Abstract
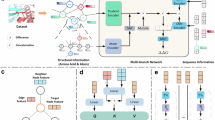
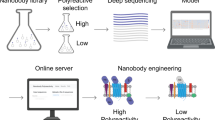
Nanobodies can provide specific binding to divergent antigens, leading to many promising therapeutic and detection applications in recent years. Traditional technologies of nanobody discovery based on alpaca immunization and phage display are very time-consuming and labour-intensive. Despite recent progress in the study of nanobodies, developing fast and accurate computational tools for nanobody–antigen interaction (NAI) prediction is urgently desirable. Here we propose an ensemble deep learning-based framework named DeepNano-seq to predict general protein–protein interaction (PPI) containing NAI from pure sequence information. Quantitative comparison results show that DeepNano-seq possesses the best cross-species generalization ability among existing PPI algorithms. Nevertheless, several of the most effective PPI methods, including DeepNano-seq, demonstrate suboptimal performance for NAI prediction due to the distinction between NAI and PPI at both the pattern and data levels. Therefore, we organize NAI data from the public database for dedicated NAI modelling. Furthermore, we enhance the prediction pipeline of DeepNano-seq by directing the model’s attention to the antigen-binding sites through a prompt-based approach to present the final DeepNano. The comprehensive evaluation demonstrates that DeepNano performs superiorly in NAI prediction and virtual screening of nanobodies. Overall, DeepNano-seq and DeepNano can offer powerful tools for nanobody discovery.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Datasets used in this study and detailed data description documentation have been deposited in our GitHub repository: https://github.com/ddd9898/DeepNano/tree/main/data (ref. 49). The raw data for several graphs in the main text can be found in Supplementary Information. Original nanobody–antigen-binding pairs exploited for training were obtained from the SAbDab-nano database at https://opig.stats.ox.ac.uk/webapps/sabdab-sabpred/sabdab/nanobodies/. The large amount of natural nanobodies adopted in the case study were downloaded from the INDI database at https://research.naturalantibody.com/nanobodies. Sequences of GST and HSA were obtained from the UniProt database at https://www.uniprot.org/. A description of the data for all PPIs and NAIs used in this study is presented in Supplementary Table 5. In addition, the analysis of differences between PPIs and NAIs21,25,50 is presented in Supplementary Note 5.
Code availability
The code repository of this study, which includes the trained models and the evaluation pipeline, is freely available at https://github.com/ddd9898/DeepNano (ref. 49). The source code based on standard Python packages (including Pymol, Biopython and numpy) to organize nanobody–antigen-binding data from SAbDab-nano can be found at https://github.com/ddd9898/DeepNano-data.
References
Hamers-Casterman, C. et al. Naturally occurring antibodies devoid of light chains. Nature 363, 446–448 (1993).
Muyldermans, S. Nanobodies: natural single-domain antibodies. Annu. Rev. Biochem. 82, 775–797 (2013).
Ingram, J. R. et al. Exploiting nanobodies’ singular traits. Annu. Rev. Immunol. 36, 695–715 (2018).
Guo, K. et al. Rapid single-molecule detection of COVID-19 and MERS antigens via nanobody-functionalized organic electrochemical transistors. Nat. Biomed. Eng. 5, 666–677 (2021).
Zhang, X. et al. Specific detection of proteins by a nanobody-functionalized nanopore sensor. ACS Nano 17, 9167–9177 (2023).
Peyvandi, F. et al. Caplacizumab for acquired thrombotic thrombocytopenic purpura. New Engl. J. Med. 374, 511–522 (2016).
Papp, K. A. et al. IL17A/F nanobody sonelokimab in patients with plaque psoriasis: a multicentre, randomised, placebo-controlled, phase 2b study. Lancet 397, 1564–1575 (2021).
Xu, J. et al. Nanobodies from camelid mice and llamas neutralize SARS-CoV-2 variants. Nature 595, 278–282 (2021).
Kourelis, J. et al. NLR immune receptor-nanobody fusions confer plant disease resistance. Science 379, 934–939 (2023).
Wilton, E. E. et al. sdAb-DB: the Single Domain Antibody Database. ACS Synth. Biol. 7, 2480–2484 (2018).
Schneider, C. et al. SAbDab in the age of biotherapeutics: updates including SAbDab-nano, the nanobody structure tracker. Nucleic Acids Res. 50, D1368–D1372 (2022).
Deszynski, P. et al. INDI-integrated nanobody database for immunoinformatics. Nucleic Acids Res. 50, D1273–D1281 (2022).
Xiong, S. et al. NanoLAS: a comprehensive nanobody database with data integration, consolidation and application. Database 2024, baae003 (2024).
Abanades, B. et al. ImmuneBuilder: deep-learning models for predicting the structures of immune proteins. Commun. Biol. 6, 575 (2023).
Ruffolo, J. A. et al. Fast, accurate antibody structure prediction from deep learning on massive set of natural antibodies. Nat. Commun. 14, 2389 (2023).
Ramon, A. et al. Assessing antibody and nanobody nativeness for hit selection and humanization with AbNatiV. Nat. Mach. Intell. 6, 74–91 (2024).
Li, S. et al. NanoBERTa-ASP: predicting nanobody paratope based on a pretrained RoBERTa model. BMC Bioinformatics 25, 122 (2024).
Soler, M. A. et al. Binding affinity prediction of nanobody-protein complexes by scoring of molecular dynamics trajectories. Phys. Chem. Chem. Phys. 20, 3438–3444 (2018).
Tam, C. et al. NbX: machine learning-guided re-ranking of nanobody-antigen binding poses. Pharmaceuticals 14, 968 (2021).
Myung, Y. et al. CSM-AB: graph-based antibody-antigen binding affinity prediction and docking scoring function. Bioinformatics 38, 1141–1143 (2022).
Yang, Y. X. et al. AREA-AFFINITY: a web server for machine learning-based prediction of protein–protein and antibody–protein antigen binding affinities. J. Chem. Inf. Model 63, 3230–3237 (2023).
Sardar, U. et al. Sequence-based nanobody-antigen binding prediction. In Proc. 19th International Symposium on Bioinformatics Research and Application Vol. 14248 (eds Xuan, G. et al.) 227–240 (ISBRA, 2023).
Sledzieski, S. et al. D-SCRIPT translates genome to phenome with sequence-based, structure-aware, genome-scale predictions of protein–protein interactions. Cell Syst. 12, 969–982.e6 (2021).
Singh, R. et al. Topsy-Turvy: integrating a global view into sequence-based PPI prediction. Bioinformatics 38, i264–i272 (2022).
Akbar, R. et al. A compact vocabulary of paratope-epitope interactions enables predictability of antibody-antigen binding. Cell Rep. 34, 108856 (2021).
Anfinsen, C. B. Principles that govern the folding of protein chains. Science 181, 223–230 (1973).
Unsal, S. et al. Learning functional properties of proteins with language models. Nat. Mach. Intell. 4, 227–245 (2022).
Elnaggar, A. et al. ProtTrans: toward understanding the language of life through self-supervised learning. IEEE Trans. Pattern Anal. Mach. Intell. 44, 7112–7127 (2022).
Gao, Z. et al. Hierarchical graph learning for protein–protein interaction. Nat. Commun. 14, 1093 (2023).
Cao, Y. et al. Ensemble deep learning in bioinformatics. Nat. Mach. Intell. 2, 500–508 (2020).
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
Kirillov, A. et al. Segment anything. In Proc. IEEE/CVF International Conference on Computer Vision 4015–4026 (IEEE, 2023).
Chen, M. et al. Multifaceted protein–protein interaction prediction based on Siamese residual RCNN. Bioinformatics 35, i305–i314 (2019).
Richoux, F. et al. Comparing two deep learning sequence-based models for protein–protein interaction prediction. Preprint at https://doi.org/10.48550/arXiv.1901.06268 (2019).
Crooks, G. E. et al. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Dunbar, J. et al. SAbDab: the structural antibody database. Nucleic Acids Res. 42, D1140–D1146 (2014).
Mitchell, L. S. et al. Comparative analysis of nanobody sequence and structure data. Proteins 86, 697–706 (2018).
Chayen, N. E. et al. Protein crystallization: from purified protein to diffraction-quality crystal. Nat. Methods 5, 147–153 (2008).
Yip, K. M. et al. Atomic-resolution protein structure determination by cryo-EM. Nature 587, 157–161 (2020).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Xiang, Y. et al. Integrative proteomics identifies thousands of distinct, multi-epitope, and high-affinity nanobodies. Cell Syst. 12, 220–234.e9 (2021).
Apweiler, R. et al. UniProt: the Universal Protein knowledgebase. Nucleic Acids Res. 32, D115–D119 (2004).
Lu, W. et al. DynamicBind: predicting ligand-specific protein-ligand complex structure with a deep equivariant generative model. Nat. Commun. 15, 1071 (2024).
Roche, R. et al. EquiPNAS: improved protein-nucleic acid binding site prediction using protein-language-model-informed equivariant deep graph neural networks. Nucleic Acids Res. 52, e27 (2024).
Wang, Y. et al. ZeroBind: a protein-specific zero-shot predictor with subgraph matching for drug-target interactions. Nat. Commun. 14, 7861 (2023).
Hie, B., Bryson, B. D. & Berger, B. Leveraging uncertainty in machine learning accelerates biological discovery and design. Cell Syst. 11, 461–477.e9 (2020).
Lishuang, L. et al. Integrating active learning strategy to the ensemble kernel-based method for protein–protein interaction extraction. Chinese J. Electron. 22, 41–45 (2013).
Reynisson, B. et al. NetMHCpan-4.1 and NetMHCIIpan-4.0: improved predictions of MHC antigen presentation by concurrent motif deconvolution and integration of MS MHC eluted ligand data. Nucleic Acids Res. 48, W449–W454 (2020).
Juntao, D. et al. ddd9898/DeepNano: DeepNano paper. Zenodo https://doi.org/10.5281/zenodo.13822580 (2024).
Mahajan, S. P. et al. Contextual protein and antibody encodings from equivariant graph transformers. Preprint at bioRxiv https://doi.org/10.1101/2023.07.15.549154 (2023).
Acknowledgements
This work was partially supported by the National Natural Science Foundation of China (grant no. 62173204 to M.L.), Beijing National Research Center for Information Science and Technology, the Fundamental Research Funds for the Central Universities (grant no. buctrc202337 to M.L.) and the Engineering Research Center of Intelligent Technology and Equipment for Saving Energy and Increasing Benefit, Ministry of Education.
Author information
Authors and Affiliations
Contributions
J.D. and M.L. conceived the original ideas for this study, designed and performed the experiments, and cowrote the manuscript. M.G. and P.Z. contributed to the model design and revision of the manuscript. M.D., T.L. and Y.Z. participated in the experimental design and algorithm programming and helped to prepare figures. M.L. was responsible for the project and guided the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Machine Intelligence thanks Sudeep Sarma and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–6, Figs. 1–10 and Tables 1–13.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Deng, J., Gu, M., Zhang, P. et al. Nanobody–antigen interaction prediction with ensemble deep learning and prompt-based protein language models. Nat Mach Intell 6, 1594–1604 (2024). https://doi.org/10.1038/s42256-024-00940-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42256-024-00940-5