Abstract
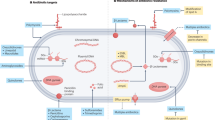
There are profound sex differences in the prevalence and outcomes of urinary tract infections (UTI). While females comprise the majority of infections, males exhibit higher morbidity and mortality with upper-tract UTI. Correspondingly, preclinical modeling has demonstrated that male and androgen-exposed female mice are highly susceptible to severe high-titer pyelonephritis, a phenotype observed in < 20% of females. Here we subject kidneys from female, male, and androgen-exposed female C3H/HeN mice with pyelonephritis and PBS-exposed control mice to single-nucleus RNA sequencing, creating (to our knowledge) the first whole-kidney single-nucleus transcriptomic dataset reflecting an infected state, comprising 248,483 nuclei. We differentiate healthy cell populations from those affected during UTI and show sex-discrepant responses that extend to kidney cell types beyond those directly interacting with bacteria. Female responses to UTI comprise a more limited range of cell types exhibiting significant upregulation of genes within KEGG pathways and pro-inflammatory transcription factor regulons. Meanwhile, males evidence predisposition to injury pathways even with control (saline) inoculation and responded to UTI with less intensity but across more cell types than females. In total, these data illuminate sex-discrepant transcriptional responses and outcomes in renal infection and enable detailed dissection of these responses at the cellular and molecular level.

Similar content being viewed by others
Data availability
Gene expression data have been submitted to the Gene Expression Omnibus (GEO) under accession number GSE296327, and Illumina reads have been submitted to NCBI’s Short Read Archive (SRA) under NCBI bioproject PRJNA1182331. Additional source data behind the figures in the paper are available online26 as Supplementary Data 1–10. Any remaining information can be obtained from the corresponding authors upon reasonable request.
References
Flores-Mireles, A. L., Walker, J. N., Caparon, M. & Hultgren, S. J. Urinary tract infections: epidemiology, mechanisms of infection and treatment options. Nat. Rev. Microbiol. 13, 269–284 (2015).
Foxman, B. Urinary tract infection syndromes: occurrence, recurrence, bacteriology, risk factors, and disease burden. Infect. Dis. Clin. North Am. 28, 1–13 (2014).
Albracht, C. D., Hreha, T. N. & Hunstad, D. A. Sex effects in pyelonephritis. Pediatr. Nephrol. 36, 507–515 (2021).
Deltourbe, L., Lacerda Mariano, L., Hreha, T. N., Hunstad, D. A. & Ingersoll, M. A. The impact of biological sex on diseases of the urinary tract. Mucosal Immunol. 15, 857–866 (2022).
O’Brien, V. P. et al. A mucosal imprint left by prior Escherichia coli bladder infection sensitizes to recurrent disease. Nat. Microbiol. 2, 16196 (2016).
Olson, P. D., Hruska, K. A. & Hunstad, D. A. Androgens enhance male urinary tract infection severity in a new model. J. Am. Soc. Nephrol. 27, 1625–1634 (2016).
Olson, P. D. et al. Androgen exposure potentiates formation of intratubular communities and renal abscesses by Escherichia coli. Kidney Int. 94, 502–513 (2018).
Hreha, T. N., Collins, C. A., Cole, E. B., Jin, R. J. & Hunstad, D. A. Androgen exposure impairs neutrophil maturation and function within the infected kidney. mBio 15, e03170 (2024).
Hreha, T. N. et al. TGFβ1 orchestrates renal fibrosis following Escherichia coli pyelonephritis. Physiol. Rep. 8, e14401 (2020).
Kirita, Y., Wu, H., Uchimura, K., Wilson, P. C. & Humphreys, B. D. Cell profiling of mouse acute kidney injury reveals conserved cellular responses to injury. Proc. Natl. Acad. Sci. USA 117, 15874–15883 (2020).
Wu, H. et al. Mapping the single cell transcriptomic response of murine diabetic kidney disease to therapies. Cell Metab. 34, 1064–1078.e6 (2022).
Gerhardt, L. M. S. et al. Lineage tracing and single-nucleus multiomics reveal novel features of adaptive and maladaptive repair after acute kidney injury. J. Am. Soc. Nephrol. 34, 554–571 (2023).
Gerhardt, L. M. S., Liu, J., Koppitch, K., Cippà, P. E. & McMahon, A. P. Single-nuclear transcriptomics reveals diversity of proximal tubule cell states in a dynamic response to acute kidney injury. Proc. Natl. Acad. Sci. USA 118, e2026684118 (2021).
Park, K. M., Kim, J. I., Ahn, Y., Bonventre, A. J. & Bonventre, J. V. Testosterone is responsible for enhanced susceptibility of males to ischemic renal injury. J. Biol. Chem. 279, 52282–52292 (2004).
Silbiger, S. R. & Neugarten, J. The impact of gender on the progression of chronic renal disease. Am. J. Kidney Dis. 25, 515–533 (1995).
Garcia-Roig, M. L. & Kirsch, A. J. Urinary tract infection in the setting of vesicoureteral reflux. F1000Research 5, F1000 Faculty Rev-1552 (2016).
Bowen, S. E., Watt, C. L., Murawski, I. J., Gupta, I. R. & Abraham, S. N. Interplay between vesicoureteric reflux and kidney infection in the development of reflux nephropathy in mice. Dis. Model. Mech. 6, 934–941 (2013).
Wilson, P. C. et al. The single-cell transcriptomic landscape of early human diabetic nephropathy. Proc. Natl. Acad. Sci. USA 116, 19619–19625 (2019).
Humphreys, B. & Kirita, Y. Nuclei isolation from human kidney for single-nucleus RNA-seq. https://www.protocols.io/view/nuclei-isolation-from-human-kidney-for-single-nucl-nahdab6 (2018).
Fleming, S. J. et al. Unsupervised removal of systematic background noise from droplet-based single-cell experiments using CellBender. Nat. Methods 20, 1323–1335 (2023).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291.e9 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Kumar, R. C. & Thakur, M. K. Androgen receptor mRNA is inversely regulated by testosterone and estradiol in adult mouse brain. Neurobiol. Aging 25, 925–933 (2004).
Kumar, N., Mishra, B., Athar, M. & Mukhtar, S. Inference of gene regulatory network from single-cell transcriptomic data using pySCENIC. Methods Mol. Biol. 2328, 171–182 (2021).
Rane, M. J., Zhao, Y. & Cai, L. Krϋppel-like factors (KLFs) in renal physiology and disease. EBioMedicine 40, 743–750 (2019).
Hunstad, D. A. & Earl, A. M. Supplementary Data 1-10. figshare https://doi.org/10.6084/m9.figshare.31411557 (2026).
Zychlinsky Scharff, A. et al. Sex differences in IL-17 contribute to chronicity in male versus female urinary tract infection. JCI Insight 5, e122998 (2019).
Rodríguez-Montes, L. et al. Sex-biased gene expression across mammalian organ development and evolution. Science 382, eadf1046 (2023).
Xiong, L. et al. Direct androgen receptor control of sexually dimorphic gene expression in the mammalian kidney. Dev. Cell 58, 2338–2358.e5 (2023).
Chen, L. et al. Renal-tubule epithelial cell nomenclature for single-cell RNA-sequencing studies. J. Am. Soc. Nephrol. 30, 1358–1364 (2019).
Saxena, V. et al. Kidney intercalated cells are phagocytic and acidify internalized uropathogenic Escherichia coli. Nat. Commun. 12, 2405 (2021).
Dumas, S. J. et al. Phenotypic diversity and metabolic specialization of renal endothelial cells. Nat. Rev. Nephrol. 17, 441–464 (2021).
Kalucka, J. et al. Single-cell transcriptome atlas of murine endothelial cells. Cell 180, 764–779.e20 (2020).
Balzer, M. S., Rohacs, T. & Susztak, K. How many cell types are in the kidney and what do they do? Annu. Rev. Physiol. 84, 507–531 (2022).
Quatredeniers, M. et al. Meta-analysis of single-cell and single-nucleus transcriptomics reveals kidney cell type consensus signatures. Sci. Data 10, 361 (2023).
Higashi, A. Y., Aronow, B. J. & Dressler, G. R. Expression profiling of fibroblasts in chronic and acute disease models reveals novel pathways in kidney fibrosis. J. Am. Soc. Nephrol. 30, 80–94 (2019).
McLellan, L. K. et al. A host receptor enables type 1 pilus-mediated pathogenesis of Escherichia coli pyelonephritis. PLoS Pathog. 17, e1009314 (2021).
Haghverdi, L., Büttner, M., Wolf, F. A., Buettner, F. & Theis, F. J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
Ding, H. et al. Kidney fibrosis molecular mechanisms Spp1 influences fibroblast activity through transforming growth factor beta Smad signaling. iScience 27, 109839 (2024).
Zhang, B., Yang, W. & Chen, L. Secreted phosphoprotein 1 exacerbates renal ischemia-reperfusion injury by inhibiting PI3K/AKT signaling pathway. Tissue Cell 83, 102154 (2023).
Bahrami, S. & Drabløs, F. Gene regulation in the immediate-early response process. Adv. Biol. Regul. 62, 37–49 (2016).
Wolf, F. A. et al. PAGA: graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biol. 20, 59 (2019).
Wee, H. N., Liu, J.-J., Ching, J., Kovalik, J.-P. & Lim, S. C. The kynurenine pathway in acute kidney injury and chronic kidney disease. Am. J. Nephrol. 52, 771–787 (2021).
Loughman, J. A. & Hunstad, D. A. Induction of indoleamine 2,3-dioxygenase by uropathogenic bacteria attenuates innate responses to epithelial infection. J. Infect. Dis. 205, 1830–1839 (2012).
Matheus, L. H. G. et al. Indoleamine 2, 3-dioxygenase (IDO) increases during renal fibrogenesis and its inhibition potentiates TGF-β1-induced epithelial to mesenchymal transition. BMC Nephrol. 18, 287 (2017).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell-cell communication from combined expression of multi-subunit ligand-receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Huang, R., Fu, P. & Ma, L. Kidney fibrosis: from mechanisms to therapeutic medicines. Signal Transduct. Target. Ther. 8, 1–20 (2023).
Humphreys, B. D. Mechanisms of renal fibrosis. Annu. Rev. Physiol. 80, 309–326 (2018).
Zhai, X. et al. Gas6/AXL pathway: immunological landscape and therapeutic potential. Front. Oncol. 13, 1121130 (2023).
Omatsu, M. et al. THBS1-producing tumor-infiltrating monocyte-like cells contribute to immunosuppression and metastasis in colorectal cancer. Nat. Commun. 14, 5534 (2023).
Gao, L. et al. THBS1/CD47 modulates the interaction of γ-catenin with E-cadherin and participates in epithelial–mesenchymal transformation in lipid nephrotoxicity. Front. Cell Dev. Biol. 8, 601521 (2021).
Huffstater, T., Merryman, W. D. & Gewin, L. S. Wnt/β-catenin in acute kidney injury and progression to chronic kidney disease. Semin. Nephrol. 40, 126–137 (2020).
Xie, Y. et al. FGF/FGFR signaling in health and disease. Signal Transduct. Target. Ther. 5, 1–38 (2020).
Baek, J. H. et al. IL-34 mediates acute kidney injury and worsens subsequent chronic kidney disease. J. Clin. Investig. 125, 3198–3214 (2015).
Buhl, E. M. et al. The role of PDGF-D in healthy and fibrotic kidneys. Kidney Int. 89, 848–861 (2016).
Feng, L. et al. Role of the SLIT-ROBO signaling pathway in renal pathophysiology and various renal diseases. Front. Physiol. 14, 1226341 (2023).
Ostendorf, T., Boor, P., van Roeyen, C. R. C. & Floege, J. Platelet-derived growth factors (PDGFs) in glomerular and tubulointerstitial fibrosis. Kidney Int. Suppl. 4, 65–69 (2014).
Zhang, M.-Z. et al. CSF-1 signaling mediates recovery from acute kidney injury. J. Clin. Investig. 122, 4519–4532 (2012).
Alikhan, M. A. et al. Colony-stimulating factor-1 promotes kidney growth and repair via alteration of macrophage responses. Am. J. Pathol. 179, 1243–1256 (2011).
Paragas, N. et al. Intercalated cells defend the urinary system from bacterial infection. J. Clin. Investig. 124, 2963–2976 (2014).
Skeldal, S. et al. Mapping of the interaction site between sortilin and the p75 neurotrophin receptor reveals a regulatory role for the sortilin intracellular domain in p75 neurotrophin receptor shedding and apoptosis. J. Biol. Chem. 287, 43798–43809 (2012).
Tran, J. et al. A role for survivin in chemoresistance of endothelial cells mediated by VEGF. Proc. Natl. Acad. Sci. USA 99, 4349–4354 (2002).
Liu, J. et al. Molecular characterization of the transition from acute to chronic kidney injury following ischemia/reperfusion. JCI Insight 2, e94716 (2017).
Cao, S. et al. Epidermal growth factor receptor activation is essential for kidney fibrosis development. Nat. Commun. 14, 7357 (2023).
Luo, A. J., Chang, F. C. & Lin, S. L. Exploring angiopoietin-2: clinical insights and experimental perspectives in kidney diseases. Kidney Int. Rep. 9, 3375–3385 (2024).
Sang, Y. et al. Role of semaphorin 3A in kidney development and diseases. Diagnostics 13, 3038 (2023).
Park, J., Liu, C.(L. inda), Kim, J. & Susztak, K. Understanding the kidney one cell at a time. Kidney Int. 96, 862–870 (2019).
Pregizer, S., Vreven, T., Mathur, M. & Robinson, L. N. Multi-omic single cell sequencing: overview and opportunities for kidney disease therapeutic development. Front. Mol. Biosci. 10, 1176856 (2023).
Hreha, T. N. et al. Androgen-influenced polarization of activin A-producing macrophages accompanies post-pyelonephritic renal scarring. Front. Immunol. 11, 1641 (2020).
McLellan, L. K., Daugherty, A. L. & Hunstad, D. A. Sex differences in population dynamics during formation of kidney bacterial communities by uropathogenic Escherichia coli. Infect. Immun. 89, e00716–e00720 (2021).
Xie, X. et al. Single-cell transcriptome profiling reveals neutrophil heterogeneity in homeostasis and infection. Nat. Immunol. 21, 1119–1133 (2020).
Mulvey, M. A., Schilling, J. D. & Hultgren, S. J. Establishment of a persistent Escherichia coli reservoir during the acute phase of a bladder infection. Infect. Immun. 69, 4572–4579 (2001).
Hannan, T. J., Mysorekar, I. U., Hung, C. S., Isaacson-Schmid, M. L. & Hultgren, S. J. Early severe inflammatory responses to uropathogenic E. coli predispose to chronic and recurrent urinary tract infection. PLoS Pathog. 6, e1001042 (2010).
Hung, C. S., Dodson, K. W. & Hultgren, S. J. A murine model of urinary tract infection. Nat. Protoc. 4, 1230–1243 (2009).
Zychlinsky Scharff, A., Albert, M. L. & Ingersoll, M. A. Urinary tract infection in a small animal model: transurethral catheterization of male and female mice. J. Vis. Exp. 130, 54432 (2017).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Li, B. et al. Cumulus provides cloud-based data analysis for large-scale single-cell and single-nucleus RNA-seq. Nat. Methods 17, 793–798 (2020).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Ge, S. X., Jung, D. & Yao, R. ShinyGO: a graphical gene-set enrichment tool for animals and plants. Bioinformatics 36, 2628–2629 (2020).
Kanehisa, M., Furumichi, M., Sato, Y., Ishiguro-Watanabe, M. & Tanabe, M. KEGG: integrating viruses and cellular organisms. Nucleic Acids Res. 49, D545–D551 (2021).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Thomas, P. D. et al. PANTHER: making genome-scale phylogenetics accessible to all. Protein Sci. 31, 8–22 (2022).
Gene Ontology Consortium et al. The gene ontology knowledgebase in 2023. Genetics 224, iyad031 (2023).
Carbon, S. et al. AmiGO: online access to ontology and annotation data. Bioinformatics 25, 288–289 (2009).
Eppig, J. T. Mouse Genome Informatics (MGI) resource: genetic, genomic, and biological knowledgebase for the laboratory mouse. ILAR J. 58, 17–41 (2017).
Acknowledgements
This work was funded by National Institutes of Health grants R01-DK126697 (to DAH and AME) and R01-AI158418 (to DAH), and by the Children’s Discovery Institute through the WashU Center for Cellular Imaging (CDI-CORE-2019-813). We acknowledge Joshua Gould, Lucas van Dijk, Mark Young, Nathan McGuire, and Patrick McGuire for assistance with analysis; and Lisa McLellan for IF images schematized within Fig. 7.
Author information
Authors and Affiliations
Contributions
T.N.H., A.L.M., A.M.E., and D.A.H. designed research; T.N.H., C.A.C., and A.L.M. performed research experiments; T.N.H., A.L.M., H.W., and C.G. performed data analysis and visualization; B.D.H., A.L.M., and D.A.H. reviewed data analyses; T.N.H. and A.L.M. drafted the manuscript; B.D.H., A.M.E., and D.A.H. edited and finalized the manuscript and figures; T.N.H., A.L.M., C.A.C., A.M.E., and D.A.H. completed revision experiments and responses to reviewers.
Corresponding authors
Ethics declarations
Competing interests
D.A.H. serves on the Board of Directors of BioVersys AG (SIX: BIOV) and has received funding for unrelated research work from BioAge Laboratories and Pfizer. All other authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Ines Ambite and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Michele Repetto and Rupali Sathe.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hreha, T.N., Manson, A.L., Collins, C.A. et al. Single-nucleus transcriptomics illuminates sex differences during murine Escherichia coli pyelonephritis. Commun Biol (2026). https://doi.org/10.1038/s42003-026-09946-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-026-09946-8