Abstract
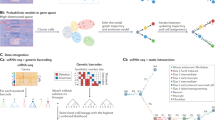
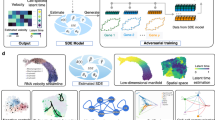
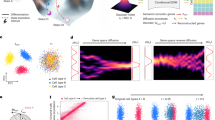
Single-cell sequencing measurements facilitate the reconstruction of dynamic biology by capturing snapshot molecular profiles of individual cells. Cell fate decisions in development and disease are orchestrated through an intricate balance of deterministic and stochastic regulatory events. Drift-diffusion equations are effective in modelling single-cell dynamics from high-dimensional single-cell measurements. While existing solutions describe the deterministic dynamics associated with the drift term of these equations at the level of cell state, diffusion is modelled as a constant across cell states. To fully understand the dynamic regulatory logic in development and disease, models explicitly attuned to the balance between deterministic and stochastic biology are required. To address these limitations, we introduce scDiffEq, a generative framework for learning neural stochastic differential equations that approximate biology’s deterministic and stochastic dynamics. Using lineage-traced single-cell data, we demonstrate that scDiffEq offers an improved reconstruction of cell trajectories and prediction of cell fate from multipotent progenitors during haematopoiesis. By imparting in silico perturbations to multipotent progenitor cells, we find that scDiffEq accurately recapitulates the dynamics of CRISPR-perturbed haematopoiesis. We generalize this approach beyond lineage-traced or multi-time-point datasets to model the dynamics of single-cell data from a single time point. Using scDiffEq, we simulate high-resolution developmental cell trajectories, which can model their drift and diffusion, enabling us to study their time-dependent gene-level dynamics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All datasets used in this study were previously published and are publicly available. The LARRY dataset is available via GitHub at https://github.com/AllonKleinLab/paper-data/tree/master/Lineage_tracing_on_transcriptional_landscapes_links_state_to_fate_during_differentiation (commit: af842ce)26. The human haematopoiesis dataset, including the final list of filtered cells was provided by the authors of ref. 9 and is available under the GEO accession number GSE193517 as well as via direct download using the ‘dynamo’ Python package9. The pancreatic endocrinogenesis dataset is available via GitHub at https://github.com/theislab/scvelo_notebooks/raw/master/data/Pancreas/endocrinogenesis_day15.h5ad (commit: 2425e7f)26,43. All data generated in this study are available via Zenodo at https://doi.org/10.5281/zenodo.17238611 (ref. 53).
Code availability
An implementation of scDiffEq is available as a Python package via GitHub at https://github.com/scDiffEq/scDiffEq and https://pypi.org/project/scdiffeq/ (ref. 54). All analysis code, notebooks and scripts to reproduce the results presented in this paper are deposited in a dedicated reproducibility repository via GitHub at https://github.com/scDiffEq/scdiffeq-analyses (ref. 53). Comprehensive documentation is accessible at https://scdiffeq.com.
References
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Chen, H. et al. Single-cell trajectories reconstruction, exploration and mapping of omics data with STREAM. Nat. Commun. 10, 1903 (2019).
Saelens, W., Cannoodt, R., Todorov, H. & Saeys, Y. A comparison of single-cell trajectory inference methods. Nat. Biotechnol. 37, 547–554 (2019).
Wang, S.-W., Herriges, M. J., Hurley, K., Kotton, D. N. & Klein, A. M. CoSpar identifies early cell fate biases from single-cell transcriptomic and lineage information. Nat. Biotechnol. 40, 1066–1074 (2022).
Schiebinger, G. et al. Optimal-transport analysis of single-cell gene expression identifies developmental trajectories in reprogramming. Cell 176, 1517 (2019).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Qiu, X. et al. Mapping transcriptomic vector fields of single cells. Cell 185, 690–711 (2022).
Lange, M. et al. CellRank for directed single-cell fate mapping. Nat. Methods 19, 159–170 (2022).
Weiler, P., Lange, M., Klein, M., Pe’er, D. & Theis, F. CellRank 2: unified fate mapping in multiview single-cell data. Nat. Methods 21, 1196–1205 (2024).
Gorin, G., Fang, M., Chari, T. & Pachter, L. RNA velocity unraveled. PLoS Comput. Biol. 18, e1010492 (2022).
Weinreb, C., Wolock, S., Tusi, B. K., Socolovsky, M. & Klein, A. M. Fundamental limits on dynamic inference from single-cell snapshots. Proc. Natl Acad. Sci. USA 115, E2467–E2476 (2018).
Hashimoto, T., Gifford, D. & Jaakkola, T. Learning population-level diffusions with generative RNNs. In Proc. 33rd International Conference on Machine Learning (eds Balcan, M. F. & Weinberger, K. Q.) Vol. 48, 2417–2426 (PMLR, 2016).
Yeo, G. H. T., Saksena, S. D. & Gifford, D. K. Generative modeling of single-cell time series with PRESCIENT enables prediction of cell trajectories with interventions. Nat. Commun. 12, 3222 (2021).
Li, Q. scTour: a deep learning architecture for robust inference and accurate prediction of cellular dynamics. Genome Biol. 24, 149 (2023).
Sha, Y., Qiu, Y., Zhou, P. & Nie, Q. Reconstructing growth and dynamic trajectories from single-cell transcriptomics data. Nat. Mach. Intell. 6, 25–39 (2024).
Tong, A., Huang, J., Wolf, G., van Dijk, D. & Krishnaswamy, S. TrajectoryNet: a dynamic optimal transport network for modeling cellular dynamics. Proc. Mach. Learn Res. 119, 9526–9536 (2020).
Huguet, G. et al. Manifold interpolating optimal-transport flows for trajectory inference. Adv. Neural Inf. Process. Syst. 35, 29705–29718 (2022).
Elowitz, M. B., Levine, A. J., Siggia, E. D. & Swain, P. S. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002).
Zechner, C., Nerli, E. & Norden, C. Stochasticity and determinism in cell fate decisions. Development 147, dev181495 (2020).
Chen, R. T. Q., Rubanova, Y., Bettencourt, J. & Duvenaud, D. K. Neural ordinary differential equations. Adv. Neural. Inf. Process. Syst. 31, 6572–6583 (2018).
Kidger, P., Foster, J., Li, X. & Lyons, T. J. Neural SDEs as infinite-dimensional GANs. In Proc. 38th International Conference on Machine Learning (eds Meila, M. & Zhang, T.) Vol. 139, 5453–5463 (PMLR, 2021).
Kidger, P. On Neural Differential Equations. DPhil thesis, University of Oxford (2021).
Jiang, Q. & Wan, L. A physics-informed neural SDE network for learning cellular dynamics from time-series scRNA-seq data. Bioinformatics 40, ii120–ii127 (2024).
Weinreb, C., Rodriguez-Fraticelli, A., Camargo, F. D. & Klein, A. M. Lineage tracing on transcriptional landscapes links state to fate during differentiation. Science 367, eaaw3381 (2020).
VanHorn, S. & Morris, S. A. Next-generation lineage tracing and fate mapping to interrogate development. Dev. Cell 56, 7–21 (2021).
Alemany, A., Florescu, M., Baron, C. S., Peterson-Maduro, J. & van Oudenaarden, A. Whole-organism clone tracing using single-cell sequencing. Nature 556, 108–112 (2018).
Biddy, B. A. et al. Single-cell mapping of lineage and identity in direct reprogramming. Nature 564, 219–224 (2018).
Bowling, S. et al. An engineered CRISPR–Cas9 mouse line for simultaneous readout of lineage histories and gene expression profiles in single cells. Cell 181, 1693–1694 (2020).
Frieda, K. L. et al. Synthetic recording and in situ readout of lineage information in single cells. Nature 541, 107–111 (2017).
Raj, B. et al. Simultaneous single-cell profiling of lineages and cell types in the vertebrate brain. Nat. Biotechnol. 36, 442–450 (2018).
Spanjaard, B. et al. Simultaneous lineage tracing and cell-type identification using CRISPR-Cas9-induced genetic scars. Nat. Biotechnol. 36, 469–473 (2018).
Bunne, C. et al. Learning single-cell perturbation responses using neural optimal transport. Nat. Methods 20, 1759–1768 (2023).
Giladi, A. et al. Single-cell characterization of haematopoietic progenitors and their trajectories in homeostasis and perturbed haematopoiesis. Nat. Cell Biol. 20, 836–846 (2018).
Feinberg, M. W. et al. The Kruppel-like factor KLF4 is a critical regulator of monocyte differentiation. EMBO J. 26, 4138–4148 (2007).
Yamanaka, R. et al. Impaired granulopoiesis, myelodysplasia, and early lethality in CCAAT/enhancer binding protein epsilon-deficient mice. Proc. Natl Acad. Sci. USA 94, 13187–13192 (1997).
Serwas, N. K. et al. Mutant specific granule deficiency correlates with aberrant granule organization and substantial proteome alterations in neutrophils. Front. Immunol. 9, 588 (2018).
Przybyla, L. & Gilbert, L. A. A new era in functional genomics screens. Nat. Rev. Genet. 23, 89–103 (2022).
Hounkpe, B. W., Chenou, F., de Lima, F. & De Paula, E. V. HRT Atlas v1.0 database: redefining human and mouse housekeeping genes and candidate reference transcripts by mining massive RNA-seq datasets. Nucleic Acids Res. 49, D947–D955 (2021).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Qiu, Q. et al. Massively parallel and time-resolved RNA sequencing in single cells with scNT-seq. Nat. Methods 17, 991–1001 (2020).
Bastidas-Ponce, A. et al. Comprehensive single cell mRNA profiling reveals a detailed roadmap for pancreatic endocrinogenesis. Development 146, dev173849 (2019).
Gulati, G. S. et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science 367, 405–411 (2020).
Liggett, L. A. & Sankaran, V. G. Unraveling hematopoiesis through the lens of genomics. Cell 182, 1384–1400 (2020).
Dahl, R., Iyer, S. R., Owens, K. S., Cuylear, D. D. & Simon, M. C. The transcriptional repressor GFI-1 antagonizes PU.1 activity through protein-protein interaction. J. Biol. Chem. 282, 6473–6483 (2007).
Orkin, S. H. & Zon, L. I. Hematopoiesis: an evolving paradigm for stem cell biology. Cell 132, 631–644 (2008).
Lipman, Y., Chen, R. T. Q., Ben-Hamu, H., Nickel, M. & Le, M. Flow matching for generative modeling. Int. Conf. Learn. Represent. (ICLR, 2023).
Tong, A. et al. Simulation-free Schrödinger bridges via score and flow matching. Proc. 27th Int. Conf. Artif. Intell. Stat. (AISTATS) 238, 1279–1287 (2024).
Virshup, I. et al. The scverse project provides a computational ecosystem for single-cell omics data analysis. Nat. Biotechnol. 41, 604–606 (2023).
Feydy, J. et al. Interpolating between optimal transport and MMD using Sinkhorn divergences. In Proc. Twenty-Second International Conference on Artificial Intelligence and Statistics (eds Chaudhuri, K. & Sugiyama, M.) Vol. 89, 2681–2690 (PMLR, 2019).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Vinyard, M. scDiffEq/scdiffeq-analyses: v.0.1.0rc0. Zenodo https://doi.org/10.5281/zenodo.17238611 (2025).
Vinyard, M. scDiffEq/scDiffEq: v0.1.10rc0. Zenodo https://doi.org/10.5281/zenodo.17238594 (2025).
Acknowledgements
We graciously thank C. Weinreb for their assistance in using the LARRY dataset. We thank S. Saksena, G. H. Ting Yeo and D. K. Gifford for their assistance in creating a benchmark comparison to PRESCIENT15. We thank X. Qiu for their assistance in comparing scDiffEq with Dynamo and using the human haematopoiesis scNT-seq dataset. We thank the entire Pinello and Getz Laboratories for thoughtful feedback and discussion throughout the preparation of this paper. We thank S. Anand for pivotal discussions that guided us towards the useful implementation of neural differential equations. We thank A. Ravi for useful conceptual design discussions. We thank D. Jia, J. Zhou and H. Levine for useful conversations and assistance in simulating synthetic datasets for proof-of-concept experiments. We thank M. Miller for editorial assistance. While working on this paper, M.E.V. was supported by the National Institutes of Health (NIH) under the Ruth L. Kirschstein National Research Service Award (grant no. 1F31CA257625) from the National Cancer Institute (NCI). L.P. was supported by the National Institutes of Health (NIH) Genomic Innovator Award (grant no. 1R35HG010717-01) and the Rappaport MGH Research Scholar Award 2024-2029. G.G. was partially funded by the Paul C. Zamecnik Chair in Oncology at the Mass General Cancer Center.
Author information
Authors and Affiliations
Contributions
All authors contributed to the conceptualization of the methodology, experiments and analyses. M.E.V. performed the M.E.V., A.W.R. and R.L. performed the fate-prediction benchmarking experiments. M.E.V. performed the time-point interpolation experiments. M.E.V. performed the gene perturbation experiments. M.E.V. and R.L. performed the investigation of model attributes and gene-level analyses. M.E.V. wrote the software package. All authors wrote the paper. A.M.K. provided guidance and edited the paper. G.G. and L.P. provided supervision and guidance in designing the experimental strategy and funded the research.
Corresponding authors
Ethics declarations
Competing interests
G.G. receives research funds from IBM, Pharmacyclics/Abbvie, Bayer, Genentech, Calico, Ultima Genomics, Inocras, Google, Kite and Novartis and is an inventor on patent applications filed by the Broad Institute related to MSMuTect, MSMutSig, POLYSOLVER, SignatureAnalyzer-GPU, MSEye, MinimuMM-seq and DLBclass. He is a founder and consultant and holds privately held equity in Scorpion Therapeutics; he is also a founder of, and holds privately held equity in, Predicta Biosciences; and holds privately held equity in Antares Therapeutics. M.E.V. is a founder of, and holds privately held equity in, Quintessence Laboratories, Inc. The other authors declare no competing interests.
Peer review
Peer review information
Nature Machine Intelligence thanks Xiuwei Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–27 and Notes 1–3.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Vinyard, M.E., Rasmussen, A.W., Li, R. et al. Learning cell dynamics with neural differential equations. Nat Mach Intell 7, 1969–1984 (2025). https://doi.org/10.1038/s42256-025-01150-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42256-025-01150-3