Abstract
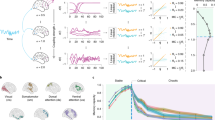
What underlies the emergence of cortex-aligned representations in deep neural network models of vision? Earlier work suggested that shared architectural constraints were a major factor, but the success of widely varied architectures after pretraining raises critical questions about the importance of architectural constraints. Here we show that in wide networks with minimal training, architectural inductive biases have a prominent role. We examined networks with varied architectures but no pretraining and quantified their ability to predict image representations in the visual cortices of monkeys and humans. We found that cortex-aligned representations emerge in convolutional architectures that combine two key manipulations of dimensionality: compression in the spatial domain, through pooling, and expansion in the feature domain by increasing the number of channels. We further show that the inductive biases of convolutional architectures are critical for obtaining performance gains from feature expansion—dimensionality manipulations were relatively ineffective in other architectures and in convolutional models with targeted lesions. Our findings suggest that the architectural constraints of convolutional networks are sufficiently close to the constraints of biological vision to allow many aspects of cortical visual representation to emerge even before synaptic connections have been tuned through experience.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The NSD is available at https://naturalscenesdataset.org/. The THINGS fMRI dataset is available at https://openneuro.org/datasets/ds004192. The monkey electrophysiology dataset is available as part of the Brain-Score GitHub package at https://github.com/brain-score. The Places dataset is available at http://places2.csail.mit.edu/index.html.
Code availability
The Expansion model, as well as code for all analyses, is available via GitHub at https://github.com/akazemian/untrained_models_of_visual_cortex (ref. 61).
References
Carandini, M. et al. Do we know what the early visual system does? J. Neurosci. 25, 10577–10597 (2005).
Jones, J. P. & Palmer, L. A. The two-dimensional spatial structure of simple receptive fields in cat striate cortex. J. Neurophysiol. 58, 1187–1211 (1987).
Movshon, J. A., Thompson, I. D. & Tolhurst, D. J. Spatial summation in the receptive fields of simple cells in the cat’s striate cortex. J. Physiol. 283, 53–77 (1978).
Riesenhuber, M. & Poggio, T. Hierarchical models of object recognition in cortex. Nat. Neurosci. 2, 1019–1025 (1999).
Agrawal, P., Stansbury, D., Malik, J. & Gallant, J. L. Pixels to voxels: modeling visual representation in the human brain. Preprint at https://arxiv.org/abs/1407.5104 (2014).
Khaligh-Razavi, S.-M. & Kriegeskorte, N. Deep supervised, but not unsupervised, models may explain IT cortical representation. PLoS Comput. Biol. 10, e1003915 (2014).
Yamins, D. L. K. et al. Performance-optimized hierarchical models predict neural responses in higher visual cortex. Proc. Natl Acad. Sci. USA 111, 8619–8624 (2014).
Chen, Z. & Bonner, M. F. Universal dimensions of visual representation. Sci. Adv. 11, eadw7697 (2025).
Conwell, C., Prince, J. S., Kay, K. N., Alvarez, G. A. & Konkle, T. A large-scale examination of inductive biases shaping high-level visual representation in brains and machines. Nat. Commun. 15, 9383 (2024).
Elmoznino, E. & Bonner, M. F. High-performing neural network models of visual cortex benefit from high latent dimensionality. PLoS Comput. Biol. 20, e1011792 (2024).
Saxe, A., Nelli, S. & Summerfield, C. If deep learning is the answer, what is the question? Nat. Rev. Neurosci. 22, 55–67 (2021).
Serre, T. Deep learning: the good, the bad, and the ugly. Annu. Rev. Vis. Sci. 5, 399–426 (2019).
Storrs, K. R., Kietzmann, T. C., Walther, A., Mehrer, J. & Kriegeskorte, N. Diverse deep neural networks all predict human inferior temporal cortex well, after training and fitting. J. Cogn. Neurosci. 10, 1–21 (2021)
Cao, R. & Yamins, D. Explanatory models in neuroscience, part 1: taking mechanistic abstraction seriously. Cogn. Syst. Res. 87, 101244 (2024).
Cao, R. & Yamins, D. Explanatory models in neuroscience, part 2: functional intelligibility and the contravariance principle. Cogn. Syst. Res. 85, 101200 (2024).
Konkle, T. & Alvarez, G. A. A self-supervised domain-general learning framework for human ventral stream representation. Nat. Commun. 13, 491 (2022).
Kriegeskorte, N. Deep neural networks: a new framework for modeling biological vision and brain information processing. Annu. Rev. Vis. Sci. 1, 417–446 (2015).
Richards, B. A. et al. A deep learning framework for neuroscience. Nat. Neurosci. 22, 1761–1770 (2019).
Yamins, D. L. K. & DiCarlo, J. J. Using goal-driven deep learning models to understand sensory cortex. Nat. Neurosci. 19, 356–365 (2016).
Bruna, J. & Mallat, S. Invariant scattering convolution networks. IEEE Trans. Pattern Anal. Mach. Intell. 35, 1872–1886 (2013).
Pogoncheff, G. et al. Explaining V1 properties with a biologically constrained deep learning architecture. Adv. Neural Inf. Process. Syst. 36, 13908–13930 (2023).
Yue, X., Pourladian, I. S., Tootell, R. B. H. & Ungerleider, L. G. Curvature-processing network in macaque visual cortex. Proc. Natl Acad. Sci. USA 111, e3467–e3475 (2014).
Yue, X., Robert, S. & Ungerleider, L. G. Curvature processing in human visual cortical areas. NeuroImage 222, 117295 (2020).
Majaj, N. J., Hong, H., Solomon, E. A. & DiCarlo, J. J. Simple learned weighted sums of inferior temporal neuronal firing rates accurately predict human core object recognition performance. J. Neurosci. 35, 13402–13418 (2015).
Allen, E. J. et al. A massive 7T fMRI dataset to bridge cognitive neuroscience and artificial intelligence. Nat. Neurosci. 25, 116–126 (2022).
Hebart, M. N. et al. THINGS-data, a multimodal collection of large-scale datasets for investigating object representations in human brain and behavior. eLife 12, e82580 (2023).
Schrimpf, M. et al. Brain-Score: which artificial neural network for object recognition is most brain-like? Preprint at bioRxiv https://doi.org/10.1101/407007 (2018).
Casper, S. Frivolous units: wider networks are not really that wide. In Proc. 35th AAAI Conference on Artificial Intelligence 6921–6929 (AAAI, 2021).
Poggio, T., Mhaskar, H., Rosasco, L., Miranda, B. & Liao, Q. Why and when can deep-but not shallow-networks avoid the curse of dimensionality: a review. Int. J. Autom. Comput. 14, 503–519 (2017).
Zhou, B., Lapedriza, A., Khosla, A., Oliva, A. & Torralba, A. Places: a 10 million image database for scene recognition. IEEE Trans. Pattern Anal. Mach. Intell. 40, 1452–1464 (2018).
Oliva, A. & Torralba, A. Modeling the shape of the scene: a holistic representation of the spatial envelope. Int. J. Comput. Vis. 42, 145–175 (2001).
Portilla, J. & Simoncelli, E. P. A parametric texture model based on joint statistics of complex wavelet coefficients. Int. J. Comput. Vis. 40, 49–70 (2000).
Cordonnier, J.-B., Loukas, A. & Jaggi, M. On the relationship between self-attention and convolutional layers. In International Conference on Learning Representations (2020).
Dosovitskiy, A. et al. An image is worth 16x16 words: transformers for image recognition at scale. In International Conference on Learning Representations (2021).
Ingrosso, A. & Goldt, S. Data-driven emergence of convolutional structure in neural networks. Proc. Natl Acad. Sci. USA 119, e2201854119 (2022).
Jarrett, K., Kavukcuoglu, K., Ranzato, M. & LeCun, Y. What is the best multi-stage architecture for object recognition? In 2009 IEEE 12th International Conference on Computer Vision 2146–2153 (IEEE, 2009).
Saxe, A. M. et al. On random weights and unsupervised feature learning. In Proc. 28th International Conference on Machine Learning 2 (2011).
Cao, Y.-H. & Wu, J. A random CNN sees objects: one inductive bias of CNN and its applications. In AAAI Conference on Artificial Intelligence 194–202 (AAAI, 2022).
Baek, S., Song, M., Jang, J., Kim, G. & Paik, S.-B. Face detection in untrained deep neural networks. Nat. Commun. 12, 7328 (2021).
Geiger, F., Schrimpf, M., Marques, T. & DiCarlo, J. J. Wiring up vision: minimizing supervised synaptic updates needed to produce a primate ventral stream. In International Conference on Learning Representations (2022).
Cadena, S. A. et al. Deep convolutional models improve predictions of macaque V1 responses to natural images. PLoS Comput. Biol. 15, e1006897 (2019).
Shi, J. et al. Comparison against task driven artificial neural networks 846 reveals functional properties in mouse visual cortex. Adv. Neural Inf. Process. Syst. 32, 5674–5774 (2019).
Chang, H. & Futagami, K. Reinforcement learning with convolutional reservoir computing. Appl. Intell. 50, 2400–2410 (2020).
Jaeger, H. The ‘Echo State’ Approach to Analysing and Training Recurrent Neural Networks—With an Erratum Note GMD Technical Report 148 (German National Research Center for Information Technology, 2001).
Mei, S. & Montanari, A. The generalization error of random features regression: precise asymptotics and the double descent curve. Commun. Pure Appl. Math. 75, 667–766 (2022).
Mei, S., Misiakiewicz, T. & Montanari, A. Generalization error of random feature and kernel methods: hypercontractivity and kernel matrix concentration. Appl. Comput. Harmon. Anal. 59, 3–84 (2022).
Rahimi, A. & Recht, B. Random features for large-scale kernel machines. Adv. Neural Inf. Process. Syst. 20, 1177–1184 (2007).
Tong, Z. & Tanaka, G. Reservoir computing with untrained convolutional neural networks for image recognition. In Proc. 24th International Conference on Pattern Recognition 1289–1294 (IEEE, 2018).
Teney, D., Nicolicioiu, A. M., Hartmann, V. & Abbasnejad, E. Neural redshift: random networks are not random functions. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 4786–4796 (IEEE, 2024).
Jacot, A., Gabriel, F. & Hongler, C. Neural tangent kernel: convergence and generalization in neural networks. Adv. Neural Inf. Process. Syst. 31, 8580–8589 (2018).
Bonner, M. F. & Epstein, R. A. Object representations in the human brain reflect the co-occurrence statistics of vision and language. Nat. Commun. 12, 4081 (2021).
Doerig, A. et al. High-level visual representations in the human brain are aligned with large language models. Nat. Mach. Intell. 7, 1220–1234 (2025).
Wang, A. Y., Kay, K., Naselaris, T., Tarr, M. J. & Wehbe, L. Better models of human high-level visual cortex emerge from natural language supervision with a large and diverse dataset. Nat. Mach. Intell. 5, 1415–1426 (2023).
Babadi, B. & Sompolinsky, H. Sparseness and expansion in sensory representations. Neuron 83, 1213–1226 (2014).
Cayco-Gajic, N. A. & Silver, R. A. Re-evaluating circuit mechanisms underlying pattern separation. Neuron 101, 584–602 (2019).
Sakai, J. How synaptic pruning shapes neural wiring during development and, possibly, in disease. Proc. Natl Acad. Sci. USA 117, 16096–16099 (2020).
Pytorch image models. timmdocs https://timm.fast.ai/ (2022).
Lin, T.-Y. et al. Microsoft COCO: common objects in context. In Proc. Computer Vision—ECCV 2014 (eds Fleet, D. et al.) 740–755 (Springer, 2014).
Kay, K. N., Rokem, A., Winawer, J., Dougherty, R. F. & Wandell, B. A. GLMdenoise: a fast, automated technique for denoising task-based fMRI data. Front. Neurosci. 7, 247 (2013).
Spearman, C. The proof and measurement of association between two things. Am. J. Psychol. 15, 72–101 (1904).
Kazemian, A. akazemian/untrained_models_of_visual_cortex: initial release. Zenodo https://doi.org/10.5281/ZENODO.16920087 (2025).
Acknowledgements
This work was supported in part by a JHU Catalyst award to M.F.B.
Author information
Authors and Affiliations
Contributions
A.K. and M.F.B. wrote the paper. E.E. provided feedback on the analyses and writing. A.K. performed the research and analysed the data. M.F.B. supervised the research. A.K., E.E. and M.F.B. conceived of the research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Machine Intelligence thanks David Klindt and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Dimensionality expansion strongly improves the encoding performance of convolutional neural networks in another large-scale fMRI dataset.
a) The effects of dimensionality expansion were examined for a convolutional neural network, a deep fully connected network, and a vision transformer. b) These plots illustrate the encoding performance for all three architectures as a function of dimensionality expansion. There was no pre-training for these networks. Encoding performance was evaluated for regions V1 to V4 from the large-scale THINGS fMRI dataset. The x-axis plots the number of random features in the output layer, and the y-axis shows the encoding score for predicting image-evoked cortical responses. The gray dashed line indicates the performance of the best performing convolutional layer of pre-trained AlexNet. The convolutional architecture without pre-training attained large performance gains as a function of dimensionality expansion. In contrast, the other two architectures showed much less improvement as the number of output features was expanded. The encoding performance plots show the mean performance across voxels from all participants, and the error bars denote 97.5% confidence intervals from 1,000 bootstrap samples. c) A summary statistic for the effect of dimensionality expansion was calculated as the difference in performance between the highest- and lowest-dimensional version of each architecture, using the mean performance across voxels from all participants. The performance of the convolutional model was strongly modulated by the dimensionality of its random feature space. In contrast, the effect of dimensionality expansions was much weaker in the other two architectures.
Extended Data Fig. 2 The benefits of dimensionality expansion depend on linear reweighting.
The effects of dimensionality expansion were examined for a convolutional neural network, a deep fully connected network, and a vision transformer. These plots illustrate representational similarity analysis (RSA) scores for all three architectures as a function of dimensionality expansion. There was no pre-training for these networks. a, RSA scores were evaluated for regions V4 and IT in the monkey electrophysiology data. The x-axis plots the number of random features in the output layer, and the y-axis shows the RSA score. The gray dashed line indicates the performance of the best performing convolutional layer of pre-trained AlexNet. As expected, the RSA scores do not systematically increase as a function of dimensionality expansion. This contrasts with the effect of dimensionality expansion observed for the encoding scores in Fig. 2, and it demonstrates that a linear reweighting procedure is needed to yield performance gains from dimensionality expansion in untrained networks. These plots show the mean RSA correlations across all participants. b, These plots show the same analyses as in panel a but for regions along the ventral stream in the human fMRI data.
Extended Data Fig. 3 Comparisons with other trained deep neural network architectures.
a) These plots show the encoding performance of state-of-the-art trained architectures for comparison with the untrained networks and trained AlexNet. For each untrained architecture, these plots show performance at both the smallest and largest levels of dimensionality. In addition to AlexNet, these plots include more modern architectures, specifically ViT and BarlowTwins, pre-trained on ImageNet. Encoding performance was evaluated for regions V4 and IT in the monkey electrophysiology data. The encoding performance plots in this panel and in panel b show the mean performance across units/voxels from all participants, and the error bars denote 97.5% confidence intervals from 1,000 bootstrap samples. The blue dashed line shows the mean noise ceiling across units/voxels. b) These plots show the same analyses as in panel a but for regions along the ventral stream in the human fMRI data.
Extended Data Fig. 4 Variance partitioning shows that the Expansion model explains the same variance as trained AlexNet.
Variance partitioning was used to determine whether the Expansion model explains the same variance in cortical responses as trained AlexNet. These analyses were performed using the largest convolutional model, and they show that in both the monkey electrophysiology data (a) and human fMRI data (b) the explained variance of the Expansion model is fully shared with AlexNet. As expected, AlexNet explains additional unique variance that is not shared with the Expansion model.
Extended Data Fig. 5 Improvements in encoding performance are linked to increases in latent dimensionality.
a) These plots illustrate the encoding performance of the networks from Fig. 2 as a function of their latent dimensionality. Encoding performance was evaluated for regions V4 and IT in the monkey electrophysiology data. Plotting conventions are the same as in Fig. 2, except that here the x-axis plots the number of principal components that account for 85% of variance in the output layer. The results show that improvements in encoding performance are closely linked to increases in the latent dimensionality of a network’s natural image representations. However, note that measures of latent dimensionality are architecture-dependent, and thus different architectures with the same level of latent dimensionality can nonetheless differ in encoding performance. b) These plots show the same analyses as in panel a but for regions along the ventral stream in the human fMRI data.
Extended Data Fig. 6 Analysis of architectural components in the final layer of the convolutional network.
a-b) These plots show the performance of the largest untrained convolutional network after altering key architectural components in the final layer only. Panel a shows results for macaque IT, and panel b shows results for the human high-level ventral stream. These plots show that ablating the nonlinearity in the final layer (No ReLU) has little effect, which means that the crucial nonlinear operations are those that occur in earlier layers. On the other hand, ablating the convolution and pooling operations (Linear Projection) reduces performance, demonstrating the importance of these operations in the final layer. They also show that removing the spatial locality of the convolutional filters in the final layer (No Spatial Continuity) results in an overall decrease in encoding performance, demonstrating that even in the highest layer of the network it is beneficial to compute spatially local representations. Plotting conventions are the same as in Fig. 2.
Extended Data Fig. 7 Effect of pre-defined wavelets in the first layer of the untrained convolutional neural network.
These plots show the effect of using pre-defined wavelets in the first layer of the untrained convolutional network. Encoding performance is shown for macaque IT (a) and the human high-level ventral stream (b). For comparison, we examined a model with 3,000 randomly initialized filters in layer 1 (the number of random filters was maximized within computational memory limits). As in Fig. 2, these plots show how encoding performance changes as a function of dimensionality expansion in the final convolutional layer. The x-axis plots the number of random features in the output layer, and the y-axis shows the encoding score for predicting image-evoked cortical responses. The gray dashed line indicates the performance of the best-performing convolutional layer of pre-trained AlexNet. There is a small, but consistent drop in performance for the fully random model across both datasets, demonstrating that overall, the network benefits from the implementation of pre-defined wavelets in its first layer.
Extended Data Fig. 8 Different activation functions yield similar encoding performance for the untrained convolutional neural network.
The effects of using different nonlinear activation functions in our untrained convolutional network were explored for monkey IT (a) and the human high-level ventral stream (b). These plots illustrate encoding performance for models with different nonlinear activation functions with otherwise identical architectures, all containing 105 features in their output layer. For comparison, this plot also includes a network without any nonlinear activation functions (the Linear model). The y-axis shows the encoding score for predicting image-evoked cortical responses. The results demonstrate that the inclusion of nonlinearities is critical, but various types of nonlinearities yield similar levels of performance. ReLU = rectified linear unit, GELU = Gaussian error linear unit, ELU = exponential linear unit.
Extended Data Fig. 9 Different random initialization methods yield similar encoding performance for the untrained convolutional neural network.
The effects of initializing the random features of our untrained convolutional network using different methods were explored for monkey IT (a) and human ventral visual stream (b). These plots illustrate encoding performance for identical architectures with different random initialization types. As in Fig. 2, the x-axis plots the number of random features in the output layer, and the y-axis shows the encoding score for predicting image-evoked cortical responses. There is variation in encoding performance for different initialization methods in models with low dimensionality (on the left side of the x-axis). However, at higher levels of dimensionality, these performance differences diminish. This indicates that the type of initialization has minimal impact on encoding performance in the presence of model expansion.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1–3.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kazemian, A., Elmoznino, E. & Bonner, M.F. Convolutional architectures are cortex-aligned de novo. Nat Mach Intell 7, 1834–1844 (2025). https://doi.org/10.1038/s42256-025-01142-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42256-025-01142-3
This article is cited by
-
Structure as an inductive bias for brain–model alignment
Nature Machine Intelligence (2025)