Abstract
Obesity is a complex, multifactorial condition with a strong genetic basis, encompassing monogenic, oligogenic and polygenic contributions. More than 1 billion people worldwide have obesity, including 150 million children. Since the discovery of leptin, over 85 monogenic forms have been identified, characterized by early-onset obesity with impaired appetite regulation, usually associated with neurodevelopmental (and other) phenotypes, making monogenic obesity mostly syndromic. However, genome-wide association studies have identified over 1,000 loci associated with weight variation. Advances in human genetics have translated into innovative therapeutic strategies. In particular, melanocortin 4 receptor agonists illustrate how genetic discoveries can target treatments, paving the way for precision medicine in obesity. These advances offer new opportunities to tailor treatments to the underlying genetic causes. In this Review, we highlight how genetic discoveries have deepened our understanding of obesity pathophysiology and accelerated precision medicine, and we discuss future strategies to enhance prevention and personalize patient care based on genetic background.
This is a preview of subscription content, access via your institution
Access options



Similar content being viewed by others
References
GBD 2021 Adult BMI Collaborators. Global, regional, and national prevalence of adult overweight and obesity, 1990–2021, with forecasts to 2050: a forecasting study for the Global Burden of Disease Study 2021. Lancet 405, 813–838 (2025).
World Health Organization. Obesity and overweight. WHO https://www.who.int/news-room/fact-sheets/detail/obesity-and-overweight?utm_source=chatgpt.com (2025).
Swinburn, B. A. et al. The global obesity pandemic: shaped by global drivers and local environments. Lancet 378, 804–814 (2011).
Maes, H. H., Neale, M. C. & Eaves, L. J. Genetic and environmental factors in relative body weight and human adiposity. Behav. Genet. 27, 325–351 (1997).
Silventoinen, K. et al. Genetic and environmental effects on body mass index from infancy to the onset of adulthood: an individual-based pooled analysis of 45 twin cohorts participating in the COllaborative project of Development of Anthropometrical measures in Twins (CODATwins) study. Am. J. Clin. Nutr. 104, 371–379 (2016).
Graff, M. et al. Genome-wide physical activity interactions in adiposity - a meta-analysis of 200,452 adults. PLoS Genet. 13, e1006528 (2017).
Han, H. Y., Masip, G., Meng, T. & Nielsen, D. E. Interactions between polygenic risk of obesity and dietary factors on anthropometric outcomes: a systematic review and meta-analysis of observational studies. J. Nutr. 154, 3521–3543 (2024).
Cromer, S. J. et al. Association and interaction of genetics and area-level socioeconomic factors on the prevalence of type 2 diabetes and obesity. Diabetes Care 46, 944–952 (2023).
O’Rahilly, S., Farooqi, I. S., Yeo, G. S. H. & Challis, B. G. Minireview: human obesity-lessons from monogenic disorders. Endocrinology 144, 3757–3764 (2003).
Saeed, S. et al. High morbidity and mortality in children with untreated congenital deficiency of leptin or its receptor. Cell Rep. Med. 4, 101187 (2023).
Gao, Y. et al. TrpC5 mediates acute leptin and serotonin effects via Pomc neurons. Cell Rep. 18, 583–592 (2017).
Doche, M. E. et al. Human SH2B1 mutations are associated with maladaptive behaviors and obesity. J. Clin. Invest. 122, 4732–4736 (2012).
Wijesuriya, T. M. et al. The Prader-Willi syndrome proteins MAGEL2 and necdin regulate leptin receptor cell surface abundance through ubiquitination pathways. Hum. Mol. Genet. 26, 4215–4230 (2017).
Davenport, J. R. et al. Disruption of intraflagellar transport in adult mice leads to obesity and slow-onset cystic kidney disease. Curr. Biol. 17, 1586–1594 (2007).
Guo, D. -F. et al. The BBSome controls energy homeostasis by mediating the transport of the leptin receptor to the plasma membrane. PLoS Genet. 12, e1005890 (2016).
Brewer, K. M., Brewer, K. K., Richardson, N. C. & Berbari, N. F. Neuronal cilia in energy homeostasis. Front. Cell Dev. Biol. 10, 1082141 (2022).
Yang, Y. et al. Steroid receptor coactivator-1 modulates the function of Pomc neurons and energy homeostasis. Nat. Commun. 10, 1718 (2019).
Xu, Y. et al. 5-HT2CRs expressed by pro-opiomelanocortin neurons regulate energy homeostasis. Neuron 60, 582–589 (2008).
Quaresma, P. G. F. et al. Growth hormone/STAT5 signaling in proopiomelanocortin neurons regulates glucoprivic hyperphagia. Mol. Cell Endocrinol. 498, 110574 (2019).
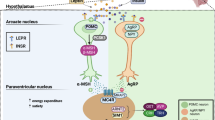
Cone, R. D. Anatomy and regulation of the central melanocortin system. Nat. Neurosci. 8, 571–578 (2005).
Asai, M. et al. Loss of function of the melanocortin 2 receptor accessory protein 2 Is associated with mammalian obesity. Science 341, 275–278 (2013).
Kempf, E. et al. Aberrant expression of agouti signaling protein (ASIP) as a cause of monogenic severe childhood obesity. Nat. Metab. 4, 1697–1712 (2022).
Michaud, J. L. et al. Sim1 haploinsufficiency causes hyperphagia, obesity and reduction of the paraventricular nucleus of the hypothalamus. Hum. Mol. Genet. 10, 1465–1473 (2001).
Schonemann, M. D. et al. Development and survival of the endocrine hypothalamus and posterior pituitary gland requires the neuronal POU domain factor Brn-2. Genes Dev. 9, 3122–3135 (1995).
Breit, A. et al. Alternative G protein coupling and biased agonism: new insights into melanocortin-4 receptor signalling. Mol. Cell Endocrinol. 331, 232–240 (2011).
Unluturk, U. et al. Molecular diagnosis and clinical characterization of pseudohypoparathyroidism type-Ib in a patient with mild Albright’s hereditary osteodystrophy-like features, epileptic seizures, and defective renal handling of uric acid. Am. J. Med. Sci. 336, 84–90 (2008).
Xu, B. et al. Brain-derived neurotrophic factor regulates energy balance downstream of melanocortin-4 receptor. Nat. Neurosci. 6, 736–742 (2003).
Arévalo, J. C. et al. The ARMS/Kidins220 scaffold protein modulates synaptic transmission. Mol. Cell Neurosci. 45, 92–100 (2010).
Bonnefond, A. et al. Loss-of-function mutations in SIM1 contribute to obesity and Prader-Willi–like features. J. Clin. Invest. 123, 3037–3041 (2013).
Stutzmann, F. et al. Prevalence of melanocortin-4 receptor deficiency in Europeans and their age-dependent penetrance in multigenerational pedigrees. Diabetes 57, 2511–2518 (2008).
Saeed, S. et al. Genetic causes of severe childhood obesity: a remarkably high prevalence in an inbred population of Pakistan. Diabetes 69, 1424–1438 (2020).
Saeed, S., Butt, T. A., Anwer, M., Arslan, M. & Froguel, P. High prevalence of leptin and melanocortin-4 receptor gene mutations in children with severe obesity from Pakistani consanguineous families. Mol. Genet. Metab. 106, 121–126 (2012).
Chami, N., Preuss, M., Walker, R. W., Moscati, A. & Loos, R. J. F. The role of polygenic susceptibility to obesity among carriers of pathogenic mutations in MC4R in the UK Biobank population. PLoS Med. 17, e1003196 (2020).
Hirschhorn, J. N. & Daly, M. J. Genome-wide association studies for common diseases and complex traits. Nat. Rev. Genet. 6, 95–108 (2005).
Carlson, C. S., Eberle, M. A., Kruglyak, L. & Nickerson, D. A. Mapping complex disease loci in whole-genome association studies. Nature 429, 446–452 (2004).
Steemers, F. J. et al. Whole-genome genotyping with the single-base extension assay. Nat. Methods 3, 31–33 (2006).
Gunderson, K. L., Steemers, F. J., Lee, G., Mendoza, L. G. & Chee, M. S. A genome-wide scalable SNP genotyping assay using microarray technology. Nat. Genet. 37, 549–554 (2005).
Uffelmann, E. et al. Genome-wide association studies. Nat. Rev. Methods Primers 1, 59 (2021).
Sudlow, C. et al. UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
GIANT consortium. Giant Consortium https://portals.broadinstitute.org/collaboration/giant/index.php/GIANT_consortium
Herbert, A. et al. A common genetic variant is associated with adult and childhood obesity. Science 312, 279–283 (2006).
Lyon, H. N. et al. The association of a SNP upstream of INSIG2 with body mass index is reproduced in several but not all cohorts. PLoS Genet. 3, e61 (2007).
Hotta, K. et al. INSIG2 gene rs7566605 polymorphism is associated with severe obesity in Japanese. J. Hum. Genet. 53, 857–862 (2008).
Rosskopf, D. et al. Comment on ‘A common genetic variant is associated with adult and childhood obesity. Science 315, 187 (2007).
Loos, R. J. F., Barroso, I., O’Rahilly, S. & Wareham, N. J. Comment on ‘A common genetic variant is associated with adult and childhood obesity’. Science 315, 187 (2007).
Dina, C. et al. Comment on ‘A common genetic variant is associated with adult and childhood obesity. Science 315, 187 (2007).
Frayling, T. M. et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 316, 889–894 (2007).
Grant, S. F. A. et al. Association analysis of the FTO gene with obesity in children of Caucasian and African ancestry reveals a common tagging SNP. PLoS ONE 3, e1746 (2008).
Dina, C. et al. Variation in FTO contributes to childhood obesity and severe adult obesity. Nat. Genet. 39, 724–726 (2007).
Yengo, L. et al. Meta-analysis of genome-wide association studies for height and body mass index in ∼700000 individuals of European ancestry. Hum. Mol. Genet. 27, 3641–3649 (2018).
Bradfield, J. P. et al. A genome-wide association meta-analysis identifies new childhood obesity loci. Nat. Genet. 44, 526–531 (2012).
Bradfield, J. P. et al. A trans-ancestral meta-analysis of genome-wide association studies reveals loci associated with childhood obesity. Hum. Mol. Genet. 28, 3327–3338 (2019).
Willer, C. J. et al. Six new loci associated with body mass index highlight a neuronal influence on body weight regulation. Nat. Genet. 41, 25–34 (2009).
Loos, R. J. F. & Yeo, G. S. H. The genetics of obesity: from discovery to biology. Nat. Rev. Genet. 23, 120–133 (2022).
Khera, A. V. et al. Polygenic prediction of weight and obesity trajectories from birth to adulthood. Cell 177, 587–596 (2019).
Smit, R. A. J. et al. Polygenic prediction of body mass index and obesity through the life course and across ancestries. Nat. Med. 31, 3151–3168 (2025).
Hüls, A. et al. Polygenic risk for obesity and its interaction with lifestyle and sociodemographic factors in European children and adolescents. Int. J. Obes. 45, 1321–1330 (2021).
Cifuentes, L. et al. Genetic and physiological insights into satiation variability predict responses to obesity treatment. Cell Metab. 37, 1655–1666 (2025).
Chami, N. et al. Genetic subtyping of obesity reveals biological insights into the uncoupling of adiposity from its cardiometabolic comorbidities. Nat. Med. 31, 3801–3812 (2025).
Kilpeläinen, T. O. et al. Physical activity attenuates the influence of FTO variants on obesity risk: a meta-analysis of 218,166 adults and 19,268 children. PLoS Med. 8, e1001116 (2011).
Andreasen, C. H. et al. Low physical activity accentuates the effect of the FTO rs9939609 polymorphism on body fat accumulation. Diabetes 57, 95–101 (2008).
Franks, P. W. et al. Assessing gene-treatment interactions at the FTO and INSIG2 loci on obesity-related traits in the Diabetes Prevention Program. Diabetologia 51, 2214–2223 (2008).
Ndiaye, F. K. et al. The expression of genes in top obesity-associated loci is enriched in insula and substantia nigra brain regions involved in addiction and reward. Int. J. Obes. 44, 539–543 (2020).
Locke, A. E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Tam, V. et al. Benefits and limitations of genome-wide association studies. Nat. Rev. Genet. 20, 467–484 (2019).
Chang, J. Y., Park, J. H., Park, S. E., Shon, J. & Park, Y. J. The fat mass- and obesity-associated (FTO) gene to obesity: lessons from mouse models. Obesity 26, 1674–1686 (2018).
Smemo, S. et al. Obesity-associated variants within FTO form long-range functional connections with IRX3. Nature 507, 371–375 (2014).
Claussnitzer, M. et al. FTO obesity variant circuitry and adipocyte browning in humans. N. Engl. J. Med. 373, 895–907 (2015).
Stratigopoulos, G. et al. Hypomorphism of Fto and Rpgrip1l causes obesity in mice. J. Clin. Invest. 126, 1897–1910 (2016).
Sobreira, D. R. et al. Extensive pleiotropism and allelic heterogeneity mediate metabolic effects of IRX3 and IRX5. Science 372, 1085–1091 (2021).
Mostafavi, H., Spence, J. P., Naqvi, S. & Pritchard, J. K. Systematic differences in discovery of genetic effects on gene expression and complex traits. Nat. Genet. 55, 1866–1875 (2023).
Cannon, M. E. et al. Open chromatin profiling in adipose tissue marks genomic regions with functional roles in cardiometabolic traits. G3 9, 2521–2533 (2019).
Littleton, S. H. et al. Variant-to-function analysis of the childhood obesity chr12q13 locus implicates rs7132908 as a causal variant within the 3’ UTR of FAIM2. Cell Genom. 4, 100556 (2024).
Joslin, A. C. et al. A functional genomics pipeline identifies pleiotropy and cross-tissue effects within obesity-associated GWAS loci. Nat. Commun. 12, 5253 (2021).
Dong, S. -S. et al. Systematic functional characterization of non-coding regulatory SNPs associated with central obesity. Am. J. Hum. Genet. 112, 116–134 (2025).
Pan, D. Z. et al. Integration of human adipocyte chromosomal interactions with adipose gene expression prioritizes obesity-related genes from GWAS. Nat. Commun. 9, 1512 (2018).
Matharu, N. et al. CRISPR-mediated activation of a promoter or enhancer rescues obesity caused by haploinsufficiency. Science 363, eaau0629 (2019).
Zhu, H. et al. CRISPRa-based activation of Fgf21 and Fndc5 ameliorates obesity by promoting adipocytes browning. Clin. Transl. Med. 13, e1326 (2023).
Dong, S. -S. et al. An intronic risk SNP rs12454712 for central obesity acts as an allele-specific enhancer to regulate BCL2 expression. Diabetes 70, 1679–1688 (2021).
Kamble, P. G. et al. Proof-of-concept for CRISPR/Cas9 gene editing in human preadipocytes: deletion of FKBP5 and PPARG and effects on adipocyte differentiation and metabolism. Sci. Rep. 10, 10565 (2020).
Bielczyk-Maczynska, E. et al. A single-cell CRISPRi platform for characterizing candidate genes relevant to metabolic disorders in human adipocytes. Am. J. Physio. Cell Physiol. 325, C648–C660 (2023).
Helsley, R. N. et al. Obesity-linked suppression of membrane-bound O-acyltransferase 7 (MBOAT7) drives non-alcoholic fatty liver disease. eLife 8, e49882 (2019).
Weng, X. et al. Deletion of CD44 promotes adipogenesis by regulating PPARγ and cell cycle-related pathways. J. Endocrinol. 262, e240079 (2024).
Ichimura, A. et al. Dysfunction of lipid sensor GPR120 leads to obesity in both mouse and human. Nature 483, 350–354 (2012).
Pearce, L. R. et al. KSR2 mutations are associated with obesity, insulin resistance, and impaired cellular fuel oxidation. Cell 155, 765–777 (2013).
Gao, W. et al. Human GLP1R variants affecting GLP1R cell surface expression are associated with impaired glucose control and increased adiposity. Nat. Metab. 5, 1673–1684 (2023).
van der Klaauw, A. A. et al. Human semaphorin 3 variants link melanocortin circuit development and energy balance. Cell 176, 729–742 (2019).
Akbari, P. et al. Sequencing of 640,000 exomes identifies GPR75 variants associated with protection from obesity. Science 373, eabf8683 (2021).
Bradshaw, A. D., Graves, D. C., Motamed, K. & Sage, E. H. SPARC-null mice exhibit increased adiposity without significant differences in overall body weight. Proc. Natl Acad. Sci. USA 100, 6045–6050 (2003).
Degerman, E. et al. From PDE3B to the regulation of energy homeostasis. Curr. Opin. Pharmacol. 11, 676–682 (2011).
Cuinat, S. et al. Loss-of-function variants in SRRM2 cause a neurodevelopmental disorder. Genet. Med. 24, 1774–1780 (2022).
Kaisinger, L. R. et al. Large-scale exome sequence analysis identifies sex- and age-specific determinants of obesity. Cell Genom. 3, 100362 (2023).
Zhao, Y. et al. Protein-truncating variants in BSN are associated with severe adult-onset obesity, type 2 diabetes and fatty liver disease. Nat. Genet. 56, 579–584 (2024).
Brenachot, X. et al. Hepatic protein tyrosine phosphatase receptor gamma links obesity-induced inflammation to insulin resistance. Nat. Commun. 8, 1820 (2017).
Zamariolli, M. et al. The impact of 22q11.2 copy-number variants on human traits in the general population. Am. J. Hum. Genet. 110, 300–313 (2023).
Farooqi, I. S. et al. Beneficial effects of leptin on obesity, T cell hyporesponsiveness, and neuroendocrine/metabolic dysfunction of human congenital leptin deficiency. J. Clin. Invest. 110, 1093–1103 (2002).
Paz-Filho, G., Wong, M. -L. & Licinio, J. Ten years of leptin replacement therapy. Obes. Rev. 12, e315–e323 (2011).
Kühnen, P. et al. Proopiomelanocortin deficiency treated with a melanocortin-4 receptor agonist. N. Engl. J. Med. 375, 240–246 (2016).
Clément, K. et al. MC4R agonism promotes durable weight loss in patients with leptin receptor deficiency. Nat. Med. 24, 551–555 (2018).
Clément, K. et al. Efficacy and safety of setmelanotide, an MC4R agonist, in individuals with severe obesity due to LEPR or POMC deficiency: single-arm, open-label, multicentre, phase 3 trials. Lancet Diabetes Endocrinol. 8, 960–970 (2020).
Ervin, C. et al. Interview-based patient- and caregiver-reported experiences of hunger and improved quality of life with setmelanotide treatment in Bardet-Biedl syndrome. Adv. Ther. 40, 2394–2411 (2023).
Haqq, A. M. et al. Efficacy and safety of setmelanotide, a melanocortin-4 receptor agonist, in patients with Bardet-Biedl syndrome and Alström syndrome: a multicentre, randomised, double-blind, placebo-controlled, phase 3 trial with an open-label period. Lancet Diabetes Endocrinol. 10, 859–868 (2022).
Argente, J. et al. Setmelanotide in patients aged 2–5 years with rare MC4R pathway-associated obesity (VENTURE): a 1 year, open-label, multicenter, phase 3 trial. Lancet Diabetes Endocrinol. 13, 29–37 (2025).
Le Collen, L. et al. Heterozygous pathogenic variants in POMC are not responsible for monogenic obesity: implication for MC4R agonist use. Genet. Med. 25, 100857 (2023).
Delplanque, J. et al. Monoallelic pathogenic variants in LEPR do not cause obesity. Am. J. Hum. Genet. 111, 2668–2674 (2024).
Folon, L. et al. Contribution of heterozygous PCSK1 variants to obesity and implications for precision medicine: a case-control study. Lancet Diabetes Endocrinol. 11, 182–190 (2023).
Collet, T. -H. et al. Evaluation of a melanocortin-4 receptor (MC4R) agonist (Setmelanotide) in MC4R deficiency. Mol. Metab. 6, 1321–1329 (2017).
Höpfner, F. et al. Evaluation of pharmacological rescue of melanocortin-4 receptor nonsense mutations by aminoglycoside. Life 12, 1793 (2022).
Mahmoud, R., Kimonis, V. & Butler, M. G. Clinical trials in Prader-Willi Syndrome: a review. Int. J. Mol. Sci. 24, 2150 (2023).
Miller, J. L. et al. Diazoxide choline extended-release tablet in people with Prader-Willi syndrome: results from long-term open-label study. Obesity 32, 252–261 (2024).
Alqahtani, A. R., Elahmedi, M. O., Al Qahtani, A. R., Lee, J. & Butler, M. G. Laparoscopic sleeve gastrectomy in children and adolescents with Prader-Willi syndrome: a matched-control study. Surg. Obes. Relat. Dis. 12, 100–110 (2016).
Bonnefond, A. et al. Eating behavior, low-frequency functional mutations in the melanocortin-4 receptor (MC4R) gene, and outcomes of bariatric operations: a 6-year prospective study. Diabetes Care 39, 1384–1392 (2016).
Cooiman, M. I. et al. Genetic obesity and bariatric surgery outcome in 1014 patients with morbid obesity. Obes. Surg. 30, 470–477 (2020).
Poitou, C. et al. Long-term outcomes of bariatric surgery in patients with bi-allelic mutations in the POMC, LEPR, and MC4R genes. Surg. Obes. Relat. Dis. 17, 1449–1456 (2021).
Samuels, J. M., Paddu, N. U., Rekulapeli, A., Madhar, A. & Srivastava, G. High prevalence of positive genetic obesity variants in postoperative bariatric surgery patients with weight regain presenting for medical obesity intervention. Obes. Surg. 34, 170–175 (2024).
Ryan, D. H. Setmelanotide: what does it mean for clinical care of patients with obesity? Lancet Diabetes Endocrinol. 8, 933–935 (2020).
Dubern, B., Lourdelle, A. & Clément, K. Beneficial effects of setmelanotide in a 5-year-old boy with POMC deficiency and on his caregivers. JCEM Case Rep. 1, luad041 (2023).
Welling, M. S., Kleinendorst, L., van Haelst, M. M. & van den Akker, E. L. T. The narrative of a patient with leptin receptor deficiency: personalized medicine for a rare genetic obesity disorder. Obes. Facts 16, 514–518 (2023).
Dubern, B., Mosbah, H., Pigeyre, M., Clément, K. & Poitou, C. Rare genetic causes of obesity: diagnosis and management in clinical care. Ann. Endocrinol. 83, 63–72 (2022).
Welling, M. S. et al. Treatment with liraglutide or naltrexone-bupropion in patients with genetic obesity: a real-world study. EClinicalMedicine 74, 102709 (2024).
Iepsen, E. W. et al. Patients with obesity caused by melanocortin-4 receptor mutations can be treated with a glucagon-like peptide-1 receptor agonist. Cell Metab. 28, 23–32 (2018).
Gokul, P. R. et al. Semaglutide, a long-acting GLP-1 analogue, for the management of early-onset obesity due to MC4R defect: a case report. Horm. Res. Paediatr. 98, 148–155 (2025).
Ali, S. et al. Glucagon-like peptide-1 analogues in monogenic syndromic obesity: real-world data from a large cohort of Alström syndrome patients. Diabetes Obes. Metab. 26, 989–996 (2024).
Styne, D. M. et al. Pediatric obesity-assessment, treatment, and prevention: an Endocrine Society Clinical Practice Guideline. J. Clin. Endocrinol. Metab. 102, 709–757 (2017).
Kohlsdorf, K. et al. Early childhood BMI trajectories in monogenic obesity due to leptin, leptin receptor, and melanocortin 4 receptor deficiency. Int. J. Obes. 42, 1602–1609 (2018).
Zorn, S. et al. Early childhood height, weight, and BMI development in children with monogenic obesity: a European multicentre, retrospective, observational study. Lancet Child Adolesc. Health 9, 297–305 (2025).
Mainieri, F., La Bella, S., Rinaldi, M. & Chiarelli, F. Rare genetic forms of obesity in childhood and adolescence, a comprehensive review of their molecular mechanisms and diagnostic approach. Eur. J. Pediatr. 182, 4781–4793 (2023).
Fitch, A. K., Malhotra, S. & Conroy, R. Differentiating monogenic and syndromic obesities from polygenic obesity: assessment, diagnosis, and management. Obes. Pillars 11, 100110 (2024).
Brittain, E. L. et al. Physical activity and incident obesity across the spectrum of genetic risk for obesity. JAMA Netw Open 7, e243821 (2024).
Azmi, S. et al. Harnessing artificial intelligence in obesity research and management: a comprehensive review. Diagnostics 15, 396 (2025).
Tauber, M. & Hoybye, C. Endocrine disorders in Prader-Willi syndrome: a model to understand and treat hypothalamic dysfunction. Lancet Diabetes Endocrinol. 9, 235–246 (2021).
Walters, R. G. et al. A new highly penetrant form of obesity due to deletions on chromosome 16p11.2. Nature 463, 671–675 (2010).
Hanssen, R. et al. Chromosomal deletions on 16p11.2 encompassing SH2B1 are associated with accelerated metabolic disease. Cell Rep. Med. 4, 101155 (2023).
Saeed, S. et al. Loss-of-function mutations in ADCY3 cause monogenic severe obesity. Nat. Genet. 50, 175–179 (2018).
Grarup, N. et al. Loss-of-function variants in ADCY3 increase risk of obesity and type 2 diabetes. Nat. Genet. 50, 172–174 (2018).
Izumi, K. et al. Germline gain-of-function mutations in AFF4 cause a developmental syndrome functionally linking the super elongation complex and cohesin. Nat. Genet. 47, 338–344 (2015).
Girard, D. & Petrovsky, N. Alström syndrome: insights into the pathogenesis of metabolic disorders. Nat. Rev. Endocrinol. 7, 77–88 (2011).
Florea, L., Caba, L. & Gorduza, E. V. Bardet–Biedl syndrome—multiple kaleidoscope images: insight into mechanisms of genotype–phenotype correlations. Genes 12, 1353 (2021).
Han, J. C. et al. Brain-derived neurotrophic factor and obesity in the WAGR syndrome. N. Engl. J. Med. 359, 918–927 (2008).
Ostrowski, P. J. et al. Null variants and deletions in BRWD3 cause an X-linked syndrome of mild-moderate intellectual disability, macrocephaly, and obesity: a series of 17 patients. Am. J. Med. Genet. C Semin. Med. Genet. 181, 638–643 (2019).
Esteghamat, F. et al. CELA2A mutations predispose to early-onset atherosclerosis and metabolic syndrome and affect plasma insulin and platelet activation. Nat. Genet. 51, 1233–1243 (2019).
Shalata, A. et al. Morbid obesity resulting from inactivation of the ciliary protein CEP19 in humans and mice. Am. J. Hum. Genet. 93, 1061–1071 (2013).
Bosch, E. et al. BDV syndrome: an emerging syndrome with profound obesity and neurodevelopmental delay resembling Prader-Willi syndrome. J. Clin. Endocrinol. Metab. 106, 3413–3427 (2021).
Tarpey, P. S. et al. Mutations in CUL4B, which encodes a ubiquitin E3 ligase subunit, cause an X-linked mental retardation syndrome associated with aggressive outbursts, seizures, relative macrocephaly, central obesity, hypogonadism, pes cavus, and tremor. Am. J. Hum. Genet. 80, 345–352 (2007).
Marenne, G. et al. Exome sequencing identifies genes and gene sets contributing to severe childhood obesity, linking PHIP variants to repressed POMC transcription. Cell Metab. 31, 1107–1119 (2020).
Tatton-Brown, K. et al. The Tatton-Brown-Rahman Syndrome: a clinical study of 55 individuals with de novo constitutive DNMT3A variants. Wellcome Open Res. 3, 46 (2018).
Folon, L. et al. Pathogenic, total loss-of-function DYRK1B variants cause monogenic obesity associated with type 2 diabetes. Diabetes Care 47, 444–451 (2024).
Bouman, A. et al. Growth, body composition, and endocrine-metabolic profiles of individuals with Kleefstra syndrome provide directions for clinical management and translational studies. Am. J. Med. Genet. A 194, e63472 (2024).
Moortgat, S. et al. Broadening the phenotypic spectrum and physiological insights related to EIF2S3 variants. Hum. Mutat. 42, 827–834 (2021).
Genovesi, M. L. et al. Case report: a novel homozygous missense variant of FBN3 supporting it is a new candidate gene causative of a Bardet-Biedl syndrome-like phenotype. Front. Genet. 13, 924362 (2022).
Laron, Z. & Werner, H. Laron syndrome - a historical perspective. Rev. Endocr. Metab. Disord. 22, 31–41 (2021).
Mendes de Oliveira, E. et al. Obesity-associated GNAS mutations and the melanocortin pathway. N. Engl. J. Med. 385, 1581–1592 (2021).
Williams, S. R. et al. Haploinsufficiency of HDAC4 causes brachydactyly mental retardation syndrome, with brachydactyly type E, developmental delays, and behavioral problems. Am. J. Hum. Genet. 87, 219–228 (2010).
He, Y. et al. Human loss-of-function variants in the serotonin 2C receptor associated with obesity and maladaptive behavior. Nat. Med. 28, 2537–2546 (2022).
Schaefer, E. et al. Identification of a novel mutation confirms the implication of IFT172 (BBS20) in Bardet-Biedl syndrome. J. Hum. Genet. 61, 447–450 (2016).
Drole Torkar, A. et al. Novel insights into monogenic obesity syndrome due to INPP5E gene variant: a case report of a female patient. Front. Endocrinol. 12, 581134 (2021).
Arnaud, M. et al. Kabuki syndrome: update and review. Arch. Pediatr. 22, 653–660 (2015).
Alstrup, M. et al. Refining the phenotype of SINO syndrome: a comprehensive cohort report of 14 novel cases. Genet. Med. 26, 101219 (2024).
Willim, J. et al. Variants in LRRC7 lead to intellectual disability, autism, aggression and abnormal eating behaviors. Nat. Commun. 15, 7909 (2024).
Schubert, T. & Schaaf, C. P. MAGEL2 (patho-)physiology and Schaaf-Yang syndrome. Dev. Med. Child Neurol. 67, 35–48 (2025).
Iskafi, R. et al. A case of Rafiq syndrome (MAN1B1-CDG) in a Palestinian child, with brief literature review of 44 cases. J. Invest. Med. High Impact. Case Rep. 13, 23247096251313732 (2025).
Twigg, S. R. F. et al. Mutations in multidomain protein MEGF8 identify a Carpenter syndrome subtype associated with defective lateralization. Am. J. Hum. Genet. 91, 897–905 (2012).
Baron, M. et al. Loss-of-function mutations in MRAP2 are pathogenic in hyperphagic obesity with hyperglycemia and hypertension. Nat. Med. 25, 1733–1738 (2019).
Coursimault, J. et al. MYT1L-associated neurodevelopmental disorder: description of 40 new cases and literature review of clinical and molecular aspects. Hum. Genet. 141, 65–80 (2022).
Yeo, G. S. H. et al. A de novo mutation affecting human TrkB associated with severe obesity and developmental delay. Nat. Neurosci. 7, 1187–1189 (2004).
Kraatari-Tiri, M. et al. HIDEA syndrome is caused by biallelic, pathogenic, rare or founder P4HTM variants impacting the active site or the overall stability of the P4H-TM protein. Clin. Genet. 102, 444–450 (2022).
Saeed, S. et al. Biallelic mutations in P4HTM cause syndromic obesity. Diabetes 72, 1228–1234 (2023).
Morava, E. et al. Impaired glucose-1,6-biphosphate production due to bi-allelic PGM2L1 mutations is associated with a neurodevelopmental disorder. Am. J. Hum. Genet. 108, 1151–1160 (2021).
Jain, V. et al. Börjeson-Forssman-Lehmann syndrome: delineating the clinical and allelic spectrum in 14 new families. Eur. J. Hum. Genet. 31, 1421–1429 (2023).
Craddock, K. E. et al. Clinical and genetic characterization of individuals with predicted deleterious PHIP variants. Cold Spring Harb. Mol. Case Stud. 5, a004200 (2019).
Stessman, H. A. F. et al. Disruption of POGZ is associated with intellectual disability and autism spectrum disorders. Am. J. Hum. Genet. 98, 541–552 (2016).
Schönauer, R. et al. Monoallelic intragenic POU3F2 variants lead to neurodevelopmental delay and hyperphagic obesity, confirming the gene’s candidacy in 6q16.1 deletions. Am. J. Hum. Genet. 110, 998–1007 (2023).
Valenzuela, I. et al. Further delineation of the phenotype caused by loss of function mutations in PRMT7. Eur. J. Med. Genet. 62, 182–185 (2019).
Deb, W. et al. PSMD11 loss-of-function variants correlate with a neurobehavioral phenotype, obesity, and increased interferon response. Am. J. Hum. Genet. 111, 1352–1369 (2024).
Jenkins, D. et al. RAB23 mutations in Carpenter syndrome imply an unexpected role for hedgehog signaling in cranial-suture development and obesity. Am. J. Hum. Genet. 80, 1162–1170 (2007).
Cipolla, C. et al. Metabolic profile of patients with Smith-Magenis syndrome: an observational study with literature review. Children 10, 1451 (2023).
Marzin, P. et al. SETD2 related overgrowth syndrome: presentation of four new patients and review of the literature. Am. J. Med. Genet. C Semin. Med. Genet. 181, 509–518 (2019).
Ramachandrappa, S. et al. Rare variants in single-minded 1 (SIM1) are associated with severe obesity. J. Clin. Invest. 123, 3042–3050 (2013).
Saeed, S. et al. Biallelic variants in SREK1 downregulating SNORD115 and SNORD116 cause a Prader-Willi-like syndrome. J. Clin. Invest. 135, e191008 (2025).
Mantovani, G. & Elli, F. M. Pseudohypoparathyroidism type Ib in 2015. Ann. Endocrinol. 76, 101–104 (2015).
Khan, S. F. et al. The roles and regulation of TBX3 in development and disease. Gene 726, 144223 (2020).
Li, Y. et al. Chronic pharmacologic manipulation of dopamine transmission ameliorates metabolic disturbance in Trappc9-linked brain developmental syndrome. JCI Insight 9, e181339 (2024).
Aerden, M. et al. The neurodevelopmental and facial phenotype in individuals with a TRIP12 variant. Eur. J. Hum. Genet. 31, 461–468 (2023).
Li, Y. et al. Loss of transient receptor potential channel 5 causes obesity and postpartum depression. Cell 187, 4176–4192 (2024).
Borman, A. D. et al. A homozygous mutation in the TUB gene associated with retinal dystrophy and obesity. Hum. Mutat. 35, 289–293 (2014).
Kolehmainen, J. et al. Delineation of Cohen syndrome following a large-scale genotype-phenotype screen. Am. J. Hum. Genet. 75, 122–127 (2004).
Turcot, V. et al. Protein-altering variants associated with body mass index implicate pathways that control energy intake and expenditure in obesity. Nat. Genet. 50, 26–41 (2018).
Sollis, E. et al. The NHGRI-EBI GWAS Catalog: knowledgebase and deposition resource. Nucleic Acids Res. 51, D977–D985 (2023).
Acknowledgements
A.B., A.M. and P.F. were supported by the European Union’s Horizon Europe Research and Innovation Programme (OBELISK grant agreement 101080465). A.B and P.F. were supported by the National Center for Precision Diabetic Medicine – PreciDIAB, which is jointly supported by the French National Agency for Research (ANR-18-IBHU-0001), by the European Union (FEDER) 20001891/NP0025517, by the Hauts-de-France Regional Council and by the European Metropolis of Lille (MEL 2019_ESR_11). A.B. was also supported by the European Research Council (OπO 101043671). W.S.B. and S.F.A.G. were funded by National Institutes of Health (NIH) award R01 HD056465. W.S.B. was also supported by NIH training grant T32 GM008638. S.F.A.G. is also supported by the Daniel B. Burke Endowed Chair for Diabetes Research.
Author information
Authors and Affiliations
Contributions
All authors contributed to the development of the Review topics, wrote specific sections, edited drafts and approved the final version of the manuscript. A.B. and P.F. coordinated the development and planning of the Review. A.B. was primarily responsible for writing the summary, introduction and the section on ‘Genetic and environmental dimensions of monogenic obesity’. A.B., W.S.B. and S.F.A.G. jointly led the writing of the ‘Identification of polygenic aetiologies of obesity and their interactions with environment and lifestyle’ section. A.M. led the writing of the ‘Lessons from obesity genetics for genomic medicine’ section. P.F. led the writing of the ‘Outlook and future directions’ section.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Paul Franks and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Revati Dewal, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bonnefond, A., Bruner, W.S., Grant, S.F.A. et al. The genetics of obesity: aetiology, prevention and therapy. Nat Metab (2026). https://doi.org/10.1038/s42255-026-01497-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42255-026-01497-w