Abstract
Soil microbial communities are central to ecosystem functioning, yet their large-scale diversity patterns and environmental sensitivities remain poorly understood. Here, we analysed a national soil microbiome dataset spanning all major Australian ecosystems to assess bacterial and fungal richness across diverse environmental gradients. Using supervised deep autoencoders—neural networks that compress complex data to identify key patterns—and structural equation models, we explained around 60% of the variance in richness. Bacteria and fungi showed contrasting spatial patterns, fungal richness was tightly constrained by moisture and organic carbon, while bacterial richness peaked in nitrogen-rich, topographically complex regions and persisted across broader conditions. We propose the bacterial-to-fungal richness ratio as a spatially explicit, scalable indicator of soil community composition and ecological condition. This ratio captures gradients of aridity, nutrient imbalance, and land-use intensification, and may help identify ecosystems vulnerable to environmental change, bridging microbial biogeography with applied soil health assessment across global biomes.

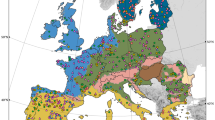
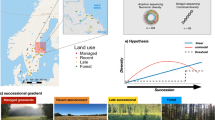
Similar content being viewed by others
Data availability
All microbial datasets analysed in this study are publicly available from the sources cited in the manuscript. Reflectance spectra from the National Geochemical Survey of Australia are openly accessible through the referenced data repository. All spatial data used in the analysis are likewise publicly available from the cited sources.
Code availability
The computer code used for the various analysis and modelling are available from the corresponding author upon request.
References
Lehmann, J. et al. Persistence of soil organic carbon caused by functional complexity. Nat. Geosci. 13, 529–534 (2020).
Schimel, J. P. & Schaeffer, S. M. Microbial control over carbon cycling in soil. Front. Microbiol. 3, 348 (2012).
Delgado-Baquerizo, M. et al. Microbial diversity drives multifunctionality in terrestrial ecosystems. Nat. Commun. 7, 10541 (2016).
Li, J. et al. Fungal richness contributes to multifunctionality in boreal forest soil. Soil Biol. Biochem. 136, 107526 (2019).
Delgado-Baquerizo, M.et al. Soil microbial communities drive the resistance of ecosystem multifunctionality to global change in drylands across the globe. Ecol. Lett. 20, 1295–1305 (2017).
Siciliano, S. D. et al. Soil fertility is associated with fungal and bacterial richness, whereas pH is associated with community composition in polar soil microbial communities. Soil Biol. Biochem. 78, 10–20 (2014).
Delgado-Baquerizo, M. et al. A global atlas of the dominant bacteria found in soil. Science 359, 320–325 (2018).
Djemiel, C. et al. Unravelling biogeographical patterns and environmental drivers of soil fungal diversity at the French national scale. Soil 10, 251–273 (2024).
Egidi, E. et al. A few ascomycota taxa dominate soil fungal communities worldwide. Nat. Commun. 10, 320–325 (2019).
Bahram, M. et al. Structure and function of the global topsoil microbiome. Nature 560, 233–237 (2018).
Labouyrie, M. et al. Patterns in soil microbial diversity across Europe. Nat. Commun. 14, 3311 (2023).
Terrat, S. et al. Mapping and predictive variations of soil bacterial richness across France. PLoS ONE 12, 1–19 (2017).
Tedersoo, L. et al. Fungal biogeography. global diversity and geography of soil fungi. Science 346, 1256688 (2014).
Orians, G. H. & Milewski, A. V. Ecology of Australia: the effects of nutrient-poor soils and intense fires. Biol. Rev. 82, 393–423 (2007).
Viscarra Rossel, R. A. et al. Environmental controls of soil fungal abundance and diversity in Australia’s diverse ecosystems. Soil Biol. Biochem. 170, 108694 (2022).
Xue, P. et al. Drivers and human impacts on topsoil bacterial and fungal community biogeography across Australia. Glob. Change Biol. 30, e17216 (2024).
Cameron, E. K. et al. Global gaps in soil biodiversity data. Nat. Ecol. Evol. 2, 1042–1043 (2018).
Bissett, A. et al. Introducing base: the biomes of Australian soil environments soil microbial diversity database. GigaScience 5, 1–11 (2016).
Egidi, E., Coleine, C., Delgado-Baquerizo, M. & Singh, B. K. Assessing critical thresholds in terrestrial microbiomes. Nat. Microbiol. 8, 2230–2233 (2023).
Bardgett, R. D. & Van Der Putten, W. H. Belowground biodiversity and ecosystem functioning. Nature 515, 505–511 (2014).
Bissett, A., Mamet, S. D., Lamb, E. G. & Siciliano, S. D. Linking niche size and phylogenetic signals to predict future soil microbial relative abundances. Front. Microbiol. 14, 1097909 (2023).
Yang, Y. et al. Soil bacterial abundance and diversity better explained and predicted with spectro-transfer functions. Soil Biol. Biochem. 129, 29–38 (2019).
Yang, Y., Shen, Z., Bissett, A. & Viscarra Rossel, R. A. Estimating soil fungal abundance and diversity at a macroecological scale with deep learning spectrotransfer functions. Soil 8, 223–235 (2022).
Chemidlin Prévost-Bouré, N. et al. Similar processes but different environmental filters for soil bacterial and fungal community composition turnover on a broad spatial scale. PLoS ONE 9, e111667 (2014).
Dobarco, M. R., Wadoux, A. & Xue, P. P. Soil and landscape grid national soil attribute maps - soil bacteria and fungi beta diversity (3” resolution) - release 1 (2022). https://data.csiro.au/collection/csiro:56612. CSIRO Data Collection.
Isbell, R., on Soil, N. C. & Terrain. The Australian Soil Classification (CSIRO Publishing, 2016).
Fierer, N. & Jackson, R. B. The diversity and biogeography of soil bacterial communities. Proc. Natl. Acad. Sci. USA 103, 626–631 (2006).
Powell, J. R. et al. Deterministic processes vary during community assembly for ecologically dissimilar taxa. Nat. Commun. 6, 8444 (2015).
Malard, L. A. & Guisan, A. Into the microbial niche. Trends Ecol. Evol. 38, 936–945 (2023).
Van Der Heijden, M. G., Bardgett, R. D. & Van Straalen, N. M. The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol. Lett. 11, 296–310 (2008).
Chen, C. et al. Fertilization regulates global thresholds in soil bacteria. Glob. Change Biol. 30, e17466 (2024).
Leff, J. W. et al. Consistent responses of soil microbial communities to elevated nutrient inputs in grasslands across the globe. Proc. Natl. Acad. Sci. USA 112, 10967–10972 (2015).
Strickland, M. S., Lauber, C., Fierer, N. & Bradford, M. A. Testing the functional significance of microbial community composition. Ecology 90, 441–451 (2009).
de Vries, F. T. & Shade, A. Controls on soil microbial community stability under climate change. Front. Microbiol. 4, 265 (2013).
Finley, B. K. et al. Soil minerals affect taxon-specific bacterial growth. ISME J. 16, 1318—1326 (2022).
Teste, F. P. et al. Mycorrhizal fungal biomass and scavenging declines in phosphorus-impoverished soils during ecosystem retrogression. Soil Biol. Biochem. 92, 119–132 (2016).
Maestre, F. T. et al. Increasing aridity reduces soil microbial diversity and abundance in global drylands. Proc. Natl. Acad. Sci. USA 112, 15684–15689 (2015).
Liu, S. et al. Decoupled diversity patterns in bacteria and fungi across continental forest ecosystems. Soil Biol. Biochem. 144, 107763 (2020).
Bengtsson-Palme, J. et al. Improved software detection and extraction of its1 and its2 from ribosomal its sequences of fungi and other eukaryotes for analysis of environmental sequencing data. Methods Ecol. Evol. 4, 914–919 (2013).
Chao, A. & Lee, S.-M. Estimating the number of classes via sample coverage. J. Am. Stat. Assoc. 87, 210–217 (1992).
Deng, Y., Umbach, A. K. & Neufeld, J. D. Nonparametric richness estimators chao1 and ace must not be used with amplicon sequence variant data. ISME J. 18, wrae106 (2024).
White, A. et al. AUSPLOTS Rangelands Survey Protocols Manual (University of Adelaide Press, 2012).
Lau, I. et al. National Geochemical Survey of Australia Reflectance Spectroscopy Measurements. v4. (CSIRO. Data Collection, 2016). https://doi.org/10.25919/5cdba18939c29.
Prescott, J. A. A climatic index for the leaching factor in soil formation. J. Soil Sci. 1, 9–19 (1950).
Gallant, J., Wilson, N., Dowling, T., Read, A. & Inskeep, C. Srtm-derived 1-second Digital Elevation Models Version 1.0. https://ecat.ga.gov.au/ (2011).
GRASS Development Team. Geographic Resources Analysis Support System (GRASS GIS) Software, Version 8.4. (Open Source Geospatial Foundation, USA, 2024). https://grass.osgeo.org.
Viscarra Rossel, R. A. et al. The Australian three-dimensional soil grid: Australia’s contribution to the globalsoilmap project. Soil Res. 53, 845–864 (2015).
ABARES. Catchment Scale Land Use of Australia - Update December 2020. Tech. Rep. (Australian Bureau of Agriculatural Resource Economics and Sciences, Canberra, 2021).
Department of Climate Change, Energy, the Environment and Water. National vegetation information system (NVIS) version 6.0 - Australia vectors (2020). Accessed 9 May 2025. www.arcgis.com/home/item.html?id=5b2f11531bd24a6a82e8fcfac8c4c5d2.
Mallat, S. G. A theory for multiresolution signal decomposition: the wavelet representation. IEEE Trans. Pattern Anal. Mach. Intell. 11, 674–693 (1989).
Daubechies, I. Ten Lectures on Wavelets (Society for Industrial and Applied Mathematics, 1992). https://doi.org/10.1137/1.9781611970104.
Webster, R. & Oliver, M. A. Geostatistics for Environmental Scientists 2nd edn (John Wiley & Sons, Chichester, UK, 2007). www.wiley.com/en-us/Geostatistics+for+Environmental+Scientists.
Le, L., Patterson, A. & White, M. Supervised autoencoders: improving generalization performance with unsupervised regularizers. In Advances in Neural Information Processing Systems, Vol. 31, 107–117 (Curran Associates, Inc., Red Hook, New York, 2018).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistical Learning: Data Mining, Inference, and Prediction 2 edn (Springer, New York, NY, 2009).
Viscarra Rossel, R. A., Webster, R., Bui, E. N. & Baldock, J. A. Baseline map of organic carbon in Australian soil to support national carbon accounting and monitoring under climate change. Glob. Change Biol. 20, 2953–2970 (2014).
Gupta, H. V., Kling, H., Yilmaz, K. K. & Martinez, G. F. Decomposition of the mean squared error and NSE performance criteria: Implications for improving hydrological modelling. J. Hydrol. 377, 80–91 (2009).
Lin, L. I.-K. A concordance correlation coefficient to evaluate reproducibility. Biometrics 45, 255–268 (1989).
MacQueen, J. Some methods for classification and analysis of multivariate observations. In Proc. Fifth Berkeley Symposium on Mathematical Statistics and Probability, Vol. 1, 281–297 (University of California Press, 1967).
Thorndike, R. L. Who belongs in the family? Psychometrika 18, 267–276 (1953).
Lundberg, S. M. & Lee, S.-I. A unified approach to interpreting model predictions. In Advances in Neural Information Processing Systems, Vol. 30, 4765–4774 (Curran Associates, Inc., 2017).
Stern, H., de Hoedt, G. & Ernst, J. Objective classification of australian climates. Aust. Meteorol. Mag. 49, 87–96 (2000).
Kline, R. B. Principles and Practice of Structural Equation Modeling 4th edn (Guilford Press, New York, NY, 2015).
Acknowledgements
This research was supported by the Australian Government through the Australian Research Council’s Discovery Projects funding scheme (project DP210100420) and the Australian Government’s Australia-China Science and Research Fund-Joint Research Centres (ACSRF-JRCs) (grant ACSRIV000077). We acknowledge the contribution of the Biomes of Australian Soil Environments (BASE) consortium in generating the data used in this publication. The BASE project is supported by funding from Bioplatforms Australia through the Australian Government National Collaborative Research Infrastructure Strategy (NCRIS).
Author information
Authors and Affiliations
Contributions
R.A.V.R. conceived the study and acquired funding. R.A.V.R. and T.B. designed the research methodology and prepared the manuscript. R.A.V.R. developed the computational tools and performed the formal analysis. A.B. conducted microbial data collection, curated and validated the data. L.W. and M.Z. prepared the spatial data and contributed to drafting of the manuscript. All authors contributed to reviewing and editing the manuscript and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Earth and Environment thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: D’Arcy Meyer-Dombard and Somaparna Ghosh. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Viscarra Rossel, R.A., Behrens, T., Bissett, A. et al. Decoding bacterial and fungal richness with autoencoders yields a unified ratio indicating soil health and ecological susceptibility. Commun Earth Environ (2026). https://doi.org/10.1038/s43247-026-03398-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43247-026-03398-y