Abstract
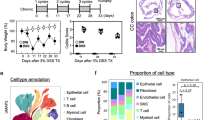
Inflammatory bowel disease (IBD), including ulcerative colitis (UC) and Crohn’s disease (CD), is closely linked to gut microbial imbalance marked by loss of beneficial microbes and overgrowth of pathogens. Christensenella intestinihominis, a bacterium that associated with intestinal homeostasis, is depleted in patients with ulcerative colitis (UC), yet its therapeutic potential against this disease remains to be elucidated. Here we demonstrate that C. intestinihominis AF73-05CM02, a strain isolated from Chinese individual, alleviates colitis. In healthy human gut, C. intestinihominis co-occurs with beneficial microbes in strongly connected networks, while these interactions are disrupted in UC. We employ the dextran sulfate sodium (DSS)-induced murine colitis model, a widely recognized preclinical model for investigating intestinal inflammation. In this model, oral gavage with AF73-05CM02 mitigates weight loss, ameliorates colonic injury, improves intestinal health markers, and reverses colon damage. It exerts these effects by reducing harmful bacteria such as Helicobacter species and increasing beneficial taxa like Akkermansia, while enhancing the intestinal epithelial barrier integrity and regulating immune responses. These findings indicate AF73-05CM02 may aid in the treatment of inflammatory bowel disease by restoring a healthy gut microbial community.
Similar content being viewed by others
Data availability
The data that support the findings of this study have been deposited into CNSA52 with accession number CNP0004539. Raw data for each graph is included in Supplementary Data File. All other data are available from the corresponding author on reasonable request.
References
Singh, S., Dulai, P. S., Zarrinpar, A., Ramamoorthy, S. & Sandborn, W. J. Obesity in IBD: epidemiology, pathogenesis, disease course and treatment outcomes. Nat. Rev. Gastroenterol. Hepatol. 14, 110–121 (2017).
Adams, A. et al. Early management of acute severe UC in the biologics era: development and international validation of a prognostic clinical index to predict steroid response. Gut 72, 433–442 (2023).
Dai, C., Dong, Z. Y., Wang, Y. N., Huang, Y. H. & Jiang, M. Development and validation of a nomogram to predict non-response to 5-aminosalicylic acid in patients with ulcerative colitis. Rev. Esp. Enferm. Dig. 116, 124–131 (2024).
Cordero, R. Y. et al. Trans-ancestry, Bayesian meta-analysis discovers 20 novel risk loci for inflammatory bowel disease in an African American, East Asian and European cohort. Hum. Mol. Genet 32, 873–882 (2023).
Sugihara, K. & Kamada, N. Metabolic network of the gut microbiota in inflammatory bowel disease. Inflamm. Regen. 44, 11 (2024).
Morotomi, M., Nagai, F. & Watanabe, Y. Description of Christensenella minuta gen. nov., sp. nov., isolated from human faeces, which forms a distinct branch in the order Clostridiales, and proposal of Christensenellaceae fam. nov. Int J. Syst. Evol. Microbiol 62, 144–149 (2012).
Braun, T. et al. Individualized Dynamics in the Gut Microbiota Precede Crohn’s Disease Flares. Am. J. Gastroenterol. 114, 1142–1151 (2019).
Loh, G. & Blaut, M. Role of commensal gut bacteria in inflammatory bowel diseases. Gut Microbes 3, 544–555 (2012).
Zuo, T. & Ng, S. C. The Gut Microbiota in the Pathogenesis and Therapeutics of Inflammatory Bowel Disease. Front Microbiol 9, 2247 (2018).
Waters, J. L. & Ley, R. E. The human gut bacteria Christensenellaceae are widespread, heritable, and associated with health. BMC Biol. 17, 83 (2019).
Relizani, K. et al. Selection of a novel strain of Christensenella minuta as a future biotherapy for Crohn’s disease. Sci. Rep. 12, 6017 (2022).
Ignatyeva, O. et al. Christensenella minuta, a new candidate next-generation probiotic: current evidence and future trajectories. Front Microbiol 14, 1241259 (2023).
Ang, W. S. et al. A Keystone Gut Bacterium Christensenella minuta-A Potential Biotherapeutic Agent for Obesity and Associated Metabolic Diseases. Foods 12, https://doi.org/10.3390/foods12132485 (2023).
Kropp, C. et al. The Keystone commensal bacterium Christensenella minuta DSM 22607 displays anti-inflammatory properties both in vitro and in vivo. Sci. Rep. 11, 11494 (2021).
Hong, H. H. et al. Multiomics Integration Reveals Microbial Gene Interactions Shaping Host Responses in a DSS-Induced Colitis Mouse Model. J. Microbiol Biotechnol. 35, e2507010 (2025).
Hodgkiss, R. & Acharjee, A. Unravelling metabolite-microbiome interactions in inflammatory bowel disease through AI and interaction-based modelling. Biochim Biophys. Acta Mol. Basis Dis. 1871, 167618 (2025).
Ji, H. et al. Prebiotics empower probiotics with gastrointestinal stress resistance for colon-targeted release to synergistically alleviate colitis. J. Control Release 380, 297–316 (2025).
Zhao, X. et al. Inflammatory Microenvironment-Responsive Microsphere Vehicles Modulating Gut Microbiota and Intestinal Inflammation for Intestinal Stem Cell Niche Remodeling in Inflammatory Bowel Disease. ACS Nano 19, 12063–12079 (2025).
Jiang, J. et al. Synergistic Genetic and Chemical Engineering of Probiotics for Enhanced Intestinal Microbiota Regulation and Ulcerative Colitis Treatment. Adv. Mater. 37, e2417050 (2025).
Barry, D., Thachuk, C. & Trink, J. Integrating gut microbiome and host transcriptomics for the personalized management of IBD. Epigenomics. 17, 1503–1512 (2025).
Banerjee, S., Schlaeppi, K. & van der Heijden, M. G. A. Keystone taxa as drivers of microbiome structure and functioning. Nat. Rev. Microbiol 16, 567–576 (2018).
Shannon, C. E. The mathematical theory of communication. 1963. MD Comput 14, 306–317 (1997).
Gower, J. C. Some Distance Properties of Latent Root and Vector Methods Used in Multivariate Analysis. Biometrika 53, 325–338 (1966).
Torres, P. J. et al. Gut Microbial Diversity in Women With Polycystic Ovary Syndrome Correlates With Hyperandrogenism. J. Clin. Endocrinol. Metab. 103, 1502–1511 (2018).
Bray, J. R. & Curtis, J. T. An Ordination of the Upland Forest Communities of Southern Wisconsin. Ecol. Monogr. 27, 326–349 (1957).
Kers, J. G. & Saccenti, E. The Power of Microbiome Studies: Some Considerations on Which Alpha and Beta Metrics to Use and How to Report Results. Front Microbiol 12, 796025 (2021).
Anderson, M. J. A new method for non-parametric multivariate analysis of variance. Austral. Ecology 26, 32–46, (2001).
Hu, Y. J. & Satten, G. A. A rarefaction-without-resampling extension of PERMANOVA for testing presence-absence associations in the microbiome. Bioinformatics 38, 3689–3697 (2022).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Fabbrini, M., Scicchitano, D., Candela, M., Turroni, S. & Rampelli, S. Connect the dots: sketching out microbiome interactions through networking approaches. Microbiome Res Rep. 2, 25 (2023).
Herren, C. M. & McMahon, K. D. Cohesion: a method for quantifying the connectivity of microbial communities. ISME J. 11, 2426–2438 (2017).
Olesen, J. M., Bascompte, J., Dupont, Y. L. & Jordano, P. The modularity of pollination networks. Proc. Natl. Acad. Sci. USA 104, 19891–19896 (2007).
Ohkusa, T. Production of experimental ulcerative colitis in hamsters by dextran sulfate sodium and changes in intestinal microflora]. Nihon Shokakibyo Gakkai Zasshi 82, 1327–1336 (1985).
Pelaseyed, T. et al. The mucus and mucins of the goblet cells and enterocytes provide the first defense line of the gastrointestinal tract and interact with the immune system. Immunol. Rev. 260, 8–20 (2014).
Odenwald, M. A. & Turner, J. R. The intestinal epithelial barrier: a therapeutic target? Nat. Rev. Gastroenterol. Hepatol. 14, 9–21 (2017).
Panwar, S., Sharma, S. & Tripathi, P. Role of Barrier Integrity and Dysfunctions in Maintaining the Healthy Gut and Their Health Outcomes. Front Physiol. 12, 715611 (2021).
Szandruk-Bender, M. et al. The Improvement of Intestinal Mucosal Epithelial Barrier Integrity by 1,3,4-Oxadiazole Derivatives of Pyrrolo[3,4-d]pyridazinone in Rat Experimental Colitis. J. Inflamm. Res 18, 9735–9753 (2025).
Peterson, L. W. & Artis, D. Intestinal epithelial cells: regulators of barrier function and immune homeostasis. Nat. Rev. Immunol. 14, 141–153 (2014).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech.: Theory Exp. 2008, P10008 (2008).
Signorino, G. et al. Role of Toll-like receptor 13 in innate immune recognition of group B streptococci. Infect. Immun. 82, 5013–5022 (2014).
Bucker, R. et al. alpha-Haemolysin of Escherichia coli in IBD: a potentiator of inflammatory activity in the colon. Gut 63, 1893–1901 (2014).
Lieberman, J. Granzyme A activates another way to die. Immunol. Rev. 235, 93–104 (2010).
Russo, I., Bubacco, L. & Greggio, E. LRRK2 as a target for modulating immune system responses. Neurobiol. Dis. 169, 105724 (2022).
Cabezudo, D., Tsafaras, G., Van Acker, E., Van den Haute, C. & Baekelandt, V. Mutant LRRK2 exacerbates immune response and neurodegeneration in a chronic model of experimental colitis. Acta Neuropathol. 146, 245–261 (2023).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28, 27–30 (2000).
Jablonska, B. & Mrowiec, S. Nutritional Status and Its Detection in Patients with Inflammatory Bowel Diseases. Nutrients 15, https://doi.org/10.3390/nu15081991 (2023).
Kropp, C. et al. Christensenella minuta protects and restores intestinal barrier in a colitis mouse model by regulating inflammation. NPJ Biofilms Microbiomes 10, 88 (2024).
Yang, C. & Merlin, D. Unveiling Colitis: A Journey through the Dextran Sodium Sulfate-induced Model. Inflamm. Bowel Dis. 30, 844–853 (2024).
Chichlowski, M., Sharp, J. M., Vanderford, D. A., Myles, M. H. & Hale, L. P. Helicobacter typhlonius and Helicobacter rodentium differentially affect the severity of colon inflammation and inflammation-associated neoplasia in IL10-deficient mice. Comp. Med 58, 534–541 (2008).
Elzinga, J. et al. Binding of Akkermansia muciniphila to mucin is O-glycan specific. Nat. Commun. 15, 4582 (2024).
Huang, L. et al. 5-Aminosalicylic acid ameliorates dextran sulfate sodium-induced colitis in mice by modulating gut microbiota and bile acid metabolism. Cell Mol. Life Sci. 79, 460 (2022).
Wang, W. et al. The China National GeneBank Sequence Archive (CNSA) 2024 update. Hortic. Res 12, uhaf036 (2025).
Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 20, 257 (2019).
Lin, X. et al. The genomic landscape of reference genomes of cultivated human gut bacteria. Nat. Commun. 14, 1663 (2023).
Kurtz, Z. D. et al. Sparse and compositionally robust inference of microbial ecological networks. PLoS Comput Biol. 11, e1004226 (2015).
Meinshausen, N. & Bühlmann, P. High-Dimensional Graphs and Variable Selection with the Lasso. Ann. Stat. 34, 1436–1462 (2006).
Liu, H., Roeder, K. & Wasserman, L. Stability Approach to Regularization Selection (StARS) for High Dimensional Graphical Models. Adv. Neural Inf. Process Syst. 24, 1432–1440 (2010).
Zou, Y. et al. Taxonomic Description and Genome Sequence of Christensenella intestinihominis sp. nov., a Novel Cholesterol-Lowering Bacterium Isolated From Human Gut. Front Microbiol 12, 632361 (2021).
He, N. et al. Stachyose Exerts Anticolitis Efficacy by Re-balancing Treg/Th17 and Activating the Butyrate-Derived PPARgamma Signaling Pathway. J. Agric Food Chem. 72, 12171–12183 (2024).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. Diet Diversity Is Associated with Beta but not Alpha Diversity of Pika Gut Microbiota. Front Microbiol 7, 1169 (2016).
Giorgi, F. M., Ceraolo, C. & Mercatelli, D. The R Language: An Engine for Bioinformatics and Data Science. Life (Basel) 12, https://doi.org/10.3390/life12050648 (2022).
Khleborodova, A. et al. lefser: implementation of metagenomic biomarker discovery tool, LEfSe, in R. Bioinformatics 40, https://doi.org/10.1093/bioinformatics/btae707 (2024).
Cantalapiedra, C. P., Hernandez-Plaza, A., Letunic, I., Bork, P. & Huerta-Cepas, J. eggNOG-mapper v2: Functional Annotation, Orthology Assignments, and Domain Prediction at the Metagenomic Scale. Mol. Biol. Evol. 38, 5825–5829 (2021).
Wagner, G. P., Kin, K. & Lynch, V. J. Measurement of mRNA abundance using RNA-seq data: RPKM measure is inconsistent among samples. Theory Biosci. 131, 281–285 (2012).
Beghini, F. et al. Integrating taxonomic, functional, and strain-level profiling of diverse microbial communities with bioBakery 3. Elife 10, https://doi.org/10.7554/eLife.65088 (2021).
Mitteer, D. R., Greer, B. D., Fisher, W. W. & Cohrs, V. L. Teaching behavior technicians to create publication-quality, single-case design graphs in graphpad prism 7. J. Appl Behav. Anal. 51, 998–1010 (2018).
Acknowledgements
This work was financially supported by the grants from the National Natural Science Foundation of China (No. 82460112), the National Key R&D Program of China (No. 2024YFA1308300), the Innovative Team Program of Yunnan Province (No. 202305AS350019), and the Shenzhen Municipal Government of China (Project No. JCYJ20241202124801003).
Author information
Authors and Affiliations
Contributions
Conceptualization: Y.Q.Z., S.Y.L., and Y.S.; Methodology: Y.Q.Z., S.Y.L., and Y.S.,and X.L.; Investigation: Z.N.W., X.F.S., X.F.L., H.Y.W., H.W.L., M.M.W, Z.Z.Y., K.Y.Z., N.N.H., and L.J.R.; Funding acquisition: Y.Q.Z., and Y.S.; Project administration: Y.Q.Z.; Writing: all authors contribute to writing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Sridhar Mani and Tobias Goris.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wu, Z., Sun, X., Li, X. et al. Amelioration of colitis through restored gut ecology using Christensenella intestinihominis AF73-05CM02 as a probiotic in mice. Commun Biol (2026). https://doi.org/10.1038/s42003-026-10031-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-026-10031-3