Abstract
Bicyclic peptides, which integrate the advantageous properties of small molecules and antibodies, have emerged as a promising class of therapeutic candidates. In particular, integrin αvβ3 serves as a critical molecular target for cancer diagnosis and therapy. However, the development of bicyclic peptide ligands specifically targeting this integrin remains inadequately explored. To address this gap, we designed and synthesized a series of RGD-containing bicyclic peptides featuring a tryptathionine bridge. Notably, bicyclic peptide 5j incorporates the non-canonical sequence norArg-Gly-Asp, exhibiting high affinity and selectivity toward integrin αvβ3. Molecular dynamics simulations provided insights into the conformational preferences and demonstrated that norArg plays a critical role in determining the selectivity between αvβ3 and αIIbβ3. Employing peptide 5j as the targeting ligand, the peptide drug conjugates P1 showed significant inhibitory effects on the A549 cell line in both, in vitro and in vivo experiments. These data provide important theoretical foundations for the development of αvβ3-targeting bicyclic peptides and offer new options for αvβ3-targeted tumor therapy.

Similar content being viewed by others
Introduction
Bicyclic peptides are a new modality that have the potential to become part of the next generation of therapeutics, combining specificity, potency, and attractive drug-like properties1,2,3,4,5. Their molecular structure is characterized by the incorporation of a constrained motif, which serves to restrict rotational freedom, thus locking the polypeptide chain into a more defined, rigid bicyclic conformation6,7. This structural attribute markedly diminishes the entropy penalties incurred during the interaction with biological targets, which in turn significantly enhances the affinity and selectivity of these peptides for challenging targets3,8.
Integrin αvβ3 serves as a pivotal target in tumor-targeted therapy, renowned for its interaction with extracellular matrix (ECM) proteins containing the Arg-Gly-Asp (RGD) tripeptide sequence9,10,11,12,13. This interaction activates critical cellular signaling pathways, such as PI3K/Akt and FAK-MAPK2-4, which play essential roles in tumor growth and progression11,14,15. Cilengitide, the most advanced inhibitor targeting αvβ316,17, however, exhibits pro-angiogenic and tumor-promoting effects at concentrations below its half-maximal inhibitory concentration (IC50), which not only led to its failure in Phase III clinical trials for glioblastoma but also raised significant concerns about the development of integrin αvβ3-targeted therapeutics18,19. Although αvβ3 as an inhibitory target still presents certain limitations, recent studies have highlighted the growing interest in its application as a target for payload delivery in the field of tumor-targeted therapy11,20,21. Furthermore, a variety of probes for tumor imaging have been extensively developed employing RGD-based strategies targeting αvβ3 integrin and utilizing radionuclides, such as 18F, 64Cu, 68Ga, and 99mTc22,23,24,25,26,27,28. Notably, two related drug conjugates, BGC0222 and VIP236, have advanced to clinical trial phases, thereby providing strong evidence for the translational potential of this target in precision oncology research29,30.
In response to this pressing need, several cyclic peptides containing novel architectures31,32,33, particularly bicyclic peptides targeting αvβ3, have been successively reported in the literature24,34. As early as 1995, Ruoslahti and co-workers utilized phage display to identify a bicyclic peptide named RGD4C, which is stabilized by two pairs of disulfide bonds and serves to constrain the spatial conformation of the peptide (see Fig. 1a). Compared to the monocyclic peptide with a single pair of disulfide bonds and the RGD-containing linear peptide, RGD4C exhibited at least a 20-fold to 200-fold increase in inhibitory activity against cell binding to vitronectin, which emphasizes the importance of conformational restriction35. However, instability of the RGD4C disulfide bonds under reducing conditions is a potential disadvantage that can significantly reduce αvβ3 binding affinity36,37. In 2019, Timmerman and co-workers successfully identified a bicyclic peptide with exceptional selectivity for the αvβ3 integrin by leveraging the CLIPS peptide platform and incorporating the “1,3,5-tris(bromomethyl)benzene” constraint motif (see Fig. 1b)38,39. It can thus be concluded that the design of bicyclic peptides by means of some high-stability scaffolds is capable of preserving the selectivity of the designed molecules for integrin αvβ3 whilst simultaneously enhancing their inhibitory activity.
a RGD4C and b CLIPS peptide CT3HPQcT3RGDcT3. c Design strategy of bicyclic peptides in this study; R* corresponds to Arg, norArg, and homoArg; G# corresponds to Gly, βAla, and GABA.
In this contribution we propose, that incorporating a tryptathionine bridge, which links the cysteine thiol group to the C-2 position of tryptophan, could serve as a novel constrained motif for the discovery of αvβ3-targeting bicyclic peptides (see Fig. 1c). This modification is expected to enhance conformational rigidity and improve binding affinity, thereby facilitating the identification of αvβ3-targeting bicyclic peptides and potentially reshaping the landscape of cancer treatment.
To validate our hypothesis, we herein report a series of tryptathionine-bridged bicyclic peptides. By employing an ELISA integrin-extracellular matrix binding assay40, we systematically optimized their structures and identified bicyclic peptide 5j, which exhibits high affinity and selectivity for αvβ3 (IC50 = 0.7 ± 0.1 nM). Molecular dynamics simulations revealed the unique binding mode of this class of bicyclic peptides. Both in vitro and in vivo experiments comprehensively confirmed the ability of peptide 5j to precisely target αvβ3, as well as its potential application value in the field of peptide-drug conjugates. Overall, our research provides significant insights into the development of αvβ3-targeting bicyclic peptides and expands the toolbox for precision targeting of tumors with elevated αvβ3 expression.
Results and discussion
Molecular design, synthesis, and integrin affinity of bicyclic peptides
As a basic molecular design, we grafted the conserved RGD motif, known for its targeting specificity to integrins, to the skeleton of the bicyclic cyclooctapeptide amanitin41. While the tryptathionine-cross-link was kept constant in order to stabilize the conformation, the original DHIle-HPro-Asn motif of the A ring was simplified to Ala-Pro-Asn, whereas the B ring was intended to carry the RGD motif. In the subsequent synthesis of an 11-membered peptide library, we allowed for ring contractions, ring expansions and amino acid exchanges of the RGD motif42. Compounds 5a-5k were synthesized using Fmoc-protected solid-phase peptide synthesis (SPPS), following the procedures outlined in Scheme 1.
Synthesis and reaction conditions of a bicyclic octapeptide library: Fmoc-AA1-n-OH can stand for Fmoc-Pro-OH, Fmoc-Ala-OH, Fmoc-Asn(Trt)-OH, Fmoc-Trp-OH, Fmoc-Asp(OtBu)-OH, Fmoc-Cys(Trt)-OH, Fmoc-homoCys(Trt)-OH or Fmoc-D-Cys(Trt)-OH, R* or G#; R* is Fmoc-Arg(Pbf)-OH, Fmoc-homoArg(Pbf)-OH or Fmoc-norArg(Pbf)-OH; G# is Fmoc-Gly-OH, Fmoc-β-Ala-OH or Fmoc-GABA-OH; a 20% piperidine in DMF for 10 min (twice); b Fmoc-AA-OH (4.0 eq.), TBTU (4.0 eq.), DIEA (12.0 eq.), DMF, 1 h; c 2 mg/ml I2 in DMF, N2, 2 h; d HFIP:DCM (3:7), 10 ml, r.t., 2 h; e HATU (2.0 eq.), DIEA (2.2 eq.), 0 oC to r.t., 12 h; f TFA:TIPS:H2O (95:2.5:2.5), 30 min.
The inhibitory activity of these peptides towards six integrins (α5β1, αvβ3, αvβ5, αvβ6, αIIbβ3, and αvβ8) was assessed employing an ELISA with integrin-extracellular matrix competing experiments (see Table 1). As reference compounds we used Kessler’s c(RGDyK) (6) and the cyclic disulfide-bridged heptapeptide Eptifibatide (7) with specificities for αvβ3 and αIIβ3, respectively40,43,44,45,46,47. The experimental results showed that our prototypic bicyclic compound 5a already exhibited potent inhibitory against αvβ3 (IC50 < 0.51 nM) while also demonstrating significant inhibitory activity against αvβ5 and αIIbβ3 (IC50 = 13.2 ± 1.2 nM and 1.9 ± 0.3 nM), respectively. Notably, the contraction to a seven-membered ring in 5b did not enhance the selectivity of the bicyclic peptide for αvβ3 and αIIbβ3 integrins; instead, it resulted in a decrease in inhibitory activity toward αvβ3 (IC50 = 7.8 ± 0.4 nM). To further optimize both specificity and inhibition, we embarked on swapping the Cys/Trp positions and rearranging the amino acid sequence (Asn-Pro-Ala) within the A ring of the bicyclic peptide, leading to the synthesis of compounds 5c and 5d. As the results demonstrate, compound 5c displayed inhibitory activity against both αvβ3 and αIIbβ3 integrins, comparable to that of compound 5a. In contrast, peptide 5d also exhibited significantly enhanced selectivity for these two integrins while maintaining favorable affinity for αvβ3. These results underscore the importance of amino acids adjacent to the RGD sequence in modulating the selectivity of the bicyclic peptide ligands for integrins, in addition to the direct interaction with the RGD motif. Subsequently, in order to decrease the conformational rigidity of the peptides, we further modulated the bicyclic core structure by slightly increasing the individual ring sizes of the tryptathionine bridge (homoCys; 5e) and of the macrolactam (β-Ala; 5f and γ-aminobutyric acid (GABA); 5g). However, these modifications led to a marked reduction in inhibitory towards all six integrins, aligning with the literature’s assertion that a more rigid conformation typically enhances the ligand-receptor affinity48. Subsequently, we synthesized the bicyclic peptide 5h incorporating D-Cys in the tryptathionine motif, however, no significant improvement in its inhibitory activity for αvβ3 was observed. These findings collectively contribute to an understanding of the structural parameters that govern the binding efficacy and specificity of bicyclic peptide ligands to integrins.
Based on a “spatial screening”, Kessler and colleagues have elucidated that in the extended state of the RGD motif, an interatomic distance of 0.7–0.9 nm between the positively charged guanidinium group of Arg and the carboxy group of Asp is in support of a higher affinity for αIIbβ3 integrin. In contrast, when the binding motif adopts a more bent conformation, the ligand exhibits enhanced affinity for other integrin subtypes, such as αvβ3 and α5β149,50. Hence, manipulating the spatial separation between the guanidinium group of the RGD motif and the carboxy group of Asp can be a critical factor in modulating the binding affinity for distinct integrins, specifically αvβ3 and αIIbβ3. To explore this modulation further, we designed and synthesized two bicyclic peptides, 5i and 5j. Peptide 5i bearing L-homoArg maintained its inhibitory activity towards αIIbβ3 (Table 1) compared to 5 d. Notably, this modification resulted in a marked reduction in its inhibitory activity against αvβ3 as well as the other four integrin subtypes. In contrast, the incorporation of L-norArg into peptide 5j led to a minimal change in its inhibitory activity towards αvβ3 but induced a decrease in its inhibitory capacity for αIIbβ3. In order to obtain an idea on the impact of bicyclic peptides versus monocyclic peptides on the inhibitory activity, we synthesized 5k. Interestingly, this peptide exhibited notable activity towards integrins αvβ3, αvβ5, and αIIbβ3, with half-maximal inhibitory concentrations (IC50) of 0.3 ± 0.003 nM, 2.2 ± 0.2 nM, and 2.9 ± 0.4 nM, respectively. However, it did not show improved selectivity among these tested integrin subtypes. Accordingly, these results support that a bicyclic conformation is vital to the selectivity and affinity of binding αvβ3.
Summarizing the above results from integrin inhibition, the eight-membered bicyclic peptide 5j was found to possess distinct advantages in terms of both inhibitory activity and selectivity for the αvβ3 integrin. Furthermore, as a novel bicyclic peptide scaffold, 5j exhibited binding characteristics that are comparable or even superior to those of the previously described αvβ3-targeting monocyclic peptide, c(RGDyK) (6). Notably, the affinity of 5j for αvβ3, with an IC50 value of 0.7 ± 0.1 nM, is in the same range as that of cilengitide, a well-known αvβ3 inhibitor, which has an IC50 of 0.61 ± 0.06 nM40. This suggests that 5j may serve as a promising candidate for further development in the realm of integrin-targeting therapeutics.
Molecular docking
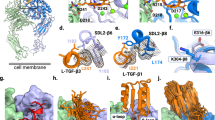
To further elucidate the influence of the guanidinium group on integrin binding activity, molecular docking (MD) studies were performed. Specifically, compound 5j was docked with the RGD binding sites of the αvβ3 (PDB: 1L5G) and αIIbβ3 (PDB: 8T2U) followed by binding pose optimization using MD simulations (see SI Methods). According to the modeling results, compound 5j interacts with both αvβ3 and αIIbβ3 in a similar manner, primarily by chelating the metal cation at the metal-ion-dependent adhesion site (MIDAS) via its Asp carboxylate group (see Fig. 2c, d). Additionally, the guanidinium group of 5j forms a cation-π interaction with the phenyl ring of Y178 in αvβ3 and F160 in αIIbβ3. A key distinction is that 5j forms a bidentate salt bridge with D218 in αvβ3 via its guanidino group, whereas this interaction is absent in its binding to αIIbβ3. In αIIbβ3, an acidic residue, D224, is located distally from the guanidinium group of 5j. However, in compound 5i, which possesses a longer homoarginine side chain, a salt bridge with D224 may be formed, potentially explaining its higher activity against αIIbβ3 compared to 5j. Furthermore, differences in hydrogen bonding patterns are observed in the interactions with αvβ3 and αIIbβ3. The backbone amide and carbonyl groups of the tryptophan moiety in 5j form two hydrogen bonds with D251 and N313 of integrin β3, respectively (see Fig. 2c). In its interaction with αIIbβ3, only the hydrogen bond with N313 is present. Nonetheless, the additional hydrogen bonds formed around the MIDAS site may compensate for this (see Fig. 2d), suggesting that variations in hydrogen bonding are unlikely to be the primary factor driving the high selectivity of 5j for αvβ3 over αIIbβ3. Overall, the unique interaction pattern of the guanidinium group of nor-Arg within the RGD binding domain of αvβ3 likely contributes to the nanomolar IC50 of 5j, reinforcing its potent and selective binding affinity (see Fig. 2a, b). Similarly, from our library, another selective αvβ3 inhibitor 5d, exhibits a comparable difference in the binding mode between the two integrins (see Fig. S2).
a Chemical structures of 5d, 5i, 5j, 6 c(RGDyK), 7 (Eptifibatide). b IC50 values and selectivity of peptides with various Arg-homologs (5d, 5i, 5j, 6, and 7) for integrin αvβ3 and αIIbβ3 (ratioa = αIIbβ3: αvβ3). MD simulations and molecular docking of the binding mode of peptide 5j with c αvβ3 (PDB: 1L5G) and d αIIbβ3 (PDB: 8T2U) integrin. Stick model: compound 5j (deep pink), protein side chains of the integrin binding pocket (green sticks, with their secondary structures illustrated as a cartoon). The interactions were visualized using ChimeraX.
Cy5-labeled compound 5j-K-Cy5 specifically binds integrin αvβ3 and is internalized by MDA-MB-231 cells
To evaluate the binding and uptake capabilities of the bicyclic peptide 5j on αvβ3-expressing cells, we labeled the peptide structure with the fluorescent dye Cy5. Since 5j inherently lacks additional conjugation sites, we replaced alanine (Ala) in 5j with an orthogonally protected lysine (Lys(Dde)), which facilitates the synthetic attachment of Cy5 in the sidechain. In contrast, Asp/Asn as an alternative attachment site is not ideal due to its unique capacity to be involved in turn structures (Asx turn)51,52. Molecular dynamics simulations reveal that the side-chain amide bond of Asn in 5j participates in intramolecular hydrogen bonding (see Fig. 2d), which most likely contributes to maintaining the spatial conformation of the bicyclic peptide. Along these lines Cy5-labeled 5j-K-Cy5 and 6-Cy5 (a validated αvβ3 ligand for tumor imaging/drug delivery as positive control22,43,44,53,54) were synthesized (Scheme S1), together with A20FMDV-Cy5 (specific to αvβ6; αvβ3-negative control) to evaluate target specificity of 5j55.
Three cell lines - MDA-MB-231 (17.03% αvβ3), A549 (13.60% αvβ3), and Huh-7 (<0.7% αvβ3/αvβ6) - were selected based on CCLE database analysis for functional assays. Flow cytometry validation (see Fig. 3a and S3) confirmed MDA-MB-231 as suitable αvβ3-positive and Huh-7 as an αvβ3-negative control cell lines, with αvβ6 expression consistently ≤0.7% across all three cell lines. The results of the cell binding experiments are shown in Fig. 3b, where red fluorescence indicates the Cy5 signal, and the blue fluorescence corresponds to the DAPI-stained nuclei. Following incubation with Cy5-labeled peptides at 4 °C for 1 h, significant binding of 5j-K-Cy5 to αvβ3-positive cells was observed. Similarly, the positive control compound S6-Cy5 exhibited comparable binding signals. In contrast, A20FMDV-Cy5, a peptide specifically binding to αvβ6, displayed negligible fluorescence signals on MDA-MB-231 cells. For Huh-7 cells, which are negative for both αvβ3 and αvβ6 expression, no significant signals were observed for any of the three Cy5-labeled peptides. These results confirm that the designed and synthesized bicyclic peptide 5j exhibits specific binding to integrin αvβ3.
a Expression of integrin αvβ3 and αvβ6 in MDA-MB-231 (αvβ3+, αvβ6−), A549 (αvβ3+, αvβ6−) and Huh-7 (αvβ3−, αvβ6−) cells. b Cell binding assay. MDA-MB-231 and Huh-7 cells were incubated with 5 μM of Cy5-labeled peptides (at 4 °C for 1 h). Cells were washed, fixed, labeled with DAPI, and imaged with confocal microscopy. c Cellular uptake and co-localization of Cy5-labeled peptides with lysosomes in MDA-MB-231 cells. d, e Co-localization curve of Cy5-labeled peptides with lysosome marker. Cells were pretreated with 5 μM Cy5-labeled peptides for 4 h, then fixed, labeled with DAPI and LAMP1. Fluorescence signals indicating the colocalization of Cy5-labeled peptide (red) with the lysosome marker (LysoTracker, green) in MDA-MB-231 cells (blue is the nucleus, green is the lysosome, and red is the fluorescence of Cy5, the scale is 10 μm).
To verify whether the bicyclic peptide 5j could serve as a targeting peptide for drug delivery in the treatment of tumors with high αvβ3 expression, we conducted cellular uptake and lysosome co-localization experiments using MDA-MB-231 cells (see Fig. 3c). Following co-incubation of Cy5-labeled peptides with cells at 37 °C for 4 h, analysis by confocal fluorescence microscopy demonstrated that the peptide 5j-K-Cy5 exhibited a marked enhancement in cellular internalization when compared to the reference compound 6-Cy5. Additionally, the Cy5 fluorescence signal displayed a high degree of co-localization with the green fluorescence signal of LAMP1 (see Fig. 3d), a validated lysosomal marker, indicating that 5j-K-Cy5 is indeed internalized by the cells via the lysosomal pathway. Since cathepsin B is highly expressed in the lysosomal environment, it is capable of efficiently cleaving the peptide-payload conjugates, thereby facilitating the release of therapeutic compounds within target cells56,57,58. These cumulative observations not only demonstrate the effective internalization and processing of 5j-K-Cy5 within lysosomes but also provide robust support for the strategic use of αvβ3-targeted peptide-drug conjugates (PDCs) as a modality for tumor-specific drug delivery.
Evaluation of the cytotoxicity of αvβ3-selective peptide-drug conjugates against CCK-8 cell lines
Before conducting the experiment, the in vitro plasma stability of the side chain-deprotected bicyclic peptide 5j-K was evaluated. The results demonstrated that the tested bicyclic peptide exhibited sufficient stability in rat plasma (t1/2 > 11 h, see Fig. S1). Then, the microtubule inhibitor monomethyl auristatin E (MMAE) was selected as the toxin payload, due to the presence of extensive data on both antibody drug conjugates (ADCs) and PDCs59,60. Accordingly, four peptide drug conjugates (PDCs), namely P1, P2, P3, and P4, were synthesized according to methods outlined in Scheme S4-6 (see Fig. 4a). All four conjugates incorporate the 6-azidohexanamido-VA-PABC-MMAE linker-drug module, which was conjugated to the peptide via a copper-catalyzed azide-alkyne cycloaddition (CuAAC) reaction. Compounds P1 and P2 utilize 5j-K as the integrin-targeting peptide, whereas P3 employs deprotected compound 8 (non-binding AGD motif) as the negative control, and compound P4 utilizes compound 6 (c(RGDyK)) as the positive control. Compounds P1, P3, and P4 feature an octaethylene glycol (PEG8) spacer, while P2 utilizes a C8 aliphatic chain as a spacer. Subsequently, the cytotoxicity of the four conjugates against the human tri-negative breast cancer cell line MDA-MB-231 and the human non-small-cell lung cancer cell line A54944 was evaluated using a standard CCK-8 cell viability assay (see Table 2, and Fig. S4a, b). Following a 72 h incubation, the CCK-8 assay results showed that the naked peptides 6 and 5j-K had no significant toxic effects (IC50 > 5 μM), whereas the drug MMAE alone exhibited potent cytotoxicity (IC50 < 1.37 nM). The positive control P4 demonstrated IC50 values of 58.5 ± 3.9 nM in MDA-MB-231 cells and 37.9 ± 8.4 nM in A549 cells. Both P1 and P2 exhibited enhanced efficacy relative to P4, with IC50 values in MDA-MB-231 cells of 41.3 ± 4.1 nM and 31.23 ± 4.3 nM, respectively, and in A549 cells, of 31.3 ± 4.3 nM and 18.5 ± 4.0 nM. The negative control P3 exhibited comparatively lower anti-proliferative activity, with IC50 values of 72.2 ± 7.9 nM in MDA-MB-231 cells and 94.1 ± 25.9 nM in A549 cells.
a Chemical structures of P1, P2, P3, P4. b–e A549 tumor-bearing mice (n = 7) were injected intravenously with 1.0 mg kg−1 P1 and 0.29 mg kg−1 MMAE every two days for three weeks. PBS was used as a vehicle. Tumor volumes and body weight were recorded. f Liver and kidney functions were evaluated at the end of the treatment. g Representative images of Ki67 staining of tumor sections in different groups. Bar = 50 μm. Statistical significances were analyzed by one-way ANOVA with Tukey’s multiple comparisons test. All the data were plotted as mean ± SD. (*): p < 0.05, (***): p < 0.001.
Meanwhile, we knocked down the mRNA associated with αvβ3 expression in A549 cells (A549-KD (αvβ3-)) through transient transfection and verified the successful knockdown of αvβ3 protein via Western blot (see Fig. S4e). In parallel, A549 cells treated with a non-targeting scrambled siRNA (si-NC) were used as the control (A549-si-NC (αvβ3+)). Subsequently, in vitro cell proliferation inhibition experiments demonstrated that, for all three conjugates, the IC50 value in A549-KD (αvβ3-) cells increased by about 2-fold compared to A549-si-NC (αvβ3+) cells (see Table 3). The IC50 of 5j-K conjugates P1 and P2 in A549-KD (αvβ3-) was 1.6-fold and 1.9-fold higher than A549-si-NC (αvβ3+), while for the positive control c(RGDyK) conjugate P4, the IC50 was 1.7-fold higher. These findings indicate that P1 and P4, to some extent, enhance cellular internalization and cytotoxicity through specific recognition of αvβ3 by peptide 5j-K and c(RGDyK).
To further elucidate the importance of targeted peptides and assess the binding capability of conjugates to integrin receptors on cell surfaces, we performed a “kiss and run” experiment61,62. The results demonstrate that conjugate P1 exhibited greater antiproliferative activity than the negative-control conjugate P3 (see Table S1). Moreover, the relative potency (RP) of the conjugates in the two tested cell lines correlates well with αvβ3 integrin expression levels, thereby confirming the specific targeting ability of bicyclic peptide 5j toward αvβ3 integrin. These findings indicate that the conjugations of MMAE to the peptide substantially reduce its cytotoxicity, likely as a result of decreased non-specific cellular uptake of MMAE. While P1 and P2 did not exhibit a marked difference in cytotoxicity compared to P4, their significant inhibitory capacity underscores the therapeutic potential of PDCs utilizing the 5j-K peptide as a targeting-αvβ3 ligand for tumor therapy.
In vivo antitumor activity of peptide-drug conjugates
To assess the in vivo antitumor activity of the PDCs, we employed an A549 xenograft mouse model (BALB/c) for efficacy studies. In preliminary CCK-8 cellular assays, P2 exhibited stronger proliferation inhibition compared to 10a. However, the solubility of P2 in a 10% DMSO-PBS solution was compromised owing to the presence of a lipid chain, leading us to select P1 for the in vivo antitumor efficacy assessment. We first evaluated two doses of P1 (2 mg kg−1 and 0.5 mg kg−1) against a control group receiving 10% DMSO in PBS (pH 7.4) (n = 6 per group). Treatment initiation occurred when tumors reached a volume of 100–150 mm3, with intravenous tail vein injections administered biologically. Tumor volume and mouse body weight were monitored prior to each injection. Notably, mice in the 2.0 mg kg−1 dose group exhibited a decrease in body weight by the third day post-administration, which intensified by the fifth and seventh days, culminating in mortality. This indicated that the 2.0 mg kg−1 dose was excessively high and inappropriate for continued in vivo experimentation (see Fig. S5b, c). In contrast, the 0.5 mg kg−1 dose group displayed minimal to no significant change in body weight, and even showed a slight increase, relative to the PBS control group. Moreover, this lower dose effectively suppressed tumor growth, with a significantly reduced tumor growth rate compared to the control (p < 0.05) (see Fig. S5b–d).
To further delineate the in vivo antitumor efficacy and targeting specificity of P1, we conducted a long-term study using a 1.0 mg kg−1 dose of P1 and an equimolar dose of MMAE (0.29 mg kg−1), with the 10% DMSO PBS solution serving as a control (n = 7 per group). Over the course of a 21-day treatment regimen, mice administered with P1 showed no significant weight loss or side effects, in marked contrast to those receiving an equimolar dose of MMAE (see Fig. 4a–d). The tumor size in the group treated with 1.0 mg kg−1 P1 was significantly controlled (p < 0.001) compared to the vehicle control group. There was also no significant difference in liver and kidney functions between 1.0 mg kg−1 P1 and the vehicle control group (see Fig. 4f). Tissue distribution studies demonstrated significant enrichment of the payload drug MMAE in tumors 4 h after administration (see Fig. S7). Moreover, IHC staining of Ki67 indicated a significantly reduced tumor proliferation in the treatment group (see Fig. 4g). Although the MMAE control group displayed comparable inhibitory effects early in the treatment, prolonged exposure resulted in significant weight loss, indicating high toxicity. These findings suggest that P1 is an effective inhibitor of αvβ3-positive tumor growth in the mouse xenograft model, with reduced side effects compared to free MMAE. Finally, we assessed the antitumor activity of conjugate P4 with the targeting peptide c(RGDyK) in A549 tumor-bearing mice (n = 6). Over the course of a 21-day treatment regimen, mice administered with 0.9 mg kg−1 P4 showed no significant weight loss or side effects (see Fig. S6a–d). The tumor progression in the group treated with 0.9 mg kg−1 P4 was significantly suppressed (p < 0.001) compared to the vehicle control group, but the antitumor effect is not superior to that of an equivalent amount of P1. The tumor growth inhibition (TGI) results indicate that P1 exhibits a comparable yet slightly superior tumor-suppressive effect (see Fig. S6e). These results underscore the specific targeting efficacy of the bicyclic peptide 5j to αvβ3 integrin and reinforce its potential value as an αvβ3-targeting ligand for drug delivery in tumor therapy.
Conclusion
In conclusion, our study has characterized a novel bicyclic peptide 5j, which exhibits high affinity and specificity for the integrin αvβ3 receptor. The molecular structure of 5j incorporates the non-canonical sequence norArg-Gly-Asp, which serves as a critical determinant in its discriminative selectivity towards αvβ3 over the αIIbβ3. In light of the molecule’s capability to be internalized by cells via the endosome-lysosome pathway, we have successfully synthesized four peptide-drug conjugates, P1, P2, P3 and P2. Our in vivo evaluation using a mouse tumor xenograft model has demonstrated that P1 induced potent tumor suppression effects while significantly reducing systemic side effects compared to free MMAE alone. These comparative efficacy and safety profiles highlight the advantages of our targeted peptide-drug conjugate strategy. The findings of this research not only validate the therapeutic potential of 5j as a novel bicyclic peptide for αvβ3-targeted tumor therapy but also provide critical theoretical and practical insights into the development of αvβ3-targeting bicyclic peptides.
Methods
Chemistry
Chemical materials
The analytical reagents were purchased from Energy Chemical and Fisher without further purification. 2-CTC resin, TBTU, HATU, and Fmoc-protected amino acids were purchased from Bide Pharmatech Ltd., DIEA, TFA, and HFIP were purchased from MACKLIN. Purity (>95%) confirming and reaction monitoring were carried out on analytical UHPLC (ThermoFisher Scientific Vanquish) using an ODS column (Welch Ultimate UHPLC XB-C18 column, 2.1 mm × 100 mm, 1.8 μm). Final compounds were purified on preparative HPLC SHIMADZU LC-20AP using an ODS column (Welch Ultimate AQ-C18 column, 21.2 mm × 250 mm, 5 μm). HR ESI–MS data were acquired using LCMS (HPLC SHIMADZU LC-40D, MS Sciex ZenoT0FTM 7600, Column SynergiTM 4 μm Fusion-RP 50 × 2 mm). 1H NMR spectra were recorded by Bruker Advance (400 MHz) using TMS as an internal Standard and DMSO-d6 as solvents, and the chemical shift was reported in parts per million (ppm). Purity assessment of synthesized compounds was performed using HPLC, NMR (see Supplementary Information and Supplementary Data 3).
The synthesis of Cy5-labeled peptides
The Dde-group-deprotected peptide S3 (15 mg, 0.01 mmol) was dissolved in DMF (5 mL), and Cy5-sulfo (7.9 mg, 0.012 mmol) was conjugated using the coupling agent HATU (2.0 eq). After reacting at room temperature for 2.0 hours, the DMF solvent was removed by rotary evaporation (See SI Scheme S1). Then a mixed solvent of TFA/TIS/H2O (95:2.5:2.5) was added to remove the remaining sidechain protecting groups at room temperature for 0.5 h. The product was precipitated with ether and purified to obtain the Cy5-labeled compound 5j-K-Cy5 as a blue solid 10.6 mg for 67.3% yield (purity > 96%). HR ESI–MS: exact m/z calculated for C73H95N16O18S3+ [M + H]+ = 1579.6167, found 1579.6125 (see Fig. S14).
Following the general protocol for solid-phase peptide synthesis, a side-chain fully protected peptide was synthesized via solid-phase peptide synthesis. Subsequently, the Dde group was removed by treating the peptide with 2% hydrazine hydrate in DMF for 10 min at room temperature (r.t.). The mixture was then concentrated and purified by preparative HPLC to afford the target product S1 as a white solid. Next, the same method was employed to get the compound 6-Cy5 as a blue solid 7.7 mg (See SI Scheme S1) for 61.2% yield (purity > 96%). HR ESI–MS: exact m/z calculated for C60H80N11O15S2+ [M + H]+ = 1258.5271, found 1258.5216 (see Fig. S12).
The synthesis of peptide-drug conjugates P1, P2, P3 and P4
Fmoc-L-Lys(Dde)-OH was used to replace Fmoc-L-Ala-OH in compound 5j. Following the general protocol provided in the Supporting Information, a side-chain fully protected bicyclic peptide was synthesized via solid-phase peptide synthesis. Subsequently, the Dde group was removed by treating the peptide with 2% hydrazine hydrate in DMF for 10 min at room temperature (r.t.). The mixture was then concentrated and purified by preparative HPLC to afford the target product as a white solid. Next, the Dde-group-deprotected peptide S3 (30 mg, 0.02 mmol) was dissolved in DMF (5 mL), followed by the addition of Propargyl-PEG8-acid (S10, 1.5 equiv), HATU (2.0 equiv), and DIPEA (2.2 equiv). The reaction mixture was stirred at rt for 2 h. Afterward, the DMF was removed under reduced pressure, and the crude product was treated with a mixture of TFA/TIS/H2O (95:2.5:2.5, v/v) at rt for 0.5 h. Following concentration, ether (30 mL) was added to precipitate the product, which was subsequently collected by centrifugation. Subsequently, the precipitate was redissolved in DMF/H2O (2:1, v/v). Then, 6-azidohexanamido-VA-PABC-MMAE S9 (35.5 mg, 0.03 mmol), CuSO4·5H2O (20 mg, 0.08 mmol), and sodium ascorbate (31.7 mg, 0.16 mmol) were added sequentially. The reaction mixture was stirred at room temperature (r.t.) for 1 h. The crude product was subsequently purified by preparative HPLC and then lyophilized to afford the final desired product P1 as a white powder with a yield of 40.2 mg (79.3%). and a purity of 98%. HR ESI–MS: exact m/z calculated for C121H188N25O32S+ (M + H)+: 2535.3567, found 2535.3541 (see SI Scheme S3, Figure S17-18).
The synthesis procedure of P2–P4 follows a methodology similar to that described previously. For detailed experimental conditions and synthetic parameters, the reader is referred to the supporting literature (See Scheme S4–S6).
Biological assays
Materials
Dulbecco’s phosphate-buffered saline (DPBS, pH 7.4), phosphate-buffered saline (PBS, pH 7.4), DMEM basic (1×), DMEM/F-12 (1:1) basic (1×), fetal bovine serum (FBS), and trypsin/EDTA were obtained from Gibco™. ECM proteins (ECM proteins: Recombinant Human Vitronectin Protein, catalog no. 2308-VN-050; Recombinant Human LAP (TGF-beta 1) Protein, catalog no. 246-LP-025; Recombinant Human fibronectin, catalog no. 4305-FNB-200), human integrin proteins (Recombinant Human Integrin αvβ3, catalog no. 3050-AV-050; Recombinant Human Integrin αvβ5, catalog no. 2528-AV-050; Recombinant Human Integrin αvβ6, catalog no. 3817-AV-050; Recombinant Human Integrin αvβ8, catalog no. 4135-AV-050; Recombinant Human Integrin αIIbβ3, catalog no. 7148-A2-025; Recombinant Human Integrin α5β1, catalog no. 3230-A5B-050), detection antibodies (Human Integrin alpha V/CD51 Biotinylated Antibody, catalog no. BAF1219; Human Integrin beta 3/CD61 Biotinylated Antibody, catalog no. BAF2266; Human Integrin beta 1/CD29 Biotinylated Antibody, BAF1778) and Streptavidin-HRP (catalog no. DY998) were purchased from R&D. Cell counting kit-8 (CCK-8, catalog no. C0039) and 4,6-diamidino-2-phenylindole DAPI (catalog no. C1002) were obtained from Beyotime. Anti-LAMP1 antibody (catalog no. ab278043) was purchased from Abcam. Cy3-conjugated AffiniPure™ Goat Anti-Rabbit IgG (H + L) (minimal cross-reaction to Human, Mouse, and Rat Serum Proteins) was purchased from Jackson (catalog no. 111-165-144). Ki67 Cell Proliferation Kit (IHC) (catalog no. E607235-0100) was obtained from Sangon Biotech (Shanghai, China). DMSO (>99.9%, HPLC) was purchased from MACKLIN. All aqueous solutions were prepared using Milli-Q purified water (18.2 MΩ). Flat bottom 96-well cell culture plates were purchased from Corning Incorporated Costar. Human breast cancer cell line MDA-MB-231, human hepatoma cell line Huh-7 cells and human lung carcinoma cell line A549 cells were initially purchased from the Chinese Academy of Science Shanghai Institute of Cell Bank (Shanghai, China). Male nude mice (SPF, 5–6 weeks) were purchased from GemPharmatech (Guangdong, China). The absorption was recorded by a microplate spectrophotometer (TECAN, Infinite E Plex). Confocal fluorescence imaging was performed using the confocal laser scanning microscope (Leica STELLARIS). The biochemical indicators (ALT, AST and UREA) testing was performed on a fully automated biochemical analyzer (Mindray, BS-240VET).
Integrin binding assay
The activity and selectivity of cyclic peptide integrin ligands were assessed through an integrin-extracellular matrix interaction assay based on ELISA detection, following the previously reported method40. Using 6 c(RGDyK) as a reference reagent, other reagents, including ECM proteins, human integrin proteins, and detection antibodies. Initially, ECM proteins were dissolved in carbonate buffer (15 mM Na2CO3, 35 mM NaHCO3, pH 9.6), then added to a flat-bottom 96-well plate at 100 μL per well, incubated overnight at 4 °C. The plate was then washed with PBS-T buffer (phosphate-buffered saline/Tween20, 137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 2 mM KH2PO4, 0.01% Tween20, pH 7.4; 3 × 200 μL), followed by blocking with TSB buffer (Tris-saline/BSA buffer, 150 μL/well, 20 mM Tris-HCl, 150 mM NaCl, 1 mM CaCl2, 1 mM MgCl2, 1 mM MnCl2, pH 7.5, 1% BSA) at room temperature for 1 h. Meanwhile, serial dilutions of the reference reagent and test compounds were prepared in a separate 96-well plate, starting from 1 μM and following a 3-fold gradient dilution across 10 concentration points. After washing the assay plate with PBS-T (200 μL) three times, 50 μL of each dilution was transferred to each well from B-G of the assay plate. Well A was filled with 100 μL of TSB solution as a blank, and well H was filled with 50 μL of TSB buffer. Then 50 μL of human integrin in TSB buffer was added to wells H-B, and the plate was incubated at room temperature for 1 h. The detection antibody was added after washing the plate with PBS-T buffer three times (100 μL per well). Following a 1 h incubation at room temperature, the plate was washed three times with PBS-T buffer. Then Streptavidin-HRP was then added (diluted 1:10, 100 μL per well) and incubated at room temperature for 1 h, after which the plate was washed three times with PBS-T buffer. Then the substrate SeramunBlau was quickly added to the plate (50 μL per well) and incubated in the dark for 5 min. Then 50 μL of 3 M H2SO4 stop solution was added to each well, and the absorbance was measured at 450 nm using a microplate reader (TECAN, Infinite E Plex). The IC50 for each compound was tested in duplicate and analyzed using Origin 2025 software.
Cell culture
MDA-MB-231 cells and Huh-7 cells were cultured in Dulbecco’s Modified Eagle Medium (DMEM), and A549 cells were cultured in DMEM/F-12 (1:1). The medium was supplemented with 10% fetal bovine serum (FBS) and 1% penicillin-streptomycin solution. The cells were grown in sterile cell culture flasks and maintained in 5% CO2 in a humidified incubator at 37 °C.
Flow cytometry analysis for cell surface integrin proteins
MDA-MB-231, A549 and Huh-7 cells were added to each centrifuge tube at a density of 2 × 105 for αvβ3 determination and 5 × 105 for αvβ6 determination. Then, 1 mL buffer (2% FBS/98% DPBS) was added to disperse cells and centrifuged (400 × g, 5 min). After discarding the supernatant, antibody (αvβ3, catalog no. 304415, Biolegend; αvβ6, catalog no. FAB4155A, R&D) was added and incubated in the dark at 0 °C (αvβ3 for 20 min, αvβ6 for 40 min). Then cells were washed with pre-chilled buffer (2 mL) twice. Dead cells were stained with PE Annexin V Apoptosis Detection Kit I (catalog no. 51-68981E, BD) for 20 min. Finally, the assay was performed on a flow cytometer (CytoFLEX LX).
Confocal microscopy of Cy5-labeled peptides binding to αvβ3-expressing cells
MDA-MB-231 (1.8 × 104 cells per well) and Huh-7 (1.6 × 104 cells per well) cells were seeded in a flat-bottom 96-well plate with transparent glass bottoms and cultured in DMEM high glucose medium for 48 h, respectively. After microscopic observation confirmed that the cells had reached an appropriate density, the culture plate was incubated at 4 °C for 20 min. Meanwhile, Cy5-labeled compounds 6-Cy5 and 5j-K-Cy5 were prepared as 5 μM test solutions in DMEM high-glucose medium and subsequently maintained in the dark at 4 °C for the same duration. After removing the old medium, the test solutions were added to the corresponding wells, then the cells were incubated with 5 μM of 6-Cy5 and 5j-K-Cy5 at 4 °C in the dark for 1 h. Following incubation, the test solutions were removed, and the cells were carefully washed twice with pre-cooled PBS (1×). Subsequently, the cells were fixed with 4% paraformaldehyde solution at room temperature for 20 min. After discarding the fixative, the cells were washed twice with PBS (1×). Subsequently, 100 μL DAPI solution (1:5000) was added to each well to stain the nuclei at room temperature for 20 min. After washing twice with PBS (1×), 55 μL of DAPI-free antifade mounting medium was added to seal the plates. Finally, the plates were observed and imaged under an inverted confocal fluorescence microscope (Leica STELLARIS).
Confocal microscopy of internalization of Cy5-labeled peptides into the αvβ3-expressing cell
MDA-MB-231 cells (1.8 × 104 cells per well) were seeded in a flat-bottom 96-well plate with transparent glass bottoms and cultured in DMEM high glucose medium for 48 h. After discarding the medium, the cells were incubated with 5 μM of pre-prepared Cy5-labeled peptide solutions in DMEM at room temperature for 4 h. Subsequently, discarding the solutions, the cells were washed twice with PBS (1×), and the cells were fixed with 4% paraformaldehyde for 20 min. After that, washed twice and permeabilized with 0.15% Triton X-100 for 15 min. Then, the cells were blocked with 10% goat serum at room temperature for 1 h and incubated with anti-LAMP1 primary antibody (1:300) at 4 °C in the dark overnight. After incubation, cells were returned to room temperature and washed three times with PBS. Next, Cy3-conjugated AffiniPure™ goat anti-rabbit (1:300) secondary antibody was incubated with the cells at room temperature for 1 h. After washing, the nuclei were stained with DAPI. Finally, the fluorescence staining was observed and photographed with an inverted confocal microscope (Leica STELLARIS).
Measurement of cytotoxicity by CCK-8 assay
The toxicity determination CCK-8 experiment was carried in 96-well plates. MDA-MB-231 (3.0 × 103 cells per well) and A549 cells (3.0 × 103 cells per well) were seeded in 96-well plates and incubated for 24 h. Then the medium was discarded and treated with 5j-K, 6, P1, P2, P3, or P4 in various concentrations (5.0 μM, 2.5 μM, 1.0 μM, 333.33 nM, 111.13 nM, 37.04 nM, 12.35 nM, 4.12 nM, 1.37 nM) for another 72 h. Following with conventional CCK-8 treatment. After 1 h, the formazan dye was quantified using a microplate reader to assess absorbance at 450 nm (measurement wavelength) and 630 nm (reference wavelength). Non-treated cells were used as controls. The percentage viability of cells was estimated as the percentage fluorescence intensity in the experimental wells relative to that in the blank and negative control wells. Each experiment was repeated three times.
RNA interference for αvβ3 and cell viability detection
A549 cells were seeded in 96-well plates at a density of 500 cells per well and allowed to adhere overnight in high-glucose DMEM supplemented with 10% FBS without antibiotics. Small interfering RNAs (siRNAs) targeting human ITGAV (sequence: 5’-3’ CCAUGUAGAUCACAAGAUAdTdT) and ITGB3 (sequence: 5′-3′ GCUACAGUCUGUGAUGAAAdTdT) were used at 1.2 pmol per well each (total 2.4 pmol/well). A non-targeting siRNA negative control (siRNA NC, same dosage; GenScript Biotech Corporation, Order No. C717J577G0) served as the control. The siRNA mixtures were complexed with 0.2 μL/well Lipofectamine RNAiMAX Reagent (Invitrogen, REF: 13778100) in OPTI-MEM (Gibco, catalog no. 31985-062) and incubated at room temperature for 20 min before being added to the cells. At 6 h post-transfection, the medium was replaced with high-glucose DMEM containing 10% FBS and 1% penicillin-streptomycin (P/S). Test drugs were administered at 48 h post-transfection, and after 72 h of co-incubation, cell viability was assessed using the CCK-8 assay (Beyotime, catalog no. C0039).
Western blot analysis for αvβ3 knockdown verification
Parallel plates of A549 cells transfected for 48 h were lysed with strong RIPA lysis buffer (Beyotime, Catalog No. P0013B). Proteins (30 µg) were separated by 4–12% SDS-PAGE (ACE Biotechnology, Cat. ET15412LGel) and transferred to PVDF membranes using 10× WB Transfer Buffer (Solarbio, Cat. No. D1060). Membranes were blocked with 5% skim milk in TBST (Servicebio, Cat: G0001-2L) for 2 h. Primary antibodies were diluted in Beyotime Western Primary Antibody Dilution Buffer (Beyotime, Catalog No. P0023A-500ml): rabbit anti-ITGB3 (ABclonal, A19073, 1:1000) or mouse anti-GAPDH (Santa Cruz, sc-32233, 1:2000), and membranes were incubated with these antibodies overnight at 4 °C. Secondary antibodies (Invitrogen, 31460 or 31430, 1:5000) were diluted in Beyotime Western Secondary Antibody Dilution Buffer (Beyotime, Catalog No. P0023D-500ml) and incubated with membranes for 1.5 h at room temperature. Bands were visualized using Clarity™ Western ECL Substrate (Bio-Rad, Cat. #170-5061) and imaged with a Tanon 5200 imaging system. Band intensities were quantified using Gel-Pro Analyzer 4 software, and ITGB3 signal was normalized to GAPDH.
In vivo assay
All animal experiments were carried out using 6-week-old male nude mice weighing 20–25 g purchased from GemPharmatech (Guangdong, China). These mice were maintained in a specific pathogen-free environment with free access to water and food. Experiments were approved by the Ethics Committee for Animal Experimentation of Greater Bay Area Institute of Precision Medicine (Guangzhou) in compliance with relevant laws and institutional rules. To establish xenograft models, A549 cells were injected (1 × 107 cells/200 μL) into the right flank of athymic nude mice. Mice were randomly divided into groups (n = 6/7) after the tumors had reached a size of 100–150 mm3. Drug solutions (100 μL each) were administered through the tail vein once every 2 days for a total of 10 injections (21 days): 2.0 mg kg−1, 1.0 mg kg−1 and 0.5 mg kg−1 per injection of P1, 0.29 mg kg−1 per injection of MMAE, and PBS as a control. The body weights and tumor sizes of each mouse were measured before drug injections. And tumor volume was estimated using the following equation: TV (mm3) = d2D/2, where d and D are the shortest and the longest diameter, respectively63. Tumors were dissected, measured, and photographed on day 21 after mice were sacrificed by cervical dislocation. Cell proliferation was determined by immunohistory chemical staining of Ki67 according to the Ki67 Cell Proliferation Kit (IHC) (E607235, Sangon Biotech, China). Image acquisition was performed using Nikon ECLIPSE Ni-U microscopes with 40× objective. The percentage of Ki67 positive cells was analyzed by ImageJ (NIH). The biological activity of P1 and the other substances was compared using one-way ANOVA. Statistical analyses were performed using Origin 2025. and p < 0.05 was considered significant. Error bars in the figures indicate standard deviations of the mean.
Plasma stability assay
Firstly, internal standard testosterone was chosen to be dissolved in ACN to be a concentration of 500 nM. Side chain-deprotected bicyclic peptide 5j-K (1.51 mg) was dissolved in 1.4 mL of DMSO to be a stock solution, then 10 μL stock solution was diluted using 190 μL MeOH, then 2.5 μL dilution was added to rat plasma (pH 7.5) at 37 °C to give final concentration for bicyclic peptide of 1 μM. The plasma was incubated for 24 h at 37 °C and 600 rpm. At 0, 0.5, 1, 2, 4, 6, 8, and 24 h, 25 μL of plasma was withdrawn and precipitated with 50 μL of cold quenching solution consisting of ACN with a final concentration of 333 nM testosterone as an internal standard. The samples were stored at 4 °C and centrifuged (12,000 × g, 4 °C, 5 min). The supernatant was analyzed by LC − MS (injection volume, 5 μL). The degradation was determined by comparing the peak area of the MS signal of the internal standard with the tested compound. And the percentage of the remaining amounts of 5j-K at each time interval was calculated based on the 0 h of peak area.
Liver and kidney functions
Blood was collected from the mice before euthanasia, then centrifuged to obtain plasma for measuring biochemical indicators of liver and kidney function. The biochemical tests (ALT, AST, and UREA) were performed on a biochemical analyzer (Mindray, BS-240VET) using ALT Kit (IFCC method) (Mindray, Lot 140124026), AST Kit (IFCC method) (Mindray, Lot 140224024), and UREA Kit (Urease-GLDH, UV method) (Mindray, Lot 141325001), respectively.
Tissue distribution
For tissue distribution tests, xenografts with tumors in the flank were prepared as mentioned previously. The compound solution of P1 (1 mg kg−1) was administered into mice (n = 3) by intravenous tail vein injection (100 μL). Blood was collected after 4 h of injection, then the mice were euthanized, and the organs (tumor, brain, heart, liver, lung, kidney, stomach, and pancreas) were collected and kept in a deep freezer until further use. The weight of each dissected tissue was measured, and PBS at a ratio of 1:3 (tissue weight: PBS volume) and ultrasonically homogenized to prepare a homogenate. The resulting homogenous solution of 25 μL was diluted with 50 μL acetonitrile solution containing internal standard testosterone, vortexed, centrifuged at 12,000 × g for 5 min, and collected the supernatant,then injected into LC-MS. The amount of payload MMAE in tissue was determined by LC-MS chromatogram equipped with a UV detector. The operation conditions of HPLC were the same as used in the calibration curve (gradient: 0.1–3.1 min, 5–95% ACN, 3.1–5.0 min, 95% ACN, 5.0–5.5 min, 95–5% ACN, 5.5–6.5 min, 5% ACN, 0.4 ml/min, equipped Phenomenex Kinetex® 2.6 μM F5 100 Å 50 × 2.1 mm column). The AUCs of each sample were obtained in triplicate. All animal tests were conducted in the Ethics Committee for Animal Experimentation of Greater Bay Area Institute of Precision Medicine (Guangzhou) in compliance with relevant laws and institutional rules.
Kiss-and-run experiment with conjugates P1 and P3
MDA-MB-231 (3.0 × 103 cells per well) and A549 cells (3.0 × 103 cells per well) were seeded in 96-well plates and incubated for 24 h. Then the medium was discarded and treated with P1 or P3 in various concentrations (40 μM, 20 μM, 10 μM, 5 μM, 2.5 μM, 1.25 μM, 0.625 μM, 0.3125 μM) for 1 h at 37 °C, followed by a washout of the treatment and a further incubation of the cells up to 72 h. Then a conventional CCK-8 treatment followed. After 1 h, the formazan dye was quantified using a microplate reader to assess absorbance at 450 nm (measurement wavelength) and 630 nm (reference wavelength). Non-treated cells were used as controls. The percentage viability of cells was estimated as the percentage fluorescence intensity in the experimental wells relative to that in the blank and negative control wells. Each experiment was repeated three times.
References
Rentero Rebollo, I. & Heinis, C. Phage selection of bicyclic peptides. Methods 60, 46–54 (2013).
Chung, B. K. & Yudin, A. K. Disulfide-bridged peptide macrobicycles from nature. Org. Biomol. Chem. 13, 8768–8779 (2015).
Rhodes, C. A. & Pei, D. Bicyclic peptides as next-generation therapeutics. Chemistry 23, 12690–12703 (2017).
Feng, D. et al. Current development of bicyclic peptides. Chin. Chem. Lett. 34, 108026 (2023).
Ullrich, S. & Nitsche, C. Bicyclic peptides: paving the road for therapeutics of the future. Pept. Sci. 116, e24326 (2023).
Thombare, V. J. & Hutton, C. A. Bridged bicyclic peptides: structure and function. Pept. Sci. 110, e24057 (2018).
Ahangarzadeh, S. et al. Bicyclic peptides: types, synthesis and applications. Drug Discov. Today 24, 1311–1319 (2019).
Heinis, C. & Winter, G. Encoded libraries of chemically modified peptides. Curr. Opin. Chem. Biol. 26, 89–98 (2015).
Danhier, F., Le Breton, A. & Preat, V. RGD-based strategies to target alpha(v) beta(3) integrin in cancer therapy and diagnosis. Mol. Pharm. 9, 2961–2973 (2012).
Bledzka, K., Smyth, S. S. & Plow, E. F. Integrin alphaIIbbeta3: from discovery to efficacious therapeutic target. Circ. Res. 112, 1189–1200 (2013).
Gu, Y. et al. The challenges and opportunities of alphavbeta3-based therapeutics in cancer: From bench to clinical trials. Pharm. Res. 189, 106694 (2023).
He, S. et al. Enhanced tumor targeting and penetration of proteolysis-targeting chimeras through iRGD peptide conjugation: a strategy for precise protein degradation in breast cancer. J. Med. Chem. 66, 16828–16842 (2023).
Zheng, J. et al. Bifunctional compounds as molecular degraders for integrin-facilitated targeted protein degradation. J. Am. Chem. Soc. 144, 21831–21836 (2022).
Montenegro, C. F. et al. Blocking alphavbeta3 integrin by a recombinant RGD disintegrin impairs VEGF signaling in endothelial cells. Biochimie 94, 1812–1820 (2012).
Liu, H. et al. The role of integrin alphavbeta3 in biphasic calcium phosphate ceramics mediated M2 Macrophage polarization and the resultant osteoinduction. Biomaterials 304, 122406 (2024).
Dechantsreiter, M. A. et al. N-Methylated cyclic RGD peptides as highly active and selective alpha(V)beta(3) integrin antagonists. J. Med. Chem. 42, 3033–3040 (1999).
Mas-Moruno, C., Rechenmacher, F. & Kessler, H. Cilengitide: the first anti-angiogenic small molecule drug candidate design, synthesis and clinical evaluation. Anticancer Agents Med. Chem. 10, 753–768 (2010).
Stupp, R. et al. Cilengitide combined with standard treatment for patients with newly diagnosed glioblastoma with methylated MGMT promoter (CENTRIC EORTC 26071-22072 study): a multicentre, randomised, open-label, phase 3 trial. Lancet Oncol. 15, 1100–1108 (2014).
Nabors, L. B. et al. Two cilengitide regimens in combination with standard treatment for patients with newly diagnosed glioblastoma and unmethylated MGMT gene promoter: results of the open-label, controlled, randomized phase II CORE study. Neuro Oncol. 17, 708–717 (2015).
Kondo, N. et al. Radioiodinated bicyclic RGD peptide derivatives for enhanced tumor accumulation. Pharmaceuticals 18, 549 (2025).
Cheng, X., Li, C., Hong, H., Zhou, Z. & Wu, Z. Macrocyclic RGD-peptides with high selectivity for alpha(v)beta(3) integrin in cancer imaging and therapy. RSC Med. Chem. https://doi.org/10.1039/d5md00280j (2025).
Chen, H., Niu, G., Wu, H. & Chen, X. Clinical application of radiolabeled RGD peptides for PET imaging of integrin alphavbeta3. Theranostics 6, 78–92 (2016).
Jin, Z. H. et al. Radiotheranostic agent (64)Cu-cyclam-RAFT-c(-RGDfK-)(4) for management of peritoneal metastasis in ovarian cancer. Clin. Cancer Res 26, 6230–6241 (2020).
Cossu, J., Thoreau, F. & Boturyn, D. Multimeric RGD-based strategies for selective drug delivery to tumor tissues. Pharmaceutics 15, 525 (2023).
Wen, X. et al. Synthesis, preclinical, and initial clinical evaluation of integrin alpha(V)beta(3) and gastrin-releasing peptide receptor (GRPR) dual-targeting radiotracer [(68)Ga]Ga-RGD-RM26-03. Eur. J. Nucl. Med. Mol. Imaging 51, 2023–2035 (2024).
Jin, Z. H. et al. Evaluation of the Gly-Phe-Lys linker to reduce the renal radioactivity of a [(64)Cu]Cu-labeled multimeric cRGD peptide. ACS Omega 10, 4102–4120 (2025).
Xiao, L. & Xin, J. Advances in clinical oncology research on (99m)Tc-3PRGD2 SPECT imaging. Front. Oncol. 12, 898764 (2022).
Xiao, L., Yu, S., Xu, W., Sun, Y. & Xin, J. 99m)Tc-3PRGD(2) SPECT/CT imaging for diagnosing lymph node metastasis of primary malignant lung tumors. Korean J. Radio. 24, 1142–1150 (2023).
Huang, Y.-Q. et al. Design, synthesis and pharmacological evaluation of a novel PEG-cRGD-conjugated irinotecan derivative as potential antitumor agent. Eur. J. Med. Chem. 158, 82–90 (2018).
Lerchen, H.-G. et al. Discovery of VIP236, an αvβ3-targeted small-molecule–drug conjugate with neutrophil elastase-mediated activation of 7-ethyl camptothecin payload for treatment of solid tumors. Cancers 15, 4381 (2023).
Kemker, I., Schnepel, C., Schroder, D. C., Marion, A. & Sewald, N. Cyclization of RGD peptides by Suzuki-Miyaura cross-coupling. J. Med. Chem. 62, 7417–7430 (2019).
Kemker, I. et al. Tuning the biological activity of RGD peptides with halotryptophans. J. Med. Chem. 64, 586–601 (2021).
Wang, P. et al. Modular synthesis of clickable peptides via late-stage maleimidation on C(7)-H tryptophan. Nat. Commun. 14, 3973 (2023).
Mitra, A. et al. Polymeric conjugates of mono- and bi-cyclic alphaVbeta3 binding peptides for tumor targeting. J. Control Release 114, 175–183 (2006).
Koivunen, E., Wang, B. & Ruoslahti, E. Phage libraries displaying cyclic peptides with different ring sizes: ligand specificities of the RGD-directed integrins. Biotechnology 13, 265–270 (1995).
Assa-Munt, N., Jia, X., Laakkonen, P. & Ruoslahti, E. Solution structures and integrin binding activities of an RGD peptide with two isomers. Biochemistry 40, 2373–2378 (2001).
Bogdanowich-Knipp, S. J., Chakrabarti, S., Williams, T. D., Dillman, R. K. & Siahaan, T. J. Solution stability of linear vs. cyclic RGD peptides. J. Pept. Res. 53, 530–541 (1999).
Bernhagen, D. et al. Bicyclic RGD peptides with exquisite selectivity for the integrin alpha(v)beta(3) receptor using a “random design” approach. ACS Comb. Sci. 21, 198–206 (2019).
Timmerman, P., Beld, J., Puijk, W. C. & Meloen, R. H. Rapid and quantitative cyclization of multiple peptide loops onto synthetic scaffolds for structural mimicry of protein surfaces. Chembiochem 6, 821–824 (2005).
Kapp, T. G. et al. A comprehensive evaluation of the activity and selectivity profile of ligands for RGD-binding integrins. Sci. Rep. 7, 39805 (2017).
Yao, G. et al. Iodine-mediated tryptathionine formation facilitates the synthesis of amanitins. J. Am. Chem. Soc. 143, 14322–14331 (2021).
Wieland, T. & Faulstich, H. Fifty years of amanitin. Experientia 47, 1186–1193 (1991).
Wu, C. et al. Novel N-methylated cyclodepsipeptide prodrugs for targeted cancer therapy. J. Med. Chem. 64, 991–1000 (2021).
Chatzisideri, T. et al. Integrin-mediated targeted cancer therapy using c(RGDyK)-based conjugates of gemcitabine. J. Med. Chem. 65, 271–284 (2022).
Liu, K. et al. Peptidic heterodimer-based radiotracer targeting fibroblast activation protein and integrin alpha(v)beta(3). Eur. J. Nucl. Med. Mol. Imaging 51, 1544–1557 (2024).
van den Kerkhof, D. L. et al. Unraveling the role of the homoarginine residue in antiplatelet drug eptifibatide in binding to the alphaIIbbeta3 integrin receptor. Thromb. Res. 217, 96–103 (2022).
Tonin, G. & Klen, J. Eptifibatide, an older therapeutic peptide with new indications: from clinical pharmacology to everyday clinical practice. Int. J. Mol. Sci. 24, 5446 (2023).
Frank, A. O. et al. Conformational control of integrin-subtype selectivity in isoDGR peptide motifs: a biological switch. Angew. Chem. Int. Ed. Engl. 49, 9278–9281 (2010).
Pfaff, M. E. A. Selective recognition of cyclic RGD peptides of NMR defined conformation by alpha IIb beta 3, alpha V beta 3, and alpha 5 beta 1 integrins. J. Biol. Chem. 269, 20233–20238 (1994).
Weide, T., Modlinger, A. & Kessler, H. Spatial screening for the identification of the bioactive conformation of integrin ligands. Top. Curr. Chem. 272, 1–50 (2007).
D’Mello, V. et al. Characterization of Asx turn types and their connate relationship with beta-Turns. Chemistry 28, e202104328 (2022).
Newell, N. E. ExploreTurns: a web tool for the exploration, analysis, and classification of beta turns and structured loops in proteins; application to beta-bulge and Schellman loops, Asx helix caps, beta hairpins, and other hydrogen-bonded motifs. Protein Sci. 34, e70046 (2025).
Chen, X., Plasencia, C., Hou, Y. & Neamati, N. Synthesis and biological evaluation of dimeric RGD peptide-paclitaxel conjugate as a model for integrin-targeted drug delivery. J. Med. Chem. 48, 1098–1106 (2005).
Li, R. et al. Asymmetric, amphiphilic RGD conjugated phthalocyanine for targeted photodynamic therapy of triple negative breast cancer. Signal Transduct. Target Ther. 7, 64 (2022).
DiCara, D. et al. Structure-function analysis of Arg-Gly-Asp helix motifs in alpha v beta 6 integrin ligands. J. Biol. Chem. 282, 9657–9665 (2007).
Park, S. H. et al. Development of a peptide-based nano-sized cathepsin B inhibitor for anticancer therapy. Pharmaceutics 15, 1131 (2023).
Li, H. et al. Cathepsin B-activated PET tracer for in vivo tumor imaging. Mol. Pharm. 21, 1382–1389 (2024).
Liu, F. et al. Cathepsin B: the dawn of tumor therapy. Eur. J. Med. Chem. 269, 116329 (2024).
Raposo Moreira Dias, A. et al. Synthesis and biological evaluation of RGD and isoDGR-monomethyl auristatin conjugates targeting integrin alpha(V) beta(3). ChemMedChem 14, 938–942 (2019).
Akaiwa, M., Dugal-Tessier, J. & Mendelsohn, B. A. Antibody-drug conjugate payloads; study of Auristatin derivatives. Chem. Pharm. Bull. 68, 201–211 (2020).
Feni, L. et al. Kiss and run: promoting effective and targeted cellular uptake of a drug delivery vehicle composed of an integrin-targeting diketopiperazine peptidomimetic and a cell-penetrating peptide. Bioconjugate Chem. 30, 2011–2022 (2019).
Zambra, M. et al. Optimizing the enzymatic release of MMAE from isoDGR-based small molecule drug conjugate by incorporation of a GPLG-PABC enzymatically cleavable linker. Front. Pharmacol. 14, 1215694 (2023).
Colombo, R. et al. Synthesis and biological evaluation (in vitro and in vivo) of cyclic arginine-glycine-aspartate (RGD) peptidomimetic-paclitaxel conjugates targeting integrin alphaVbeta3. J. Med. Chem. 55, 10460–10474 (2012).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (grants 82204189) and Guangdong Basic and Applied Basic Research Foundation (2022A1515111207).
Author information
Authors and Affiliations
Contributions
H.Y., H.P., and T. R. contributed equally to this work. H.Y. synthesized the target compounds, completed HPLC analysis and performed partial biological assays. H.P., W.D., and J.T. completed most biological assays and analyzed part of the data. W.P. performed the pharmacokinetic experiments. T. R. performed the theoretical calculations and molecular docking. G.Y., W.S., R.D.S., and J.S. proposed the project. G.Y., R.D.S., and H.Y. performed data analysis and contributed to the writing-review and editing of the manuscript. All authors have given approval to the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Chemistry thanks Didier Boturyn and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, H., Pan, H., Ran, T. et al. Identification of an αvβ3-targeting bicyclic peptide with atypical norArg-Gly-Asp sequence. Commun Chem 9, 83 (2026). https://doi.org/10.1038/s42004-026-01886-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42004-026-01886-y