Abstract
Both regulatory B (Breg) and myeloid cells in tumors and lymph nodes drive immune suppression in pancreatic cancer. Current strategies to counter immune suppression emphasize myeloid cells but overlook Breg cells. We discovered that STING agonist expanded Breg cells depended on PI3Kγ but not PI3Kδ in pancreatic cancer, whereas activating myeloid cells were independent of PI3Kγ. Inhibition of PI3Kγ, but not PI3Kδ, decreased STING-induced IRF3 phosphorylation and Breg cell expansion in pancreatic cancer, while sustaining STING-induced IRF3 phosphorylation to activate myeloid cells. We developed a dual targeting compound and its albumin nanoformulation Nano-273, which stimulated STING to activate myeloid cells and inhibited PI3Kγ to decrease STING-induced Breg cell expansion. Nano-273 delivered the drug to tumors and lymph nodes to overcome myeloid cell- and Breg cell-mediated immune suppression in pancreatic cancer. Nano-273, combined with anti-programmed cell death protein 1, achieved durable efficacy in transgenic KPC mice with pancreatic cancer, offering potential for pancreatic cancer treatment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All raw and processed RNA-seq and single-cell RNA-seq data are available via the GEO repository under accession numbers GSE275780 and GSE275785, respectively. Data are available within the article and Supplementary Information, and/or from the corresponding authors upon request. Source data are provided with this paper.
Code availability
The code for RNA-seq analysis is available at https://github.com/Anycl/Dual_targeting_STING_PI3K/tree/main.
References
Corn, B. W. & Feldman, D. B. Cancer statistics, 2025: a hinge moment for optimism to morph into hope?. CA Cancer J. Clin. 75, 7–9 (2025).
Conroy, T. et al. FOLFIRINOX versus Gemcitabine for metastatic pancreatic cancer. N. Engl. J. Med. 364, 1817–1825 (2011).
Von Hoff, D. D. et al. Gemcitabine plus nab-paclitaxel is an active regimen in patients with advanced pancreatic cancer: a phase I/II trial. J. Clin. Oncol. 29, 4548–4554 (2011).
Bear, A. S., Vonderheide, R. H. & O’Hara, M. H. Challenges and opportunities for pancreatic cancer immunotherapy. Cancer Cell 38, 788–802 (2020).
Macherla, S. et al. Emerging role of immune checkpoint blockade in pancreatic cancer. Int. J. Mol. Sci. 19, 3505 (2018).
Leinwand, J. & Miller, G. Regulation and modulation of antitumor immunity in pancreatic cancer. Nat. Immunol. 21, 1152–1159 (2020).
Ho, W. J., Jaffee, E. M. & Zheng, L. The tumour microenvironment in pancreatic cancer—clinical challenges and opportunities. Nat. Rev. Clin. Oncol. 17, 527–540 (2020).
Mirlekar, B. et al. Balance between immunoregulatory B cells and plasma cells drives pancreatic tumor immunity. Cell Rep. Med. 3, 100744 (2022).
Senturk, Z. N., Akdag, I., Deniz, B. & Sayi-Yazgan, A. Pancreatic cancer: emerging field of regulatory B-cell-targeted immunotherapies. Front. Immunol. 14, 1152551 (2023).
Tong, D.-N. et al. Characterization of B cell-mediated PD-1/PD-L1 interaction in pancreatic cancer patients. Clin. Exp. Pharmacol. Physiol. 47, 1342–1349 (2020).
Li, S. et al. STING-induced regulatory B cells compromise NK function in cancer immunity. Nature 610, 373–380 (2022).
Balachandran, V. P., Beatty, G. L. & Dougan, S. K. Broadening the impact of immunotherapy to pancreatic cancer: challenges and opportunities. Gastroenterology 156, 2056–2072 (2019).
Chamma, H., Vila, I. K., Taffoni, C., Turtoi, A. & Laguette, N. Activation of STING in the pancreatic tumor microenvironment: a novel therapeutic opportunity. Cancer Lett. 538, 215694 (2022).
Le Naour, J., Zitvogel, L., Galluzzi, L., Vacchelli, E. & Kroemer, G. Trial watch: STING agonists in cancer therapy. Oncoimmunology 9, 1777624 (2020).
Wang, Y., Geller, A. E. & Yan, J. Unexpected Breg-NK crosstalk in STING agonist therapy. Cell. Mol. Immunol. 19, 1330–1332 (2022).
Thorpe, L. M., Yuzugullu, H. & Zhao, J. J. PI3K in cancer: divergent roles of isoforms, modes of activation and therapeutic targeting. Nat. Rev. Cancer 15, 7–24 (2015).
Kaneda, M. M. et al. PI3Kγ is a molecular switch that controls immune suppression. Nature 539, 437–442 (2016).
De Henau, O. et al. Overcoming resistance to checkpoint blockade therapy by targeting PI3Kγ in myeloid cells. Nature 539, 443–447 (2016).
Okkenhaug, K. & Vanhaesebroeck, B. PI3K in lymphocyte development, differentiation and activation. Nat. Rev. Immunol. 3, 317–330 (2003).
Zhao, H. Y. et al. An oral tricyclic STING agonist suppresses tumor growth through remodeling of the immune microenvironment. Cell Chem. Biol. 32, 280–290 e214 (2025).
Song, Y. et al. Albumin nanoparticle containing a PI3Kγ inhibitor and paclitaxel in combination with α-PD1 induces tumor remission of breast cancer in mice. Sci. Transl. Med. 14, eabl3649 (2022).
Vanpouille-Box, C., Hoffmann, J. A. & Galluzzi, L. Pharmacological modulation of nucleic acid sensors—therapeutic potential and persisting obstacles. Nat. Rev. Drug Discov. 18, 845–867 (2019).
Jneid, B. et al. Selective STING stimulation in dendritic cells primes antitumor T cell responses. Sci. Immunol. 8, eabn6612 (2023).
Sun, X., Zhou, X., Lei, Y. L. & Moon, J. J. Unlocking the promise of systemic STING agonist for cancer immunotherapy. J. Control. Release 357, 417–421 (2023).
Brinkmann, V. et al. Fingolimod (FTY720): discovery and development of an oral drug to treat multiple sclerosis. Nat. Rev. Drug Discov. 9, 883–897 (2010).
Lu, Y. et al. Complement signals determine opposite effects of B cells in chemotherapy-induced immunity. Cell 180, 1081–1097 (2020).
Pan, B.-S. et al. An orally available non-nucleotide STING agonist with antitumor activity. Science 369, eaba6098 (2020).
Li, W. et al. cGAS-STING-mediated DNA sensing maintains CD8+ T cell stemness and promotes antitumor T cell therapy. Sci. Transl. Med. 12, eaay9013 (2020).
Zhu, Y. et al. STING: a master regulator in the cancer-immunity cycle. Mol. Cancer 18, 152 (2019).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Ishikawa, H., Ma, Z. & Barber, G. N. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature 461, 788–792 (2009).
Barber, G. N. STING: infection, inflammation and cancer. Nat. Rev. Immunol. 15, 760–770 (2015).
Hopfner, K.-P. & Hornung, V. Molecular mechanisms and cellular functions of cGAS–STING signalling. Nat. Rev. Mol. Cell Biol. 21, 501–521 (2020).
DeNardo, D. G., Andreu, P. & Coussens, L. M. Interactions between lymphocytes and myeloid cells regulate pro-versus anti-tumor immunity. Cancer Metastasis Rev. 29, 309–316 (2010).
Xiong, J., Wang, H. & Wang, Q. Suppressive myeloid cells shape the tumor immune microenvironment. Adv. Biol. 5, 1900311 (2021).
Mantovani, A., Allavena, P., Marchesi, F. & Garlanda, C. Macrophages as tools and targets in cancer therapy. Nat. Rev. Drug Discov. 21, 799–820 (2022).
Werner, M., Hobeika, E. & Jumaa, H. Role of PI3K in the generation and survival of B cells. Immunol. Rev. 237, 55–71 (2010).
Olayinka-Adefemi, F., Hou, S. & Marshall, A. J. Dual inhibition of phosphoinositide 3-kinases delta and gamma reduces chronic B cell activation and autoantibody production in a mouse model of lupus. Front. Immunol. 14, 1115244 (2023).
Lucas, C. L., Chandra, A., Nejentsev, S., Condliffe, A. M. & Okkenhaug, K. PI3Kδ and primary immunodeficiencies. Nat. Rev. Immunol. 16, 702–714 (2016).
Beer-Hammer, S. et al. The catalytic PI3K isoforms p110γ and p110δ contribute to B cell development and maintenance, transformation, and proliferation. J. Leukoc. Biol. 87, 1083–1095 (2010).
Lanahan, S. M., Wymann, M. P. & Lucas, C. L. The role of PI3Kgamma in the immune system: new insights and translational implications. Nat. Rev. Immunol. 22, 687–700 (2022).
Lanahan, S. M., Wymann, M. P. & Lucas, C. L. The role of PI3Kγ in the immune system: new insights and translational implications. Nat. Rev. Immunol. 22, 687–700 (2022).
Lanahan, S. M. et al. PI3Kγ in B cells promotes antibody responses and generation of antibody-secreting cells. Nat. Immunol. 25, 1422–1431 (2024).
Takeda, A. J. et al. Human PI3Kγ deficiency and its microbiota-dependent mouse model reveal immunodeficiency and tissue immunopathology. Nat. Commun. 10, 4364 (2019).
Lee, J. W., Komar, C. A., Bengsch, F., Graham, K. & Beatty, G. L. Genetically engineered mouse models of pancreatic cancer: the KPC model (LSL-Kras(G12D/+);LSL-Trp53(R172H/+);Pdx-1-Cre), its variants, and their application in immuno-oncology drug discovery. Curr. Protoc. Pharmacol. 73, 14.39.11–14.39.20 (2016).
Paster, E. V., Villines, K. A. & Hickman, D. L. Endpoints for mouse abdominal tumor models: refinement of current criteria. Comp. Med. 59, 234–241 (2009).
Workman, P. et al. Guidelines for the welfare and use of animals in cancer research. Br. J. Cancer 102, 1555–1577 (2010).
Lutz, M. B. et al. An advanced culture method for generating large quantities of highly pure dendritic cells from mouse bone marrow. J. Immunol. Methods 223, 77–92 (1999).
Li, J. et al. Tumor cell-intrinsic factors underlie heterogeneity of immune cell infiltration and response to immunotherapy. Immunity 49, 178–193 (2018).
Grigorova, I. L., Panteleev, M. & Cyster, J. G. Lymph node cortical sinus organization and relationship to lymphocyte egress dynamics and antigen exposure. Proc. Natl Acad. Sci. USA 107, 20447–20452 (2010).
Renier, N. et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014).
Below, C. R. et al. A microenvironment-inspired synthetic three-dimensional model for pancreatic ductal adenocarcinoma organoids. Nat. Mater. 21, 110–119 (2022).
Boj, S. F. et al. Organoid models of human and mouse ductal pancreatic cancer. Cell 160, 324–338 (2015).
Huch, M. et al. Unlimited in vitro expansion of adult bi-potent pancreas progenitors through the Lgr5/R-spondin axis. EMBO J. 32, 2708–2721 (2013).
Li, C. et al. Antigen-clustered nanovaccine achieves long-term tumor remission by promoting B/CD 4 T cell crosstalk. ACS Nano 18, 9584–9604 (2024).
Zhang, Y. et al. Small circular RNAs as vaccines for cancer immunotherapy. Nat. Biomed. Eng. 9, 249–267 (2025).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Acknowledgements
We thank Y. Bu (Department of Pharmaceutical Sciences, College of Pharmacy, the University of Michigan) for instruction in the western blotting procedure. We thank the shared resource facilities at the University of Michigan (Pharmacokinetics, Flow cytometry, Microscopy, Advanced Genomics and In Vivo Animal cores) for support. This study is partially supported by The Michigan Translational Research and Commercialization (MTRAC) grant and internal funding from the University of Michigan (N035800, D.S.).
Author information
Authors and Affiliations
Contributions
C.L. designed and conducted the experiments, analyzed the data and wrote the paper. S.M., H.Z. and M.D. designed and synthesized the dual functional compound. F.K., B.W., M.H. and M.W. assisted with the pharmacokinetics and toxicity studies. F.K. and M.W. contributed to the characterization of the albumin nanoformulation. F.K., Z.Q., A.R.X., C.L.L. and M.P.W. assisted with the animal studies. D.N., V.S., L.H., N.P. and M.N. assisted with the processing of human samples and related experiments. Z.Y. developed the PDAC organoid models and conducted the distribution experiments. H.W. and Z.L. measured the binding assays for PI3K isoforms and cellular STING potency. Z.Q., A.R.X., C.L.L. and M.P.W. provided PI3Kγ knockout mice and help with studies using cells in these mice. L.F. guided study design and edited the paper. D.S. and W.G. designed the experiments, analyzed the data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The University of Michigan has submitted a patent application (PCT/US2024/053672, Dual functioning immune modulating compounds, formulations, and uses thereof), in which some authors (D.S., C.L., W.G., B.W., M.W., M.H., H.Z., S.M., M.D., Z.L., H.W. and F.K.) are listed as inventors. The other authors declare no competing interests.
Peer review
Peer review information
Nature Cancer thanks David Barbie, Kris Wood and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Immunosuppressive microenvironment and antitumor efficacy of STING agonists in PDAC mice.
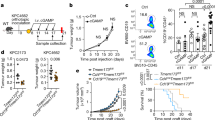
(a, b) Quantification of flow cytometry analysis of M2 macrophages in KPC mice pancreatic tumor and C57BL/6 mice pancreas tissues (a), KPC mice and C57BL/6 mice lymph node (b). (c) Antitumor efficacy of STING agonists in C57BL/6 mice inoculated with pancreatic tumor (KPC 6422 cell line). Average and individual tumor volumes change after treatment with PBS (control), anti-PD1 antibody (100 µg, i.p.), diABZi (i.t., 20 µg/mouse) with or without anti-PD1 antibody (100 µg, i.p.) or diABZi (i.v., 1.5 mg/kg) with or without anti-PD1 antibody (100 µg, i.p.). (d) Quantification of flow cytometry analysis of IL-35+ (Ebi3+) and IL-10+ Breg cells in lymph node and tumor from mice inoculated with KPC 6422 cell after treatment with PBS (control), anti-PD1 antibody (100 µg, i.p.), diABZi (i.t., 20 µg/mouse or i.v., 1.5 mg/kg) with or without anti-PD1 antibody (100 µg, i.p.) or diABZi (i.v., 1.5 mg/kg) with or without anti-PD1 antibody (100 µg, i.p.). (e) Antitumor efficacy of STING agonist (MSA-2, i.t., or i.v., 34.0 μmol/kg) with or without anti-PD1 antibody (100 µg, i.p.) in C57BL/6 mice subcutaneously inoculated with pancreatic tumor (KPC 6422 cell line). (f, g) Quantification of flow cytometry analysis of IL-35+ (Ebi3+) and IL-10+ T cells in lymph node (f) and tumor (g) from syngeneic mouse model subcutaneously inoculated with KPC 6422 cell after treatment with PBS (control), anti-PD1 antibody (100 µg, i.p.), or MSA-2 (i.t., or i.v., 34.0 μmol/kg) with or without anti-PD1 antibody (100 µg, i.p.). (h) Antitumor efficacy of STING agonists in μMt mice or C57BL/6 mice subcutaneously inoculated with pancreatic tumor (KPC 6422 cell line). (i) Antitumor efficacy of STING agonists in C57BL/6 mice with or without CD4+ T cells subcutaneously inoculated with pancreatic tumor (KPC 6422 cell line). Individual tumor volumes change after treatment with PBS (control), anti-PD1 antibody (100 µg, i.p.), or MSA-2 (i.t., or i.v., 34.0 μmol/kg) with or without anti-PD1 antibody (100 µg, i.p.). The data represents two independent experiments (a-i) with mean ± SD, and n = 3 biologically independent samples (a, b), n = 4 biologically independent samples (d, f, g), n = 5 mice (e, h, i) and n = 10 mice (c). Statistical comparisons are based on unpaired two-tailed Student’s t-tests (a, b), one-way analysis of variance (d, f, g) or two-way analysis of variance (c). P values are indicated.
Extended Data Fig. 2 Expression of PI3K isoform subunits and time courses of phosphorylation of IRF3 with or without treatments of STING agonist and PI3Kγ inhibitor.
(a, b) Gene expected count of Pik3cg, Pik3r5 and Pik3r6 in splenic B cells (a) and in BMDC (b) from KPC mice after treatment with or without treatment of MSA-2 (5 µg/mL). (c) Western blot analysis of p110γ, p101, p87, p110δ, phosphorylation of AKT, AKT, and GAPDH expression in splenic B cells and BMDC from KPC transgenic mice treated with or without MSA-2 (5 µg/mL). (d, e) Gene expected count of Pik3cd, Pik3r1 and Pik3r2 in splenic B cells (d) and BMDC (e) from KPC mice treated with or without MSA-2 (5 µg/mL). (f, g) Western blot analysis of p110γ, p110δ, and STING expression in lymph node tissues and pancreatic tumor whole tissues (including tumor immune infiltrated cells) from KPC transgenic mice and lymph node and pancreas from C57BL/6 mice (f) and B cell, BMDCs, and BMDMs (g) from KPC transgenic mice and C57BL/6 mice. (h) Western blot of phosphorylation of IRF3 in B cells, CD4+ T cells, CD8+ T cells, bone marrow derived dendritic cells and bone marrow derived macrophages after treatments with MSA-2 (5 µg/mL) or MSA-2 (5 µg/mL) &IPI-549 (5 μM) at different time points (0.5 – 12 hrs). The data represents two independent experiments (a-h) with mean ± SD, and n = 3 biologically independent samples (a,b,d,e). Statistical comparisons are based on two-way analysis of variance (a,b,d,e). P values are indicated.
Extended Data Fig. 3 Phosphorylation of IRF3 after treatment with PI3K isoform inhibitors or knock down of PI3K isoform subunits under STING agonist treatment.
(a) Phosphorylation of IRF3 at B cells from KPC transgenic mice after treatments with MSA-2 (5 µM) or combination of MSA-2 (5 µM) with IPI-549 (20 nM – 500 nM). (b) Phosphorylation of IRF3 at Human CD4+ T cells, Human CD8+ T cells and Human NK cells after treatments with MSA-2 (5 µg/mL) or combination of MSA-2 (10 µg/mL) with IPI-549 (10 μM) or SH-273 (10 μM). (c, d) Phosphorylation of IRF3 at B cells from KPC transgenic mice after treatments with MSA-2 (5 µM) or combination of MSA-2 (5 µM) with SH-344 (100 nM – 500 nM, c), or combination of MSA-2 (5 µM) with TGR-1202 (50 nM – 500 nM, d). (e, f) Western blot of B cells from KPC transgenic mice after knocking down of p110γ (e), p101 (e) or p110δ (f) by shRNA. The data represents two independent experiments (a-f).
Extended Data Fig. 4 STING agonist expands Breg cells, while selective inhibition of PI3Kγ reduces STING-induced Breg cell expansion.
(a) Quantification flow cytometry analysis of IL-35+ (Ebi3+) and IL-10+ Breg cells treated with MSA-2 (5 µg/mL), or IPI-549 (5 µM), or MSA-2 (5 µg/mL) and IPI-549 (5 µM) in splenic B cells from KPC transgenic mice. (b) Quantification flow cytometry analysis of IL-35+ (Ebi3+) and IL-10+ Breg cells treated with MSA-2 (5 µg/mL), or IPI-549 (5 µM), or MSA-2 (5 µg/mL) and IPI-549 (5 µM) in splenic B cells from STING knock out (C57BL/6J-Sting1gt/J) mice. (c) Quantification flow cytometry analysis of IL-35+ (Ebi3+) and IL-10+ Breg cells after pretreated with anti-IgM & anti-CD40 (5 µg/mL) and then treated with MSA-2 (5 µg/ml) or MSA-2 (5 µg/mL) and IPI-549 (2 µM), or MSA-2 (5 µg/mL) and SH-273 (5 µM) in splenic B cells from STING knock out (C57BL/6J-Sting1gt/J) mice. (d-f) Quantification flow cytometry analysis of IL-35+ (Ebi3+) and IL-10+ Breg cells treated with MSA-2 (5 µM) or combination of MSA-2 (5 µM) with different concentrations of IPI-549 (20 nM – 500 nM, d), or SH-344 (100 nM – 500 nM, e), or TGR-1202 (50 nM – 500 nM, f). (g, h) Quantification flow cytometry analysis of IL-35+ (Ebi3+) and IL-10+ BMDCs (g) and BMDMs (h) (from KPC mice) after treatments with or without MSA-2 (5 µg/mL), with or without IPI-549 (5 µM). (i, j) Quantification flow cytometry analysis of IL35+ (Ebi3+) and IL10+ THP-1 wild type (i) and THP-1 hSTINGR232 (j) cells after treatments with or without MSA-2 (5 µg/mL), with or without IPI-549 (5 µM). (k) Quantification flow cytometry analysis of CD80 intensity of BMDC after treatments with or without MSA-2 (2.95 µg/mL), with or without IPI-549 (10 µM). (l) Quantification flow cytometry analysis of M1 macrophage polarization of RAW264.7 by CD80 intensity after treatments with or without MSA-2 (2.95 µg/mL), with or without IPI-549 (10 µM). The data represents two independent experiments (a-l) with mean ± SD, and n = 3 biologically independent samples (a-l). Statistical comparisons are based on one-way analysis of variance. P values are indicated.
Extended Data Fig. 5 Dual targeting compound SH-273 and its albumin nanoformulation inhibit the PI3Kγ isoform and reduce STING-induced Breg cell expansion.
(a) Quantification flow cytometry analysis of DID+ (DiIC18(5) solid (1,1’-Dioctadecyl-3,3,3’,3’-Tetramethylindodicarbocyanine, 4-Chlorobenzenesulfonate Salt)) cells from in vitro uptake of Nano-273&DID or solvent based formulations of SH-273&DID at splenocytes. (b) Cellular EC50 of STING activation after MSA-2 or ZSA-52 (released active moiety of SH-273 to activate STING) treatments in THP-1-BlueTM ISG cells. (c) Phosphorylation of AKT (473) at BMDCs and BMDMs after treatments with different concentrations of TGR-1202 (1 µM, 150 nM, from left to right lane), SH-273 (2 µM, 400 nM, from left to right lane) or IPI-549 (1 µM, 150 nM, from left to right lane) in the presence of SDF-1α for 1.5 hours. (d) Phosphorylation of AKT (473) at BMDC or BMDM after treatments with different concentrations of IPI-549 (1 µM, 150 nM, or 20 nM, from left to right lane) or SH-273 (2 µM, 400 nM, or 40 nM, from left to right lane) in the presence of SDF-1α for 1.5 hours. (e) Phosphorylation of AKT (473) and AKT (308) at SK-OV-3 or 786-O cells after treatments with Alpelisib (1 µM, SK-OV-3 cells, PI3Kα), TGX-211 (1 µM, 786-O cells, PI3Kβ), or SH-273 (2 µM) in the presence of SDF-1α for 1.5 hours. (f) Phosphorylation of IRF3 in BMDCs from KPC transgenic mice after treatments with MSA-2 (2.95 µg/mL), IPI-549 (10 µM) or SH-273 (10 µM), respectively. (g) Quantification flow cytometry analysis of IL-35+ (Ebi3+) and IL-10+ Breg cells after pretreated with anti-IgM&anti-CD40 (5 µg/mL) and then treated with MSA-2 (5 µg/mL), IPI-549 (2 µM), or MSA-2 (5 µg/mL) and IPI-549 (2 µM), or SH-273 (2 µM) in B cells from STING knock out (C57BL/6J-Sting1gt/J) mice. (h) Phosphorylation of IRF3 at B cells from KPC transgenic mice after treatments with MSA-2 (5 µM) or combination of MSA-2 (5 µM) with Nano-273 (50 nM – 500 nM). (i) Quantification flow cytometry analysis of IL-35+ (Ebi3+) and IL-10+ Breg cells treated with MSA-2 (5 µM) or combination of MSA-2 (5 µM) with different concentrations of Nano-273 (50 nM – 500 nM). The data represents two independent experiments (a-i) with mean ± SD, and n = 3 biologically independent samples (a, b, g, i). Statistical comparisons are based on one-way analysis of variance (g, i) or two-way analysis of variance (a). P values are indicated.
Extended Data Fig. 6 Albumin nanoparticles of the dual targeting compound (Nano-273) extend survival in KPC transgenic and syngeneic mouse models.
(a) Antitumor efficacy in transgenic KPC (LSL-KrasG12D, LSL-Trp53R172H, Pdx1cre/+) PDAC mice with different treatments. Survival rate of KPC mice after treatment with Paclitaxel in albumin nanoformulation (i.v., 11.7 μmol/kg) and IPI-549 (i.p., 28.4 μmol/kg) plus anti-PD1 antibody (100 µg, i.p.), MSA-2 (oral dose, 204 μmol/kg) and IPI-549 (oral dose, 28.4 μmol/kg) plus anti-PD1 antibody (100 µg, i.p.) and SH-273 (i.v., 17.6 μmol/kg) plus anti-PD1 antibody (100 µg, i.p.). The experiments are conducted at the same time with Fig. 7a with different groups (Nano-P + IPI-549+anti-PD-1, MSA-2 + IPI-549+anti-PD-1 and SH-273+anti-PD-1) for comparison. (b) Antitumor efficacy in C57BL/6 mice inoculated with KPC 6422 tumor after different treatments. Individual tumor volume after treatment with PBS (control), MSA-2 (i.v., 34.0 μmol/kg) with or without anti-PD1 antibody (i.p., 100 µg), Nano-273 (i.v., 17.6 μmol/kg) with or without anti-PD1 antibody (i.p., 100 µg). (c, d) The serum concentration of IL35, IL10 and IL27 (c) and IFNβ and TNF (d) of syngeneic C57BL/6 mice subcutaneous inoculated with KPC 6422 tumor after treatment with PBS (control), MSA-2 (i.v., 34.0 μmol/kg) with or without anti-PD1 antibody (100 µg, i.p.), Nano-273 (i.v., 17.6 μmol/kg) with or without anti-PD1 antibody (100 µg, i.p.). (e) Antitumor efficacy in syngeneic pancreatic cancer (KPC 6422 cell line) model with different treatments. Average and individual tumor volume after treatment with PBS (control), MSA-2 (i.v., 34.0 μmol/kg) plus anti-PD1 antibody (100 µg, i.p.), SH-273 (i.v., 17.6 μmol/kg) plus anti-PD1 antibody (100 µg, i.p.) or Nano-273 (i.v., 17.6 μmol/kg) plus anti-PD1 antibody (100 µg, i.p.). (f) Quantification flow cytometry analysis of IL-35+ (Ebi3+) and IL-10+ Breg cells at tumor and lymph node in mice from antitumor efficacy study at (e). The data represents two (c-f) or three (a, b) independent experiments with mean ± SD, and n = 3 biologically independent samples (f), n = 4 biologically independent samples (c, d), n = 5 mice (e) and n = 10 mice (a, b). Statistical comparisons are based on log-rank (Mantel–Cox) test (a) one-way analysis of variance (f), or two-way analysis of variance (c, d, e). P values are indicated.
Extended Data Fig. 7 Systemic administration of albumin nanoparticles of the dual targeting compound (Nano-273) induce systemic antitumor immune responses that drive anticancer efficacy.
(a, b) Antitumor efficacy for pancreatic cancer (KPC 6422 cell line) with Nano-273 treatment in different administration routes. Individual tumor volume (a, local tumor; b, distal tumor) after treatment with or without Fingolimod (FTY720, 5 mg/kg, oral, daily), or with PBS (control), Nano-273 (17.6 μmol/kg i.t., s.c. or i.v. injection, respectively) plus anti-PD1 antibody (100 µg, i.p.). (c, d) Antitumor efficacy for pancreatic cancer (KPC 6422 cell line) with Nano-273 treatment in subcutaneously injection. Average tumor volume (c, local tumor; d, distal tumor) after treatment with Fingolimod (FTY720, 5 mg/kg, oral, daily) with PBS (control) or Nano-273 (17.6 μmol/kg s.c. injection) plus anti-PD1 antibody (100 µg, i.p.). The experiments are conducted at the same time with Fig. 7i with different groups (Control + FTY-720 and Nano-273 s.c.+anti-PD-1 + FTY-720) for comparison. (e-g) Quantification and representative flow cytometry analysis of tumor immune infiltration (e) and M1 macrophage ratios in uninvolved lymph node (f) and distal tumor (g) after treatments from efficacy study from Fig. 7i and (c, d). The data represents two independent experiments (a-g) with mean ± SD, and n = 3 biologically independent samples (e-g) and n = 5 mice (a-d). Statistical comparisons are based on one-way analysis of variance (e-g) or two-way analysis of variance (c, d). P values are indicated.
Extended Data Fig. 8 Complete blood count, clinical chemistry and body weight of mice after treatment with Nano-273 or SH-273.
Liver enzymes (a), kidney function (a), and complete blood counts (CBCs) (b) in CD1 mice after treatments with SH-273 and Nano-273 every 3 days for 5 doses (17.6 μmol/kg) through intravenous injection. Alanine aminotransferase, ALT; aspartate aminotransferase, AST; Alkaline phosphatase, ALKP; total bilirubin, TBIL; creatine, CREA; blood urine nitrogen, BUN; albumin, ALB. White blood cells, WBC; neutrophils, NEU; lymphocytes, LYM; monocytes, MONO; eosinophils, EOS; basophils, BAS; red blood cells, RBC; hemoglobin, HGB; hematocrit, HCT; mean corpuscular volume, MCV; mean corpuscular hemoglobin, MCH; mean corpuscular hemoglobin concentration, MCHC; red cell distribution width, RDW; platelets, PLT; mean platelet volume, MPV. (c) Body weight measurements of C57BL/6 mice after a single intravenous dose of Nano-273 (200 mg/kg). (d-f) Body weight measurements of Es1−/− mice (B6.Cg-Ces1ctm1.1Loc/J) after a single intravenous dose of Nano-273 (200 mg/kg, d), Nano-273 (150 mg/kg, e) or Nano-273 (100 mg/kg, f). (g) Body weight measurements of Es1−/− mice (B6.Cg-Ces1ctm1.1Loc/J) after multiple intravenous doses of Nano-273. In the multiple-dose study, Nano-273 was administered intravenously at escalating doses of 15, 50, and 100 mg/kg every three days for 28 days. The data represents two independent experiments (a-g) with mean ± SD, and n = 3 biologically independent samples (a, b) and n = 3 mice (c-g).
Extended Data Fig. 9 Complete blood count and clinical chemistry in Es1−/− mice after Nano-273 treatment.
(a) Complete blood cell count (CBC) after multiple doses of Nano-273 by intravenous administration (15-100 mg/kg in Es1−/− mice (B6.Cg-Ces1ctm1.1Loc/J). In the multiple-dose study, Nano-273 was administered intravenously at escalating doses of 15, 50, and 100 mg/kg every three days for 28 days. White blood cells, WBC; monocytes, MONO; lymphocytes, LYM; eosinophils, EOS; basophils, BAS; neutrophils, NEU; red blood cells, RBC; mean corpuscular volume, MCV; hematocrit, HCT; red cell distribution width, RDW; hemoglobin, HGB; mean corpuscular hemoglobin, MCH; mean corpuscular hemoglobin concentration, MCHC; platelets, PLT; mean platelet volume, MPV. (b) Blood chemistry after multiple intravenous doses of Nano-273 (15-100 mg/kg) in Es1−/− mice (B6.Cg-Ces1ctm1.1Loc/J). In the multiple-dose study, Nano-273 was administered intravenously at escalating doses of 15, 50, and 100 mg/kg every three days for 28 days. Alanine aminotransferase, ALT; total bilirubin, TBIL; Alkaline phosphatase, ALKP; creatine, CREA; blood urine nitrogen, BUN; total protein, TPRO; albumin, ALB; glucose, GLUC; calcium, CA. The data represents two independent experiments (a,b) with mean ± SD, and n = 3 biologically independent samples.
Extended Data Fig. 10 STING-induced Breg cells compromise NK and CD8+ T cells functions in LL/2 lung cancer model but not in MC-38 colon cancer model.
(a) Quantification flow cytometry analysis of IL-35+ (Ebi3+) or IL-10+ Breg cells in the lymph node from KPC transgenic mice after treatments with PBS (control), anti-PD1 antibody (100 µg, i.p.), MSA-2 (i.v., 34.0 μmol/kg) plus anti-PD1 antibody (100 µg, i.p.), or Nano-273 (i.v., 17.6 μmol/kg) plus anti-PD1 antibody (100 µg, i.p.). (b) Individual plots for antitumor efficacy in C57BL/6 mice inoculated with LL/2 tumor cells with treatments of MSA-2 (i.v., 34.0 μmol/kg) plus anti-PD1 antibody (100 µg, i.p.), Nano-273 (i.v., 17.6 μmol/kg), or Nano-273 (i.v., 17.6 μmol/kg) plus anti-PD1 antibody (100 µg, i.p.). (c) Quantification flow cytometry analysis of IL10+ Bregs at lymph node and tumor from (b). (d) Antitumor efficacy in C57BL/6 mice inoculated with MC-38 tumor cells with treatments of MSA-2 (i.v., 34.0 μmol/kg) plus anti-PD1 antibody (100 µg, i.p.), Nano-273 (i.v., 17.6 μmol/kg), or Nano-273 (i.v., 17.6 μmol/kg) plus anti-PD1 antibody (100 µg, i.p.). (e, f) Quantification flow cytometry analysis of IL-35+ (Ebi3+) and IL-10+ Breg cells at lymph node (e) and tumor (f) from (d). (g) Antitumor efficacy in immunodeficient (NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ, NSG) mice inoculated with KPC 6422 tumor with or without Nano-273 (i.v., 17.6 μmol/kg) with anti-PD1 antibody (100 µg, i.p.). (h) Antitumor efficacy in C57BL/6 mice with or without NK cell depletion inoculated with KPC 6422 tumor with Nano-273 (i.v., 17.6 μmol/kg) plus anti-PD1 antibody (i.p., 100 µg) treatment. (i) Quantification flow cytometry analysis of GzmB+ NK cells in lymph node from KPC transgenic mice after treatments with anti-PD1 antibody (i.p., 100 µg), MSA-2 (i.v., 34.0 μmol/kg) plus anti-PD1 antibody (i.p., 100 µg) or Nano-273 (i.v., 17.6 μmol/kg) plus anti-PD1 antibody (i.p., 100 µg). (j) Antitumor efficacy in C57BL/6 mice with or without CD8+ T cell depletion inoculated with KPC 6422 tumor with Nano-273 (i.v., 17.6 μmol/kg) plus anti-PD1 antibody (100 µg, i.p.) treatment. (k) Antitumor efficacy in C57BL/6 mice with or without CD4+ T cell depletion inoculated with KPC 6422 tumor with Nano-273 (i.v., 17.6 μmol/kg) plus anti-PD1 antibody (100 µg, i.p.) treatment. The data represents two (a, c-f, i) or three (b, g, h, j, k) independent experiments with mean ± SD, n = 3 biologically independent samples (a, c, e, f, i), n = 5 mice (d), n = 6 mice (k), n = 8 mice (b) and n = 10 mice (g, h, j). Statistical comparisons are based on one-way analysis of variance (a, c, e, f, i) and two-way analysis of variance (d). P values are indicated.
Supplementary information
Supplementary Information (download PDF )
Supplementary Table 1 and Figs. 1–11.
Supplementary Video 1 (download MP4 )
3D imaging of the distribution of albumin nanoformulation co-encapsulated with SH-273 and fluorescent dye (DiD) in tumor tissue from KPC transgenic mice. Albumin nanoformulation (magenta), macrophages (F4/80, red), B cells (B220, blue) and tumor cells (YFP, green).
Supplementary Video 2 (download MP4 )
Section view of the distribution of albumin nanoformulation co-encapsulated with SH-273 and fluorescent dye (DiD) in tumor tissue from KPC transgenic mice. Albumin nanoformulation (magenta), macrophages (F4/80, red), B cell (B220, blue) and tumor cells (YFP, green).
Supplementary Video 3 (download MP4 )
3D imaging of the distribution of solvent-based SH-273 mixed with fluorescent dye (DiD) in tumor tissue from KPC transgenic mice. Solvent-based formulation (magenta), macrophages (F4/80, red), B cell (B220, blue) and tumor cells (YFP, green).
Supplementary Video 4 (download MP4 )
3D view (merged) of the distribution of albumin nanoformulation co-encapsulated with SH-273 and fluorescent dye (DiD) in lymph node from KPC transgenic mice. Albumin nanoformulation (magenta), macrophages (F4/80, red), B cell (B220, blue) and T cells (CD3, green).
Supplementary Video 5 (download MP4 )
3D view of T cells (CD3, green) in lymph node from KPC transgenic mice administrated with albumin nanoformulation co-encapsulated with SH-273 and fluorescent dye (DiD).
Supplementary Video 6 (download MP4 )
3D view of Macrophages (F4/80, red) in lymph node from KPC transgenic mice administrated with albumin nanoformulation co-encapsulated with SH-273 and fluorescent dye (DiD).
Supplementary Video 7 (download MP4 )
3D view of B cells (B220, blue) in lymph node from KPC transgenic mice administrated with albumin nanoformulation co-encapsulated with SH-273 and fluorescent dye (DiD).
Supplementary Video 8 (download MP4 )
3D view of albumin nanoformulation (DiD, magenta) distribution in lymph node from KPC transgenic mice administrated with albumin nanoformulation co-encapsulated with SH-273 and fluorescent dye (DiD).
Supplementary Video 9 (download MP4 )
Section view (merged) of the distribution of albumin nanoformulation co-encapsulated with SH-273 and fluorescent dye (DiD) in lymph node from KPC transgenic mice. Albumin nanoformulation (magenta), macrophages (F4/80, red), B cell (B220, blue) and T cells (CD3, green).
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download PDF )
Unprocessed western blots.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Fig. 8 (download XLSX )
Statistical source data
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, C., Ke, F., Zhao, H. et al. Dual targeting of PI3Kγ and STING overcomes regulatory B cell- and myeloid cell-driven immune suppression in pancreatic cancer. Nat Cancer (2026). https://doi.org/10.1038/s43018-026-01158-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43018-026-01158-7