Abstract
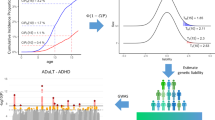
With the growing availability of time-stamped electronic health records linked to genetic data in large biobanks and cohorts, time-to-event phenotypes are increasingly studied in genome-wide association studies. Although numerous Cox-regression-based methods have been proposed for a large-scale genome-wide association study, case ascertainment in time-to-event phenotypes has not been well addressed. Here we propose a computationally efficient Cox-based method, named WtCoxG, that accounts for case ascertainment by fitting a weighted Cox proportional hazards null model. A hybrid strategy incorporating saddlepoint approximation largely increases its accuracy when analyzing low-frequency and rare variants. Notably, by leveraging external minor allele frequencies from public resources, WtCoxG further boosts statistical power. Extensive simulation studies demonstrated that WtCoxG is more powerful than ADuLT and other Cox-based methods, while effectively controlling type I error rates. UK Biobank real data analysis validated that leveraging external minor allele frequencies contributes to the power gains of WtCoxG compared with ADuLT in the analysis of type 2 diabetes and coronary atherosclerosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
This work used genotype and phenotype data from the UK Biobank (http://www.ukbiobank.ac.uk/). Source data are provided with this paper. The source data are also available via Zenodo at https://doi.org/10.5281/zenodo.16314727 (ref. 53).
Code availability
The method WtCoxG is available in the R package GRAB (version 0.2.0) via GitHub at https://github.com/GeneticAnalysisinBiobanks/GRAB. This package is also available via Zenodo at https://doi.org/10.5281/zenodo.16316316 (ref. 54).
References
McGuire, A. L. et al. The road ahead in genetics and genomics. Nat. Rev. Genet. 21, 581–596 (2020).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
All of Us Research Program Investigators. The “All of Us” research program. N. Engl. J. Med. 381, 668–676 (2019).
Wei, C.-Y. et al. Genetic profiles of 103,106 individuals in the Taiwan Biobank provide insights into the health and history of Han Chinese. NPJ Genomic Med. 6, 10 (2021).
Green, M. S. & Symons, M. J. A comparison of the logistic risk function and the proportional hazards model in prospective epidemiologic studies. J. Chronic Dis. 36, 715–723 (1983).
Callas, P. W., Pastides, H. & Hosmer, D. W. Empirical comparisons of proportional hazards, Poisson, and logistic regression modeling of occupational cohort data. Am. J. Ind. Med. 33, 33–47 (1998).
Staley, J. R. et al. A comparison of Cox and logistic regression for use in genome-wide association studies of cohort and case-cohort design. Eur. J. Hum. Genet. 25, 854–862 (2017).
Lin, D. Y. & Wei, L.-J. The robust inference for the Cox proportional hazards model. J. Am. Stat. Assoc. 84, 1074–1078 (1989).
Cox, D. R. Regression models and life-tables. J. R. Stat. Soc. Series B 34, 187–202 (1972).
Rizvi, A. A. et al. gwasurvivr: an R package for genome-wide survival analysis. Bioinformatics 35, 1968–1970 (2019).
Bi, W., Fritsche, L. G., Mukherjee, B., Kim, S. & Lee, S. A fast and accurate method for genome-wide time-to-event data analysis and its application to UK Biobank. Am. J. Hum. Genet. 107, 222–233 (2020).
Dey, R. et al. Efficient and accurate frailty model approach for genome-wide survival association analysis in large-scale biobanks. Nat. Commun. 13, 5437 (2022).
Wojcik, G. L. et al. Opportunities and challenges for the use of common controls in sequencing studies. Nat. Rev. Genet. 23, 665–679 (2022).
Pedersen, E. M. et al. ADuLT: an efficient and robust time-to-event GWAS. Nat. Commun. 14, 5553 (2023).
Pedersen, C. B. et al. The iPSYCH2012 case–cohort sample: new directions for unravelling genetic and environmental architectures of severe mental disorders. Mol. Psychiatry 23, 6–14 (2018).
Kurki, M.I. et al. FinnGen: Unique genetic insights from combining isolated population and national health register data. Preprint at medrxiv https://doi.org/10.1101/2022.03.03.22271360 (2022)
Kurki, M. I. et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 613, 508–518 (2023).
Pedersen, E. M. et al. Accounting for age of onset and family history improves power in genome-wide association studies. Am. J. Hum. Genet. 109, 417–432 (2022).
Neale, B. Liability threshold models. In Encyclopedia of Statistics in Behavioral Science (eds Everitt, B. S. & Howell, D. C.) (2005); https://doi.org/10.1002/0470013192.bsa343
Hujoel, M. L., Gazal, S., Loh, P.-R., Patterson, N. & Price, A. L. Liability threshold modeling of case–control status and family history of disease increases association power. Nat. Genet. 52, 541–547 (2020).
Leffondre, K. et al. A weighted Cox model for modelling time‐dependent exposures in the analysis of case–control studies. Stat. Med. 29, 839–850 (2010).
Sitlani, C. M. et al. Incorporating sampling weights into robust estimation of Cox proportional hazards regression model, with illustration in the Multi-Ethnic Study of Atherosclerosis. BMC Med. Res. Method. 20, 1–10 (2020).
Wu, W. et al. Retrospective association analysis of longitudinal binary traits identifies important loci and pathways in cocaine use. Genetics 213, 1225–1236 (2019).
Jiang, D., Mbatchou, J. & McPeek, M. S. Retrospective association analysis of binary traits: overcoming some limitations of the additive polygenic model. Hum. Heredity 80, 187–195 (2016).
Xu, H. et al. SPAGRM: effectively controlling for sample relatedness in large-scale genome-wide association studies of longitudinal traits. Nat. Commun. 16, 1413 (2025).
Ma, Y., Zhao, Y., Zhang, J.-F. & Bi, W. Efficient and accurate framework for genome-wide gene–environment interaction analysis in large-scale biobanks. Nat. Commun. 16, 3064 (2025).
Hayeck, T. J. et al. Mixed model association with family-biased case–control ascertainment. Am. J. Hum. Genet. 100, 31–39 (2017).
Jakobsdottir, J. & McPeek, M. S. MASTOR: mixed-model association mapping of quantitative traits in samples with related individuals. Am. J. Hum. Genet. 92, 652–666 (2013).
Wu, X. & McPeek, M. S. L-gator: genetic association testing for a longitudinally measured quantitative trait in samples with related individuals. Am. J. Hum. Genet. 102, 574–591 (2018).
The UK10K Consortium. The UK10K project identifies rare variants in health and disease. Nature 526, 82–90 (2015).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Cong, P.-K. et al. Genomic analyses of 10,376 individuals in the Westlake BioBank for Chinese (WBBC) pilot project. Nat. Commun. 13, 2939 (2022).
Chen, D. et al. A data harmonization pipeline to leverage external controls and boost power in GWAS. Hum. Mol. Genet. 31, 481–489 (2022).
Hendricks, A. E. et al. ProxECAT: Proxy External Controls Association Test. A new case–control gene region association test using allele frequencies from public controls. PLoS Genet. 14, e1007591 (2018).
Lee, S., Kim, S. & Fuchsberger, C. Improving power for rare-variant tests by integrating external controls. Genet. Epidemiol. 41, 610–619 (2017).
Li, Y. & Lee, S. Integrating external controls in case–control studies improves power for rare-variant tests. Genet. Epidemiol. 46, 145–158 (2022).
Zhu, L., Yan, S., Cao, X., Zhang, S. & Sha, Q. Integrating external controls by regression calibration for genome-wide association study. Genes 15, 67 (2024).
Li, Y. & Lee, S. Novel score test to increase power in association test by integrating external controls. Genet. Epidemiol. 45, 293–304 (2021).
Hu, Y.-J., Liao, P., Johnston, H. R., Allen, A. S. & Satten, G. A. Testing rare-variant association without calling genotypes allows for systematic differences in sequencing between cases and controls. PLoS Genet. 12, e1006040 (2016).
Arntzen, V. H. et al. A new inverse probability of selection weighted Cox model to deal with outcome‐dependent sampling in survival analysis. Biometrical J. 67, e70056 (2025).
Buchanan, A. L. et al. Worth the weight: using inverse probability weighted Cox models in AIDS research. AIDS Res. Hum. Retroviruses 30, 1170–1177 (2014).
Zou, K. H., Fielding, J. R., Silverman, S. G. & Tempany, C. M. Hypothesis testing I: proportions. Radiology 226, 609–613 (2003).
Nielsen, R., Paul, J. S., Albrechtsen, A. & Song, Y. S. Genotype and SNP calling from next-generation sequencing data. Nat. Rev. Genet. 12, 443–451 (2011).
Hamet, P. et al. PROX1 gene CC genotype as a major determinant of early onset of type 2 diabetes in slavic study participants from Action in Diabetes and Vascular Disease: Preterax and Diamicron MR Controlled Evaluation study. J. Hypertens. 35, S24–S32 (2017).
Steinthorsdottir, V. et al. A variant in CDKAL1 influences insulin response and risk of type 2 diabetes. Nat. Genet. 39, 770–775 (2007).
Mueller, P. A. et al. Coronary artery disease risk-associated Plpp3 gene and its product lipid phosphate phosphatase 3 regulate experimental atherosclerosis. Arter. Thromb. Vasc. Biol. 39, 2261–2272 (2019).
Leslie, S. et al. The fine-scale genetic structure of the British population. Nature 519, 309–314 (2015).
McCaw, Z. R., Gao, J., Lin, X. & Gronsbell, J. Synthetic surrogates improve power for genome-wide association studies of partially missing phenotypes in population biobanks. Nat. Genet. 56, 1527–1536 (2024).
Zhu, J. et al. Temporal trends in the prevalence of Parkinson’s disease from 1980 to 2023: a systematic review and meta-analysis. Lancet Healthy Longevity 5, e464–e479 (2024).
The Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Gummesson, A. et al. A genome-wide association study of imaging-defined atherosclerosis. Nat. Commun. 16, 2266 (2025).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Li, Y. Applying weighted Cox regression to genome-wide association studies of time-to-event phenotypes. Zenodo https://doi.org/10.5281/zenodo.16314727 (2025).
Li, Y., Bi, W. & Miao, L. GeneticAnalysisinBiobanks/GRAB: WtCoxG (0.2.0). Zenodo https://doi.org/10.5281/zenodo.16316316 (2025).
Acknowledgements
This research was supported by Beijing Natural Science Foundation (grant no. F251049, W.B.) and National Natural Science Foundation of China (grant nos. 62273010 and 82441004, W.B.; 82441005, W.Y.). UK Biobank data were accessed under accession number 78795 (https://biobank.ndph.ox.ac.uk/crystal/app.cgi?id=78795). This research is supported by high-performance computing platform of Peking University. We acknowledge L. Miao’s significant contributions to the development of the R package GRAB.
Author information
Authors and Affiliations
Contributions
Y.L., W.Z. and W.B designed the experiments. Y.L. and W.B performed the experiments. Y.M. gave valuable suggestions on the retrospective idea. H.X. and W.Z. contributed to analysis using GATE. Y.S. and M.Z. contributed to the code programming. Y.L. and W.B. wrote the manuscript with the assistance of W.Y. and W.Z. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Computational Science thanks Tomas Fitzgerald and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Kaitlin McCardle, in collaboration with the Nature Computational Science team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Manhattan plot for Type 2 diabetes.
Manhattan plots of GWAS results for WtCoxG0k, SPACoxALL, SPACox, GATE and GLM, with the numbers on the top left in each subplot representing the total count of detected loci, significant level α = 5 × 10−8.
Extended Data Fig. 2 Manhattan plot for coronary atherosclerosis.
Manhattan plots of GWAS results for WtCoxG0k, SPACoxALL, SPACox, GATE and GLM, with the numbers on the top left in each subplot representing the total count of detected loci, significant level α = 5 × 10−8.
Supplementary information
Supplementary Information (download PDF )
Supplementary Sections 1–7, Figs. 1–46 and Tables 1–11.
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download CSV )
Statistical source data.
Source Data Fig. 6 (download CSV )
Statistical source data.
Source Data Extended Data Fig. 1 (download CSV )
Statistical source data.
Source Data Extended Data Fig. 2 (download CSV )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Y., Ma, Y., Xu, H. et al. Applying weighted Cox regression to genome-wide association studies of time-to-event phenotypes. Nat Comput Sci 5, 1064–1079 (2025). https://doi.org/10.1038/s43588-025-00864-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43588-025-00864-z