Abstract
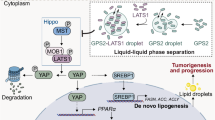
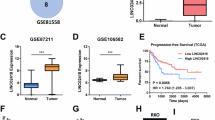
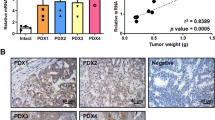
RNA-binding proteins play critical roles in RNA processing and are aberrantly expressed in colorectal cancer (CRC). Through comprehensive analysis of multiple gene expression datasets (GSE20916, GSE18105, GSE21510, TCGA-COAD and TCGA-READ), NCBP2 was identified as a potential tumorigenic gene in CRC. NCBP2 expression was significantly elevated in CRC tissues, correlated with tumour invasion and metastasis, and associated with poor patient survival outcomes. Furthermore, overexpression of NCBP2 in CRC cells was shown to increase cell proliferation, migration, and tumour invasion in both in vitro and in vivo models. Mechanistically, the NCBP2 protein stabilised LIPG mRNA via direct binding to the m7G motif in the 5’-cap structure of LIPG mRNA, thereby increasing LIPG expression. Additionally, NCBP2 promoted lipid droplet accumulation in CRC cells in a LIPG-dependent manner. These findings collectively suggest that the NCBP2-LIPG-lipid droplet axis represents a novel mechanism underlying CRC progression and metastasis, providing a promising therapeutic target for CRC treatment.
Similar content being viewed by others
Data availability
All uncropped blots are provided in Supplementary information. The source data behind the graphs in the manuscript can be found in Supplementary Data 1. Data related to mRNA sequence analysis have been deposited in the GEO, and the accession number is GSE270402. All other data supporting the findings of this study are available from the corresponding author upon reasonable request.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74, 229–263 (2024).
Bingfeng Han, R. Z. et al. Cancer incidence and mortality in China, 2022. J. Natl. Cancer Cent. 4, 47–53 (2024).
Group, C. C. Colorectal Endoscopic Stenting Trial (CReST) for obstructing left-sided colorectal cancer: randomized clinical trial. Br. J. Surg. 109, 1073–1080 (2022).
van Hooft, J. E. et al. Colonic stenting versus emergency surgery for acute left-sided malignant colonic obstruction: a multicentre randomised trial. Lancet Oncol. 12, 344–352 (2011).
Talboom, K. et al. Ferric carboxymaltose infusion versus oral iron supplementation for preoperative iron deficiency anaemia in patients with colorectal cancer (FIT): a multicentre, open-label, randomised, controlled trial. Lancet Haematol. 10, e250–e260 (2023).
Chandra, R. et al. The colorectal cancer tumor microenvironment and its impact on liver and lung metastasis. Cancers 13, 6206 (2021).
Shin, A. E., Giancotti, F. G. & Rustgi, A. K. Metastatic colorectal cancer: mechanisms and emerging therapeutics. Trends Pharmacol. Sci. 44, 222–236 (2023).
Rumpold, H. et al. Prediction of mortality in metastatic colorectal cancer in a real-life population: a multicenter explorative analysis. BMC Cancer 20, 1149 (2020).
Müller-McNicoll, M. & Neugebauer, K. M. How cells get the message: dynamic assembly and function of mRNA-protein complexes. Nat. Rev. Genet. 14, 275–287 (2013).
Finan, J. M., Sutton, T. L., Dixon, D. A. & Brody, J. R. Targeting the RNA-binding protein HuR in cancer. Cancer Res. 83, 3507–3516 (2023).
Lewis, J. D. & Izaurralde, E. The role of the cap structure in RNA processing and nuclear export. Eur. J. Biochem. 247, 461–469 (1997).
Sonenberg, N., Guertin, D., Cleveland, D. & Trachsel, H. Probing the function of the eucaryotic 5’ cap structure by using a monoclonal antibody directed against cap-binding proteins. Cell 27, 563–572 (1981).
Topisirovic, I., Svitkin, Y. V., Sonenberg, N. & Shatkin, A. J. Cap and cap-binding proteins in the control of gene expression. Wiley Interdiscip. Rev. RNA 2, 277–298 (2011).
Rambout, X. & Maquat, L. E. The nuclear cap-binding complex as choreographer of gene transcription and pre-mRNA processing. Genes Dev. 34, 1113–1127 (2020).
Ishigaki, Y., Li, X., Serin, G. & Maquat, L. E. Evidence for a pioneer round of mRNA translation: mRNAs subject to nonsense-mediated decay in mammalian cells are bound by CBP80 and CBP20. Cell 106, 607–617 (2001).
Kim, S. et al. Two cap-binding proteins CBP20 and CBP80 are involved in processing primary MicroRNAs. Plant cell Physiol. 49, 1634–1644 (2008).
Arora, R. et al. NCBP2 and TFRC are novel prognostic biomarkers in oral squamous cell carcinoma. Cancer gene Ther. 30, 752–765 (2023).
Xie, J. et al. The m(7)G reader NCBP2 promotes pancreatic cancer progression by upregulating MAPK/ERK signaling. Cancers 15, 5454 (2023).
Xu, X. et al. m7G-related genes-NCBP2 and EIF4E3 determine immune contexture in head and neck squamous cell carcinoma by regulating CCL4/CCL5 expression. Mol. Carcinogenesis 62, 1091–1106 (2023).
Li, J. et al. TRIB3 promotes the progression of renal cell carcinoma by upregulating the lipid droplet-associated protein PLIN2. Cell Death Dis. 15, 240 (2024).
Slebe, F. et al. FoxA and LIPG endothelial lipase control the uptake of extracellular lipids for breast cancer growth. Nat. Commun. 7, 11199 (2016).
Qiu, B. et al. HIF2α-dependent lipid storage promotes endoplasmic reticulum homeostasis in clear-cell renal cell carcinoma. Cancer Discov. 5, 652–667 (2015).
Cruz, A. L. S., Barreto, E. A., Fazolini, N. P. B., Viola, J. P. B. & Bozza, P. T. Lipid droplets: platforms with multiple functions in cancer hallmarks. Cell Death Dis. 11, 105 (2020).
Liu, Q., Luo, Q., Halim, A. & Song, G. Targeting lipid metabolism of cancer cells: a promising therapeutic strategy for cancer. Cancer Lett. 401, 39–45 (2017).
Sainero-Alcolado, L. et al. Targeting MYC induces lipid droplet accumulation by upregulation of HILPDA in clear cell renal cell carcinoma. Proc. Natl. Acad. Sci. USA 121, e2310479121 (2024).
Zhou, L. et al. ACSS3 represses prostate cancer progression through downregulating lipid droplet-associated protein PLIN3. Theranostics 11, 841–860 (2021).
Cotte, A. K. et al. Lysophosphatidylcholine acyltransferase 2-mediated lipid droplet production supports colorectal cancer chemoresistance. Nat. Commun. 9, 322 (2018).
Zhang, X. L. et al. CTHRC1 promotes liver metastasis by reshaping infiltrated macrophages through physical interactions with TGF-β receptors in colorectal cancer. Oncogene 40, 3959–3973 (2021).
Wang, C. et al. Prognostic and biological function value of OSBPL3 in colorectal cancer analyzed by multi-omic data analysis. BMC Gastroenterol. 23, 270 (2023).
Xu, C. et al. SPP1, analyzed by bioinformatics methods, promotes the metastasis in colorectal cancer by activating EMT pathway. Biomed. Pharmacother. 91, 1167–1177 (2017).
Bu, H. et al. Diagnostic and prognostic potential of the novel biomarker nuclear cap binding protein subunit 2 (NCBP2) in colon adenocarcinoma. J. Gastrointest. Oncol. 13, 1782–1792 (2022).
Calero, G. et al. Structural basis of m7GpppG binding to the nuclear cap-binding protein complex. Nat. Struct. Biol. 9, 912–917 (2002).
Cheng, X. et al. Targeting DGAT1 ameliorates glioblastoma by increasing fat catabolism and oxidative stress. Cell Metab. 32, 229–242.e228 (2020).
Saito, Y. et al. NOVA1 acts as an oncogenic RNA-binding protein to regulate cholesterol homeostasis in human glioblastoma cells. Proc. Natl. Acad. Sci. USA 121, e2314695121 (2024).
Dang, H. et al. Oncogenic activation of the RNA binding protein NELFE and MYC signaling in hepatocellular carcinoma. Cancer Cell 32, 101–114.e108 (2017).
Xu, Z. et al. Suppression of DDX39B sensitizes ovarian cancer cells to DNA-damaging chemotherapeutic agents via destabilizing BRCA1 mRNA. Oncogene 39, 7051–7062 (2020).
Liu, W. et al. A signature of five 7-methylguanosine-related genes is a prognostic marker for lung squamous cell carcinoma. J. Thorac. Dis. 15, 6265–6278 (2023).
Mazza, C., Ohno, M., Segref, A., Mattaj, I. W. & Cusack, S. Crystal structure of the human nuclear cap binding complex. Mol. Cell 8, 383–396 (2001).
Huang, X., Zhu, B., Qian, C. & Feng, Y. The prognostic index of m(7)G-related genes in CRC correlates with immune infiltration. Sci. Rep. 12, 21282 (2022).
Jin, Y., Tan, Y., Wu, J. & Ren, Z. Lipid droplets: a cellular organelle vital in cancer cells. Cell Death Discov. 9, 254 (2023).
Cui, H. et al. Targeting DGAT1 inhibits prostate cancer cells growth by inducing autophagy flux blockage via oxidative stress. Oncogene 43, 136–150 (2024).
Wilcock, D. J. et al. Oxidative stress from DGAT1 oncoprotein inhibition in melanoma suppresses tumor growth when ROS defenses are also breached. Cell Rep. 39, 110995 (2022).
Riederer, M., Köfeler, H., Lechleitner, M., Tritscher, M. & Frank, S. Impact of endothelial lipase on cellular lipid composition. Biochim. Biophys. Acta 1821, 1003–1011 (2012).
Cadenas, C. et al. LIPG-promoted lipid storage mediates adaptation to oxidative stress in breast cancer. Int. J. Cancer 145, 901–915 (2019).
Lo, P. K., Yao, Y. & Zhou, Q. Inhibition of LIPG phospholipase activity suppresses tumor formation of human basal-like triple-negative breast cancer. Sci. Rep. 10, 8911 (2020).
Skrzypczak, M. et al. Modeling oncogenic signaling in colon tumors by multidirectional analyses of microarray data directed for maximization of analytical reliability. PloS ONE 5, e13091 (2010).
Tsukamoto, S. et al. Clinical significance of osteoprotegerin expression in human colorectal cancer. Clin. Cancer Res. 17, 2444–2450 (2011).
Matsuyama, T. et al. MUC12 mRNA expression is an independent marker of prognosis in stage II and stage III colorectal cancer. Int. J. Cancer 127, 2292–2299 (2010).
Marisa, L. et al. Gene expression classification of colon cancer into molecular subtypes: characterization, validation, and prognostic value. PLoS Med. 10, e1001453 (2013).
Alves, C. M. L., Noyszewski, A. K. & Smith, A. G. Structure and function of class III pistil-specific extensin-like protein in interspecific reproductive barriers. BMC Plant Biol. 19, 118 (2019).
Acknowledgements
This study was supported by the USTC Research Funds of the Double First-Class Initiative (No. YD9110002066), the Research Funds of the Centre for Leading Medicine and Advanced Technologies of IHM (2023IHM01092), and the Natural Science Foundation of Anhui Province (2308085QC84).
Author information
Authors and Affiliations
Contributions
Liu Liu, X.X. Wang and Lin Liu designed the study; Liu Liu, W. Lu, S.Y. Miao, Y.Yu and X.B. Zhang conducted the experiments and analysed the data; Liu Liu, W. Lu, Y. Yu and S.Y. Miao prepared the figures and drafted the manuscript. S.D. Ye and Lin Liu reviewed and revised the manuscript. All the authors have read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Georgios Giamas and Johannes Stortz. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, L., Lu, W., Miao, S. et al. NCBP2 drives colorectal cancer growth and metastasis through LIPG-mediated lipid droplet accumulation. Commun Biol (2026). https://doi.org/10.1038/s42003-026-09903-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-026-09903-5