Abstract
Deep learning foundation models are becoming increasingly popular for use in bioactivity prediction. Recently, Feng et al. developed ActFound, a bioactive foundation model that jointly uses pairwise learning and meta-learning. By utilizing these techniques, the model is capable of being fine-tuned to a more specific bioactivity task with only a small amount of new data. Here, to investigate the generalizability of the model, we looked to fine-tune the foundation model on an antibacterial natural products (NPs) dataset. Large, labelled NPs datasets, which are needed to train traditional deep learning methods, are scarce. Therefore, the bioactivity prediction of NPs is an ideal task for foundation models. We studied the performance of ActFound on the NPs dataset using a range of few-shot settings. Additionally, we compared ActFound’s performance with those of other state-of-the-art models in the field. We found ActFound was unable to reach the same level of accuracy on the antibacterial NPs dataset as it did on other cross-domain tasks reported in the original publication. However, ActFound displayed comparable or better performance compared to the other models studied, especially at the low-shot settings. Our results establish ActFound as a useful foundation model for the bioactivity prediction of tasks with limited data, particularly for datasets that contain the bioactivities of similar compounds.
Similar content being viewed by others
Main
The bioactivity of compounds plays a key role in drug discovery. Bioactivity refers to the effect, either beneficial or adverse, a compound has on a biological process1. It encompasses the efficacy, potency and selectivity of a compound and is important in the identification of hits in a drug campaign and subsequent lead optimization2. Deep learning (DL) approaches have shown promise in their ability to predict the bioactivity of compounds3,4,5,6,7. However, DL models require large, high-quality datasets to accurately identify patterns within the data8. Many bioactivity tasks do not have an adequate number of labelled data sufficient for training. One solution is to use foundation models, which are pretrained on large, general datasets. These models serve as a ‘foundation’ that can be fine-tuned for more specific tasks9.
Recently, Feng et al. introduced ActFound10, a meta-learning foundation model trained to predict the bioactivity of compounds. To accomplish this, ActFound jointly utilizes meta-learning and pairwise learning. Meta-learning, or ‘learning to learn’, is a commonly used algorithm to develop foundation models11,12. Meta-learning models are trained on a variety of tasks with the intention of creating a model that can quickly adapt to new tasks from only a small amount of new data13. By using meta-learning, Feng et al.10 were able to pretrain their model on a wide range of diverse assays, leveraging the information to develop a general foundation model capable of few-shot learning. Pairwise learning was used to address the incompatibility of information within training assays differing in metrics, units and value ranges. Instead of directly predicting the bioactivity of compounds, pairwise learning allows ActFound to predict the difference in bioactivity between two compounds within the same assay14. During the fine-tuning stage, ActFound utilized the algorithm k-nearest neighbours model-agnostic meta-learning (kNN-MAML), which identifies assays within the training set that are similar to the fine-tuning assay. Leveraging information from similar assays allows for rapid fine-tuning to a new, unseen assay.
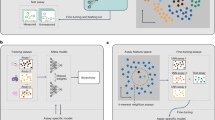
In this Reusability Report, we looked to study ActFound’s performance on a natural products (NPs) dataset that contains plant-derived compounds with antibacterial activity15. NPs are an abundant source of antibiotics, with many approved antibiotics being NPs or NP derivatives16. However, antibacterial NPs have historically been plagued by the need for a dereplication process, to avoid frequent rediscovery of known NPs17. The lack of new antibiotics being identified has led to a rising interest in DL models for antibacterial activity prediction18,19,20,21. However, there is a lack of large, labelled bioactivity datasets in the NPs field22. This makes it an ideal task for a pairwise meta-learning model like ActFound. In this study, we fine-tuned both the ActFound model pretrained on assays from ChEMBL23 as well as the model pretrained on assays from BindingDB24. We investigated the use of the few-shot setting, fine-tuning the models on differing numbers of NPs within the dataset. The shot settings ranged between 8 and 128 fine-tuning compounds (Fig. 1). We then compared the performance of the ActFound fine-tuned models with other conventional meta-learning models: MAML and ProtoNet as well as transfer learning variants of ActFound and MAML13,25.
Growth-inhibitory assays were used to fine-tune ActFound and create bacteria-specific models.
Fine-tuning ActFound on an NPs dataset
To investigate ActFound’s ability to generalize to new domains not explored in the original publication, we fine-tuned the model on an antibacterial NPs dataset. This dataset was curated by Porras et al. in an extensive literature review spanning from 2012 to 2019 and contains the growth-inhibitory activity of NPs against a range of bacteria. A t-distributed stochastic neighbour embedding (t-SNE) comparing the compounds within the NPs dataset and the ChEMBL training data shows overlap, indicating the two datasets likely contain similar compounds (Fig. 2e). To determine the extent of the overlap, we computed the Tanimoto similarities between compounds in the two datasets and found 248 identical compounds (Supplementary Fig. 1). We also analysed how molecular properties, specifically molecular weight, calculated LogP, total polar surface area, hydrogen bond donors, hydrogen bond acceptors and negative log of bioactivity, compare between NPs and the ChEMBL and BindingDB datasets and found that although the distribution differs for some properties, the NP distribution always overlaps the distributions for the other datasets (Supplementary Table 1 and Supplementary Fig. 2). We acknowledge that overlapping compounds between the fine-tuning and training datasets could have caused data leakage that inflated the performance of the model pretrained on ChEMBL assays. Given that this dataset was the result of manual literature curation, it is unlikely that the exact assays in this dataset were deposited into ChEMBL. We investigated the overlap between the bioactivities in the NPs dataset and the ChEMBL database and identified 324 instances where identical molecules were tested in similar assays across the two datasets. When fine-tuning, we found that removing these bioactivities from the dataset had no significant impact on the performance of ActFound (Supplementary Fig. 3). Due to this, we chose to include the overlapping bioactivities in the fine-tuning dataset. In addition, considering that the NPs dataset contained only growth-inhibitory assays, there should be no identical assays in the BindingDB training dataset.
a,b, Bar plots comparing ActFound and ActFound Transfer’s performance on the NPs dataset in terms of (a) r2 and (b) RMSE. Plots show the mean ± standard error of the mean (s.e.m.) performance values across the n = 14 assays used to fine-tune the ChEMBL pretrained models and the BindingDB pretrained models. Before plotting, the performance values for the assays were averaged across each shot setting used to fine-tune the models. c,d, Bar plots comparing the models’ performances in terms of r2 at the (c) 8- (n = 14), 16- (n = 14), 32- (n = 10), 64- (n = 4) and 128- (n = 1) shot settings and when (d) 20% (n = 14), 40% (n = 14), 60% (n = 14) and 80% (n = 14) of the assays were used for fine-tuning. Plots show the mean ± s.e.m. performance values across the n number of assays used to fine-tune the pretrained models. Statistical significance between the performance of ActFound and ActFound Transfer was assessed using a one-sided Wilcoxon test. e, t-SNE plot comparing the molecules in the NPs dataset and a random selection of 50% of the ChEMBL training dataset.
When fine-tuning, we considered each bacterial strain to be its own assay and assessed the performance of ActFound across a range of shot settings. This included using 8–128 fine-tuning compounds as well as using 20–80% of the compounds within each assay for fine-tuning. Averaging the r2 value across all shot settings for ActFound and ActFound Transfer, a transfer learning variant of ActFound, showed that overall ActFound Transfer had a higher r2 value than ActFound on the NPs dataset (Fig. 2a). However, ActFound was found to have the lowest root mean square error (RMSE) value (Fig. 2b). When looking at the performance for each shot setting, ActFound and ActFound Transfer performed the best in the 16-shot setting, with performance dropping as the shot setting increased (Fig. 2c and Supplementary Fig. 4). This is in contrast with the original publication, where the performance of ActFound increased with the number of compounds used for fine-tuning. Because providing more compounds for fine-tuning should inherently improve performance, we investigated using a percentage of the assays for fine-tuning instead of supplying a specific shot setting. In this case, ActFound performed as expected, with the r2 values increasing as the percentage of compounds used for fine-tuning increased (Fig. 2d). We attribute this behaviour to the fact that only four assays had enough compounds to be used for fine-tuning in the 64- and 128-shot settings. When looking at the performance on each assay, the four largest assays (Bacillus subtilis, Escherichia coli, Staphylococcus aureus and S. aureus (MRSA)) yielded the worst model performance (Fig. 3a). This caused the average performance of ActFound across the assays to decrease as the shot setting increased. The performance of ActFound on the four largest assays across the 8- to 128-shot settings showed that the r2 value did increase as the shot setting increased, following the expected trend (Supplementary Fig. 5).
a, Bar plot comparing ActFound and ActFound Transfer’s performance on each assay in the NPs dataset in terms of r2. Plot shows the mean ± s.e.m. performance value across the n number of shot settings used to fine-tune the ChEMBL pretrained models and the BindingDB pretrained models. The n value for each assay is as follows: E. aerogenes, P. stuartii, S. mutans, S. pyogenes, n = 6; B. cereus, E. faecalis, K. pneumoniae, M. tuberculosis, P. aeruginosa, S. epidermidis, n = 7; B. subtilis, E. coli, S. aureus (MRSA), n = 8; S. aureus, n = 9. b, Scatter plot comparing the first step loss and r2 value for every fine-tuning iteration for each assay for the ChEMBL pretrained ActFound and the BindingDB pretrained ActFound. ρP is the Pearson correlation between r2 and the first step loss.
ActFound was found to have varying degrees of performance across the 14 assays, with average r2 values ranging from 0.01 to 0.13. For reference, in the cross-domain setting, the original publication found two kinase inhibitor datasets to have average r2 values between 0.15 and 0.25, so the model performs only slightly worse on some of the NP antibacterial datasets and much worse on others. We also examined fine-tuning the model on a scaffold split, which was defined as a ‘realistic split’ by Feng et al.10. In a scaffold split, the molecules within each assay were split so that the compounds within the fine-tuning set were dissimilar to those in the testing set. We found there to be no significant difference in r2 values for ActFound or ActFound Transfer when a scaffold split was performed instead of a random split (Supplementary Fig. 6).
ActFound utilizes pairwise learning and works under the assumption that similar compounds will have similar bioactivities. This is typically advantageous, as many assay datasets are assembled to investigate structure–activity relationships (SARs), and thus the compounds will be similar to one another. However, one disadvantage to this approach, and what we believe is causing such a range in performance for the NPs dataset, is that if the assay does not contain similar compounds, the pairwise learning function will cause large errors. In the original publication, Feng et al.10 removed what they called ‘orphan compounds’ from the assays. These orphan compounds are those that have a Tanimoto similarity less than 0.2 to the other compounds in the assay. When we performed the same procedure, none of the assays had enough compounds left for fine-tuning. This is likely due to how we defined our ‘assays’. Although Porras et al. provided references in the NPs dataset, there were not enough bioactivities per bacterium in each reference to fine-tune the model. Therefore, we combined compounds from multiple references to treat each bacterial strain as its own assay, which likely led to many dissimilar compounds in each assay. Because removing the orphan compounds left only one assay available for fine-tuning, we decided not to remove them. Additionally, further analysis showed the NPs dataset having a large number of scaffolds that were not present in the training sets, as well as having more scaffold diversity than the training sets (Supplementary Tables 2 and 3). We acknowledge that this likely played a part in how well ActFound was able to perform on the NPs dataset. However, it also reveals an important limitation of the ActFound method. NP datasets will often lack closely related pairs of compounds, as NPs are highly diverse26, and congeners of a primary product may be difficult to discover and isolate without the use of specialized techniques27. This limitation may also pose a problem for high-throughput screening assay datasets that use compound libraries assembled for compound diversity over preliminary SAR, as is sometimes the case28.
The original paper found a correlation between a small loss value for the first optimization and a large r2 value. This was identified as a way to determine how well ActFound will perform on the fine-tuning assay because assays with small loss values will likely result in high r2 values. However, we did not find this correlation to hold on the NPs dataset. Although most of the assays had larger loss values, even the assays with small loss values had small r2 values (Fig. 3b).
Evaluation against other state-of-the-art models
In addition to ActFound and ActFound Transfer, we also used the NPs dataset to fine-tune three other conventional models: MAML, ProtoNet and TransferQSAR. MAML and ProtoNet are meta-learning models, whereas TransferQSAR is a transfer learning variant of MAML. However, none of the models incorporate pairwise learning to learn the relative bioactivities as ActFound does. ActFound Transfer outperformed all other models with both the ChEMBL and BindingDB versions, but ActFound performed the worst (Fig. 4a,b), a trend that holds when identical references or similar assays are moved (Supplementary Figs. 7 and 8). Even though we found ActFound to perform worse on the NPs dataset compared to the results in the original publication, ActFound and ActFound Transfer have a higher r2 value than the other three models when the fewest compounds were used for fine-tuning (Fig. 4c,d and Supplemental Fig. 9). This is indicative of ActFound’s ability to quickly adapt to new assays with only a small number of fine-tuning compounds. Additionally, at this setting, the median number of compounds used for fine-tuning was nine, which is a smaller shot setting than was studied in the original paper. At each shot setting, ActFound Transfer outperformed all other models. The performance of ActFound was more varied, with it having a higher r2 value than the non-ActFound variants at the 20% and 60% shot settings but having a lower r2 value than most of the models at the 40% shot setting and all of the models at the 80% shot settings. Although we identified disadvantages to using pairwise learning with our dataset, these results also indicate the effectiveness of utilizing pairwise learning to learn the relative bioactivity values.
a,b, Bar plots comparing each model’s performance on the NPs dataset in terms of (a) r2 and (b) RMSE. Plots show the mean ± s.e.m. performance values across n = 14 assays used to fine-tune the ChEMBL pretrained models and the BindingDB pretrained models. Before plotting, the performance values for the assays were averaged across each shot setting used to fine-tune the models. c,d, Bar plots comparing the models’ performances when 20%, 40%, 60% and 80% of the assays were used for fine-tuning in terms of (c) r2 and (d) RMSE. Plots show the mean ± s.e.m. performance values across the n = 14 assays used to fine-tune the pretrained models. Statistical significance between the performance of ActFound, ActFound Transfer and the best comparison approach was assessed using a one-sided Wilcoxon test.
Discussion and conclusion
To investigate ActFound’s reusability, we fine-tuned the model on an antibacterial NPs dataset, studying its performance on a range of shot settings. With the availability of Feng et al.’s10 Google Colab, we found ActFound to be easy to use and fine-tune on our dataset. In contrast to the results of the original publication, ActFound Transfer was found to perform better than the meta-learning variant of ActFound. We also found both variants of ActFound to have diminished performance compared to the cross-domain setting in the original paper. This was likely due to the incompatibility of ActFound with the NPs dataset, as the assays in this dataset contained dissimilar compounds. This meant the pairwise learning function of ActFound was not able to be fully advantageous for our dataset. Another possible explanation for the relatively low accuracy is that NPs have different chemical properties than synthetic compounds, which likely make up most of the training data. However, given that the t-SNE analysis and Tanimoto similarity distribution show that the chemical space of the NP dataset overlaps with the training set, we believe that the lack of suitable pairs of compounds for pairwise learning contributes more to lowering the accuracy. Despite the poorer performance on our chosen dataset, we found both variants of ActFound to perform better than other state-of-the-art models at the lower shot settings. Therefore, we believe ActFound to be a very useful framework for those who do not have enough labelled data to train a task-specific DL model—especially those whose datasets consist of structure–activity relationship studies, as these datasets will contain the bioactivities of similar compounds, which will increase the capabilities of the pairwise learning function. However, it is important to note that the more challenging problem of accurate activity prediction for compounds in unassayed areas of chemical space remains unsolved by ActFound and other DL methods.
Methods
Dataset preparation
The NPs dataset used in this Rreusability Report was obtained from ref. 15. To prepare the dataset for fine-tuning, we followed a pipeline similar to that used by Feng et al.10 when evaluating ActFound on two kinase inhibitor datasets, KIBA29 and Davis30. In this cross-domain setting, they considered each kinase to be its own assay. Similarly, we considered each bacterial strain as a separate assay. The NPs dataset contains 1,439 growth-inhibitory values of 472 unique compounds against 115 bacterial strains. We considered resistant strains and subspecies as separate assays from the original strain. Assays with fewer than 20 compounds were removed. If an assay contained duplicate compounds, the growth-inhibitory values were averaged across the duplicated compounds. Additionally, we considered only compounds with minimum inhibitory concentration (MIC) values and ug ml−1 units. This left us with 14 assays with an average of 64 compounds per assay.
Model fine-tuning
All of the foundation models (ActFound, ActFound Transfer, MAML, ProtoNet and TransferQSAR) trained by Feng et al. were obtained from their figshare at https://figshare.com/articles/dataset/ActFound_data/24452680 (ref. 31). We fine-tuned the models using the public code of ActFound from its GitHub repository at https://github.com/BFeng14/ActFound.git (ref. 32). During the fine-tuning process, the hyperparameters and architecture for each model were the same as used by Feng et al.10. The input for the models were 2,048-dimensional Morgan fingerprints, which were computed using RDKit33 and the negative log of the MIC values p(MIC) = −log10(MIC).
Each model was fine-tuned using the 8-, 16-, 32-, 64- and 128-shot settings. Additionally, each model was fine-tuned on different proportions of the data. During this fine-tuning stage, we used 20%, 40%, 60% and 80% of the assay data for fine-tuning. Each assay was randomly split 40 times into the fine-tuning and testing sets, and the models were fine-tuned with each random split. The results of the models are an average across each iteration.
t-SNE
To study the similarities between the compounds in the ChEMBL training set and the NPs dataset, we performed a t-SNE analysis using scikit-learn’s t-SNE34. A t-SNE reduces the dimensionality of the Morgan fingerprints from 2,048 to 2. We used the default values for each parameter except the distance metric, which we set to ‘jaccard’. Because the Jaccard distance, or the Tanimoto distance, is equal to 1 − Tanimoto similarity, the distance between points is directly related to the similarity between compounds. The ChEMBL training set is large (1.4 million datapoints), and running a t-SNE on the entire dataset would be computationally expensive. Therefore, to speed up the computation, we opted to randomly select 50% of the training set to perform the t-SNE.
Data availability
The original antibacterial NPs dataset used in this report is available in ref. 15. Our processed dataset is available via figshare at https://doi.org/10.6084/m9.figshare.30334318.v2 (ref. 35).
Code availability
The original ActFound code is available via GitHub at https://github.com/BFeng14/ActFound.git (ref. 32). All of the foundation model checkpoints are available via figshare at https://figshare.com/articles/dataset/ActFound_data/24452680 (ref. 31). Our modified code, along with the split speeds we used to fine-tune the NPs dataset on the models, is available via GitHub at https://github.com/caitlinbutt04/Actfound_reusability_report.git (ref. 36).
References
Jackson, C. M., Esnouf, M. P., Winzor, D. J. & Duewer, D. L. Defining and measuring biological activity: applying the principles of metrology. Accredit. Qual. Assur. 12, 283–294 (2007).
Dougall, I. G. & Unitt, J. in The Practice of Medicinal Chemistry 4th edn (eds Wermuth, C. G. et al.) Ch. 2 (Academic, 2015).
Wang, S., Guo, Y., Wang, Y., Sun, H. & Huang, J. SMILES-BERT: large scale unsupervised pre-training for molecular property prediction. In Proc. 10th ACM International Conference on Bioinformatics, Computational Biology and Health Informatics 429–436 (ACM, 2019).
Zhang, Z., Liu, Q., Wang, H., Lu, C. & Lee, C.-K. Motif-based graph self-supervised learning for molecular property prediction. In Proc. 35th International Conference on Neural Information Processing Systems (eds Ranzato, M. et al.) 15870–15882 (Curran, 2021).
Fang, X. et al. Geometry-enhanced molecular representation learning for property prediction. Nat. Mach. Intell. 4, 127–134 (2022).
Rong, Y. et al. Self-supervised graph transformer on large-scale molecular data. In 34th Conference on Neural Information Processing Systems (NeurIPS 2020) https://proceedings.neurips.cc/paper_files/paper/2020/file/94aef38441efa3380a3bed3faf1f9d5d-Paper.pdf (2020).
Yang, K. et al. Analyzing learned molecular representations for property prediction. J. Chem. Inf. Model. 59, 3370–3388 (2019).
Sun, C., Shrivastava, A., Singh, S. & Gupta, A. Revisiting unreasonable effectiveness of data in deep learning era. In Proc. IEEE International Conference on Computer Vision 843–852 (IEEE, 2017).
Zhou, C. et al. A comprehensive survey on pretrained foundation models: a history from BERT to ChatGPT. Int. J. Mach. Learn. Cyber 16, 9851–9915 (2025).
Feng, B. et al. A bioactivity foundation model using pairwise meta-learning. Nat. Mach. Intell. 6, 962–974 (2024).
Hospedales, T., Antoniou, A., Micaelli, P. & Storkey, A. Meta-learning in neural networks: a survey. IEEE Trans. Pattern Anal. Mach. Intell. 44, 5149–5169 (2022).
Bommasani, R. et al. On the opportunities and risks of foundation models. Preprint at https://arxiv.org/abs/2108.07258 (2022).
Finn, C., Abbeel, P. & Levine, S. Model-agnostic meta-learning for fast adaptation of deep networks. In Proc. 34th International Conference on Machine Learning (eds Precup, D. & Teh, Y.) 70, 1126–1135 (JMLR, 2017).
Koch, G., Zemel, R. & Salakhutdinov, R. Siamese neural networks for one-shot image recognition. In Proc. 32nd International Conference on Machine Learning (eds Bach, F. & Blei, D.) https://www.cs.cmu.edu/~rsalakhu/papers/oneshot1.pdf (JMLR, 2015).
Porras, G. et al. Ethnobotany and the role of plant natural products in antibiotic drug discovery. Chem. Rev. 121, 3495–3560 (2021).
Rossiter, S. E., Fletcher, M. H. & Wuest, W. M. Natural products as platforms to overcome antibiotic resistance. Chem. Rev. 117, 12415–12474 (2017).
Hutchings, M. I., Truman, A. W. & Wilkinson, B. Antibiotics: past, present and future. Curr. Opin. Microbiol. 51, 72–80 (2019).
Wong, F., de la Fuente-Nunez, C. & Collins, J. J. Leveraging artificial intelligence in the fight against infectious diseases. Science 381, 164–170 (2023).
Stokes, J. M. et al. A deep learning approach to antibiotic discovery. Cell 180, 688–702 (2020).
Wong, F. et al. Discovery of a structural class of antibiotics with explainable deep learning. Nature 626, 177–185 (2024).
Ma, Y. et al. Identification of antimicrobial peptides from the human gut microbiome using deep learning. Nat. Biotechnol. 40, 921–931 (2022).
Sorokina, M. & Steinbeck, C. Review on natural products databases: where to find data in 2020. J. Cheminform. 12, 20 (2020).
Gaulton, A. et al. ChEMBL: a large-scale bioactivity database for drug discovery. Nucleic Acids Res. 40, D1100–D1107 (2012).
Liu, T., Lin, Y., Wen, X., Jorissen, R. N. & Gilson, M. K. BindingDB: a web-accessible database of experimentally determined protein–ligand binding affinities. Nucleic Acids Res. 35, D198–D201 (2007).
Snell, J., Swersky, K. & Zemel, R. S. Prototypical networks for few-shot learning. In Proc. 31st International Conference on Neural Information Processing Systems (eds Guyon, I. et al.) 4080-4090 (Curran, 2017).
Hong, J. Role of natural product diversity in chemical biology. Curr. Opin. Chem. Biol. 15, 350–354 (2011).
Herath, K. B. et al. Rapid, selective, and sensitive method for semitargeted discovery of congeneric natural products by liquid chromatography tandem mass spectrometry. J. Nat. Prod. 84, 814–823 (2021).
Dandapani, S., Rosse, G., Southall, N., Salvino, J. M. & Thomas, C. J. Selecting, acquiring, and using small molecule libraries for high-throughput screening. Curr. Protoc. Chem. Biol. 4, 177–191 (2012).
Tang, J. et al. Making sense of large-scale kinase inhibitor bioactivity data sets: a comparative and integrative analysis. J. Chem. Inf. Model. 54, 735–743 (2014).
Davis, M. I. et al. Comprehensive analysis of kinase inhibitor selectivity. Nat. Biotechnol. 29, 1046–1051 (2011).
Feng, B. The data and checkpoint for ActFound. figshare https://doi.org/10.6084/m9.figshare.24452680 (2023).
Feng, B. Bfeng14/actfound: Actfound v0.0. Zenodo https://doi.org/10.5281/zenodo.11800155 (2024).
Landrum, G. et al. RDKit. Zenodo https://doi.org/10.5281/zenodo.18428170 (2025).
Pedregosa, F. et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Butt, C. ActFound NPs reusability report. Dataset. figshare https://doi.org/10.6084/m9.figshare.30334318 (2025).
Butt, C. caitlinbutt04/Actfound_reusability_report: ActFound NPs reusability report (v0.0). Zenodo https://doi.org/10.5281/zenodo.17873072 (2025).
Acknowledgements
Research reported in this publication was supported by the National Institute of General Medical Sciences under grant no. R35GM146987 (A.S.W.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. We would like to acknowledge Vanderbilt’s ACCRE computing cluster for computational resources.
Author information
Authors and Affiliations
Contributions
C.M.B. performed all evaluations of the model, wrote and modified the code for this work, analysed the data and wrote the paper and Supplementary Information. A.S.W. conceived and supervised the study and edited the paper and Supplementary Information.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Machine Intelligence thanks Tunca Dogan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods, Tables 1–4 and Figs. 1–9.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Butt, C.M., Walker, A.S. Reusability Report: Evaluating the performance of a meta-learning foundation model on predicting the antibacterial activity of natural products. Nat Mach Intell 8, 270–275 (2026). https://doi.org/10.1038/s42256-026-01187-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42256-026-01187-y