Abstract
Marine microorganisms drive the Earth’s biogeochemical cycles, yet most cannot be cultured in laboratories, severely limiting in-depth studies of their ecological roles and application potential. In a decade-long study, we isolated 16,931 microbial strains from 1516 sampling sites across seven oceanic regions. By integrating omics and environmental data, we found that light conditions—such as long-term variations in sunlight at different latitudes and depths—are key factors shaping microbial biogeography and metabolic strategies. Moreover, 71.67% of the strains exhibited strict light-pattern specificity, a trait linked to differences in metabolic strategies adapted to their native environments. Based on this, we developed a database to predict optimal culture conditions for uncultured microorganisms, clarified the core reasons for their culturability challenges, and provided a practical pathway for exploring the potential of unknown microbes.
Similar content being viewed by others
Introduction
Marine microorganisms constitute a profoundly diverse and ecologically vital component of Earth’s biosphere, underpinning global biogeochemical cycles and the sustained functioning of marine ecosystems. Despite their substantial ecological significance and biotechnological potential, the majority of marine microbial diversity remains uncultured and genetically uncharacterized, a challenge colloquially termed “microbial dark matter”1. While contemporary advancements in metagenomic and single-cell sequencing technologies have dramatically expanded our understanding of microbial diversity, these approaches are inherently limited in their capacity to elucidate microbial physiology, functional plasticity, and intricate environmental interactions, insights that only be definitively obtained from cultured isolates2,3.
The persistent gap between molecular detection and successful cultivation is multifaceted. It is estimated that over 99% of marine microorganisms cannot be cultured under conventional laboratory conditions4. This vast uncultured majority severely constrains our understanding of their metabolic capabilities and ecological contributions. Moreover, the prevailing research paradigm frequently overlooks intraspecies genetic and phenotypic variation. Subtle genotypic differences, such as variations in genes associated with environmental adaptation or metabolic pathways, can manifest as considerable functional divergence, profoundly influencing survival strategies and subsequent culturing outcomes5. Consequently, reliance on a singular “model strain” is insufficient to accurately represent the full spectrum of functional diversity and adaptive mechanisms within natural populations, which necessitates the cultivation of a vast array of strains to achieve a comprehensive understanding of microbial functions.
A key unresolved inquiry pertains to whether microbial cultivability is indeed modulated by biogeographical and environmental factors, with particular emphasis on light exposure, a parameter that exhibits pronounced variability across latitudes and ocean depths. Solar irradiation is a primary driver of photosynthesis and directly influences energy availability, thereby potentially exerting selective pressure for specific metabolic strategies (MS). Although previous studies have suggested that environmental parameters are instrumental in shaping microbial community composition6,7, the mechanistic connections between light regimes, MS differentiation, and cultivation potential of global ocean microbes remain poorly understood.
In addition to established methodologies that infer optimal microbial culture media based on phylogenetic affiliations, recent efforts have integrated multi-omics data with focused cultivation experiments to specifically target previously uncultured lineages8,9. However, these integrated approaches possess inherent limitations. Phylogeny-guided cultivation strategies frequently prove ineffectual for exhibiting significant phylogenetic distances, and genome-informed media design is frequently hampered by incomplete functional annotations10,11. Adoption of a functional trait-based framework, one that strategically aligns environmental conditions with pertinent MS, may offer a potentially more effective pathway to overcoming existing cultivation bottlenecks.
This study systematically investigates the influence of light patterns on microbial biogeography and cultivability by analyzing a comprehensive global dataset comprising 16,931 microbial strains isolated from 1516 sites across seven oceanic regions, combined with metagenomic functional profiling. We propose that light-driven metabolic diversification is a fundamental driver of the observed biogeographical structuring of culturable phenotypes and that the recognition of these patterns can serve as a valuable guide for the targeted isolation of hitherto uncultured microbial taxa. Consequently, our study establishes a novel framework for predicting marine microbial culture media by drawing explicit links between metabolic features and optimal cultivation conditions, thereby providing a scalable strategy to access microbial dark matter of the ocean. Moreover, we have compiled a reference database containing detailed cultivation conditions for over 16,931 physical strains, offering a resource applicable for predicting appropriate cultivation protocols for 57 marine microbial phyla.
Results
Global marine microbial diversity and biogeographical patterns
This research endeavors to elucidate the influence of solar irradiance on the cultivability of marine microorganisms. This objective is pursued through large-scale, global microbial isolation utilizing various defined seawater media, complemented by metagenomic analyses to characterize metabolic functions and adaptive strategies.
An investigation into the global marine microbial diversity profile was conducted using 8308 representative genomes derived from 1038 seawater samples collected worldwide (Fig. 1a). This analysis revealed that Proteobacteria, Bacteroidota, Actinobacteriota, Marinisomatota, and Thermoplasmatota constitute the most prevalent microbial phyla across the global oceans (Supplementary Fig. 1). Furthermore, autoencoder neural networks identified significant roles of drug resistance mechanisms, efflux transporters/pumps, and two-component regulatory systems in shaping marine microbial diversity. These functional categories collectively explained over half of the variance in differentiation observed among marine microbial taxa (Fig. 1b), indicating microbial adaptation to diverse niches and environmental challenges7,12,13.
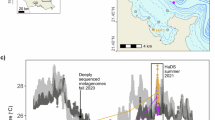
The impact of five zones on the differentiation of marine microbial diversity. a Distribution of genomic and culturable strain sites. b Functional differences shaping marine microbial diversity differentiation. c Microbial diversity distribution in the epipelagic (EPI) layer of global marine microorganisms (Phylum level), with the highest diversity in the near-equatorial southern Pacific. d Microbial diversity distribution in the dark ocean (Phylum level), with the highest diversity in mid-latitude regions of the northern Pacific and mid-high latitude regions of the Atlantic. e Diversity distribution of cultured strains in the EPI layer (Phylum level), with the mid-latitude North Pacific having the most Phyla that have been isolated and cultured. f Diversity distribution of cultured strains in the dark ocean (Phylum level), with the highest cultured microbial diversity near the equator and mid-high latitude regions of the southern hemisphere.
Analysis of the global dataset demonstrated distinct biogeographical distributions for different marine microbial phyla (Supplementary Fig. 1). These patterns were less sensitive to specific oceanic regions, but more profoundly influenced by latitudinal zones, correlating with solar illumination patterns, and by distinct water layers with depths (Fig. 1c, d and Supplementary Fig. 1). For example, sunlit, near-equatorial regions exhibited higher marine microbial diversity (Fig. 1c, d, Supplementary Fig. 1). Specific phyla showed marked abundance in the EPI layer of the tropical Pacific (N = 33), the mesopelagic (MES) layer of the tropical Indian Ocean (N = 23), the EPI layer of the tropical Red Sea (N = 22), the EPI layer of the Arctic Ocean (N = 20), and the MES layer of the tropical Pacific (N = 20) (Supplementary Fig. 1). These findings underscore the critical role of solar illumination patterns in dictating the biogeographical distribution of marine microorganisms6,14.
Biogeographic distribution features of marine microbes under different light conditions based on pure cultures from global oceans
To address the inherent limitations in strain diversity observed in previous cultivation-dependent studies, we established a comprehensive global collection of cultured marine microbial strains. This dataset encompasses microbial isolates from over 1516 sampling sites spanning seven oceanic regions, with collected samples representing a broad latitudinal range of 159.3° and depths up to 11,034 m. From these diverse seawater samples, we isolated 16,931 strains in the past two decades, representing 708 microbial genera, thereby ensuring a robust and representative collection of cultivable marine microorganisms (Supplementary Fig. 1 and Supplementary Data 1). Detailed information regarding strain entities and their respective cultivation conditions is curated and accessible via the Marine Culture Collection of China database (MCCC, https://www.mccc.org.cn/).
Taxonomic annotations of these pure cultured strains revealed their affiliation with 17 major microbial phyla, including but not limited to: Pseudomonadota, Bacillota, Verrucomicrobiota, Actinomycetota, Bacteroidota, Streptophyta, Planctomycetota, Deinococcota, Balneolota, Thermodesulfobacteriota, Fusobacteriota, Rhodothermota, Campylobacterota, delta/epsilon subdivisions, Bdellovibrionota, Methanobacteriota, Ascomycota. Consistent with our hypothesis, these cultured strains exhibited differential distribution patterns with respect to solar illumination; for example, strains belonging to Balneolota and Deinococcota were exclusively recovered from aphotic ocean environments. Conversely, strains of Ascomycota, Bdellovibrionota, Fusobacteriota, Proteobacteria, and Thermodesulfobacteriota were predominantly, if not exclusively, obtained from photic (Supplementary Data 1).
When analyzing their distribution across five recognized biogeographical zones, it became evident that the global microbial isolates demonstrated distinct preferences influenced by specific sunlight illumination patterns, rather than solely by the presence or absence of light. In sunlit oceanic environments, isolates from seven phyla—Ascomycota and Fusobacteriota (Tropical zone), Planctomycetota and Thermodesulfobacteriota (Tropical zone), Campylobacterota and Proteobacteria (North Temperate zone), and Verrucomicrobiota (South Temperate zone)—were restricted to particular climatic zones, collectively representing approximately 43.75% of these geographically constrained phyla. In contrast, the influence of direct sunlight was less pronounced in aphotic environments, leading to weaker climatic zone preferences among isolated phyla. For instance, only four phyla—Deinococcota, Planctomycetota, and Verrucomicrobiota (Tropical zone), and Methanobacteriota (North Temperate zone)—were exclusively isolated from specific climatic zones, accounting for approximately 33.33% of these less geographically defined phyla. The heightened preference for specific climatic zones observed in cultivated strains from sunlit oceans, compared to those from dark oceans, further reinforces the significant role of illumination patterns in shaping the global distribution of marine microorganisms.
This disparity in distribution, influenced by light conditions, was not confined to the phylum level but was also evident at the species level. Approximately 45.10% of all identified species were exclusively isolated from illuminated marine environments, whereas 23.68% were exclusively recovered from dark marine habitats. Considering the five climatic zones, 66.28% of all species exhibited a preference for isolation within specific climatic zones. This proportion increased to 71.67% for species exclusively isolated from illuminated marine environments. Notably, a substantial proportion of species confined to specific climatic zones also shared the same higher taxonomic ranks: 50.00% belonged to the same phylum and 27.50% to the same genus. These findings collectively illustrate the profound impact of sunlight patterns on the cultivation acquisition of novel microbial taxa, suggesting that sunlight directly influences the metabolic functional differentiation and strategic adaptation selection of these marine species.
Distribution of the major MS of marine microbes in global oceans
We have identified eight major MS (MS1-MS8) employed by marine microorganisms based on the similarity of their metabolic functions, and observed that individual species can exhibit diverse strategy profiles. (Supplementary Data 2). These distinct MS displayed differential preferences for varying illumination conditions (Fig. 2a, b). For instance, within tropical regions, the type MS7 demonstrated higher abundance in sunlit oceans compared to aphotic environments, while MS8 showed an inverse trend. Additionally, MS1, MS2, MS3, and MS4 were predominantly found in northern polar regions, whereas MS8 exhibited a stronger association with tropical zones (Fig. 2b, c). This suggests a compelling relationship between the biogeographical patterns of marine microorganisms, shaped by environmental factors such as solar illumination, and the influence of these factors on their metabolic functions, thereby manifesting distinct MS.
a Functional preferences for different MS. b Light environment preferences of marine microorganisms with different MS types. c Functional differences and cultivation potential of different MS. d Important functions shaping marine microbial MS, predicted based on autoencoder neural networks. e Differences in key functions of different MS. We observed that different MS exhibit unique functional preferences, which may reflect the adaptability of marine microorganisms to their environments and could also be influenced by the combinations of their functions.
As expected, clustering analysis of metabolic functions confirmed that different MS align with specific functional preferences (Fig. 2a), illustrating how the integration of metabolic functions shapes and delimits these strategies. In terms of carbon fixation, all MS, with the exception of MS2 and MS8, demonstrated a preference for particular carbon fixation pathways (Supplementary Fig. 2). Specifically, MS1 favored the C4-dicarboxylic acid cycle (NADP-malic enzyme type) (M00172) and the dark reactions of Crassulacean acid metabolism (CAM) (M00168). MS3 showed a preference for the 3-Hydroxypropionate bi-cycle (M00376). MS4 favored the phosphate acetyltransferase-acetate kinase pathway (M00579) and the reductive citrate cycle (Arnon-Buchanan cycle) (M00173). MS5 preferred the reductive acetyl-CoA pathway (Wood-Ljungdahl pathway) (M00377) and the reductive pentose phosphate cycle (Calvin cycle) (M00165). MS6 favored the C4-dicarboxylic acid cycle (NADP-malic enzyme type) (M00172) and the light reactions of CAM (M00169). Finally, MS7 prefers the Incomplete reductive citrate cycle (acetyl-CoA => oxoglutarate) (M00620).
To exemplify how MS diverge in their underlying functional frameworks, we analyzed molecular requirements of carbon fixation—a process directly coupled to light availability. This analysis revealed that framework-specific differences dictate functional specialization across microbes. These frameworks encompass the essential reaction products, enzymes, substrates, and energy sources required for the integration of novel functionalities. For example, the differences in connectivity and linked functions between reaction products, substrates, and enzymes of primary functions (those with relative abundance >1%) and carbon fixation pathways contributed to the observed preferences of MS1 (M00168, n = 6, Rank = 3), MS5 (M00165, n = 21, Rank = 1; M00377, n = 2, Rank = 2), MS6 (M00169, n = 8, Rank = 1; M00172, n = 8, Rank = 1), and MS7 (M00620, n = 39, Rank = 1) for specific carbon fixation pathways (Supplementary Data 3 and 4). Furthermore, the strong preference for carbon fixation pathways in MS3 (M00376, n = 74, Rank = 1) and MS4 (M00173, n = 108, Rank = 2; M00579, n = 8, Rank = 1) might be attributable to the extensive connectivity between reaction products, substrates, and enzymes of certain secondary metabolic functions (those with relative abundance >0.1%) (Supplementary Data 3 and 4). To contextualize these pathway preferences within microbial trophic strategies, we note that carbon fixation not only supports autotrophy but also intersects with mixotrophy in heterotrophic-dominated communities. For instance, pathways like the reductive citrate cycle (M00173) may provide metabolic flexibility under fluctuating light conditions, explaining their association with specific MS (e.g., MS4) in energy-limited environments. For MS2 and MS8, which did not exhibit a distinct preference for carbon fixation pathways, different underlying reasons were identified. While MS2 possessed substantial and diverse potential for developing carbon fixation capabilities, evidenced by a high functional investment in both the dark reactions of CAM (M00168), only second to MS1, and the light reactions of CAM (M00169), ranking behind MS6 and MS3 (Supplementary Data 3–5), this broad potential might inadvertently limit its specialization in specific carbon fixation pathways. In contrast, MS8 appeared to have a diminished capacity to establish connections between reaction products, substrates, and enzymes through carbon fixation pathways, consequently resulting in a lower potential for developing these functions (Supplementary Data 3 and 4).
To further investigate how light regimes shape MS differentiation, we employed autoencoder neural networks to predict crucial functions influenced by solar illumination gradients. The model identified core cellular systems—including transport systems, secretion systems, ribosome synthesis, RNA processing, and DNA polymerases—as key determinants of metabolic specialization (Fig. 2d, e and Supplementary Fig. 3). We propose that light availability serves as a primary selective pressure, fine-tuning resource allocation to these systems: energy-replete, high-light environments favor investment in rapid-growth machinery (e.g., ribosome synthesis), while energy-limited conditions select for efficient nutrient acquisition and conservation strategies15,16,17. The divergence among MS1-8 stems from distinct functional trajectories shaped by light-driven selection on key metabolic gene sets. Specifically, varying light regimes create differential selection pressures on: (1) energy investment strategies (ribosome synthesis, RNA processing), (2) nutrient acquisition systems (transporters), and (3) environmental interaction mechanisms (secretion systems). These genomic investment patterns, captured by our deep learning model, ultimately manifest as the global biogeography of MS here observed. (Fig. 2d, e). For instance, MS3, MS4, and MS6, characterized by a lower proportion of key genes, displayed preferences for distinct supplementary functions. MS3 tends to utilize microbial secretion systems and ABC-2 type transport systems, MS4 favors mineral and organic ion transport systems, and MS6 favors a broader repertoire of RNA processing functions (enabling rapid sensing and response) over the maintenance of diverse phthalate transport systems.
Database of marine microbial cultivation conditions designed based on MS
We observed that the distribution patterns of cultured strains in illuminated and aphotic oceanic environments displayed distinct biogeographical characteristics: planar and zonal (Fig. 1e, f). This finding suggests that the diversity in MS not only reflects adaptation to natural environmental conditions but also informs potential artificial cultivation parameters.
Leveraging the culture media employed by cultured strains with specific MS, we established a framework of reference media for uncultured microbes exhibiting the same strategy. This approach facilitates the creation of a rapid matching and query database for predicting optimal cultivation conditions for marine microbes (Fig. 3, https://mccc.org.cn/about/CultureConditions). This database enables users to retrieve recommended cultivation conditions for both cultured and uncultured marine microorganisms. Researchers can query generalized culture media compositions and reference temperature ranges based on GTDB classification names. By integrating in situ environmental data from sampling sites and available metagenomic sequencing data, researchers can further personalize predicted MS-based general culture media for specific targets. Additionally, we provide an MS recommendation feature, offering a prioritized reference for metabolic functions when customizing media for target strains. The application of these principles holds potential for the isolation and cultivation of microorganisms from additional 57 marine phyla.
This figure illustrates the analytical workflow for elucidating potential culture protocols based on the MS of marine microorganisms. 1. Acquire genomes and cultured strains of microorganisms from global oceanic sources. 2. Analyze MS through genomic sequencing and functional annotation. 3. Align culture protocols with identified MS. 4. Develop a searchable database of recommended culture protocols for marine. 5. Validate the actual matching rate of the predicted culture protocols. 6. Assess potential application scenarios and translational value.
To validate the efficacy of our approach, we conducted extensive isolation and cultivation experiments utilizing the culture media generated from this database. The results demonstrated a high accuracy of 99.57% (n = 231) for predicted culture media in successfully cultivating strains isolated from seawater samples. For strains originating from other water bodies, the accuracy was 79.11% (n = 316) (Supplementary Data 6). The diminished accuracy observed for samples from non-seawater environments may be attributed to the fact that the genomic and strain data underpinning our database were primarily derived from seawater isolates.
These findings underscore the specificity and feasibility of employing metabolically predictive media for the isolation and cultivation of novel marine microbial strains. They also highlight how variations in MS directly influence the success rate and efficiency of acquiring.
Metabolic differences between the cultured and uncultured phyla of each MS type
Over the past two decades, our efforts have yielded cultured strains representing six types of MS (MS1, MS2, MS3, MS4, MS5, and MS8). By comparing the abundance of functional modules between cultured phyla (those for which we have obtained cultured strains) and uncultured phyla (those currently lacking cultured representatives) across these different MS (Fig. S5 and Supplementary Data 7), we have identified significant functional deficiencies or dependencies in uncultured microbial phyla. These deficiencies are particularly evident in areas related to energy metabolism, substance transport, and environmental adaptation.
Generally, uncultured phyla exhibit limited intrinsic energy metabolic capabilities and display a pronounced reliance on diverse substance transport systems (Fig. S5 and Supplementary Data 7). For instance, within MS1, uncultured phyla showed a mere 3.76% abundance in the cytochrome b6f complex (M00162), a significant deficit compared to the 44.94% observed in cultured phyla (a difference of 41.18%), indicating constrained energy synthesis capacity. Uncultured MS2 phyla demonstrated a substantial abundance of 69.14% in the iron complex transport system (M00240), vastly exceeding the 7.51% found in cultured phyla (a difference of 61.63%), suggesting a strong dependence on exogenous metal ions. Similarly, uncultured MS3 phyla presented with a 52.63% abundance in the cobalt-nickel transport system (M00245), in stark contrast to the 7.53% in cultured phyla (a difference of 45.10%), further corroborating their specific metal ion requirements.
We also observed that uncultured MS4 phyla demonstrated exceptionally high abundance of 89.54% in coenzyme A biosynthesis (M00120), significantly surpassing the 6.45% in cultured phyla (a difference of 83.09%). This suggests a potential dependence on abundant environmental cofactors, which may contribute to their cultivation challenges. For uncultured MS5 phyla, abundances of 54.04% and 53.02% were observed in photosystem I (M00163) and photosystem II (M00161), respectively, while cultured phyla showed no such representation. This highlights nutritional constraints that likely limit the cultivation of MS5-type marine microbes. Notably, limitations in gene transformation mechanisms may also restrict the cultivability of certain groups. Uncultured MS8 phyla accounted for 91.03% in the DNA transformation transport system (M00429), a significant increase from the 26.67% in cultured phyla (a difference of 64.36%). This pronounced genetic functional variability may impede their cultivation under fixed laboratory conditions or lead to the isolation of only specific subpopulations within this MS.
This comprehensive functional comparison elucidates the metabolic and ecological factors underlying the cultivation barriers of marine microbial phyla, providing valuable insights for future cultivation efforts and ecological studies.
Discussion
Our decade-long global investigation establishes that solar illumination gradients fundamentally govern marine microbial biogeography and functional diversification by acting as a selective filter on core cellular functions. The key genes identified by our deep learning model—involved in transport, secretion, ribosome synthesis, and RNA processing—represent the molecular mechanisms through which light regimes shape MS differentiation. By integrating 16,931 cultured microbial strains from global 1516 sites with metagenomic functional profiles, we demonstrate that light-exposure patterns, influenced by latitude, depth, and climatic zones, driven MS differentiation even within phylogenetically identical populations. This establishes a mechanistic link that addresses a critical knowledge gap regarding the pronounced biogeographical structuring of marine microbial cultivability. Specifically, 71.67% species were isolated exclusively from specific climatic zones (e.g., Proteobacteria in temperate photic zones, Fusobacteriota in tropical photic zones, Supplementary Data 1), a finding that challenges conventional assumptions of widespread microbial cosmopolitanism implying uniform cultivability.
The phenomenon is underscored by the dominance of eight core MS (MS1–MS8). Solar illumination regimes appear to function as evolutionary filters, preferentially selecting for energy-acquisition tactics optimized for local environmental conditions. For instance, MS1, MS2, MS3, and MS4 are predominantly found in Arctic regions, whereas MS8 is more abundant in tropical areas (Fig. 2b, c). This distribution strongly suggests that MS differentiation reflects microbial adaptations to distinct environmental selection pressures. Further analysis of the metabolic frameworks underlying these strategies (Supplementary Data 3 and 4) indicates variability in the energetic and biosynthetic costs associated with evolving new metabolic functions15. For example, MS4-type microbes likely necessitate less complexity to develop the reductive citrate cycle (Arnon-Buchanan cycle, M00173) compared to other metabolic types, as they possess more reactions that share substrates, products, or enzymes with the reductive citrate cycle. This is supported by a greater abundance of genes related to substrates, products, and enzymes necessary for its completion (Supplementary Data 3 and 4). The association of the type MS4 with carbon fixation highlights the existence of diversified pathways for energy acquisition and material metabolism in marine microbes and explains how varying MS optimizes ecological adaptability through functional synergy, thereby shaping global distribution patterns of marine microbial communities.
Furthermore, the prevalence of specific carbon fixation pathways (e.g., Calvin cycle in MS5) underscores their significance beyond obligatory autotrophy. In mixotrophic microbes, which oscillate between heterotrophic and autotrophic modes, these pathways serve as adaptive tools for energy optimization under variable light regimes. For example, the investment in carbon fixation by heterotrophic-dominated lineages may enhance survival in oligotrophic sunlit zones, where light-driven plasticity allows for resource partitioning. This functional synergy not only elucidates the biogeographical patterns we observed but also may affect the isolation and cultivation of microorganisms. Remarkably, even our extensive, albeit historically random, cultivation efforts yielded clear affinities to solar illumination patterns, with 66.28% of cultured species found exclusively within specific climatic zones. This observation suggests that artificial cultivation conditions can be regarded as distinct ecological niches. Consequently, functional variations in MS, largely shaped by solar illumination, likely enhance the amenability of microbes from the same phylum but different climatic zones to cultivation due to their divergent MS. For example, widely distributed marine microorganisms such as Proteobacteria and Fusobacteriota, although common in general (Supplementary Fig. 1), were exclusively isolated from northern temperate and tropical photic zones, respectively, in our study (Supplementary Data 1). Cultivation designs informed by these MS subsequently achieved a high validation match rate of 99.57% in experimental verification (n = 231, Supplementary Data 6), underscoring the critical role of MS-informed cultivation in effectively sampling marine microbial diversity aligned with environmental factors.
It is important to note that our MS analysis relies on MAGs, which, despite rigorous quality control, may exhibit incompleteness or contamination. These limitations could affect the annotation of certain metabolic pathways (e.g., those with low coverage or horizontal gene transfer events), potentially influencing the clustering of MS. Future studies incorporating complete genomes or long-read sequencing could further validate these findings.
The cultivation condition design approach based on MS presents a novel paradigm for guiding the large-scale isolation of previously uncultured microorganisms. This methodology circumvents the limitations of traditional phylogeny-based medium recommendations, which often exhibit suboptimal performance for distantly related taxa8,9, and partially mitigates challenges posed by emerging metabolism-modeling-based cultivation designs susceptible to confounding by unknown functions10,11. It is crucial to note that uncultured phyla associated with different MS generally exhibit reduced energy metabolism capabilities and a heightened dependence on diverse substance transport systems compared to cultured phyla (Supplementary Data 7). For example, the genomes of the uncultured phyla belonging to MS2 show a 61.63% higher abundance in the iron complex transport system (M00240) than their cultured counterparts. Therefore, when designing new culture media, these metabolic features related to nutrient requirement must be considered, referencing the recommended key functional modules specific to each MS in the database. Moreover, our study identified several representative functional modules within MS1 directly or indirectly involved in cell wall synthesis, including lipopolysaccharide biosynthesis, CMP-KDO biosynthesis, and lipopolysaccharide export systems (Fig. 2a), highlighting the critical role of cell wall synthesis for MS1-type marine microorganisms. Notably, the sole isolated Marinisomatota strain IA91 exhibits a metabolic profile consistent with the MS1 strategy (best match r = 0.7292, P < 0.05) and was successfully cultivated via co-culture, which compensated for its dependence on exogenous peptidoglycan for cell wall synthesis18. Although the database developed here achieved a 99.57% validation match rate in seawater cultivation experiments, the validation match rate dropped to 79.11% (n = 316) when applied to isolates from other aquatic environments (Supplementary Data 6). This decrease may be attributed to the database’s primary construction from seawater-based media and associated genomes and isolates.
This framework not only applies to marine systems but also holds potential for broader environments. A universally predictive understanding of microbial cultivation requires systematic expansion and integration of strain-culture data derived from diverse environmental samples. We propose to extend this framework beyond marine systems by incorporating data from a broader range of biomes, thereby testing the generalizability of the “light–metabolism–cultivation” principle. This approach will not only enable forecasting of future ecological dynamics but also allow a reinterpretation of the evolutionary past—elucidating how historical variations in light regimes have acted as a selective force in shaping microbial MS. Such efforts are expected to bridge deep-time evolutionary history with predictive ecology, providing a foundational framework for understanding microbial responses to environmental change.
Conclusion
Our global investigation establishes solar illumination gradients as fundamental drivers of marine microbial biogeography and metabolic specialization. We demonstrate that phylogenetically conserved populations diverge in their energy acquisition strategies, influenced by localized light exposure histories. This plasticity explains the observed strict climatic-zone specificity in over 70% of cultured species. The discovery of eight core MS (MS1-MS8) reveals how varying light regimes optimize functional trade-offs between energetic efficiency and environmental adaptability. Based on these principles, we developed the predictive cultivation framework, achieving 99.57% accuracy in seawater strain cultivation. This framework enables targeted acquisition of previously unculturable lineages across the 57 microbial phyla, thereby transitioning the exploration of microbial dark matter from a serendipitous endeavor to a predictive scientific pursuit. Consequently, these findings present paradigm-shifting implications for microbial ecology. Light emerges as a significant, non-phylogenetic determinant of biogeographic distribution, underscoring the importance of environmental context in cultivation strategies. The establishment of metabolic plasticity necessitates an environment-first approach to microbial isolation. The comprehensive database (https://mccc.org.cn/about/CultureConditions) provides a universally applicable blueprint for unlocking microbial diversity. This resource holds transformative potential for both the bioprospecting of novel bioactive compounds and the development of climate resilience solutions.
Methods
Metagenomic data selection
The metagenomic data originates from large-scale marine surveys, offering sufficient sequencing depth and temporal series information. These data aim to comprehensively cover marine microbial communities across various global basins, different depth layers, and multiple time points. This dataset has been published by Paoli et al.19 and is interactively accessible via the Ocean Microbiomics Database (OMD, https://www.microbiomics.io/ocean/). The dataset includes 1038 metagenomes, comprising samples collected from Tara Oceans (virus-enriched, n = 190; prokaryote-enriched, n = 180)20,21, BioGEOTRACES (n = 480), the Hawaiian Ocean Time-series project (HOT, n = 68), the Bermuda-Atlantic Time-series Study (BATS, n = 62)22, and samples from Malaspina (n = 58)23 (Supplementary Data 2). To this end, we placed microbial genomes in the standardized bacterial and archaeal phylogenomic trees of the GTDB24.
Quality evaluation of genomes and identification of representative genomes
The quality of each genome was assessed by CheckM (v.1.0.13)25 and Anvi’o (v.5.5.0)26. If CheckM or Anvi’o report completeness ≥50% and contamination ≤10%, the metagenomic box and external genomes are reserved for downstream analysis. Although stringent quality filters (completeness ≥50% and contamination ≤10%) were applied to minimize biases, the inherent incompleteness and potential contamination of MAGs may still introduce uncertainties in metabolic pathway annotations, particularly for low-abundance or complex pathways. This limitation should be considered when interpreting the MS clustering results. These indicators are then aggregated into mean completeness (mcpl) and mean contamination (mctn). Genome quality was classified according to the following criteria: high quality (mcpl ≥ 90%, mctn ≤ 5%), good quality (mcpl ≥ 70%, mctn ≤ 10%)27, medium quality (mcpl ≥ 50%, mctn ≤ 10%), and average quality (mcpl ≤ 90% or mctn ≥ 10%). Evaluate the quality of the filtered genome using dRep (v.2.5.4)28 to generate a quality score (Q and Q’):
(1) Q = mcpl − 5 × mctn;
(2)Q′ = mcpl − 5 × mctn + mctn × (strain heterogeneity)/100 + 0.5 × log[N50].
Subsequently, the genome was deduplicated using dRep (v.2.5.4, parameter -comp 0-con 1000-sa 0.95-nc 0.2)28 at 95% ANI threshold29,30, and using single-copy marker gene from SpecI31. This process aims to provide species-level clustering of the genome. A representative genome was selected for each dRep cluster based on the previously defined maximum mass score (Q ‘).
Functional annotation of genome
Each genome is functionally annotated using prokka (v.1.14.5)32 and the “Bacteria” parameter is specified, which also reports genomic features such as CRISPR regions. Universal single-copy marker genes (uscMGs) were identified using fetchmg (v.1.2)33. Based on eggNOG (v.5.0)34, emapper (v.2.0.1)35 was used to assign homologous groups. Gene prediction was performed by querying the KEGG database (release 2020-02-10) with DIAMOND (v.0.9.30)36. Proteins were aligned with the database, and the query coverage was required to be ≥70%. After NCBI prokaryotic genome annotation pipeline37, further filtering was performed based on the maximum expected bit fraction (reference-self), with a cut-off value of ≥50%.
Gene-level profiling
Similar to the method described previously20,21, the proteins coding for genes are clustered using CD-HIT (v.4.8.1)38, with shorter genes clustered with 95% consistency and 90% coverage. The longest sequence is selected as the representative gene for each gene cluster. The mapping was then performed using BWA (v.0.7.17-r1188, -a)39, and the BAM file was filtered to preserve only aligners with ≥95% consistency and ≥45 base pair aligns. According to the gene abundance normalized by length, the number of insertions for the best unique comparison is first calculated, and then the number of ambiguous insertions is calculated and added to the respective target genes in proportion.
Metabolic reconstitution
Metabolic(v4.0)40 software was used to reconstruct key metabolic pathways representing the genome. Specifically, in metaboly-G mode, with the genome (Supplementary Data 2, Supplementary Data 5) file in fasta format as input, the parameter is “-m-cutoff 0.75-kofam-db full”.
Metabolic strategies analysis
We calculated the Pearson’s correlation of different metabolic function (KEGG module) qualitative matrix in 8306 marine microbial genomes (encompassing only bacteria and archaea) using the R-package “Hmsic,” and retained data with correlation coefficient > 0.6 and P < 0.05 (https://cran.r-project.org/). Then we used Louvain algorithm (https://github.com/topics/louvain-algorithm) from the interactive platform Gephi 0.10.1 (https://gephi.org/) to identify MS41. The Louvain algorithm is a hierarchical clustering algorithm, that recursively merges communities into a single node and executes the modularity clustering on the condensed graphs. The relationships that connect the nodes in each component have a property weight which determines the strength of the relationship.
Dimensionality reduction and feature importance analysis in species functional data using autoencoder neural networks
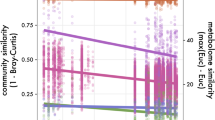
In this study, we initiated our analysis by importing the Phylum metabolic dataset into a Python-based analytical environment, utilizing the Pandas42 library. Subsequently, the dataset was transformed into a NumPy43 array to facilitate subsequent computational procedures. We designed an autoencoder neural network to derive a compressed representation of the species functional data. The architecture of the model incorporated an input layer, which was dimensioned to match the number of features present in the dataset. This was followed by an encoded layer with a reduced dimensionality of 64, and a decoded layer tasked with reconstructing the original input data. To mitigate the risk of overfitting, a Dropout layer was integrated subsequent to the encoded layer. Training of the model was executed employing the Adam optimization algorithm in conjunction with a binary cross-entropy loss function. The dataset was partitioned into training and testing subsets, adhering to an 80:20 split. The training process spanned 100 epochs and was conducted with a batch size of 32. To further counteract overfitting, an early stopping mechanism was implemented, which would halt training should the validation loss fail to exhibit improvement over a sequence of 5 epochs. We evaluated the significance of individual features in ascertaining species functional similarity by examining the weights within the encoded layer. The mean absolute weight for each feature was computed and subsequently ordered to pinpoint the most salient features. The encoding segment of the autoencoder was utilized to compress the functional data into a lower-dimensional space. We then calculated the Euclidean distance between the encoded representations of species pairs, which served as a metric for similarity assessment.
For the MS dataset, encompassing 8 MS clusters and 559 features, the model was structured with an input layer equivalent to the feature count, an encoded layer with a dimensionality reduction to 32, and a decoded layer to reconstruct the input data. A Dropout layer was incorporated post-encoding to avert overfitting. The model underwent training with the Adam optimizer and binary cross-entropy as the loss function. The dataset was divided into training and testing sets with a 90:10 ratio. The training was carried out over 50 epochs with a batch size of 16. An early stopping protocol was applied to prevent overfitting, terminating training if no improvement in validation loss was observed over 10 consecutive epochs. The encoded layer’s weights were analyzed to ascertain the importance of each feature in determining species functional similarity. The mean absolute weight of each feature was calculated and ranked to discern the most influential features. The encoder component of the autoencoder was employed to encode the functional data into a condensed dimensional space. The Euclidean distance between the encoded representations of species pairs was computed to yield a similarity score, which is detailed in the Supplementary Files under the filename EcoEncoderSimScore_Phylum.py and EcoEncoderSimScore_MS.py.
Diversity and data selection of cultured strains
We established a comprehensive strain and 16S rRNA resource for culturable microorganisms from global ocean seawater. While the primary focus is on bacterial and archaeal composition, the collection also encompasses eukaryotic lineages (e.g., Streptophyta, Chordata, Arthropoda, Ascomycota). The prokaryotic (bacterial and archaeal) data were utilized for cultivation condition prediction. In contrast, the eukaryotic data served to investigate the influence of light exposure on random isolation outcomes and to provide a valuable, accessible resource for future eukaryotic isolation efforts. This integrated dataset, available through the Marine Culture Collection of China (MCCC; https://www.mccc.org.cn/), comprises 16,931 cultured strains isolated using over 125 media types, with accompanying culture temperatures and habitat information (Supplementary Data 2).
Potential culture conditions for marine microorganisms
The potential Culture conditions of Marine microorganisms (encompassing only bacteria and archaea) are identified by MCCC The strain annotation information included was matched with genome annotation information in different MS.
Statistical analysis
Use Chiplot44 (https://www.chiplot.online/) for deep learning prediction of functional similarity visualization of MS, heatmap clustering using the complete linkage method, and based on Euclidean distance to calculate. Use Chiplot44 (https://www.chiplot.online/) to visualize the metabolic functions of different MS. The metabolic functions were clustered using the weighted method and calculated based on Euclidean distance and standardized using StandardScaler.
Data availability
The metagenomic data used in this study were downloaded from the European Nucleotide Archive (ENA) and a summary of their accessions is provided in Supplementary Data 2. Publicly available genomes were downloaded from Figshare (https://doi.org/10.6084/m9.figshare.4902923) for manually curated MAGs from Tara Oceans, from ENA using the project accession PRJEB33281 for GORG and from https://mmp2.sfb.uit.no/databases/ for MarDB. The GEM MAGs were downloaded from https://portal.nersc.gov/GEM/. MAGs contained in the GTDB r89 were downloaded from https://data.gtdb.ecogenomic.org/releases/release89/. The MIBiG and BiG-FAM databases can be accessed at https://mibig.secondarymetabolites.org/ and https://bigfam.bioinformatics.nl/, respectively. Cultivating strains and culture conditions can be obtained by MCCC query.
Code availability
The code supporting the findings of this study is publicly available at FigShare: https://figshare.com/articles/software/AE-code/28668908.
References
Lewis, W. H., Tahon, G., Geesink, P., Sousa, D. Z. & Ettema, T. J. G. Innovations to culturing the uncultured microbial majority. Nat. Rev. Microbiol. 19, 225–240 (2021).
Sunagawa, S. et al. Tara Oceans: towards global ocean ecosystems biology. Nat. Rev. Microbiol. 18, 428–445 (2020).
Robinson, S. L., Piel, J. & Sunagawa, S. A roadmap for metagenomic enzyme discovery. Nat. Prod. Rep. 38, 1994–2023 (2021).
Lloyd, K. G., Steen, A. D., Ladau, J., Yin, J. & Crosby, L. Phylogenetically novel uncultured microbial cells dominate earth microbiomes. mSystems 3, e00055–18 (2018).
Newman, D. J. & Cragg, G. M. Natural products as sources of new drugs over the nearly four decades from 01/1981 to 09/2019. J. Nat. Prod. 83, 770–803 (2020).
Hanson, C. A. Temporal and Spatial Patterns in Marine Cyanophage Communities. PhD thesis, University of California, Irvine (2012).
Rappaport, H. B. & Oliverio, A. M. Extreme environments offer an unprecedented opportunity to understand microbial eukaryotic ecology, evolution, and genome biology. Nat. Commun. 14, 4959 (2023).
Zhang, J. et al. High-throughput cultivation and identification of bacteria from the plant root microbiota. Nat. Protoc. 16, 988–1012 (2021).
Oberhardt, M. A. et al. Harnessing the landscape of microbial culture media to predict new organism–media pairings. Nat. Commun. 6, 8493 (2015).
Dukovski, I. et al. A metabolic modeling platform for the computation of microbial ecosystems in time and space (COMETS). Nat. Protoc. 16, 5030–5082 (2021).
Rodríguez del Río, Á et al. Functional and evolutionary significance of unknown genes from uncultivated taxa. Nature 626, 377–384 (2024).
Du, D. et al. Multidrug efflux pumps: structure, function and regulation. Nat. Rev. Microbiol. 16, 523–539 (2018).
Shu, W.-S. & Huang, L.-N. Microbial diversity in extreme environments. Nat. Rev. Microbiol. 20, 219–235 (2022).
Burnett, W. J. Longitudinal variation in algal symbionts (zooxanthellae) from the Indian Ocean zoanthid Palythoa caesia. Mar. Ecol. Prog. Ser. 234, 105–109 (2002).
Malik, A. A. et al. Defining trait-based microbial strategies with consequences for soil carbon cycling under climate change. ISME J. 14, 1–9 (2020).
Duchêne, C. et al. Diatom phytochromes integrate the underwater light spectrum to sense depth. Nature 637, 691–697 (2025).
Lu, Y. et al. Role of an ancient light-harvesting protein of PSI in light absorption and photoprotection. Nat. Commun. 12, 679 (2021).
Katayama, T. et al. A marine group A isolate relies on other growing bacteria for cell wall formation. Nat. Microbiol. 9, 736–750 (2024).
Paoli, L. et al. Biosynthetic potential of the global ocean microbiome. Nature 607, 111–118 (2022).
Sunagawa, S. et al. Structure and function of the global ocean microbiome. Science 348, 1261359 (2015).
Salazar, G. et al. Gene expression changes and community turnover differentially shape the global ocean metatranscriptome. Cell 179, 1068–1083.e21 (2019).
Biller, S. J. et al. Marine microbial metagenomes sampled across space and time. Sci. Data 5, 180176 (2018).
Acinas, S. G. et al. Deep ocean metagenomes provide insight into the metabolic architecture of bathypelagic microbial communities. Commun. Biol. 4, 1055 (2021).
Parks, D. H. et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat. Biotechnol. 36, 996–1004 (2018).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Eren, A. M. et al. Anvi’o: an advanced analysis and visualization platform for ‘omics data. PeerJ 3, e1319 (2015).
Bowers, R. M. et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat. Biotechnol. 35, 725–731 (2017).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J 11, 2864–2868 (2017).
Jain, C., Rodriguez-R, L. M., Phillippy, A. M., Konstantinidis, K. T. & Aluru, S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9, 5114 (2018).
Olm, M. R. et al. Consistent metagenome-derived metrics verify and delineate bacterial species boundaries. mSystems 5, e00731–20 (2020).
Mende, D. R., Sunagawa, S., Zeller, G. & Bork, P. Accurate and universal delineation of prokaryotic species. Nat. Methods 10, 881–884 (2013).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Sunagawa, S. et al. Metagenomic species profiling using universal phylogenetic marker genes. Nat. Methods 10, 1196–1199 (2013).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
Huerta-Cepas, J. et al. Fast genome-wide functional annotation through orthology assignment by eggNOG-mapper. Mol. Biol. Evol. 34, 2115–2122 (2017).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2014).
Tatusova, T. A. et al. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 44, 6614–6624 (2016).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Zhou, Z. et al. METABOLIC: high-throughput profiling of microbial genomes for functional traits, metabolism, biogeochemistry, and community-scale functional networks. Microbiome 10, 33 (2022).
Blondel, V., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. 2008, P10008 (2008).
McKinney, W. Pandas: a foundational Python library for data analysis and statistics. Python High Perform. Sci. Comput. 14, 1–9 (2011).
Harris et al. Array programming with NumPy. Nature 585, 357–362 (2020).
Xie, J. et al. Tree visualization by one table (tvBOT): a web application for visualizing, modifying and annotating phylogenetic trees. Nucleic Acids Res. 51, W587–W592 (2023).
Acknowledgements
This research was supported by the National Natural Science Foundation of China (Grant Nos. 42030412 and 91851203 to Z.Z.S.) and the Discipline Breakthrough Precursor Project of the Ministry of Education of China (JYB2025XDXM803). We would like to express our gratitude to the individuals and teams responsible for generating the metagenomic data (Tara Oceans, Malaspina, as well as Biogeotraces, HOT, and BATS) and for producing the publicly available genomes (GORG SAGs, The MAR Databases, and manually curated MAGs).
Author information
Authors and Affiliations
Contributions
Shizheng Xiang: Conceptualization, methodology, formal analysis, investigation, writing— original draft. Guangyu Li: Investigation, data curation, validation. Yifei Huang: Investigation, resources. Linfeng Gong: Data curation, validation. Jianyang Li: Resources, validation. Guizhen Li: Resources, validation. Liping Wang: Resources. Wei Ye: Visualization. Libo Yu: Data curation. Zhen Chen: Formal analysis, visualization. Hongchen Jiang: Supervision, conceptualization, writing— review and editing. Zongze Shao: Supervision, project administration, funding acquisition, writing —review and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Earth and Environment thanks the anonymous reviewers for their contribution to the peer review of this work. Primary handling editors: Haihan Zhang and Alice Drink.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xiang, S., Li, G., Huang, Y. et al. Light exposure patterns shape marine microbial biogeography and metabolic strategy. Commun Earth Environ 7, 270 (2026). https://doi.org/10.1038/s43247-026-03289-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43247-026-03289-2