Abstract
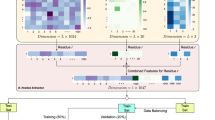
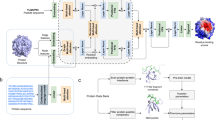
Accurate prediction of protein–peptide interactions is critical for peptide drug discovery. However, due to the limited number of protein–peptide structures in the Protein Data Bank, it is challenging to train an accurate scoring function for protein–peptide interactions. Here, addressing this challenge, we propose an interaction-derived graph neural network model for scoring protein–peptide complexes, named GraphPep. GraphPep models protein–peptide interactions instead of traditional atoms or residues as graph nodes, and focuses on residue–residue contacts instead of a single peptide root mean square deviation in the loss function. Therefore, GraphPep can not only efficiently capture the most important protein–peptide interactions, but also mitigate the problem of limited training data. Moreover, the power of GraphPep is further enhanced by the ESM-2 protein language model. GraphPep is extensively evaluated on diverse decoy sets generated by various protein–peptide docking programs and AlphaFold, and is compared against state-of-the-art methods. The results demonstrate the accuracy and robustness of GraphPep.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The raw data of the evaluation results are provided in the Article and its Supplementary Information. The protein–peptide structures for testing in this Article were taken from the PDB. The training decoy set and test decoy sets are available via Zenodo at https://doi.org/10.5281/zenodo.17097750 (ref. 68). Source data are provided with this paper.
Code availability
The GraphPep package is freely available for academic or non-commercial users at http://huanglab.phys.hust.edu.cn/GraphPep/or via Zenodo at https://doi.org/10.5281/zenodo.17099863 (ref. 69).
Change history
17 February 2026
Since the version of the article initially published, the Major Project of Guangzhou National Laboratory grant no. has been corrected to GZNL2023A03007 in the HTML and PDF versions of the article.
References
Petsalaki, E. & Russell, R. B. Peptide-mediated interactions in biological systems: new discoveries and applications. Curr. Opin. Biotechnol. 19, 344–350 (2008).
Muttenthaler, M., King, G. F., Adams, D. J. & Alewood, P. F. Trends in peptide drug discovery. Nat. Rev. Drug Discov. 20, 309–325 (2021).
Lei, Y. et al. A deep-learning framework for multi-level peptide–protein interaction prediction. Nat. Commun. 12, 5465 (2021).
Zhao, Z., Peng, Z. & Yang, J. Improving sequence-based prediction of protein–peptide binding residues by introducing intrinsic disorder and a consensus method. J. Chem. Inf. Model. 58, 1459–1468 (2018).
Taherzadeh, G., Zhou, Y., Liew, A. W. & Yang, Y. Structure-based prediction of protein–peptide binding regions using Random Forest. Bioinformatics 34, 477–484 (2018).
Weng, G. et al. Comprehensive evaluation of fourteen docking programs on protein–peptide complexes. J. Chem. Theory Comput. 16, 3959–3969 (2020).
Neduva, V. et al. Systematic discovery of new recognition peptides mediating protein interaction networks. PLoS Biol. 3, e405 (2005).
Ciemny, M. et al. Protein–peptide docking: opportunities and challenges. Drug Discov. Today 23, 1530–1537 (2018).
Zhou, P., Jin, B., Li, H. & Huang, S. Y. HPEPDOCK: a web server for blind peptide–protein docking based on a hierarchical algorithm. Nucleic Acids Res. 46, W443–W450 (2018).
Zhou, P. et al. Hierarchical flexible peptide docking by conformer generation and ensemble docking of peptides. J. Chem. Inf. Model. 58, 1292–1302 (2018).
Zhang, Y. & Sanner, M. F. AutoDock CrankPep: combining folding and docking to predict protein–peptide complexes. Bioinformatics 35, 5121–5127 (2019).
Schindler, C. E., de Vries, S. J. & Zacharias, M. Fully blind peptide–protein docking with pepATTRACT. Structure 23, 1507–1515 (2015).
Lee, H., Heo, L., Lee, M. S. & Seok, C. GalaxyPepDock: a protein–peptide docking tool based on interaction similarity and energy optimization. Nucleic Acids Res. 43, W431–W435 (2015).
Yan, C., Xu, X. & Zou, X. Fully blind docking at the atomic level for protein–peptide complex structure prediction. Structure 24, 1842–1853 (2016).
Kurcinski, M. et al. CABS-dock standalone: a toolbox for flexible protein–peptide docking. Bioinformatics 35, 4170–4172 (2019).
Raveh, B., London, N. & Schueler-Furman, O. Sub-angstrom modeling of complexes between flexible peptides and globular proteins. Proteins 78, 2029–2040 (2010).
London, N., Raveh, B., Cohen, E., Fathi, G. & Schueler-Furman, O. Rosetta FlexPepDock web server—high resolution modeling of peptide–protein interactions. Nucleic Acids Res. 39, W249–W253 (2011).
Trellet, M., Melquiond, A. S. & Bonvin, A. M. A unified conformational selection and induced fit approach to protein–peptide docking. PLoS ONE 8, e58769 (2013).
Honorato, R. V. et al. The HADDOCK2.4 web server for integrative modeling of biomolecular complexes. Nat. Protoc. 19, 3219–3241 (2024).
Huang, S. Y. & Zou, X. An iterative knowledge-based scoring function for protein–protein recognition. Proteins 72, 557–579 (2008).
Feliu, E., Aloy, P. & Oliva, B. On the analysis of protein–protein interactions via knowledge-based potentials for the prediction of protein–protein docking. Protein Sci. 20, 529–541 (2011).
Liu, S. & Vakser, I. A. DECK: distance and environment-dependent, coarse-grained, knowledge-based potentials for protein–protein docking. BMC Bioinf. 12, 280 (2011).
Fink, F., Hochrein, J., Wolowski, V., Merkl, R. & Gronwald, W. PROCOS: computational analysis of protein–protein complexes. J. Comput. Chem. 32, 2575–2586 (2011).
Geng, C. et al. iScore: a novel graph kernel-based function for scoring protein–protein docking models. Bioinformatics 36, 112–121 (2020).
Jung, Y., Geng, C., Bonvin, A. M., Xue, L. C. & Honavar, V. G. MetaScore: a novel machine-learning-based approach to improve traditional scoring functions for scoring protein–protein docking conformations. Biomolecules 13, 121 (2023).
Renaud, N. et al. DeepRank: a deep learning framework for data mining 3D protein–protein interfaces. Nat. Commun. 12, 7068 (2021).
Rèau, M., Renaud, N., Xue, L. C. & Bonvin, A. M. DeepRank-GNN: a graph neural network framework to learn patterns in protein–protein interfaces. Bioinformatics 39, btac759 (2023).
McFee, M. & Kim, P. M. GDockScore: a graph-based protein–protein docking scoring function. Bioinform. Adv. 3, vbad072 (2023).
Wang, X., Terashi, G., Christoffer, C. W., Zhu, M. & Kihara, D. Protein docking model evaluation by 3D deep convolutional neural networks. Bioinformatics 36, 2113–2118 (2020).
Wang, X., Flannery, S. T. & Kihara, D. Protein docking model evaluation by graph neural networks. Front. Mol. Biosci. 8, 647915 (2021).
Mastropietro, A., Pasculli, G. & Bajorath, J. Learning characteristics of graph neural networks predicting protein–ligand affinities. Nat. Mach. Intell. 5, 1427–1436 (2023).
Hamilton, W., Ying, Z. & Leskovec, J. Inductive representation learning on large graphs. Adv. Neural Inf. Process. Syst. 30, 1024–1034 (2017).
Xu, K., Hu, W., Leskovec, J. & Jegelka, S. How powerful are graph neural networks? In Proc. 7th International Conference on Learning Representations https://openreview.net/pdf?id=ryGs6iA5Km (ICLR, 2019).
Johansson-åkhe, I., Mirabello, C. & Wallner, B. InterPepRank: assessment of docked peptide conformations by a deep graph network. Front. Bioinform. 1, 763102 (2021).
Johansson-åkhe, I. & Wallner, B. InterPepScore: a deep learning score for improving the FlexPepDock refinement protocol. Bioinformatics 38, 3209–3215 (2022).
Linsley, D. et al. Learning long-range spatial dependencies with horizontal gated recurrent units. Adv. Neural Inf. Process Syst. 31, 152–164 (2018).
Satorras, V. G., Hoogeboom, E. & Welling, M. E(n) equivariant graph neural networks. In International Conference on Machine Learning 9323–9332 (PMLR, 2021).
Fuchs, F., Worrall, D., Fischer, V. & Welling, M. SE(3)-transformers: 3D roto-translation equivariant attention networks. In 34th Conference on Neural Information Processing Systems (NeurIPS 2020) https://papers.neurips.cc/paper/2020/file/15231a7ce4ba789d13b722cc5c955834-Paper.pdf (NeurIPS, 2020).
Baek, M. et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 373, 871–876 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Evans, R. et al. Protein complex prediction with AlphaFold-Multimer. Preprint at bioRxiv https://doi.org/10.1101/2021.10.04.463034 (2021).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
Madani, A. et al. Large language models generate functional protein sequences across diverse families. Nat. Biotechnol. 41, 1099–1106 (2023).
Brandes, N., Ofer, D., Peleg, Y., Rappoport, N. & Linial, M. ProteinBERT: a universal deep-learning model of protein sequence and function. Bioinformatics 38, 2102–2110 (2022).
Yang, K. K., Fusi, N. & Lu, A. X. Convolutions are competitive with transformers for protein sequence pretraining. Cell Syst. 15, 286–294 (2024).
Xu, X. & Bonvin, A. M. DeepRank-GNN-esm: a graph neural network for scoring protein–protein models using protein language model. Bioinform. Adv. 4, vbad191 (2024).
Mariani, V., Biasini, M., Barbato, A. & Schwede, T. lDDT: a local superposition-free score for comparing protein structures and models using distance difference tests. Bioinformatics 29, 2722–2728 (2013).
Zhang, L. et al. ComplexQA: a deep graph learning approach for protein complex structure assessment. Brief. Bioinform. 24, bbad287 (2023).
Basu, S. & Wallner, B. DockQ: a quality measure for protein–protein docking models. PLoS ONE 11, e0161879 (2016).
Chen, X., Morehead, A., Liu, J. & Cheng, J. A gated graph transformer for protein complex structure quality assessment and its performance in CASP15. Bioinformatics 39, i308–i317 (2023).
Yang, Z., Zhong, W., Lv, Q. & Dong, T. Geometric Interaction Graph Neural Network for predicting protein–ligand binding affinities from 3D structures (GIGN). J. Phys. Chem. Lett. 14, 2020–2033 (2023).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Bresson, X. & Laurent, T. Residual gated graph ConvNets. Preprint at https://arxiv.org/abs/1711.07553 (2017).
Hauser, A. S. & Windshügel, B. LEADS-PEP: a benchmark data set for assessment of peptide docking performance. J. Chem. Inf. Model. 56, 188–200 (2015).
London, N., Movshovitz-Attias, D. & Schueler-Furman, O. The structural basis of peptide–protein binding strategies. Structure 18, 188–199 (2010).
Shanker, S. & Sanner, M. F. Predicting protein–peptide interactions: benchmarking deep learning techniques and a comparison with focused docking. J. Chem. Inf. Model. 63, 3158–3170 (2023).
Lee, J. H., Yin, R., Ofek, G. & Pierce, B. G. Structural features of antibody–peptide recognition. Front. Immunol. 13, 910367 (2022).
Su, M. et al. Comparative assessment of scoring functions: the CASF-2016 update. J. Chem. Inf. Model. 59, 895–913 (2019).
Buttenschoen, M., Morris, G. M. & Deane, C. M. PoseBusters: AI-based docking methods fail to generate physically valid poses or generalise to novel sequences. Chem. Sci. 15, 3130–3139 (2023).
Santos, K. B., Guedes, I. A., Karl, A. L. & Dardenne, L. E. Highly flexible ligand docking: benchmarking of the DockThor program on the LEADS-PEP protein–peptide data set. J. Chem. Inf. Model. 60, 667–683 (2020).
Loshchilov, I. & Hutter, F. Decoupled weight decay regularization. Preprint at https://arxiv.org/abs/1711.05101 (2017).
Janin, J. et al. CAPRI: a critical assessment of predicted interactions. Proteins 52, 2–9 (2003).
Shen, C. et al. Boosting protein–ligand binding pose prediction and virtual screening based on residue-atom distance likelihood potential and graph transformer. J. Med. Chem. 65, 10691–10706 (2022).
Lin, T. Y., Goyal, P., Girshick, R., He, K. & Dollar, P. Focal loss for dense object detection. IEEE Trans. Pattern Anal. Mach. Intell. 42, 318–327 (2020).
Wen, Z., He, J., Tao, H. & Huang, S. Y. PepBDB: a comprehensive structural database of biological peptide–protein interactions. Bioinformatics 35, 175–177 (2019).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Tao, H., Wang, X. & Huang, S. Y. An interaction-derived graph learning framework for scoring protein–peptide complexes. Zenodo https://doi.org/10.5281/zenodo.17097750 (2025).
Tao, H., Wang, X. & Huang, S. Y. GraphPep program. Zenodo https://doi.org/10.5281/zenodo.17099863 (2025).
Acknowledgements
This work is supported by the National Natural Science Foundation of China (grant nos. 32430020, 32161133002 and 62072199), the Major Project of Guangzhou National Laboratory (GZNL2023A03007) and the startup grant of Huazhong University of Science and Technology.
Author information
Authors and Affiliations
Contributions
S.-Y.H. conceived and supervised the project. H.T. performed the experiments. S.-Y.H. and H.T. analysed the data. H.T. and X.W. tested the program. H.T. and S.-Y.H. wrote the manuscript. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Machine Intelligence thanks Jianyi Yang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–7.
Supplementary Tables (download XLSX )
Supplementary Tables 1–11.
Supplementary Data 1 (download XLSX )
Source data for supplementary figures.
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tao, H., Wang, X. & Huang, SY. An interaction-derived graph learning framework for scoring protein–peptide complexes. Nat Mach Intell 7, 1858–1869 (2025). https://doi.org/10.1038/s42256-025-01136-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42256-025-01136-1
This article is cited by
-
Multi-AOP: a lightweight multi-view deep learning framework for antioxidant peptide discovery
Bioresources and Bioprocessing (2026)
-
AFP-GFuse: an antifungal peptide identification model with structural information fusion via multi-graph neural networks and cross-attention mechanism
Molecular Diversity (2025)