Abstract
Denitrification, a major source of gaseous nitrogen emissions from agricultural soils, is influenced by management. Practices promoting belowground diversity are suggested to support sustainable agriculture, but how they modulate nitrogen losses via denitrification remains inconclusive. Here we sampled 106 cereal fields spanning a 3000 km North-South gradient across Europe and compiled 56 associated climatic, soil, microbial and management variables. We show that increased denitrification potential was associated with higher proportion of time with crop cover over the last ten years and was best predicted by microbial biomass and microbial functional guilds involved in nitrogen cycling, in particular denitrification. We also demonstrate that several diversification practices affect the variation in denitrification potential predictors, suggesting a trade-off between agricultural diversification and nitrogen losses via denitrification. However, increased crop diversity in rotations improved yield-scaled denitrification, highlighting the potential of this practice to minimize nitrogen losses while contributing to sustainable food production.
Similar content being viewed by others
Introduction
Globally, there is a surplus of reactive nitrogen (N) in the environment due to the widespread use of fertilizers in agriculture1. Nitrogen-use efficiency in cropping systems is low, with only half of the N inputs recovered in the harvested crop2, and the loss of N from fertilized soils represents a major threat to the integrity of both terrestrial and aquatic ecosystems3. Denitrification, an anaerobic microbial respiratory pathway, is the main source of the potent greenhouse gas nitrous oxide (N2O) and dinitrogen (N2) from fertilized soils, returning 48 ± 10 Tg N to the atmosphere annually at the global scale4. These agricultural soils also account for nearly half of the anthropogenic N2O production worldwide5, making reactive N surplus a key driver of climate change, which could create undefined feedbacks on terrestrial N cycling6,7. Identifying and promoting management practices that minimize gaseous N losses is therefore critical for sustainable agricultural intensification8 and climate change mitigation9.
Cropping systems with more diverse crop rotations and longer periods with crop cover, as well as management practices such as reduced tillage and the application of organic amendments, have been advocated to increase belowground biodiversity and promote soil multifunctionality10,11,12. Diverse cropping can also provide higher grain yields13 and support a variety of ecosystem services14. Although the abundance, composition, and activity of denitrifying microbial communities are influenced by agricultural management practices15,16, most studies cover a limited range of environmental conditions with only a few variables measured and the relative importance of management versus other environmental drivers on denitrification remains unclear17. Characterising the underlying factors explaining the effects of agricultural management practices on denitrification is thus key to develop effective strategies aimed at minimizing gaseous N losses through denitrification in cropping systems.
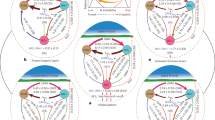
Building on our previous study conducted along a 3000 km North-South gradient across Europe, where we showed that crop cover promotes soil multifunctionality10, we investigate here whether a trade-off exists between N losses via denitrification and various diversification practices, including long-term phylogenetic and functional crop diversification, the proportion of time with crop cover, tillage frequency and the application of organic and mineral fertilizers. For this purpose, we used a dataset comprising 56 climatic, soil, microbial and management variables associated to 106 soils along the European gradient (Fig. 1). We hypothesized that management practices enhancing belowground diversity would lead to a greater abundance of denitrifiers, as these facultative anaerobes make up a substantial portion of the overall microbial community18. This, in turn, would increase the potential for denitrification activity, measured here as the potential production of N2 and N2O under standardized conditions using a common-garden approach (hereafter, denitrification). We then expressed denitrification in relation to the harvested yield (yield-scaled denitrification, hereafter, ydenitrification) to determine whether any management practice can off-set denitrification activity with higher yields. Finally, we identified the best predictors of denitrification and ydenitrification from our dataset and assessed how much of the variation in denitrification predictors could be explained by management. We demonstrate that enhanced denitrification activity is associated with a greater proportion of time that the fields are cultivated with crops over a decade-long period, and the best predictors of denitrification were various microbial factors. However, the yield-scaled denitrification capacity decreased with increasing phylogenetic diversity of crops, suggesting that the implementation of diverse crop rotations can mitigate yield-scaled emissions of gaseous nitrogen.
a Map of Europe showing sampling sites. b The nitrogen cycle with genes encoding the respective enzymes that catalyze the reactions and that were used in this study. In the nitrification process, amoA genes in both ammonia-oxidizing archaea (AOA) and bacteria (AOB) were quantified. For the abundance of prokaryotic and fungal communities, we quantified the 16S rRNA gene and the ITS2 region, respectively. Products and a selection of intermediates are shown by their chemical names. c Additional variables included were associated with management, climate, soil properties and microbial diversity, biomass, and activity. See Supplementary Table 1 for units.
Results
Effects of management practices on denitrification and ydenitrification
The generalized additive models including the six management practices as explanatory variables (Figs. 2 and 3) explained 47.3% and 37.6% of the deviance for denitrification and ydenitrification, respectively. We found a non-linear relationship between the proportion of time with crop cover during the last 10 years and denitrification (Fig. 2b). Denitrification was highest in soils at 100% time under cover, meaning crops were grown all year-round (including cover crops), and lowest at 50–60% time under crop cover. By contrast, lower ydenitrification, or higher efficiency, was associated with higher phylogenetic diversity of the crops in the rotations (Fig. 3a). Reduced tillage was also associated with lower ydenitrification (Fig. 3f), but none of the other management practices, including N fertilization rates in the sampling year, affected denitrification or ydenitrification (Figs. 2 and 3).
Responses to a phylogenetic diversity of the crops, b proportion of time with crop cover and c presence of ley in the rotation, all during the last 10 years, and d amount of applied nitrogen (N) fertilizer the year of sampling (including both organic and inorganic N), e amount of organic fertilizers during the last 4 years, and f number of tillage occurrences the year of sampling. The blue line represents the estimated effect of each variable conditional upon the other terms in the model. The grey dots denote the partial residuals, and the shaded area indicates 95% confidence intervals. P-values are indicated on the corresponding plots. See Supplementary Table 2 for summary of the model statistics.
Responses to a phylogenetic diversity of the crops, b proportion of time with crop cover and c presence of ley in the rotation, all during the last 10 years, and d amount of applied nitrogen (N) fertilizer the year of sampling (including both organic and inorganic N), e amount of organic fertilizers during the last 4 years, and f number of tillage occurrences the year of sampling. The blue line represents the estimated effect of each variable conditional upon the other terms in the model, the grey dots denote the partial residuals, and the shaded area indicates 95% confidence intervals. P-values are indicated on the corresponding plots. See Supplementary Table 3 for summary of the model statistics.
Predictors of denitrification activity
We then examined the relative importance of climatic, microbial, soil and management variables to predict denitrification and ydenitrification in our dataset (Fig. 1b, c). Ten predictors of denitrification were retained in the final random forest model, all displaying non-linear relationships with denitrification (Fig. 4). The majority of predictors were microbial, with several related to the overall size of the microbial communities, as reflected by the strong positive effect of microbial biomass, the abundance of several functional guilds involved in N cycling, more specifically denitrifiers (nirS, nosZ clade I) and bacteria performing nitrate ammonification (nrfA), as well as the composition of the microbial communities (Fig. 4b–d). Soil predictors, and in particular total N in the range 0–3000 mg N kg−1 soil, were also positively associated with denitrification activity, but to a lower extent than microbial predictors (Fig. 4a). Regarding ydenitrification, the final random forest model comprised a subset of the best predictors of denitrification. Higher ydenitrification, or higher gaseous N losses in relation to the harvested yield, was mainly associated with increasing total N, particularly in the range 1500–3000 mg N kg−1 soil, microbial biomass and abundance of nirS-denitrifiers (Fig. 5). Neither climatic variables (mean annual temperature and mean annual precipitation over the past 10 years) nor management practices were selected as important predictors of denitrification or ydenitrification by the random forest models.
Predictor variables selected by VSURF among a soil properties, b microbial biomass, c microbial community composition, and d abundances of nitrogen-cycling genes were used to generate accumulated local effects plots. They show the differences in prediction of denitrification rates compared to the mean prediction along the range of each predictor (x-axis), while accounting for potential correlations amongst predictor values. The effect is centered so that the mean effect is zero. The random forest model was built with 500 trees, 5 features considered at each split and a tree depth set to 6 (variance explained: 75 %, root mean square error: 69.9).
Predictor variables selected by VSURF among a soil properties, b microbial biomass, and c abundances of nitrogen-cycling genes were used to generate accumulated local effects plots. They show the differences in prediction of ydenitrification (y-axis) compared to the mean prediction along the range of each predictor (x-axis), while accounting for potential correlations amongst predictor values. The effect is centered so that the mean effect is zero. The random forest model was built with 500 trees, 2 features considered at each split and a tree depth set to 9 (variance explained: 58%, root mean square error: 4.3).
Combining management and denitrification predictors
Finally, we assessed whether management practices could indirectly influence denitrification and ydenitrification by affecting their best predictors. The redundancy analysis models, including the best predictors as response variables (Figs. 4 and 5) and the six management practices as explanatory variables, showed that management explained 37.5% and 32.3% of the variance in the predictors of denitrification and ydenitrification, respectively (p = 0.001). Forward selection of the management variables in the full model identified three management practices as drivers, namely proportion of time with crop cover, application of organic fertilizers and tillage frequency, but their individual contribution was relatively small (3.0–5.5%, Fig. 6). These practices correlated positively with all denitrification predictors, except the bacterial and cercozoan non-metric multidimensional scaling (NMDS) axes (Supplementary Fig. 1). The inclusion of climate (mean annual temperature and mean annual precipitation over the past 10 years) and soil texture (clay and silt content) variables improved the models, which explained 56.9% and 53.9% of the variance in the predictors of denitrification and ydenitrification, respectively (p = 0.001; Supplementary Fig. 2). Although clay content and mean annual precipitation over the past 10 years were the two variables explaining most of the variation (9.3–12.3%) in both models, the collective contribution of the different management, climate and soil texture variables was comparable (9–13%).
Variance explained by the full redundancy analysis model (all management practices, MP) and individual management practices after accounting for the remaining management practices in partial models for both a denitrification and b yield-scaled potential denitrification activity (ydenitrification = denitrification/raw crop yield) predictors. Crop cov., proportion of time with crop cover (past 10 years); Org. fert., organic fertilizer application (past 4 years); Till. occ., tillage occurrences (sampling year); Leys in rot., presence of ley (past 10 years); PD crops, phylogenetic diversity of the crops (past 10 years); Appl. N, applied N fertilizer (sampling year).
Discussion
Increased time with crop cover has positive effects on soil biodiversity and functioning, including the accrual and accumulation of organic matter12,19 and the capacity to provide multiple ecosystems functions and services10,12. However, our study shows that crop cover also increases the risk for gaseous N losses through denitrification, which aligns with studies showing higher N2O emissions from fields with cover crops20,21. This is possibly explained by the increased carbon input through higher primary production supporting the growth of heterotrophic denitrifiers combined with crop residue degradation generating anoxic hotspots that favor denitrification as well as N2O emissions22,23. Higher crop diversity, measured here as the phylogenetic diversity of the crops in the rotations over 10 years prior to sampling, and fewer tillage occasions were associated with lower denitrification in relation to the harvested yield (ydenitrification). Both practices have been shown to promote multiple ecosystem services, including soil fertility and carbon sequestration but differ in their influence on yield12. In contrast to reduced tillage, crop diversification can enhance cereal yields (as shown earlier with this dataset)10, particularly at low N fertilization rates13,24, and can also buffer against yield loss from global warming and changes in precipitation25,26. Moreover, diverse crop rotations can increase N-use efficiency in arable soils27,28, as supported here for ydenitrification, thereby decreasing the amount of reactive N available for denitrification. We found that the presence of leys in the rotations had no effect on denitrification, suggesting that the capacity of such rotations to decrease gaseous N losses from soils depends on a combination of the local environment and management, which contrasts with site and region-specific reports indicating that cropping systems including perennials emit less N2O compared to systems with annual crops29,30,31. Overall, we conclude that agronomic management practices aiming to increase belowground diversity and support multiple ecosystems functions come with both trade-offs and co-benefits.
Denitrification was best predicted by the cumulative effect of N inputs, as reflected by total soil N, rather than the amount of N fertilizer applied in the sampling year. The growing surplus of reactive N in arable soils causes profound shifts in both the composition and functioning of soil microbial communities32,33,34, and this aligns with the acceleration of yearly increases in global N2O emission rates35. Surprisingly, there was no direct effect of addition of organic fertilizers during the past 4 years36. Nevertheless, organic fertilizer amendment affected the best predictors of both denitrification and ydenitrification, indicating an indirect effect of organic amendments on denitrification, possibly by increasing soil C and thereby supporting microbial biomass accumulation. Apart from soil N content, several microbial variables, including microbial biomass, the composition of microbial communities and abundance of specific N-cycling guilds, predicted denitrification capacity across European croplands. The abundance of nirS and nosZ clade I, two genes commonly found in combination in the genome of complete denitrifiers37, best explained effects of the functional guilds. We also found that denitrification was predicted by the abundance of nitrate ammonifiers (i.e., nrfA), even though nitrate ammonification leads to N retention rather than loss38. This is likely because denitrifiers and nitrate ammonifiers share the same substrates (nitrate and nitrite, and organic compounds), making nrfA abundance an indirect predictor of denitrification. However, the genetic potential for nitrate ammonification was one to two orders of magnitude lower than that of denitrification (average nrfA:nir abundance ratio = 0.04 ± 0.02). This suggests a limited mitigating effect of ammonification on N losses in arable soils across Europe and aligns with recent findings showing that denitrification capacity dominates over nitrate ammonification in terrestrial biomes39. Regarding the bacterial OTUs associated with an increase in the prediction of denitrification activity, the high diversity of physiological traits at the class level40 and the limited congruence between denitrification capacity and organismal phylogeny41 prevented further inferences on specific OTUs for denitrification capacity. The absence of variables related to fungal abundance and diversity in the set of the best denitrification predictors suggests that, relative to bacteria, denitrifying fungi42 contribute less to denitrification in arable soils. This could be due to the sieving and freezing of the soils before performing the assays, which could disrupt fungal hyphae and reduce fungal activity, or, more likely, because fungal denitrifiers make up only a small proportion of the overall fungal community as well as of the total denitrifying community43. At any rate, the existence of a threshold for total N but not for gene abundances of N cycling guilds indicates that microbial factors are an important control of denitrification in arable soils. Future research on how resource availability shapes the interactions and niche partitioning within and between the different N-cycling guilds may provide additional insight into the mechanisms that link community composition and gaseous N losses in arable soils. We also acknowledge that our findings are based on a one-time point sampling and future research should confirm our conclusions.
Conclusions
Our analysis of the relationships between denitrification and climatic, soil, microbial, and management variables associated with 106 agricultural soils across Europe revealed that denitrification activity was directly and indirectly influenced by management practices aiming to promote belowground diversity, particularly the proportion of time with crop cover. However, when denitrification was linked to crop yield by using yield-scaled denitrification rates, increased phylogenetic diversity of the crops in the rotations during the last 10 years was associated with lower gaseous N losses. This supports the widespread adoption of diverse crop rotations as a promising way to contribute to food security while minimizing N loss through denitrification and possibly negative impacts on the environment14. However, although identifying how management modulates the N2O:N2 end-product ratio of denitrification and net N2O emissions was beyond the scope of this work, we believe this merits further investigation in future research efforts. Overall, our study shows that management strategies should be implemented with consideration of trade-offs as well as co-benefits that could affect sustainability and productivity goals.
Methods
Sampling and associated data
The sampling procedure and part of the data used in this study have been described in detail in Garland et al.10. Briefly, soil samples were collected in fields under cereal cultivation across Europe (in Sweden, Germany, Switzerland, France and Spain) around flowering time between May and August 2017, depending on the country and site. In this study, we used a subset of the samples (n = 106) from Garland et al.10 for which information on management practices, crop yield, soil properties and microbial communities were available (Fig. 1b, c, and Supplementary Table 1). Management practices and crop yield were obtained by surveying the farmers and farm managers at each site via a questionnaire. Physical and chemical soil properties were measured using the Swiss standard protocols44. Potential rates for mineralization were determined following Gregorich and Carter45. Mean annual air temperature and precipitation (2007–2016) were obtained using the GPS coordinates of each sampling site and the historical monthly weather data from the WorldClim database (https://worldclim.org), using the R package ‘raster’ v. 3.6–26 (ref. 46) and the R software v. 4.4.1 (ref. 47). A weighed measure of Faith’s phylogenetic diversity in the crop rotations at each site, incorporating both the evolutionary distance between crops (in My) and the period of time each crop was grown over the last 10 years prior to sampling (in month), was calculated with the weighted.faith function (https://rdrr.io/github/NGSwenson/lefse_0.5/), using the GBOTB.extended.tre phylogeny of vascular plants available in the ‘V.PhyloMaker’ v. 0.1.0 package48 (scenario 3) as backbone.
OTU tables for archaea, bacteria, cercozoan, and fungi obtained from Garland et al.10 were rarefied by averaging the OTU counts over 1000 computations using the rrarefy function in ‘vegan’ v. 2.6–4 (ref. 49). Shannon’s diversity and Pielou’s evenness were determined using ‘vegan’ on the rarefied OTU tables with the diversity function. The species abundance distribution patterns of the OTUs were then examined to partition the rarefied table between frequent and rare community members50. This was achieved by calculating an index of dispersion for each OTU, which corresponded to the ratio of the variance to the mean abundance multiplied by the occurrence51. The index was then used to model whether OTUs followed a stochastic (Poisson) distribution. The rare community members were defined as the OTUs falling below the 2.5% confidence limit of the χ2 distribution52 and were discarded. The remaining OTUs (or frequent community members), which distribution was not random, were used to compute NMDS ordinations using the metaMDS function. This approach minimizes the risk of sampling artefacts that can bias the distribution of the rare OTUs and thus increases the likelihood to detect relevant associations with the potential for denitrification.
Potential denitrification activity
Potential denitrification activity was determined on soil samples kept at −20 °C using the acetylene inhibition technique modified from Pell et al.53. Thawed soil samples (10 g fresh weight) were placed in 125 mL Duran flasks and kept at room temperature overnight. The following morning, a slurry was prepared for each sample by adding 20 ml of water. The bottles were tightly capped, and the headspace was exchanged by flushing with N2 to obtain anoxic conditions. After 30 min of pre-incubation at 25 °C on a shaker (175 rpm), acetylene was added equivalent to 10% (v/v) of the headspace to inhibit the reduction of N2O to N2. Then, 1 ml of substrate was added to reach a final concentration of 3 mM KNO3, 1.5 mM succinate, 1 mM glucose, and 3 mM acetate54. Gas samples were taken from the headspace after 30, 75, 120, 150 and 180 min. Nitrous oxide concentrations were determined using a gas chromatograph (Clarus-500, Elite-Q PLOT phase capillary column, Perkin-Elmer, Hägersten, Sweden) equipped with a 63Ni electron-capture detector and the rate of N2O accumulation was determined in each bottle by non-linear regression.
We also adapted the widely used “yield-scaled N2O emissions” metric (e.g., ref. 55) to “yield-scaled denitrification” (ydenitrification), corresponding to the potential activity divided by the crop yield for the corresponding field, to determine whether any management practice can off-set denitrification activity with higher yields.
Quantitative PCR analyses
The abundance of the 16S rRNA gene, fungal ITS and a set of nine genes involved in various steps in the N cycle (i.e., nitrogen fixation, nitrification, denitrification and ammonification; Fig. 1b) were measured by quantitative real-time PCR based on SYBR green detection. The qPCR reactions were carried out in duplicate runs on either the CFX Connect Real-Time System (Bio-Rad, Hercules, CA, USA) or ViiA7 (Life Technologies, Carlsbad, CA, USA) machine. Standard curves were obtained by serial dilutions of linearized plasmids with cloned fragments of the specific genes. The amplifications were validated by melting curve analyzes and gel electrophoresis. Potential inhibition of PCR reactions was checked for all samples by amplifying a known amount of the pGEM-T plasmid (Promega, Madison, WI, USA) with the plasmid-specific T7 and SP6 primers when added to the DNA extracts or non-template controls. No inhibition was detected with the amount of DNA used. Primer sequences and concentration, qPCR conditions and amplification efficiencies can be found in Supplementary Table 4.
Relationships between denitrification and management
Relationships between management practices and denitrification or ydenitrification rates were assessed using generalized additive models. The models were fitted using the ‘mgcv’ package v. 1.9-1 (ref. 56) with a gamma distribution for the response (denitrification or ydenitrification) and the log link function. Fertilizer amount, phylogenic diversity of the crops and proportion of time with crop cover were modeled as thin plate splines (with k = 10) and tillage occurrences, application of organic fertilizer and presence of leys in the rotations as parametric terms. Country was included as a random factor. The gam.check function was used to ensure that enough basis functions (k) were specified for each smooth and that model residuals were normally distributed56. Concurvity between smooth terms, the non-linear equivalent of co-linearity, was assessed using the concurvity function implemented in the ‘mgcv’ package. Concurvity estimates were <0.6 for all pair-wise comparisons. Model coefficients were estimated using restricted marginal likelihood57,58 and the goodness-of-fit of each model was calculated as the percentage of deviance explained. Effects were visualized on the response scale using the ‘visreg’ package v. 2.7.0 (ref. 59).
Identification of the predictors of denitrification activity
Random forest modeling was used to determine the relationships between denitrification and ydenitrification and their best predictors among climatic, microbial, soil and management variables. The best predictors were identified by variable selection using the ‘VSURF’ package v. 1.2.0 (ref. 60) with default parameters and denitrification or ydenitrification as response variable. The algorithm was run 100 times, and only the variables selected in more than 95% of the runs were retained. Then, a grid search was conducted to find the optimal combination of tuning parameters for the models containing the best predictors (n = 10 for denitrification and n = 5 for ydenitrification): the number of variables to randomly sample as candidates at each split, the minimal number of samples within the terminal nodes and the fraction of samples to train the model on (with n = 500 trees; ‘randomForest’ package v. 4.7-1.1 (ref. 61)). The search was run 100 times and the combination of parameters corresponding to the best model fit (i.e., lowest out-of-bag root-mean-square error) was selected. The relationship between each variable and denitrification or ydenitrification was visualized using accumulated local effects plots (grid.size = 25) implemented in the ‘iml’ package v. 0.11.3 ref. 62. These plots show how the prediction changes on average over the range of each individual explanatory variable, while accounting for potential correlations amongst explanatory variables63.
Relationships between denitrification predictors and management, climate and soil texture
Redundancy analysis models were used to explore the relationships between management practices and denitrification and ydenitrification predictors. Two separate models were run for the two sets of predictors, using the ‘vegan’ package, and significance testing was done using permutation tests. Forward selection (ordiR2step function with default settings) and significance testing with permutation tests were applied to identify the management practices that were statistically significant for explaining the variation in the denitrification and ydenitrification predictors. Partial models were implemented to test the amount of variation explained by each individual management practice (constrained variance) while accounting for the variation explained by the other management practices (conditioned variance). The same procedure was used to determine the relative importance of management, climate (mean annual temperature and mean annual precipitation over the past 10 years) and soil texture (silt and clay) on the variation in denitrification and ydenitrification predictors.
Data availability
Data and OTU tables used in this study as well as source data for the figures are available at Zenodo (https://doi.org/10.5281/zenodo.14760398).
Code availability
The R code used in this study is available at Zenodo (https://doi.org/10.5281/zenodo.14760398).
References
Sutton, M. A. et al. Too much of a good thing. Nature 472, 159–161 (2011).
Ludemann, C. I. et al. A global FAOSTAT reference database of cropland nutrient budgets and nutrient use efficiency (1961–2020): nitrogen, phosphorus and potassium. Earth Syst. Sci. Data 16, 525–541 (2024).
Richardson, K. et al. Earth beyond six of nine planetary boundaries. Sci. Adv. 9, eadh2458 (2023).
Scheer, C. et al. Estimating global terrestrial denitrification from measured N2O:(N2O + N2) product ratios. Curr. Opin. Environ. Sustain. 47, 72–80 (2020).
Tian, H. et al. A comprehensive quantification of global nitrous oxide sources and sinks. Nature 586, 248–256 (2020).
Bowles, T. M. et al. Addressing agricultural nitrogen losses in a changing climate. Nat. Sustain. 1, 399–408 (2018).
Sinha, E. et al. Eutrophication will increase during the 21st century as a result of precipitation changes. Science 357, 405–408 (2017).
Springmann, M. et al. Options for keeping the food system within environmental limits. Nature 562, 519–525 (2018).
You, L. et al. Optimized agricultural management reduces global cropland nitrogen losses to air and water. Nat. Food 5, 995–1004 (2024).
Garland, G. et al. Crop cover is more important than rotational diversity for soil multifunctionality and cereal yields in European cropping systems. Nat. Food 2, 28–37 (2021).
Kim, N. et al. Do cover crops benefit soil microbiome? A meta-analysis of current research. Soil Biol. Biochem. 142, 107701 (2020).
Tamburini, G. et al. Agricultural diversification promotes multiple ecosystem services without compromising yield. Sci. Adv. 6, eaba1715 (2020).
Smith, M. E. et al. Increasing crop rotational diversity can enhance cereal yields. Commun. Earth Environ. 4, 89 (2023).
Kremen, C. et al. Ecosystem services in biologically diversified versus conventional farming systems: benefits, externalities, and trade-offs. Ecol. Soc. 17, 40 (2012).
Domeignoz-Horta, L. A. et al. Peaks of in situ N2O emissions are influenced by N2O-producing and reducing microbial communities across arable soils. Glob. Change Biol. 24, 360–370 (2017).
Wang, J. et al. No-till increases soil denitrification via its positive effects on the activity and abundance of the denitrifying community. Soil Biol. Biochem. 142, 107706 (2020).
Pan, B. et al. A global synthesis of soil denitrification: driving factors and mitigation strategies. Agric. Ecosyst. Environ. 327, 107850 (2022).
Philippot, L. et al. Ecology of denitrifying prokaryotes in agricultural soil. Adv. Agron. 96, 249–305 (2007).
Fageria, N. K. et al. Role of cover crops in improving soil and row crop productivity. Commun. Soil Sci. Plant Anal. 36, 2733–2757 (2005).
Duan, Y. F. et al. Catch crop residues stimulate N2O emissions during spring, without affecting the genetic potential for nitrite and N2O reduction. Front. Microbiol. 9, 2629 (2018).
Olesen, J. E. et al. Challenges of accounting nitrous oxide emissions from agricultural crop residues. Glob. Change Biol. 29, 6846–6855 (2023).
Kravchenko, A. N. et al. Hotspots of soil N2O emission enhanced through water absorption by plant residue. Nat. Geosci. 10, 496–500 (2017).
Højberg, O. et al. Denitrification in soil aggregates analyzed with microsensors for nitrous oxide and oxygen. Soil Sci. Soc. Am. J. 58, 1691–1698 (1994).
MacLaren, C. et al. Long-term evidence for ecological intensification as a pathway to sustainable agriculture. Nat. Sustain. 5, 770–779 (2022).
Costa, A. et al. Crop rotational diversity can mitigate climate-induced grain yield losses. Glob. Change Biol. 30, e17298 (2024).
Renard, D. et al. National food production stabilized by crop diversity. Nature 571, 257–260 (2019).
Fageria, N. K. et al. Enhancing nitrogen use efficiency in crop plants. Adv. Agron. 88, 97–185 (2005).
Yi, B. et al. Diversified cropping systems with limited carbon accrual but increased nitrogen supply. Nat. Sustain. 8, 152–161 (2025).
Gelfand, I. et al. Long-term nitrous oxide fluxes in annual and perennial agricultural and unmanaged ecosystems in the upper Midwest USA. Glob. Change Biol. 22, 3594–3607 (2016).
Putz, M. et al. Relative abundance of denitrifying and DNRA bacteria and their activity determine nitrogen retention or loss in agricultural soil. Soil Biol. Biochem. 123, 97–104 (2018).
Thompson, K. A. et al. Soil microbial communities as potential regulators of in situ N2O fluxes in annual and perennial cropping systems. Soil Biol. Biochem. 103, 262–273 (2016).
Dai, Z. et al. Long-term nitrogen fertilization decreases bacterial diversity and favors the growth of Actinobacteria and Proteobacteria in agro-ecosystems across the globe. Glob. Change Biol. 24, 3452–3461 (2018).
Jones, C. M. et al. Reactive nitrogen restructures and weakens microbial controls of soil N2O emissions. Commun. Biol. 5, 273 (2022).
Isobe, K. et al. Phylogenetic conservation of bacterial responses to soil nitrogen addition across continents. Nat. Commun. 10, 2499 (2019).
Thompson, R. L. et al. Acceleration of global N2O emissions seen from two decades of atmospheric inversion. Nat. Clim. Change 9, 993–998 (2019).
Charles, A. et al. Global nitrous oxide emission factors from agricultural soils after addition of organic amendments: a meta-analysis. Agric. Ecosyst. Environ. 236, 88–98 (2017).
Graf, D. R. H. et al. Intergenomic comparisons highlight modularity of the denitrification pathway and underpin the importance of community structure for N2O emissions. PLoS ONE 9, e114118 (2014).
Saghaï, A. et al. Diversity and ecology of NrfA-dependent ammonifying microorganisms. Trends Microbiol. 32, 602–613 (2024).
Saghaï, A. et al. Phyloecology of nitrate ammonifiers and their importance relative to denitrifiers in global terrestrial biomes. Nat. Commun. 14, 8249 (2023).
Morrissey, E. M. et al. Phylogenetic organization of bacterial activity. ISME J. 10, 2336–2340 (2016).
Jones, C. M. et al. Phylogenetic analysis of nitrite, nitric oxide, and nitrous oxide respiratory enzymes reveal a complex evolutionary history for denitrification. Mol. Biol. Evol. 25, 1955–1966 (2008).
Shoun, H. et al. Fungal denitrification and nitric oxide reductase cytochrome P450nor. Philos. Trans. R. Soc. B Biol. Sci. 367, 1186–1194 (2012).
Bösch, Y. et al. Distribution and environmental drivers of fungal denitrifiers in global soils. Microbiol. Spectr. 11, e00061–23 (2023).
FAL et al. Referenzmethoden Der Eidg. Landwirtschaftlichen Forschungsanstalten. 1. Bodenuntersuchung Zur Düngeberatung (Zürich- Reckenholz, 1996).
Gregorich, E. G. et al. Soil Sampling and Methods of Analysis (CRC Press, 2007).
Hijmans, R. J. Raster: Geographic Data Analysis and Modeling https://doi.org/10.32614/CRAN.package.raster (2024).
R Core Team. R: a language and environment for statistical computing. R Foundation for Statistical Computing https://www.R-project.org/ (Vienna, Austria 2024).
Jin, Y. et al. V.PhyloMaker: an R package that can generate very large phylogenies for vascular plants. Ecography 42, 1353–1359 (2019).
Oksanen, J. et al. Vegan: community ecology package. https://doi.org/10.32614/CRAN.package.vegan (2018).
Magurran, A. et al. Explaining the excess of rare species in natural species abundance distributions. Nature 422, 714–716 (2003).
Hubbell, S. The Unified Neutral Theory of Biodiversity and Biogeography (Princeton University Press, Princeton, NJ, USA, 2001).
Krebs, C. J. Ecological Methodology (Addison-Wesley Educational Publishers, Inc: New York, NY, USA, 1999).
Pell, M. et al. Potential denitrification activity assay in soil - With or without chloramphenicol? Soil Biol. Biochem. 28, 393–398 (1996).
Philippot, L. et al. Importance of denitrifiers lacking the genes encoding the nitrous oxide reductase for N2O emissions from soil. Glob. Change Biol. 17, 1497–1504 (2011).
van Groenigen, J. W. et al. Towards an agronomic assessment of N2O emissions: a case study for arable crops. Eur. J. Soil Sci. 61, 903–913 (2010).
Wood, S. N. Generalized Additive Models: An Introduction with R (Chapman and Hall/CRC, 2017).
Marra, G. et al. Practical variable selection for generalized additive models. Comput. Stat. Data Anal. 55, 2372–2387 (2011).
Wood, S. N. Fast stable restricted maximum likelihood and marginal likelihood estimation of semiparametric generalized linear models. J. R. Stat. Soc. B. 73, 3–36 (2011).
Breheny, P. et al. Visualization of regression models using visreg. R. J. 9, 56–71 (2017).
Genuer, R. et al. VSURF: an R package for variable selection using random forests. R. J. 7, 19–33 (2015).
Liaw, A. et al. Classification and regression by randomForest. R. N. 2, 18–22 (2002).
Molnar, C. et al. iml: an R package for interpretable machine learning. J. Open Source Softw. 3, 786 (2018).
Apley, D. W. et al. Visualizing the effects of predictor variables in black box supervised learning models. J. R. Stat. Soc. Ser. B Stat. Methodol. 82, 1059–1086 (2020).
Acknowledgements
The Digging Deeper project was funded through the 2015–2016 BiodivERsA call, with national funding from the Swiss National Science Foundation (grant 31BD30-172466 to M.G.A.v.d.H), the Deutsche Forschungsgemeinschaft (grant 317895346 to M.C.R.), the Swedish Research Council Formas (grant 2016-0194 to S.H. and 2018-02321 to R.B.), the Spanish Ministerio de Economía y Competitividad (grant PCIN-2016-028 to F.T.M.) and the Agence Nationale de la Recherche (grant ANR-16-EBI3-0004-01 to L.P.). We thank Claudia von Brömssen (Swedish University of Agricultural Sciences) for advice on the generalized additive models.
Funding
Open access funding provided by Swedish University of Agricultural Sciences.
Author information
Authors and Affiliations
Contributions
S.H., M.G.A.v.d.H., F.T.M., L.P., and M.C.R. initiated the study, planned the field work, and contributed materials. A.S., S.B., F.D., A.E., P.G-P., G.G., C.H., D.S.P., and S.R. contributed to data collection. A.S. and M.E.S. performed the analyses, and A.S., M.E.S., G.V., R.B., and S.H. interpreted the results. A.S., M.E.S., and S.H. drafted the manuscript. All authors commented on and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Earth and Environment thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Jinfeng Chang and Mengjie Wang. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Saghaï, A., Smith, M.E., Vico, G. et al. Diverse crop rotations offset yield-scaled nitrogen losses via denitrification. Commun Earth Environ 7, 73 (2026). https://doi.org/10.1038/s43247-025-03116-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43247-025-03116-0