Abstract
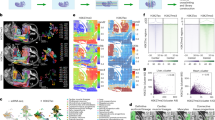
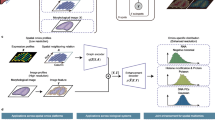
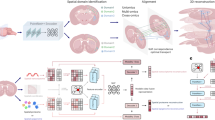
Advancements in spatial omics permit spatially resolved measurements across several biological modalities. The high cost of acquiring co-profiled multimodal data limits the analysis. This underscores the necessity for computational methods to integrate unpaired spatial multi-omics data and perform cross-modal predictions on single-modality data. The integration of spatial omics is challenging due to typically low signal-to-noise ratios. Here we introduce SWITCH (Spatially Weighted Multi-omics Integration and Cross-modal Translation with Cycle-mapping Harmonization), a deep generative model for spatial multi-omics integration. SWITCH presents a cycle-mapping mechanism that produces dependable cross-modal translations without requiring additional paired data. These cross-modal translations function as pseudo-pairs to provide supplementary signals. Systematic evaluations demonstrate that SWITCH outperforms existing methods in terms of integration accuracy and achieves more precise spatial domain delineation, resolving brain cortical structures at higher resolution. The reliability of cross-modal translations was validated, facilitating various downstream analyses such as differential analysis, trajectory inference and gene regulatory network inference.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Source data are available with this paper. All datasets used in this study are publicly available. Detailed information about the datasets, as well as the accessible code and links, can be found in Supplementary Table 2. The processed datasets are freely available at https://doi.org/10.5281/zenodo.15602076 (ref. 43).
Code availability
The source code of SWITCH, along with Jupyter notebooks for reproducing the results in this study, is available at https://github.com/zzli123/SWITCH/ and https://doi.org/10.5281/zenodo.16522594 (ref. 44).
References
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Rodriques, S. G. et al. Slide-seq: a scalable technology for measuring genome-wide expression at high spatial resolution. Science 363, 1463–1467 (2019).
Liu, Y. et al. High-spatial-resolution multi-omics sequencing via deterministic barcoding in tissue. Cell 183, 1665–1681 (2020).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 (2022).
Deng, Y. et al. Spatial-CUT&tag: spatially resolved chromatin modification profiling at the cellular level. Science 375, 681–686 (2022).
Deng, Y. et al. Spatial profiling of chromatin accessibility in mouse and human tissues. Nature 609, 375–383 (2022).
Lu, T., Ang, C. E. & Zhuang, X. Spatially resolved epigenomic profiling of single cells in complex tissues. Cell 185, 4448–4464 (2022).
Goltsev, Y. et al. Deep profiling of mouse splenic architecture with codex multiplexed imaging. Cell 174, 968–981 (2018).
He, S. et al. High-plex imaging of RNA and proteins at subcellular resolution in fixed tissue by spatial molecular imaging. Nat. Biotechnol. 40, 1794–1806 (2022).
Liu, Y. et al. High-plex protein and whole transcriptome co-mapping at cellular resolution with spatial CITE-seq. Nat. Biotechnol. 41, 1405–1409 (2023).
Zhang, D. et al. Spatial epigenome–transcriptome co-profiling of mammalian tissues. Nature 616, 113–122 (2023).
Jiang, F. et al. Simultaneous profiling of spatial gene expression and chromatin accessibility during mouse brain development. Nat. Methods 20, 1048–1057 (2023).
Vandereyken, K., Sifrim, A., Thienpont, B. & Voet, T. Methods and applications for single-cell and spatial multi-omics. Nat. Rev. Genet. 24, 494–515 (2023).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Welch, J. D. et al. Single-cell multi-omic integration compares and contrasts features of brain cell identity. Cell 177, 1873–1887 (2019).
Dou, J. et al. Bi-order multimodal integration of single-cell data. Genome Biol. 23, 112 (2022).
Cao, Z. J. & Gao, G. Multi-omics single-cell data integration and regulatory inference with graph-linked embedding. Nat. Biotechnol. 40, 1458–1466 (2022).
Xiong, L. et al. Online single-cell data integration through projecting heterogeneous datasets into a common cell-embedding space. Nat. Commun. 13, 6118 (2022).
Chen, H., Ryu, J., Vinyard, M. E., Lerer, A. & Pinello, L. SIMBA: single-cell embedding along with features. Nat. Methods 21, 1003–1013 (2024).
Chen, S. et al. Integration of spatial and single-cell data across modalities with weakly linked features. Nat. Biotechnol. 42, 1096–1106 (2024).
Samaran, J., Peyré, G. & Cantini, L. scConfluence: single-cell diagonal integration with regularized inverse optimal transport on weakly connected features. Nat. Commun. 15, 7762 (2024).
Tang, Z. et al. Modal-nexus auto-encoder for multi-modality cellular data integration and imputation. Nat. Commun. 15, 9021 (2024).
You, Y. et al. Systematic comparison of sequencing-based spatial transcriptomic methods. Nat. Methods 21, 1743–1754 (2024).
Cohen Kalafut, N., Huang, X. & Wang, D. Joint variational autoencoders for multimodal imputation and embedding. Nat. Mach. Intell. 5, 631–642 (2023).
Ashuach, T. et al. MultiVI: deep generative model for the integration of multimodal data. Nat. Methods 20, 1222–1231 (2023).
Cao, Y. et al. scButterfly: a versatile single-cell cross-modality translation method via dual-aligned variational autoencoders. Nat. Commun. 15, 2973 (2024).
Wu, K. E., Yost, K. E., Chang, H. Y. & Zou, J. Babel enables cross-modality translation between multiomic profiles at single-cell resolution. Proc. Natl Acad. Sci. USA 118, e2023070118 (2021).
Liu, J., Huang, Y., Singh, R., Vert, J. P. & Noble, W. S. Jointly embedding multiple single-cell omics measurements. Algorithms Bioinform. 143, 10 (2019).
Sunkin, S. M. et al. Allen Brain Atlas: an integrated spatio-temporal portal for exploring the central nervous system. Nucleic Acids Res. 41, D996–D1008 (2012).
Ma, N. X., Puls, B. & Chen, G. Transcriptomic analyses of NeuroD1-mediated astrocyte-to-neuron conversion. Dev. Neurobiol. 82, 375–391 (2022).
Zhou, Y. et al. Cooperative integration of spatially resolved multi-omics data with COSMOS. Nature Commun. 16, 27 (2025).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Englund, C. et al. Pax6, Tbr2, and Tbr1 are expressed sequentially by radial glia, intermediate progenitor cells, and postmitotic neurons in developing neocortex. J. Neurosci. 25, 247–251 (2005).
Bayam, E. et al. Genome-wide target analysis of NEUROD2 provides new insights into regulation of cortical projection neuron migration and differentiation. BMC Genom. 16, 681 (2015).
Bormuth, I. et al. Neuronal basic helix–loop–helix proteins Neurod2/6 regulate cortical commissure formation before midline interactions. J. Neurosci. 33, 641–651 (2013).
Hahn, M. A. et al. Reprogramming of DNA methylation at NEURO2-bound sequences during cortical neuron differentiation. Sci. Adv. 5, eaax0080 (2019).
Zu, S. et al. Single-cell analysis of chromatin accessibility in the adult mouse brain. Nature 624, 378–389 (2023).
Chai, H et al. Tri-omic single-cell mapping of the 3D epigenome and transcriptome in whole mouse brains throughout the lifespan. Nat. Methods 22, 994–1007 (2025).
Bogutz, A. B. et al. Transcription factor ASCL2 is required for development of the glycogen trophoblast cell lineage. PLoS Genet. 14, e1007587 (2018).
Kunke, M. et al. SOX10-mediated regulation of enteric glial phenotype in vitro and its relevance for neuroinflammatory disorders. J. Mol. Neurosci. 75, 26 (2025).
Forrest, M. P. et al. The psychiatric risk gene transcription factor 4 (TCF4) regulates neurodevelopmental pathways associated with schizophrenia, autism, and intellectual disability. Schizophr. Bull. 44, 1100–1110 (2018).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
Li, Z. Benchmark datasets for SWITCH. Zenodo https://doi.org/10.5281/zenodo.15602076 (2025).
Zhongzhan, L. SWITCH: a deep generative model for spatial multi-omics integration and cross-modal prediction. Zenodo https://doi.org/10.5281/zenodo.16522594 (2025).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2022YFA1103102), the National Natural Science Foundation of China (32170589, 32370616, 32488101, 32330030), Xiaomi Young Talents Program and Shanghai Eastern Youth Talent Program.
Author information
Authors and Affiliations
Contributions
G.C., Y.Z. and S.G. conceived and supervised the study. Z.L. and S.Q. designed the method with help of X.Z. and F.L. Z.L. conducted the experiment with the help of J.Y., R.T., H.L. and Z.G. Z.L. designed and created all main figures. Z.L., Y.Z., S.Q. and G.C. wrote the paper. All authors read and approved the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Computational Science thanks Christopher A. Jackson, Michelle Y. Y. Lee and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Ananya Rastogi, in collaboration with the Nature Computational Science team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes, Tables 1 and 2, and Figs. 1–24.
Source data
Source Data Fig. 2 (download ZIP )
Statistical source data.
Source Data Fig. 3 (download ZIP )
Statistical source data.
Source Data Fig. 4 (download ZIP )
Statistical source data.
Source Data Fig. 5 (download ZIP )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Z., Qu, S., Liang, H. et al. Integrative deep learning of spatial multi-omics with SWITCH. Nat Comput Sci 5, 1051–1063 (2025). https://doi.org/10.1038/s43588-025-00891-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43588-025-00891-w