Abstract
Soybean farming—providing protein-rich feed for farm animals worldwide—is the third largest driver of tropical deforestation and expanding. Importing economies are considering regulating the trade of soybeans and other deforestation-driving commodities, and trading companies will be required to conduct due diligence to ensure compliance. However, complex supply chains obscure provenance, and origin declarations may be falsified. Here, leveraging Gaussian Process modelling and a georeferenced dataset of isotopic and elemental composition of soybeans from across the main soy growing areas of South America, we identify soybean origin to within 192.52 ( ± 23.51) kilometres from the true harvest location. The average 95% Credible Regions reduces prediction uncertainty to within 3.8% of the area considered for prediction. Our spatially explicit model is a leap forward in commodity traceability, enabling both origin determination and verification of origin claims in true geographical space. Applicable to many commodities, this framework provides transparency regardless of supply-chain complexity, and facilitates effective regulation of commodity supply chains to tackle illegal deforestation.
Similar content being viewed by others
Introduction
Despite multi-national, governmental and corporate pledges, global deforestation remains alarmingly high, driven primarily by agricultural expansion1,2. Tropical forests are hardest hit by agricultural encroachment, and have lost 3.7 million hectares (Mha) to agricultural expansion in 2023 alone3. The demand for seven internationally traded “deforestation-driving commodities” (DDCs) has driven the loss of 71.6 Mha of tropical forest (twice the size of Germany) between 2001–2015, of which 63% were cleared for beef pastures, 14.5% for oil palm plantations, 11.5% for soy, and 11% for cocoa, coffee, rubber and wood plantations combined1,2,4,5. Agriculture-driven deforestation disrupts regional and global climate patterns6,7,8, imperils biodiversity9,10,11 and threatens livelihoods12,13,14,15, which undermines international sustainability efforts such as the Paris Agreement, Kunming-Montreal Global Biodiversity Framework and multiple UN Sustainable Development Goals, and others.
Soybean is a cornerstone in global protein production, used mainly as feed for farmed poultry, pork and fish16,17. Soybean production is export-driven, providing crucial protein for consumer countries and dependable income for producers13,18,19,20. Brazil, the world’s largest soybean producer, exported more than 80% of its soybean harvest in 2023/24, generating US$61 billion in revenue—the country’s largest export and 19.3% of its total exports20,21. Soybean farms covered 140 Mha in 2023/24—a 9.6% increase in area in 1 year, the largest consumers being China and the European Union importing 120 million tons (Mt) and 33 Mt, respectively—30% and 8% of the global production16,20. The trade tariffs newly imposed by the US—the second largest soybean exporter—are already increasing demand for South American soybeans, which is not expected to subside22. The ever-increasing demand for soybeans is not met by yield improvement and drives rapid farmland expansion, particularly in the tropics5,23. Despite efforts to reduce soybean-driven deforestation, such as the Amazon Soy Moratorium24 and Cerrado Manifesto25—whereby traders sourcing soy from these regions pledged to eliminate deforestation from their supply chains, forest loss remains unsustainably high23,26,27,28,29,30,31.
To address commodity-driven deforestation, consumer economies are considering introducing restrictions on trading in and importing DDCs, timber, and their products32,33,34. Building on lessons learnt from efforts to regulate the illegal timber trade, the European Union developed Regulation (EU) 1115/2023 (known as the European Union Deforestation Regulation—EUDR), which requires large companies trading in seven DDCs to prove that the products they trade do not originate from areas deforested after 2020 or contribute to forest degradation. To do this, the EUDR requires the companies to carry out due diligence, including collecting and reporting precise geolocation coordinates and time of harvest for DDC provenance34. However, DDC supply chains are notoriously complex, which obscures the commodities’ provenance35, and declarations of origin are liable to fraud and misrepresentation36,37,38,39. The recent US trade tariffs may provide additional financial incentive for origin misdeclaration. Competent Authorities are therefore tasked with conducting compliance checks using “any technical and scientific means adequate to determine” the exact place of production34.
Analytic techniques for origin traceability are based on the correlation between the chemical composition (“fingerprint”) of tissues and the environmental conditions in which the organism grew40,41,42, and may be used for verification of a declaration of origin, if one exists, or for origin determination if not. Recent advances in traceability techniques have achieved impressive prediction accuracy and error reduction, largely due to high-precision measurement of stable isotope ratios (SIR) and trace elements (TE) in the examined material39,43,44,45. Subsequently, Machine Learning algorithms have proven to be instrumental for origin identification efforts based on such high-dimensional data39,43,45. The distributions of the spatially informative stable isotopes (2H, 13C, 15N, 18O, 34S) in plants are governed by atmospheric fluctuations in temperature and precipitation, and vary over large (tens of kilometres) spatial scales40,42, whereas informative trace elements have patchier distributions (ranging from metres to kilometres in scale) that vary with the physical properties of the soil46,47. Thus, combining stable isotope ratios and trace elements enhances the accuracy of origin estimates44, but extracting both is costly, and analysing such complex data requires powerful models.
Nearly all Machine Learning models to date have posed the origin determination problem as a classification task, where the predicted origin is selected from a short list of pre-determined locations, most often countries45,48. Classification studies at sub-national level are increasing38,43,49, but remain constrained by the inherent shortcomings of classification models. First, classification cannot generalise to unfamiliar data: prediction is limited to the classes (=locations) that appear in the training data50, which restricts the spatial scope of the model and precludes intermediate locations. Second, the compositional similarity between two plants decreases with the distance between them40,42,46,47,51, but classification models are oblivious to real-world distances and relative positioning. These make classification models poorly equipped for tackling traceability problems where accurate origin estimates are needed.
Gaussian Process (GP) models estimate the covariance between observations as a function of the geographical distance between them50,52, so that the study area is explicitly considered as contiguous rather than a few pre-selected locations. This enables computation of the probability of observing a given chemical fingerprint at any point in that area, from which the harvest location of the sample can be estimated using Bayes’ theorem. This approach has been applied to origin tracing of wood from long-lived tree species within their natural range39,52, but not to short-lived annual crops cultivated globally, for which traceability still relies on non-spatial classification models38,45,48,49.
Here, we develop a powerful GP regression model and apply it in a Bayesian framework to a set of georeferenced stable isotope ratios and trace elements data, to determine the harvest location of soybeans (Glycine max (L.) Merr.) from across the soy-producing regions of Brazil, Argentina and Bolivia. These countries collectively account for 60% of global soybean exports and >90% of soybean-linked deforestation20,53. South America encompasses most of the global soybean production, in multiple neighbouring countries—a setup that is the most challenging for provenance prediction, and therefore the most appropriate for testing a new model. Classification models based on chemical composition data can successfully distinguish between soybeans from different continents45,48, but plants from nearby origins are much more challenging to tease apart. We show that, despite predicting onto a large study area, this method matches or exceeds the origin determination accuracy of classification models; it outperforms them by featuring spatial quantification of uncertainty, and its predictions can be explained to identify the most informative stable isotope ratios and trace elements for soybean origin traceability. This framework can support companies in upholding environmental commitments and empower national and international authorities to effectively enforce supply chain regulations.
Results
Origin determination models
We measured the stable isotope ratios (SIR) and trace elements (TE) profiles of 267 soybean samples from across its region of cultivation in several South American countries (Fig. 1) and generated three datasets: SIR-only, TE-only, and SIR + TE data. We fit the origin identification model to each dataset using a 5-fold cross-validation (CV) study design. The SIR-only model generally inferred harvest origin (revealed by the allocation of posterior probability) over large areas, whereas the TE-only model generated more concentrated spatial predictions. We quantified the models’ prediction quality using two performance metrics: the distance between true- and predicted harvest location (prediction error hereafter), as an index of accuracy (lower error indicates higher accuracy), and the 95% Credible Region (CR)—the smallest area that includes 95% of the posterior probability—as an index of precision. The combined SIR + TE model surpassed both other models in prediction accuracy and precision (Fig. 2).
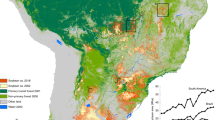
The colour of each 1° x 1° grid cell represents the number of samples collected within its boundaries. The 1° x 1° grid cells are only used in this figure to protect farmers identity; modelling and spatial prediction were carried out on 0.133° x 0.133° grid cells. BACS stands for Best Available Crop-Specific masks (given at a 0.05° resolution). Green shading represents relative abundance of soybean farming. Soybean production map from the USDA20, accessed on 19/01/2026 at: https://ipad.fas.usda.gov/cropexplorer/cropview/commodityView.aspx?cropid=2222000.
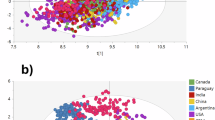
Performance metrics calculated from a model based on SIR data only (pink), a model based on Trace Elements data only (blue), and a model combining SIR and TE data (yellow). Boxplots show median values, quartiles, and outliers of the prediction error index (a) and the area of 95% Credible Region as prediction precision index (b) across all samples. Diamonds denote median values of individual cross-validation folds. Outliers (green dots) are values greater than 2*IQR. Distribution curves corresponding to the boxplots are colour-coded accordingly and illustrate the density of the indices. Insets corresponding to the dashed boxes show the distribution of median values of the cross-validation folds. The distributions of medians from the SIR + TE model are significantly different from the SIR-only or TE-only models (two-sided Kolmogorov-Smirnov test; p < 0.008). Lower median values of both performance indices across all cross-validation folds demonstrate the superior predictive ability of the SIR + TE model.
The cross-validated (mean) prediction error was 301.24 (sd: ±87.61) km based on the SIR-only model, 316.82 (±105.52) km for the TE-only model, and 192.52 (±23.51) km for the SIR + TE model. The prediction precision was 6.9 (±1.3) * 106 km2 in the SIR-only model, 4.5 (±1.7) * 106 km2 in the TE-only model, and 0.44 ( ± 0.28) * 106 km2 in the SIR + TE model. In all three models, prediction accuracy and precision varied widely across samples, but the models differed in how that variation was distributed. The prediction error medians (one per CV fold) of the SIR + TE model differed significantly from the medians of either the SIR-only or TE-only model (two-sided Kolmogorov-Smirnov test, p < 0.008), which were statistically indistinguishable from each other (p = 0.873). A similar pattern was observed in the extent of the 95% CR, where the medians of the SIR + TE model were significantly smaller than those of the SIR-only or TE-only models (two-sided KS test; p < 0.008 in both cases). Across all CV folds, the performance metrics of the SIR + TE model were always the lowest of the three models, demonstrating its consistently superior predictive ability.
We present six prediction maps from the SIR + TE model to illustrate the range of prediction results in a spatial context (Fig. 3). These maps show the posterior probability distribution for samples that achieved the highest-, median-, and lowest score on each of the performance indices. The similarity between the median prediction and the highest scoring prediction on either index scale, and the dissimilarity of the respective lowest scoring prediction, illustrate the pronounced skew of the SIR + TE prediction quality distributions towards high accuracy and high precision (Fig. 2).
Maps show the most- (a, d), median- (b, e), and least accurate (c, f) predictions of the SIR + TE origin determination model, based on the prediction error metric (a–c), and on the size of the 95% Credible Region (d–f). True harvest location is marked by a green star, and the location of the posterior mode is marked by a red “+”. Regions where expected soybean yield (FAO and IIASA, 2021), as a proxy for suitability for commercial soybean farming, is lower than 2000 kg dry weight per hectare were excluded from the prior (shown in white). For both indices, the median values (b, e) are closer to the most accurate prediction (a, d) than to the least accurate prediction (c, f), demonstrating the model’s high predictive power for most soybean samples.
Quasi-classification for comparison with other studies
To enable comparison between the GP model here and classification models in other studies, we assigned each prediction to the country indicated by the coordinates of the posterior mode (“Quasi-classification tests” section; Supplementary File 1). For each data type (SIR, TE, SIR + TE) we used a “One-vs.-Many” framework to assess the classification results and calculated the accuracy, precision, sensitivity, specificity and F1 scores (Table 1). The TE and the SIR + TE models achieved the same overall prediction accuracy of 0.884, whereas that of the SIR model was 0.798. Country-specific classification accuracy for Argentina was highest with the TE model (0.944), whereas for Bolivia and Brazil it was highest with the SIR + TE model (0.951 and 0.929, respectively). The TE model scored highest for precision, sensitivity, and F1 scores for Argentina, and for precision, specificity, and F1 scores for Brazil. For Bolivia, the SIR + TE model achieved the highest precision, specificity, and F1 scores, whereas for sensitivity the highest score came from the SIR model—the only instance of the SIR model outperforming the other two (Table 1).
We mapped the quasi-classification results onto geographical space to assess whether there are specific regions associated with erroneous prediction. Expectedly, all models show higher rates of erroneous predictions near political borders (Fig. 4a–c), where a small error could place a prediction across the border from its country of origin. The error rates of the SIR + TE model were lower than those of the SIR-only or TE-only models, and more concentrated near the Brazil-Argentina border, where the borders of Paraguay and Uruguay also make misclassification more likely. For further considerations on quasi-classification see Supplementary File 1.
Predictions are based on SIR data (a, d), TE data (b, e), or SIR + TE data (c, f). The origin prediction was considered correct if the predicted harvest coordinates pointed to the country from which the sample was taken. Grid cell colour in a–c indicates the proportion of false predictions out of all samples harvested in that pixel. Grid cell colour in d–f indicates the proportion of false predictions out of all samples predicted to that pixel. We pooled data into 1° x 1° grid cells to protect farmer identity. All models show higher rates of incorrect country assignment near borders, but this effect is less widespread in the SIR + TE model.
Predictor contribution and model explainability
We performed five-fold cross-validated SHAP analysis to assess the unique contribution of each predictor to the origin prediction of each sample (Fig. 5). The three most important predictors of both longitude and latitude are δ2H in lipids, 60Ni and 137Ba, although the order of these varied between longitude and latitude. The next three most important predictors were 27Al, 59Co and δ34S (in varying order as well). The least informative predictors for both longitude and latitude are 55Mn and 85Rb, and 88Sr is third- and fourth least informative for latitude and longitude, respectively. The ranking of the remaining predictors was less consistent between longitude and latitude.
Explanatory values for longitude (left) and latitude (right) prediction are ordered by the magnitude of impact on model output (largest at the top). Dots denote the cross-validated SHAP value for the prediction of the origin of each sample. Colours represent the value of the feature (predictor variable) in each sample, and SHAP values indicate the effect of the feature’s value on the resulting prediction of harvest origin.
Discussion
This study is, to our knowledge, the first to provide a probabilistic estimation of the harvest origin of an annual crop. We applied a continent-wide model based on georeferenced isotopic and elemental datasets of soybeans from across the main soy growing areas, representing >94% of South American soy production. Our analytic pipeline, using georeferenced stable isotope ratios and elemental composition measurements with a powerful geostatistical model, provides spatially explicit origin predictions with probabilistic indicators of uncertainty. It enables recent and upcoming policies to advance DDC traceability and supply-chain transparency and will support verifying commodity provenance as supply chains undergo increased scrutiny to enforce trade tariffs.
The spatial accuracy- and precision- of prediction that we present are not comparable with previous efforts as no other soybean origin tracing approach considers contiguous space54 and classification models cannot produce spatial predictions. The prediction accuracy and precision here are similar to recent findings for European tree species, where an SIR + TE model reduced prediction error from ~300 km to ~200 km compared to SIR- or TE-based models39. Using a comparable dataset (267 soybeans vs. 174–302 tree samples), our predictions are similarly accurate, indicating the model’s robustness and the approach’s economic efficiency as a longer-term policy tool.
Explainability of Machine Learning models facilitates greater public trust, which is paramount given the magnitude of economic and societal implications of regulating DDC trade55,56. Using SHAP analysis we identified the most informative SIR and TE predictors in the data, towards establishing a minimal set of predictors for accurate soybean origin determination. We verified the selection of predictors through models with various permutations of the set of predictors, ranging from all informative predictors (those with >5 different values across all samples) to models with fewer predictors, but the selected 22 predictors that we report here performed best in terms of prediction accuracy and precision. Limiting subsequent data generation to that minimal set will reduce the per-sample cost of analysis, testing time and technical skills required, and facilitate rapid testing with minimal disruption to trade.
Benchmarking against other approaches
To facilitate comparison of model performance with previous studies, we carried out quasi-classification experiments to obtain True/False prediction results. This ad hoc addition is meant only for contextualising our study. Traceability has so far focused on identifying country of origin because most regulations were satisfied with country-level declarations of origin. We, however, aim to identify harvest location independent of country boundaries, in line with EUDR. We have included here a quasi-classification experiment, to compare the performance of this method against previous (classification-based) studies on agricultural commodities, which do not provide spatially indicative information.
Soybean harvest origin is currently estimated using classification models, which often consider sets of non-adjacent countries—for example, Brazil-Canada-USA-Vietnam48 or Argentina-Brazil-China-USA45—arguably detecting continent of origin rather than country. Plants’ chemical composition varies widely between distant locations40,42,51, inflating classification accuracy in long-range studies. Attempting classification over small distances, particularly across arbitrary land borders, yields much higher misclassification rates49,57, highlighting the trade-off between spatial generalisability and prediction accuracy. Our spatially explicit model overcomes this trade-off, as demonstrated by achieving quasi-classification accuracy and precision scores (both >88%) that are comparable to classification-based soybean traceability studies, while additionally providing an intuitive probabilistic estimate of prediction uncertainty. Unlike classification models, our model is spatially generalisable irrespective of arbitrary borders, as demonstrated by predictions in Paraguay and Uruguay, neither of which are represented in the dataset. Border-related prediction error is evident in all our models (Fig. 4), whether it is a sample’s true origin that is near a border (error of type “false negative”) or the predicted origin (“false positive”). These errors mostly overlap in the SIR + TE model, meaning that samples harvested near a border are also predicted to come from near that border, although not always on the correct side of it. The difficulty to correctly predict the origin of samples near borders underscores the need for a shift to spatially explicit models.
The quasi-classification results support previous findings that soybean elemental profiles are more indicative of country of origin than stable isotope ratios54. Expectedly, however, the quasi-classification prediction accuracy dropped near borders, illustrating a fundamental shortcoming of classification-based traceability. As elemental availability depends on soil properties rather than atmospheric factors, it may be more stable year-on-year, making trace elements particularly useful for predicting harvest origin of annual crops, although intra-annual variation has been found in beech trees58. We hope that future studies examine seasonal and annual variability in isotopic and elemental content of soybeans and other crops in detail, to provide useful information for fine-tuning traceability models. We expect that even with heavy inputs from farming practices, trace element levels should not fluctuate radically as the farmer aims for the same optimum soil conditions year on year. As different varieties may differentially take up soil elements due to divergent root traits, for example, a more industrialised crop would likely have a smaller pool of agrobiodiversity which may make it more difficult rather than easier to trace. Although genetic uniformity of industrial crops may reduce among-individual variation in soybean chemical composition, annual environmental variations may still necessitate adjusting for year of harvest in origin determination efforts.
The analytic pipeline here relies on initially creating a reference dataset that powers the model’s predictions. When generating predictions in regions that are very far from reference data, the model may require additional data such as environmental predictors to support its predictions. For example, as heavy Hydrogen and Oxygen isotopes mostly emanate from the oceans, the ratio of these isotopes in rainwater decreases with elevation and distance from the coast42,51, governed by precipitation patterns, soil water retention and so on. Such auxiliary information may help predicting stable isotope ratios far from training data points, but has its own uncertainty which is propagated into the soybean origin prediction. We constructed extended models, including climate and soil variables as predictors of the isotopic and elemental distributions used for the origin determination model. For each stable isotope and elemental predictor included in the model we specified an independent GP model with up to three environmental variables that are hypothesised (based on literature) to drive its spatial distribution.
We constructed one SIR model with climatic predictors and one SIR + TE model with climatic and soil predictors (Supplementary File 1, Sections 3 and 4). Although more complex, the extended models demonstrated lower prediction accuracy and precision compared to the SIR + TE model (Supplementary File 1, Section 3). This is similar to the findings of a recent study on oak trees from the USA, where a model including climate variables was outperformed by a model based on stable isotope ratios alone52. It may be due to several factors, such as mis-specified statistical links between environmental predictors and isotopic/elemental content, as detailed literature for soybean is lacking. Another possibility is that high variability in the environmental data caused high uncertainty in the modelled relationships, resulting in less stable predictions. Nonetheless, we believe that environmental data could be useful for traceability, but further research is needed for identifying how to best incorporate it into the modelling framework.
Deforestation regulations enforcement
This study provides, to the best of our knowledge, the only tool to date for accurate identification of crop harvest origin at large scales. Tools such as this are instrumental in enabling the effective enforcement of regulations designed to improve supply chain transparency and traceability, and in supporting operators preparing for the introduction of precise provenance requirements such as in the EUDR.
While our GP model predicts provenance with high accuracy, precision, and transparency, it has limitations. First, nearly half of the model’s predictions are >200 km from the true harvest location. In some regions, the distinction between recently deforested and previously cleared blocks of land requires higher prediction accuracy than is currently achieved by the model23,27. Increasing prediction accuracy near deforestation fronts—where legally- and illegally-cleared plots often intersperse—may require additional data inputs, such as from remote sensing, or denser sample collection. However, sampling near deforestation fronts is logistically and financially challenging, farmers in such areas are often unwilling to donate samples, and collectors may face hostility. Although high precision traceability is likely to be more effective when carried out closer to the harvest origin, random sampling of shipments and imports (provided that rigorous sampling is undertaken) may be sufficient to identify soybeans from areas flagged for illegal deforestation. The deforestation legislation therefore aims to employ multiple tools to verify the origin of commodities rather than relying on a single source of information. Second, this model can indicate the harvest origin of a sample by direct prediction (origin determination) or by discounting unlikely locations (origin verification). Accuracy could also be improved by incorporating into the priors high-resolution remote-sensing data on land-use59 and forest clearing (Global Forest Watch: https://www.globalforestwatch.org/map/), to delimit the range of plausible harvest locations. Additionally, local sources of information may help, as demonstrated by the ‘Soy Working Group’ underpinning Brazil’s Amazon Soy Moratorium, who keeps a register of soy-growing farms, and combines it with satellite imagery, field visits and production records to assess violations of the moratorium27. Each additional source of information will further increase the overall prediction quality of the model. The technology we describe here is not meant as a “silver bullet” but is a powerful technique to be added to the suite of tools available to law enforcement authorities and other stakeholders.
There are, however, obstacles on the way to adopting this technology, such as funding and logistics of the testing operations or managing commodity admixture in shipping, and adaptations will need to take place along the supply chain. Trace elements analysis currently costs 50–120 USD per sample, for stable isotope ratios it is 200–500 USD, depending on how many samples are analysed, how many variables are measured, and which variables those are. Strontium isotopes ratio analysis (87Sr/86Sr; not performed in this study) costs about 250 USD per sample as well, and may be particularly useful for traceability47, unlike Sr content per se. Although each sample is costly to obtain and analyse, the model we present can minimise the size of the reference dataset by predicting beyond the data points, which effectively replaces physical samples with statistical inference.
Naturally, high precision traceability is likely more effective when testing closer to the harvest origin, but sampling at local or regional aggregation facilities is logistically complex. However, random sampling of shipments and imports (provided that rigorous sampling is undertaken) may be sufficient to identify soybeans from areas flagged for illegal deforestation. Bad actors may try to blend commodities from legal and illegal sources, but random sampling may force them to keep the blending to a very low ratio to remain undetected. Additionally, the EUDR operates on an individual “product” level not a shipment/container level, so an origin claim for a container must list all harvest locations that contributed to its content. Similar legislation for the timber trade (European Union Timber Regulation; EUTR) achieved broad uptake by the timber industry of chemical testing for origin traceability. Ultimately, achieving traceability in currently-opaque supply chains will require operational modifications (hence the industry’s strong pushback against EUDR), but regardless of where along the chain the testing is done—all that our method requires is the bean itself and the origin claim. The analysis of whole beans in our study ensures a scalable approach from the start to the end of the supply chain.
The utility and consequences of DDC origin determination tools remain uncertain as the legal frameworks are not yet implemented and the issues they address change rapidly. The EUDR currently refers to deforestation where canopy cover exceeds 10%, excluding other wooded at-risk ecosystems like the Chaco and the Cerrado biomes. The Cerrado—a mega-biodiverse savanna with varying levels of tree cover—has been undergoing devastating land clearing in the two decades since the Amazon Soy Moratorium was introduced60. The moratorium placed effective protection against deforestation in the Brazilian Amazon, which inadvertently diverted deforestation pressure for the expanding cattle and soy industries onto the Cerrado24,28. Since 2017, the Cerrado Manifesto aims to protect the Cerrado against further deforestation25, yet Cerrado deforestation rates continued to increase until 202426,61. The Chaco, a semi-arid forest spanning Paraguay, Bolivia and Argentina, has too undergone accelerated deforestation in recent decades31,62, and lacks effective protection. Trade agreements and regulations must keep up with rapid landscape modification, which highlights the urgent need to include all impacted biomes in anti-deforestation legislation10,24,60,63.
The potential problem of losing predictive power as production expands to new frontiers is mitigated by additional scrutiny of commodities from uncertain origin as they carry higher deforestation risk by definition, which the legislation is designed to disincentivise. Unlike previous methods, the method we describe here expresses prediction uncertainty in geographically meaningful units (area, distance, etc.), facilitating better assessment of the deforestation risk in a tested product. Updates to the reference dataset—ensuring it remains comprehensive—will be required irrespective of which traceability method, model (classification vs. predictive), or conceptual framework is used (origin verification vs. determination), so this expense is not unique to our method. Future changes to the spatial scope of origin prediction can be readily accommodated by our model, and its Bayesian framework facilitates integrating additional data sources into the prediction.
To conclude, the modelling framework we present here is a powerful tool for harvest origin determination and verification. It revolutionises commodity traceability by circumventing arbitrary classification decisions, facilitates prediction accuracy and precision through considering real-world geography and represents a leap forward in DDC supply chain transparency, supporting deforestation regulations.
Methods and materials
Study area
In this study we consider mainland South America, between latitudes 13°N and 41°S, and longitudes 34°W and 82°W. The study area measures 6000 km north to south by approximately 5343–4033 km east to west (depending on latitude). This area includes all soy growing regions in South America, which account for more than half of global soybean production20. We divide the study area to 406 × 361 pixels, retaining its latitude:longitude ratio so that each pixel is 14.84 km × 14.84 km ≈ 220 km2 at the equator, tapering to 14.84 km × 11.2 km ≈ 166 km2 at the southern edge of the study area. The number of pixels is the result of optimising the models’ spatial resolution and available computing resources. The original latitude:longitude proportions of the study area were retained to maintain the integrity of distance and area measurements. By defining this broad spatial scope, we aim to reduce the amount of unintended information passed to the model by the user, emulating real-world origin identification problems. Defining a small study area necessarily directs origin predictions to that area, i.e., the user informs the model about the origin of the sample, which is unlikely to happen in real-world use cases. The low information content of the priors better simulates real-world use cases to test the geospatial prediction capabilities of this technique. Once reference data becomes available from other continents, those regions could be simply added to the geographic scope of the model.
Sample collection
Soybean seed samples were collected by in-country teams in Argentina, Bolivia, and Brazil during the harvest months (Bolivia: February 2022; Brazil: February–May 2022; Argentina: May–June 2022). These countries were selected because they are the top soy producers—and exporters—in the continent, accounting for >94% of the soybeans grown in South America, and form a contiguous landmass to enable rigorous testing of our model. As our study is centred on identifying deforestation embedded in exported soybeans, our sampling strategy aimed to maximise both spatial coverage and soybean production levels, as well as proximity to deforestation fronts, as safely feasible. This was achieved by prioritising sampling locations in areas of higher soybean production (equal to or greater than 2000 kg per hectare, dry weight) across maximal longitude and latitude within individual countries. This also ensured sampling points close to political borders and natural forests. Because of budget limitations and minimal supply chain and export power of the lowest production areas (those producing less than 2000 kg per hectare, dry weight), these were excluded from the sampling strategy. We did not pre-allocate the number of samples or sampling locations per country because of the great discrepancy in country size, extent of soybean-producing regions, and quantities produced. The collectors then identified farms and contacted farmers as close as possible to the desired sampling locations; if a farmer chose not to be included in the study, the collectors contacted the next nearest farm, and so on. Sampling locations were selected as a result of optimising these and other factors, including keeping within budget for both field expeditions and downstream sample analyses, avoiding areas that suffered damage to crops (e.g. through disease and droughts), and designing a sampling route via roads that coincided with the harvest period in the various regions so that seeds were collected at the correct developmental stage (dried seed-filled pods). Wherever possible, soybeans were collected dry so that post-harvest drying was minimal. Dry collection mimics agronomic pod harvesting in the field, making our methodology effective for testing whole beans in the supply chain.
Location-level sampling strategy was as follows: Each sampling location was formed of five sampling points, arranged (approximately) at the centre and corners of an imaginary square with a 10 km diagonal. One sample was taken at each sampling point, made of soybeans from 2–5 plants to meet a minimum total mass criterion. This design is optimising for several objectives: capturing natural variation (farther is better), travel time and accessibility to collectors (nearer is better), reducing the number of farms sampled (nearer is better), and keeping to sample quotas (farther is better). See collection protocols at: https://learn.worldforestid.org/. Due to the multi-dimensionality of the Gaussian Process models, estimating the optimal size of the reference dataset and the ideal locations for sampling would require its own study. We expect that the minimal number of samples required per location is variable and context-dependent (e.g., on species, terrain, farming practices and history, etc.), meaning that different samples may carry different importance to the model’s predictions. To assess the model’s robustness to systematic bias in the data, we carried out experiments where we omitted data based on arbitrary latitude and longitude values to compare the predictions from partial data to those based on the full dataset (see Supplementary File 1, Section 2).
To clarify, the purpose of this study is to develop a model that can identify the origin of soybeans wherever they may be from in South America. As our reference dataset includes hundreds of samples taken from a polygon that spans thousands of kilometres in all directions, the influence of idiosyncratic reference data is strongly reduced.
Stable isotope ratio (SIR) and trace elements (TE) analyses
Isotope Ratio Mass Spectrometry (IRMS) was used for Stable Isotope Ratio Analysis of 267 soybean samples, to quantify the relative abundance of stable isotopes of Oxygen (δ18O), Hydrogen (δ2H), Carbon (δ13C), Nitrogen (δ15N), and Sulphur (δ34S). These isotopes vary naturally in the environment due to atmospheric and climatic processes and are incorporated into plant tissues through a range of metabolic pathways40,42. To account for elemental sequestration through alternative biochemical pathways, δ18O, δ2H, and δ13C were measured in both the lipid- and the non-lipid phase of the sample material. Nitrogen and Sulphur were measured in the non-lipid phase only (Supplementary Fig. S1). Further details on the IRMS protocol are in Supplementary File 1.
For Trace Elements Analysis, Inductively Coupled Plasma Mass-Spectrometry (ICP-MS) with a collision cell was used, to quantify the presence of 69 chemical elements in each of the 267 soybean samples (Supplementary Fig. S2). In 19 of the 69 elements (45Sc, 52Cr, 72Ge, 93Nb, 101Ru, 103Rh, 107Ag, 118Sn, 121Sb, 125Te, 178Hf, 181Ta, 189Os, 193Ir, 195Pt, 197Au, 201Hg, 209Bi, and 238U), we found very little variation—fewer than 10 different values across all samples. We do not consider these 19 elements in subsequent analyses. We log-transformed the TE data before further analysis because the raw measurements span eight orders of magnitude across elements, which may hinder spatial signal detection by downstream models. For details on the ICP-MS protocol, see Supplementary File 2. All analytic data is at https://zenodo.org/records/18786976.
We used three criteria to select appropriate elements for spatial analysis: higher numbers of unique measured values, a large range of measured values, and higher median values were all preferred. Unique measurement values make the samples distinguishable, a large range of measured values makes the spatial variation among samples distinguishable, and a high median value facilitates greater robustness to error as the measurements move away from detection limits. The levels of 24Mg, 31P, 39K, 44Ca, 56Fe, and 66Zn were very narrowly distributed across samples. The small range of values decreases the signal:noise ratio, so we excluded these elements from the models. Consequently, 14 elements were used in the geostatistical analyses.
Geostatistical models
The origin estimation algorithm uses Gaussian Process (GP) regression models to estimate the expected value of each predictor (isotope ratio or trace element) at every pixel in the study area. The likelihood function derived from these values is then used with the prior in a Bayesian framework to compute for every pixel the probability that it overlaps with the harvest location of that sample39. To quantify and compare the predictive ability between models, we assessed used two complimentary metrics: prediction error, measured as the distance between the posterior mode (point of highest posterior probability) and the true harvest location; and prediction precision, quantified as the 95% Credible Region, which is the land area of the 95% highest posterior density (HPD) region.
Due to the localised nature of farming and sample availability in the field, some reference samples form spatial clusters, whereas others are further apart. Elements and stable isotopes are also distributed by natural process as well as agronomic management practices, such as crop or soil treatments, so high spatial autocorrelation is expected among samples from nearby localities. If adjacent samples are split between the training and test datasets, the high spatial autocorrelation between them would cause “contamination” of the training data with information about the test data, known as data leakage64. Data leakage falsely improves model performance metrics, thereby inflating confidence in the model—which would not hold up in real-world tasks. To avoid data leakage due to spatial autocorrelation, we assigned the samples into spatial clusters, where all samples that are <30 km apart are assigned to the same cluster. We explored cluster cut-offs ranging from 15–180 km, but the value of 30 km resulted in most clusters having exactly 5 samples, adequately reflecting the sampling design. This ensured that no cluster was represented in both the training and testing datasets simultaneously, which would have caused data leakage and inflated the model’s predictive ability. Too large a cut-off would have reduced the model’s predictive power by grouping multiple spatial clusters into one, thereby masking their respective characteristic values and the spatial signal that the model relies on. We then split the data into training and test sets by cluster, which ensured that adjacent samples always fall in the same set. Most of the resulting clusters consist of five samples, but some clusters contain fewer samples.
We trained three separate GP models to compare their relative performance: the SIR model, TE model and SIR + TE model. For training the SIR model we included δ2H, δ13C, and δ18O data (each measured on the lipid and non-lipid fractions separately), and δ15N and δ34S measured on the non-lipid fraction only. For training the TE model we used the following 14 elements: 27Al, 28Si, 55Mn, 59Co, 60Ni, 63Cu, 78Se, 85Rb, 88Sr, 89Y, 95Mo, 111Cd, 133Cs, 137Ba. The SIR + TE model was trained on all 8 SIR and 14 TE included in the other two sets.
We used a map of average attainable soybean yields65 to delimit origin prediction to regions where expected soybean yield is at least 2000 kg per hectare (dry weight). This threshold is below the lowest estimates of soybean yields currently produced in any of the main soy-producing biomes in Brazil66. We used this threshold provisionally to support our spatial analysis (Supplementary Fig. S3), because it suggests a greater range suitable for soybean farming than estimated previously24. A greater area of suitability makes the prior distribution less informative because the prior probability is divided over more pixels. A less informative prior, in turn, reduces the impact of researcher assumptions on the model’s output.
We used five-fold cross-validation, so that each of the five iterations of the model was trained on 213–215 (~80%) of the reference data, and the remaining samples were used as the test set. The variation in size of the training and test sets was due to the different number of samples in some spatial clusters. Each model was trained until it reached stationarity of the loss function, which we established by observing no decrease in loss value over at least 2000 consecutive training iterations.
We used two indices to assess the accuracy and precision of the estimates of origin generated by the different GP models:
-
(1)
“Prediction error”, as an index of prediction accuracy, is the great circle distance between the point of highest posterior probability (the posterior mode) and the true origin of the sample in question. Smaller prediction error indicates a more accurate prediction of the true origin.
-
(2)
“95% Credible Region (CR)”, indicating prediction precision, is the combined area of the smallest number of pixels that together contain 95% of the posterior probability. Hence, the CR contains the true origin of a given sample with 95% certainty. A smaller CR indicates higher prediction precision.
-
(3)
These two indices quantify complementary aspects of the deviation of the predicted origin from the true origin of a sample. We used the median values (rather than means, due to strong skew in the index distributions) of the five cross-validation folds to obtain a cross-validated index of model performance for each of SIR, TE, or SIR + TE models.
Quasi-classification tests
To facilitate comparison of the performance of our Gaussian Process model with standard classification models (whether for origin determination or verification), we converted the maximum posterior probability predictions (i.e., coordinates) into the country names where those coordinates fall in. We then summarised the categorised results in confusion matrices (one per model per country) and used a one-vs.-rest approach to calculate the following performance metrics:
where TP stands for True Positive, i.e. correct prediction of the focal country for a given confusion matrix, FN is False Negative, and so on. The balanced accuracy metric addresses biases in the accuracy metric due to differences in the number of samples from each country in the data.
SHAP analysis
We performed SHAP analysis67 to identify the most informative predictors for soybean origin determination. SHAP approximates the determination model to calculate the marginal contribution of each predictor (SIR or TE) to each prediction made, thereby indicating the key variables that drive the origin predictions. The SHAP analysis followed the same five-fold cross-validation framework described above, with the same data partitions and respective GP models as input. We calculated SHAP values for every point in the training sets and averaged over the five folds to obtain the cross-validated SHAP values.
Data processing and visualisation were done in R v4.3.068, using the packages tidyverse v2.0.069, terra v1.7.7170, sf v1.0.1571,72, raster v3.6.2673, ggspatial v1.1.974, and fields v15.275. The GP regression modelling is implemented in Python v3.10.12, using Pytorch v1.12.176, Gpytorch v1.9.077, Shap v0.44.067, Scikit-learn v1.3.078, Geopandas v0.11.179.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Due to the sensitive nature of the data, the SIRA and TEA data are available upon request and provided without geolocation information at https://zenodo.org/records/18786976. Deforestation, soybean traceability and the EUDR are highly contentious subjects in parts of South America, and some farmers have explicitly conditioned their participation on remaining anonymous. To avoid compromising collector and donor security, we do not provide coordinates or display exact farm locations in the figures.
Materials availability
The soybean seed samples are part of the World Forest ID Georeferenced Sample Collection held at the Royal Botanic Gardens, Kew. Requests for materials should be addressed to C.C.: c.chater@kew.org.
Code availability
The computer code developed for this study is available upon request at https://zenodo.org/records/18786976.
References
Curtis, P. G., Slay, C. M., Harris, N. L., Tyukavina, A. & Hansen, M. C. Classifying drivers of global forest loss. Science 361, 1108–1111 (2018).
FAO and UNEP. The State of the World’s Forests 2020. Forests, Biodiversity and People (FAO and UNEP, 2020).
Weisse, M., Goldman, E. & Carter, S. Tropical forest loss drops steeply in Brazil and Colombia, but high rates persist overall. https://research.wri.org/gfr/latest-analysis-deforestation-trends (2024).
Goldman, E., Weisse, M., Harris, N. & Schneider, M. Estimating the role of seven commodities in agriculture-linked deforestation: oil palm, soy, cattle, wood fiber, cocoa, coffee, and rubber. https://www.wri.org/research/estimating-role-seven-commodities-agriculture-linked-deforestation-oil-palm-soy-cattle (2020).
Pendrill, F. et al. Disentangling the numbers behind agriculture-driven tropical deforestation. Science 377, eabm9267 (2022).
Bochow, N. & Boers, N. The South American monsoon approaches a critical transition in response to deforestation. Sci. Adv. 9, eadd9973 (2023).
Gatti, L. V. et al. Amazonia as a carbon source linked to deforestation and climate change. Nature 595, 388–393 (2021).
Smith, C., Baker, J. C. A. & Spracklen, D. V. Tropical deforestation causes large reductions in observed precipitation. Nature 615, 270–275 (2023).
Gibson, L. et al. Primary forests are irreplaceable for sustaining tropical biodiversity. Nature 478, 378–381 (2011).
Hoang, N. T. et al. Mapping potential conflicts between global agriculture and terrestrial conservation. Proc. Natl. Acad. Sci. USA 120, e2208376120 (2023).
Moreira, H. et al. Threats of land use to the global diversity of vascular plants. Divers. Distrib. 29, 688–697 (2023).
Albert, J. S. et al. Human impacts outpace natural processes in the Amazon. Science 379, eabo5003 (2023).
Correia, J. E. Soy states: resource politics, violent environments and soybean territorialization in Paraguay. J. Peasant Stud. 46, 316–336 (2019).
Leite-Filho, A. T., Soares-Filho, B. S., Davis, J. L., Abrahão, G. M. & Börner, J. Deforestation reduces rainfall and agricultural revenues in the Brazilian Amazon. Nat. Commun. 12, 2591 (2021).
McKay, B. & Colque, G. Bolivia’s soy complex: the development of ‘productive exclusion’. J. Peasant Stud. 43, 583–610 (2016).
De Maria, M. et al. Global soybean trade—the geopolitics of a bean. https://tradehub.earth/reports/; https://doi.org/10.34892/7YN1-K494 (2020).
Ritchie, H. Drivers of deforestation. Our World in Data. https://ourworldindata.org/drivers-of-deforestation (2024).
Giraudo, M. E. & Grugel, J. Imaginaries of soy and the costs of commodity-led development: reflections from Argentina. Dev. Change 53, 796–826 (2022).
Lende, S. G. & Velázquez, G. Soybean agribusiness in Argentina (1990–2015): socio- economic, territorial, environmental, and political implications. in Agricultural Value Chain (ed. Egilmez, G.) (InTech, 2018).
USDA. Soybean Explorer (2024).
MDIC. Comex Stat (2024).
Antonelli, A., Strassburg, B. & Balmford, A. Trade tariffs could worsen deforestation in South America. Nature 640, 881 (2025).
Song, X.-P. et al. Massive soybean expansion in South America since 2000 and implications for conservation. Nat. Sustain. 4, 784–792 (2021).
Gibbs, H. K. et al. Brazil’s Soy Moratorium. Science 347, 377–378 (2015).
The Cerrado Manifesto. https://www.fairr.org/investor-statements/cerrado (2017).
Federal Government of Brazil. Federal Government announces Amazon, Cerrado deforestation drop; concludes prevention pact. (2024).
Heilmayr, R., Rausch, L. L., Munger, J. & Gibbs, H. K. Brazil’s Amazon Soy Moratorium reduced deforestation. Nat. Food 1, 801–810 (2020).
Rausch, L. L. et al. Soy expansion in Brazil’s Cerrado. Conserv. Lett. 12, e12671 (2019).
Richards, P. D., Walker, R. T. & Arima, E. Y. Spatially complex land change: the indirect effect of Brazil’s agricultural sector on land use in Amazonia. Glob. Environ. Change 29, 1–9 (2014).
Silva Junior, C. H. L. et al. The Brazilian Amazon deforestation rate in 2020 is the greatest of the decade. Nat. Ecol. Evol. 5, 144–145 (2020).
Fehlenberg, V. et al. The role of soybean production as an underlying driver of deforestation in the South American Chaco. Glob. Environ. Change 45, 24–34 (2017).
US Department of State. A demand-side policy framework to combat commodity-driven illegal deforestation. (2024).
UKTR report: 2022 to 2025. The Department for Environment, Food & Rural Affairs, UK Government. https://www.gov.uk/government/publications/timber-regulations-reports/uktr-report-2022-to-2025 (2026).
Regulation (EU) No. 1115/2023 of the European Parliament and of the Council of 31 May 2023 on the making available on the Union market and the export from the Union of certain commodities and products associated with deforestation and forest degradation and repealing Regulation (EU) No 995/2010. https://doi.org/10.5040/9781782258674 (2023).
Lyons-White, J. & Knight, A. T. Palm oil supply chain complexity impedes implementation of corporate no-deforestation commitments. Glob. Environ. Change 50, 303–313 (2018).
Adombila, M. A. Ghana lost 160,000 tons of cocoa to smuggling in 2023/24 season, Cocobod official says. https://www.reuters.com/world/africa/ghana-lost-160000-tons-cocoa-smuggling-202324-season-cocobod-official-says-2024-09-16/ (2024).
Cerniauskas, S., Ratmirova, O., Viaznikoutsava, K., Yarashevich, A. & Dauksza, J. Traders are sneaking banned Russian and Belarusian wood into the EU by pretending it’s from Central Asia. (2022).
Cui, D., Liu, Y., Yu, H., Wang, Z. & Mao, X. Geographical traceability of soybean based on elemental fingerprinting and multivariate analysis. J. Consum. Prot. Food Saf. 16, 323–331 (2021).
Mortier, T. et al. A framework for tracing timber following the Ukraine invasion. Nat. Plants https://doi.org/10.1038/s41477-024-01648-5 (2024).
Siegwolf, R. T. W., Brooks, J. R., Roden, J. & Saurer, M. Stable Isotopes in Tree Rings: Inferring Physiological, Climatic and Environmental Responses, Vol. 8 (Springer International Publishing, 2022).
Werner, C. et al. Progress and challenges in using stable isotopes to trace plant carbon and water relations across scales. Biogeosciences 9, 3083–3111 (2012).
West, J. B., Bowen, G. J., Dawson, T. E. & Tu, K. P. Isoscapes: Understanding Movement, Pattern, and Process on Earth through Isotope Mapping (Springer, 2010).
Boeschoten, L. E. et al. A new method for the timber tracing toolbox: applying multi-element analysis to determine wood origin. Environ. Res. Lett. 18, 054001 (2023).
Hong, Y. et al. Data fusion and multivariate analysis for food authenticity analysis. Nat. Commun. 14, 3309 (2023).
Zhou, X. et al. Towards verifying the imported soybeans of China using stable isotope and elemental analysis coupled with chemometrics. Foods 12, 4227 (2023).
Boeschoten, L. E. et al. Clay and soil organic matter drive wood multi-elemental composition of a tropical tree species: implications for timber tracing. Sci. Total Environ. 849, 157877 (2022).
Silva, C. et al. Spatial distribution of strontium and neodymium isotopes in South America: a summary for provenance research. Environ. Earth Sci. 82, 348 (2023).
Nguyen-Quang, T., Bui-Quang, M. & Truong-Ngoc, M. Rapid identification of geographical origin of commercial soybean marketed in Vietnam by ICP-MS. J. Anal. Methods Chem. 2021, 1–9 (2021).
Hidalgo, M. J., Fechner, D. C., Ballabio, D., Marchevsky, E. J. & Pellerano, R. G. Traceability of soybeans produced in Argentina based on their trace element profiles. J. Chemom. 34, e3252 (2020).
Rasmussen, C. E. & Williams, C. K. I. Gaussian Processes for Machine Learning (MIT Press, 2006).
Pederzani, S. & Britton, K. Oxygen isotopes in bioarchaeology: principles and applications, challenges and opportunities. Earth Sci. Rev. 188, 77–107 (2019).
Truszkowski, J. et al. A probabilistic approach to estimating timber harvest location. Ecol. Appl. 35, e3077 (2025).
Goldman, E. & Weisse, M. Deforestation Linked to Agriculture Indicator (World Resources Institute, 2024).
Soni, K., Frew, R. & Kebede, B. A review of conventional and rapid analytical techniques coupled with multivariate analysis for origin traceability of soybean. Crit. Rev. Food Sci. Nutr. 1–20 https://doi.org/10.1080/10408398.2023.2171961 (2023).
Amarasinghe, K., Rodolfa, K. T., Lamba, H. & Ghani, R. Explainable machine learning for public policy: use cases, gaps, and research directions. Data Policy 5, e5 (2023).
Diprose, W. K. et al. Physician understanding, explainability, and trust in a hypothetical machine learning risk calculator. J. Am. Med. Inform. Assoc. 27, 592–600 (2020).
Lee, E. M. et al. Highly geographical specificity of metabolomic traits among Korean domestic soybeans (Glycine max). Food Res. Int. 120, 12–18 (2019).
Hagemeyer, J., Lülfsmann, A., Perk, M. & Breckle, S.-W. Are there seasonal variations of trace element concentrations (Cd, Pb, Zn) in wood of Fagus trees in Germany?. Vegetatio 101, 55–63 (1992).
Slagter, B. et al. Monitoring direct drivers of small-scale tropical forest disturbance in near real-time with Sentinel-1 and -2 data. Remote Sens. Environ. 295, 113655 (2023).
Strassburg, B. B. N. et al. Moment of truth for the Cerrado hotspot. Nat. Ecol. Evol. 1, 0099 (2017).
Global Witness. The Cerrado crisis: Brazil’s deforestation frontline. https://www.globalwitness.org/en/campaigns/forests/the-cerrado-crisis-brazils-deforestation-frontline/ (2024).
Henderson, J., Godar, J., Frey, G. P., Börner, J. & Gardner, T. The Paraguayan Chaco at a crossroads: drivers of an emerging soybean frontier. Reg. Environ. Change 21, 72 (2021).
Soterroni, A. C. et al. Expanding the Soy Moratorium to Brazil’s Cerrado. Sci. Adv. 5, eaav7336 (2019).
Apicella, A., Isgrò, F. & Prevete, R. Don’t push the button! Exploring data leakage risks in machine learning and transfer learning. Artif. Intell. Rev. 58, 339 (2025).
FAO and IIASA. Global Agro-Ecological Zones (GAEZ v4) (FAO and IIASA, 2021).
Marin, F. R. et al. Protecting the Amazon forest and reducing global warming via agricultural intensification. Nat. Sustain. 5, 1018–1026 (2022).
Lundberg, S. M. & Lee, S.-I. A unified approach to interpreting model predictions. In Proceedings of the 31st International Conference on Neural Information Processing Systems (NIPS'17). Curran Associates Inc., Red Hook, NY, USA, 4768–4777.
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2023).
Wickham, H. et al. Welcome to the tidyverse. J. Open Source Softw. 4, 1686 (2019).
Hijmans, R. J. Terra: Spatial Data Analysis. R package version 1.7-71. https://CRAN.R-project.org/package=terra (2024).
Pebesma, E. Simple features for R: standardized support for spatial vector data. R J 10, 439–446 (2018).
Pebesma, E. & Bivand, R. Spatial Data Science: With Applications in R (Chapman and Hall/CRC, 2023).
Hijmans, R. J. Raster: geographic data analysis and modeling. (2023).
Dunnington, D. Ggspatial: spatial data framework for Ggplot2. (2023).
Nychka, D., Furrer, R., Paige, J. & Sain, S. fields: tools for spatial data. (2021).
Paszke, A. et al. Automatic differentiation in PyTorch. (2017).
Gardner, J. R., Pleiss, G., Bindel, D., Weinberger, K. Q. & Wilson, A. G. GPyTorch: blackbox matrix-matrix Gaussian process inference with GPU acceleration. In Advances in Neural Information Processing Systems (2018).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Jordahl, K. et al. geopandas/geopandas: v0.11.1. Zenodo https://doi.org/10.5281/zenodo.6894736 (2022).
Acknowledgements
We extend our gratitude and appreciation to all the soybean farmers who donated the material that was used in this work. We are deeply grateful to the soybean collectors Francisco Mereles and Pamela Arza (Argentina), Raul Aguire (Bolivia), and the Brazilian team, who have requested to remain unnamed, for carrying out the field collections from the farms. We also thank Matthew Clarke and the Kew HPC services for supporting the computationally intensive analyses. A.A. acknowledges financial support from the Swedish Research Council (2024-04303), the Swedish Foundation for Strategic Environmental Research MISTRA (Project BioPath) and RBG Kew Development.
Author information
Authors and Affiliations
Contributions
C.C., J.S., P.W., A.A. and V.D.: conceptualisation and project administration; R.M., H.J. and F.A.: data curation; R.M. and J.T.: software development, formal analysis, investigation and methodology; R.M., H.J., F.A., C.C., J.S., V.D., I.M.-B., A.A., H.W., J.D., R.C., M.J.-A., L.Ph. and L.Pr.: sample collection planning, subsampling, accessioning, curation and preparation; J.S., M.J.-A., P.W. and C.C.: funding acquisition; R.M., C.C. and V.D.: writing—original draft; R.M., C.C., V.D., M.N., J.S. and A.A.: writing—review and editing. All authors reviewed the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Earth and Environment thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Nandita Basu. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Maor, R., Truszkowski, J., Ablett, F. et al. High-resolution soybean tracing for deforestation-free supply chains. Commun Earth Environ 7, 310 (2026). https://doi.org/10.1038/s43247-026-03380-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43247-026-03380-8