seq. BMC Bioinformatics 12, 15 (2011).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol 9, R137 (2008).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102, 15545–15550 (2005).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol 37, 907–915 (2019).
Anders, S., Pyl, P. T. & Huber, W. HTSeq-a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15, 550 (2014).
Robinson, J. T. et al. Integrative genomics viewer. Nat Biotechnol 29, 24–26 (2011).
Acknowledgements
This study was supported by the National Institute for Health Research (DRM, OL, SH & VL; NIHR-RP-2016-06-004), King Abdulaziz University, Ministry of High Education for Saudi Arabia (DRM & PJR) the NIH-Oxford-Cambridge Scholars Program (JDK), the Ludwig Institute for Cancer Research (PJR), the Wellcome Trust (PJR; 106241/Z/14/Z) and the Francis Crick Institute (PJR), which receives its core funding from Cancer Research UK (FC001501, the UK Medical Research Council (FC001501), and the Wellcome Trust (FC001501) and the Intramural Research Program, National Institutes of Health, Center for Cancer Research, National Cancer Institute, Center for Cancer Research (CHC & WDF; ZIA BC 010453). The computational aspects of this research were supported by the Wellcome Trust Core Award Grant Number 203141/Z/16/Z and the NIHR Oxford BRC. We thank Chris Pugh for scientific discussions. The views expressed are those of the author(s) and not necessarily those of the NHS, the NIHR or the Department of Health.
Author information
Authors and Affiliations
Contributions
Author contributions were as follows: conception and design of the work - JDK, CHC, WDF, PJR & DRM; acquisition, analysis, and interpretation of data – JDK, OL, SH, VNL & DRM, drafting and revising the work – JDK & DRM.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing interests: P. J. Ratcliffe reports grants from Ludwig Institute for Cancer Research and Francis Crick Institute during the conduct of the study and personal fees from Immunocore plc outside the submitted work. No disclosures were reported by the other authors.
Peer review
Peer review information
Communications Biology thanks Sonia Rocha, Sanjeev Shukla and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Eirini Trompouki and Mengtan Xing. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kindrick, J.D., Lombardi, O., Halim, S. et al. Hypoxic regulation of chromatin and gene transcription. Commun Biol (2026). https://doi.org/10.1038/s42003-026-09875-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-026-09875-6