Abstract
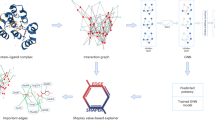
Developing robust methods for evaluating protein–ligand interactions has been a long-standing problem. Data-driven methods may memorize ligand and protein training data rather than learning protein–ligand interactions. Here we show a scoring approach called EquiScore, which utilizes a heterogeneous graph neural network to integrate physical prior knowledge and characterize protein–ligand interactions in equivariant geometric space. EquiScore is trained based on a new dataset constructed with multiple data augmentation strategies and a stringent redundancy-removal scheme. On two large external test sets, EquiScore consistently achieved top-ranking performance compared to 21 other methods. When EquiScore is used alongside different docking methods, it can effectively enhance the screening ability of these docking methods. EquiScore also showed good performance on the activity-ranking task of a series of structural analogues, indicating its potential to guide lead compound optimization. Finally, we investigated different levels of interpretability of EquiScore, which may provide more insights into structure-based drug design.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The PDBscreen dataset supporting this study’s findings is available via Zenodo at https://doi.org/10.5281/zenodo.8049380 (ref. 67). The test dataset supporting this study’s findings is available via Zenodo at https://doi.org/10.5281/zenodo.8047224 (ref. 68). Original data and supplementary information supporting this study’s findings are available via Zenodo at https://doi.org/10.5281/zenodo.10812637 (ref. 69). Source data are provided with this paper.
Code availability
The code used to generate the results shown in this study is available under an MIT License via GitHub at https://github.com/CAODH/EquiScore (ref. 70) and via Zenodo at https://doi.org/10.5281/zenodo.10812534 (ref. 71).
References
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Baek, M. et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 373, 871–876 (2021).
Muller, S. et al. Target 2035—update on the quest for a probe for every protein. RSC Med. Chem. 13, 13–21 (2022).
Kaplan, A. L. et al. Bespoke library docking for 5-HT(2A) receptor agonists with antidepressant activity. Nature 610, 582–591 (2022).
Lyu, J. et al. Ultra-large library docking for discovering new chemotypes. Nature 566, 224–229 (2019).
Shen, C. et al. Beware of the generic machine learning-based scoring functions in structure-based virtual screening. Brief. Bioinform. 22, bbaa070 (2021).
Guedes, I. A., Pereira, F. S. S. & Dardenne, L. E. Empirical scoring functions for structure-based virtual screening: applications, critical aspects, and challenges. Front. Pharmacol. 9, 411637 (2018).
Shen, C. et al. Accuracy or novelty: what can we gain from target-specific machine-learning-based scoring functions in virtual screening? Brief. Bioinform. 22, bbaa410 (2021).
Zhu, H., Yang, J. & Huang, N. Assessment of the generalization abilities of machine-learning scoring functions for structure-based virtual screening. J. Chem. Inf. Model. 62, 5485–5502 (2022).
Francoeur, P. G. et al. Three-dimensional convolutional neural networks and a cross-docked data set for structure-based drug design. J. Chem. Inf. Model. 60, 4200–4215 (2020).
Ragoza, M., Hochuli, J., Idrobo, E., Sunseri, J. & Koes, D. R. Protein-ligand scoring with convolutional neural networks. J. Chem. Inf. Model. 57, 942–957 (2017).
Li, S. et al. Structure-aware interactive graph neural networks for the prediction of protein-ligand binding affinity. In Proc. 27th ACM SIGKDD Conference on Knowledge Discovery & Data Mining (eds Feida, Z. et al.) 975–985 (ACM, 2021); https://doi.org/10.1145/3447548.3467311
Lim, J. et al. Predicting drug-target interaction using a novel graph neural network with 3D structure-embedded graph representation. J. Chem. Inf. Model. 59, 3981–3988 (2019).
Moon, S., Zhung, W., Yang, S., Lim, J. & Kim, W. Y. PIGNet: a physics-informed deep learning model toward generalized drug-target interaction predictions. Chem. Sci. 13, 3661–3673 (2022).
Shen, C. et al. Boosting protein-ligand binding pose prediction and virtual screening based on residue-atom distance likelihood potential and graph transformer. J. Med. Chem. 65, 10691–10706 (2022).
Jiang, D. et al. InteractionGraphNet: a novel and efficient deep graph representation learning framework for accurate protein-ligand interaction predictions. J. Med. Chem. 64, 18209–18232 (2021).
Méndez-Lucio, O., Ahmad, M., del Rio-Chanona, E. A. & Wegner, J. K. A geometric deep learning approach to predict binding conformations of bioactive molecules. Nat. Mach. Intell. 3, 1033–1039 (2021).
Li, Y. & Yang, J. Structural and sequence similarity makes a significant impact on machine-learning-based scoring functions for protein–ligand interactions. J. Chem. Inf. Model. 57, 1007–1012 (2017).
Chen, L. et al. Hidden bias in the DUD-E dataset leads to misleading performance of deep learning in structure-based virtual screening. PLoS ONE 14, e0220113 (2019).
Chatterjee, A. et al. Improving the generalizability of protein-ligand binding predictions with AI-Bind. Nat. Commun. 14, 1989 (2023).
Geirhos, R. et al. Shortcut learning in deep neural networks. Nat. Mach. Intell. 2, 665–673 (2020).
Sastry, G. M., Dixon, S. L. & Sherman, W. Rapid shape-based ligand alignment and virtual screening method based on atom/feature-pair similarities and volume overlap scoring. J. Chem. Inf. Model. 51, 2455–2466 (2011).
Volkov, M. et al. On the frustration to predict binding affinities from protein–ligand structures with deep neural networks. J. Med. Chem. 65, 7946–7958 (2022).
Li, S. et al. MONN: a multi-objective neural network for predicting compound-protein interactions and affinities. Cell Syst. 10, 308–322 (2020).
Cain, S., Risheh, A. & Forouzesh, N. Calculation of protein-ligand binding free energy using a physics-guided neural network. In Proc. IEEE International Conference on Bioinformatics and Biomedicine (BIBM) (eds Chen, Y. et al.) 2487–2493 (IEEE, 2021); https://doi.org/10.1109/bibm52615.2021.9669867
Stärk, H., Ganea, O., Pattanaik, L., Barzilay, R. & Jaakkola, T. Equibind: geometric deep learning for drug binding structure prediction. In Proc. 39th International Conference on Machine Learning (eds Chaudhuri, K. et al.) 20503–20521 (PMLR, 2022); https://doi.org/10.48550/arXiv.2202.05146
Batzner, S. et al. E(3)-equivariant graph neural networks for data-efficient and accurate interatomic potentials. Nat. Commun. 13, 2453 (2022).
Atz, K., Grisoni, F. & Schneider, G. Geometric deep learning on molecular representations. Nat. Mach. Intell. 3, 1023–1032 (2021).
Thurlemann, M., Boselt, L. & Riniker, S. Learning atomic multipoles: prediction of the electrostatic potential with equivariant graph neural networks. J. Chem. Theory Comput. 18, 1701–1710 (2022).
Batool, M., Ahmad, B. & Choi, S. A structure-based drug discovery paradigm. Int. J. Mol. Sci. 20, 2783 (2019).
Imrie, F., Bradley, A. R. & Deane, C. M. Generating property-matched decoy molecules using deep learning. Bioinformatics 37, 2134–2141 (2021).
Mysinger, M. M., Carchia, M., Irwin, J. J. & Shoichet, B. K. Directory of useful decoys, enhanced (DUD-E): better ligands and decoys for better benchmarking. J. Med. Chem. 55, 6582–6594 (2012).
Bauer, M. R., Ibrahim, T. M., Vogel, S. M. & Boeckler, F. M. Evaluation and optimization of virtual screening workflows with DEKOIS 2.0—a public library of challenging docking benchmark sets. J. Chem. Inf. Model. 53, 1447–1462 (2013).
Wang, L. et al. Accurate and reliable prediction of relative ligand binding potency in prospective drug discovery by way of a modern free-energy calculation protocol and force field. J. Am. Chem. Soc. 137, 2695–2703 (2015).
Sieg, J., Flachsenberg, F. & Rarey, M. In need of bias control: evaluating chemical data for machine learning in structure-based virtual screening. J. Chem. Inf. Model. 59, 947–961 (2019).
Berman, H. M. et al. The protein data bank. Nucleic Acids Res. 28, 235–242 (2000).
Adeshina, Y. O., Deeds, E. J. & Karanicolas, J. Machine learning classification can reduce false positives in structure-based virtual screening. Proc. Natl Acad. Sci. USA 117, 18477–18488 (2020).
Bouysset, C. & Fiorucci, S. ProLIF: a library to encode molecular interactions as fingerprints. J. Cheminform. 13, 72 (2021).
Satorras, V. G., Hoogeboom, E. & Welling, M. E(n) equivariant graph neural networks. In Proc. 38th International Conference on Machine Learning (eds Meila, M. & Zhang, T.) 9323–9332 (PMLR, 2021); https://doi.org/10.48550/arXiv.2102.09844
Yun, S., Jeong, M., Kim, R., Kang, J. & Kim, H. J. Graph transformer networks. In Advances in Neural Information Processing Systems 32 (eds Wallach, H. et al.) 11983–11993 (NeurIPS, 2019); https://doi.org/10.48550/arXiv.1911.06455
Friesner, R. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 47, 1739–1749 (2004).
Mastropietro, A., Pasculli, G. & Bajorath, J. Learning characteristics of graph neural networks predicting protein–ligand affinities. Nat. Mach. Intell. 5, 1427–1436 (2023).
Yu, Y., Lu, S., Gao, Z., Zheng, H. & Ke, G. Do deep learning models really outperform traditional approaches in molecular docking? Preprint at https://doi.org/10.48550/arXiv.2302.07134 (2023).
Sastry, G. M., Adzhigirey, M., Day, T., Annabhimoju, R. & Sherman, W. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J. Comput. Aided Mol. Des. 27, 221–234 (2013).
Harder, E. et al. OPLS3: a force field providing broad coverage of drug-like small molecules and proteins. J. Chem. Theory Comput. 12, 281–296 (2016).
Tuccinardi, T., Poli, G., Romboli, V., Giordano, A. & Martinelli, A. Extensive consensus docking evaluation for ligand pose prediction and virtual screening studies. J. Chem. Inf. Model. 54, 2980–2986 (2014).
Westbrook, J. D. et al. The chemical component dictionary: complete descriptions of constituent molecules in experimentally determined 3D macromolecules in the Protein Data Bank. Bioinformatics 31, 1274–1278 (2015).
UniProt, C. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res. 49, D480–D489 (2021).
Wierbowski, S. D., Wingert, B. M., Zheng, J. & Camacho, C. J. Cross‐docking benchmark for automated pose and ranking prediction of ligand binding. Protein Sci. 29, 298–305 (2020).
Shen, C. et al. The impact of cross-docked poses on performance of machine learning classifier for protein–ligand binding pose prediction. J. Cheminform. 13, 1–18 (2021).
Zhang, X. et al. TocoDecoy: a new approach to design unbiased datasets for training and benchmarking machine-learning scoring functions. J. Med. Chem. 65, 7918–7932 (2022).
Su, M., Feng, G., Liu, Z., Li, Y. & Wang, R. Tapping on the black box: how is the scoring power of a machine-learning scoring function dependent on the training set? J. Chem. Inf. Model. 60, 1122–1136 (2020).
Scantlebury, J. et al. A small step toward generalizability: training a machine learning scoring function for structure-based virtual screening. J. Chem. Inf. Model. 63, 2960–2974 (2023).
Ying, C. et al. Do transformers really perform bad for graph representation? In Advances in Neural Information Processing Systems 34 (eds Ranzato, M. et al.) 28877–28888 (NeurIPS, 2021); https://doi.org/10.48550/arXiv.2106.05234
Gilmer, J., Schoenholz, S. S., Riley, P. F., Vinyals, O. & Dahl, G. E. Neural message passing for quantum chemistry. In Proc. 34th International Conference on Machine Learning (eds Precup, D. & Teh, Y. W.) 1263–1272 (PMLR, 2017); https://doi.org/10.5555/3305381.3305512
Jiao, Q. et al. Edge-gated graph neural network for predicting protein-ligand binding affinities. In Proc. IEEE International Conference on Bioinformatics and Biomedicine (BIBM) (eds Huang, Y. et al.) 334–339 (IEEE, 2021); https://doi.org/10.1109/bibm52615.2021.9669846
Shang, C. et al. Edge attention-based multi-relational graph convolutional networks. Preprint at https://doi.org/10.48550/arXiv.1802.04944 (2018).
Gong, L. & Cheng, Q. Exploiting edge features for graph neural networks. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition (eds Michael S. B. et al.) 9203–9211 (IEEE, 2019); https://doi.org/10.1109/CVPR.2019.00943
Dwivedi, V. P. & Bresson, X. A generalization of transformer networks to graphs. Preprint at https://doi.org/10.48550/arXiv.2012.09699 (2020).
Bradley, A. P. The use of the area under the roc curve in the evaluation of machine learning algorithms. Pattern Recognit. 30, 1145–1159 (1997).
Xue, Y., Tong, Y. & Neri, F. An ensemble of differential evolution and Adam for training feed-forward neural networks. Inf. Sci. 608, 453–471 (2022).
Lu, W. et al. Tankbind: Trigonometry-aware neural networks for drug-protein binding structure prediction. In Advances in Neural Information Processing Systems 35 (eds Koyejo, S. et al.) 7236–7249 (NeurIPS, 2022); https://doi.org/10.1101/2022.06.06.495043
Mendez, D. et al. ChEMBL: towards direct deposition of bioassay data. Nucleic Acids Res. 47, D930–D940 (2019).
Liu, T., Lin, Y., Wen, X., Jorissen, R. N. & Gilson, M. K. BindingDB: a web-accessible database of experimentally determined protein-ligand binding affinities. Nucleic Acids Res. 35, D198–D201 (2007).
Irwin, J. J. & Shoichet, B. K. ZINC—a free database of commercially available compounds for virtual screening. J. Chem. Inf. Model. 45, 177–182 (2005).
Truchon, J. F. & Bayly, C. I. Evaluating virtual screening methods: good and bad metrics for the ‘early recognition’ problem. J. Chem. Inf. Model. 47, 488–508 (2007).
Cao, D., Chen, G., Jiang, J. & Zheng, M. PDBscreen with multiple data augmentation strategies suitable for training protein-ligand interaction prediction methods. Zenodo https://doi.org/10.5281/zenodo.8049380 (2023).
Cao, D., Chen, G., Jiang, J., Yu, J. & Zheng, M. TEST dataset pocket for EquiScore. Zenodo https://doi.org/10.5281/zenodo.8047224 (2023).
Cao, D. & Chen, G. Original data and supplementary information for ‘EquiScore is a generic protein–ligand interaction scoring method integrating physical prior knowledge with data-augmentation modeling’. Zenodo https://doi.org/10.5281/zenodo.10812637 (2023).
Cao, D. Code for ‘EquiScore is a generic protein–ligand interaction scoring method integrating physical prior knowledge with data-augmentation modeling’. GitHub https://github.com/CAODH/EquiScore (2023).
Cao, D. Code for ‘EquiScore is a generic protein–ligand interaction scoring method integrating physical prior knowledge with data-augmentation modeling’. Zenodo https://doi.org/10.5281/zenodo.10812534 (2023).
Acknowledgements
We gratefully acknowledge financial support from the National Natural Science Foundation of China (T2225002 and 82273855 to M.Z., 82204278 to X. Li), the National Key Research and Development Program of China (2023YFC2305904 to M.Z.), the Shanghai Municipal Science and Technology Major Project (to M.Z.), the SIMM-SHUTCM Traditional Chinese Medicine Innovation Joint Research Program (E2G805H to M.Z.) and the Youth Innovation Promotion Association CAS (2023296 to S.Z.). We also acknowledge the Shanghai Supercomputer Center for providing computing resources.
Author information
Authors and Affiliations
Contributions
M.Z. designed the study. D.C. developed the method and implemented the code. G.C. and D.C. collected and processed training data. D.C., G.C., J.J. and J.Y. benchmarked the methods. All authors contributed to the analysis of the results. D.C., G.C. and M.Z. wrote the paper. All authors read and approved the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Machine Intelligence thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Ablation results for VS and analogs ranking tasks.
a: VS performance is measured by 1.0% EF on DEKOIS2.0 (number of data points n = 81). The white points in the violin plots represent the means for each bin. b: Analogs ranking is measured by Spearman’s coefficient on LeadOpt (number of data points n = 8). The white points represent the average of coefficient values weighted by the number of ligands in each group (details on sample size for each group in Table 1).
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1–4, Figs. 1–5 and other supplementary content.
Supplementary Data 1 (download XLSX )
Raw data for Supplementary Fig. 3.
Supplementary Data 2 (download XLSX )
Raw data for Supplementary Fig. 4.
Source data
Source Data Fig. 3 (download XLSX )
Raw data for Fig. 3 and Supplementary Fig. 1.
Source Data Fig. 4 (download XLSX )
Raw data for Fig. 4 and Supplementary Fig. 2.
Source Data Fig. 5 (download XLSX )
Raw data for Fig. 5.
Source Data Extended Data Fig. 1 (download XLSX )
Raw data for Extended Data Fig. 1.
Source Data Extended Data Table 2 (download XLSX )
Raw data for Extended Data Table 2.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cao, D., Chen, G., Jiang, J. et al. Generic protein–ligand interaction scoring by integrating physical prior knowledge and data augmentation modelling. Nat Mach Intell 6, 688–700 (2024). https://doi.org/10.1038/s42256-024-00849-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42256-024-00849-z
This article is cited by
-
The algebraic extended atom-type graph-based model for precise ligand–receptor binding affinity prediction
Journal of Cheminformatics (2025)
-
Knowledge-guided diffusion model for 3D ligand-pharmacophore mapping
Nature Communications (2025)
-
Resolving data bias improves generalization in binding affinity prediction
Nature Machine Intelligence (2025)
-
SurfDock is a surface-informed diffusion generative model for reliable and accurate protein–ligand complex prediction
Nature Methods (2025)
-
Benchmarking AI-powered docking methods from the perspective of virtual screening
Nature Machine Intelligence (2025)