Abstract
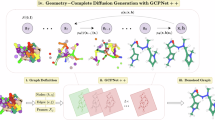
Molecule generation is advancing rapidly in chemical discovery and drug design. Flow-matching methods have recently set the state of the art (SOTA) in unconditional molecule generation, surpassing score-based diffusion models. However, diffusion models still lead in property-guided generation. In this work, we introduce PropMolFlow, an approach for property-guided molecule generation based on geometry-complete SE(3)-equivariant flow matching. Integrating five different property embedding methods with a Gaussian expansion of scalar properties, PropMolFlow achieves competitive performance against previous SOTA diffusion models in conditional molecule generation while maintaining high structural stability and validity. Additionally, it enables higher sampling speed with fewer time steps compared with baseline models. We highlight the importance of validating the properties of generated molecules through density functional theory calculations. Furthermore, we introduce a task to assess the model’s ability to propose molecules with under-represented property values, assessing its capacity for out-of-distribution generalization.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Data availability
Source data for Fig. 1d and Extended Data Figs. 1, 2b–d, 3 and 4a,c are available with this Brief Communication. All data can be found in the Zenodo repository36, including revised QM9 SDF data, SDF files for generated raw molecules, DFT-calculated structures and their molecular properties saved in the extxyz format, full property MAE and structural-validity results in CSV format for all saved PropMolFlow checkpoint models, sampled structures and their full PoseBusters results for all baseline models, and EGNN property-predictor pretrained models and notebook examples to use the property predictors.
Code availability
Our PropMolFlow implementation is available at https://github.com/Liu-Group-UF/PropMolFlow and on Zenodo40. A web interface is also provided for a simple demonstration at https://propmolflow-website.vercel.app.
References
Sanchez-Lengeling, B. & Aspuru-Guzik, A. Inverse molecular design using machine learning: generative models for matter engineering. Science 361, 360–365 (2018).
Hoogeboom, E., Satorras, V. G., Vignac, C. & Welling, M. Equivariant diffusion for molecule generation in 3D. Proc.Mach. Learn. Res. 162, 8867–8887 (2022).
Satorras, V. G., Hoogeboom, E. & Welling, M. E(n) equivariant graph neural networks. Proc. Mach. Learn. Res. 139, 9323–9332 (2021).
Xu, M., Powers, A. S., Dror, R. O., Ermon, S. & Leskovec, J. Geometric latent diffusion models for 3D molecule generation. Proc. Mach. Learn. Res. 202, 38592–38610 (2023).
Liu, X., Gong, C. & Liu, Q. Flow straight and fast: learning to generate and transfer data with rectified flow. In Eleventh International Conference on Learning Representations (ICLR, 2023); https://openreview.net/forum?id=XVjTT1nw5z
Lipman, Y., Chen, R. T. Q., Ben-Hamu, H., Nickel, M. & Le, M. Flow matching for generative modeling. In Eleventh International Conference on Learning Representations (ICLR, 2023); https://openreview.net/forum?id=PqvMRDCJT9t
Albergo, M. S. & Vanden-Eijnden, E. Building normalizing flows with stochastic interpolants. In Eleventh International Conference on Learning Representations (ICLR, 2023); https://openreview.net/forum?id=li7qeBbCR1t
Höllmer, P. et al. Open materials generation with stochastic interpolants. In International Conference on Machine Learning 23417–23450 (PMLR, 2025).
Campbell, A., Yim, J., Barzilay, R., Rainforth, T. & Jaakkola, T. Generative flows on discrete state-spaces: enabling multimodal flows with applications to protein co-design. In International Conference on Machine Learning 5453–5512 (PMLR, 2024).
Dunn, I. & Koes, D. R. Exploring discrete flow matching for 3D de novo molecule generation. Preprint at https://arxiv.org/abs/2411.16644 (2024).
Song, Y. et al. Equivariant flow matching with hybrid probability transport for 3D molecule generation. Adv. Neural Inf. Process. Syst. 36, 549–568 (2023).
Dumitrescu, A. et al. E(3)-equivariant models cannot learn chirality: field-based molecular generation. In Thirteenth International Conference on Learning Representations (ICLR, 2025); https://openreview.net/forum?id=mXHTifc1Fn
Vignac, C., Osman, N., Toni, L. & Frossard, P. MiDi: mixed graph and 3D denoising diffusion for molecule generation. In Machine Learning and Knowledge Discovery in Databases: Research Track (eds Koutra, D. et al.) 560–576 (Springer, 2023).
Morehead, A. & Cheng, J. Geometry-complete diffusion for 3D molecule generation and optimization. Commun. Chem. 7, 150 (2024).
Ramakrishnan, R., Dral, P. O., Rupp, M. & Von Lilienfeld, O. A. Quantum chemistry structures and properties of 134 kilo molecules. Sci. Data 1, 140022 (2014).
Wu, Z. et al. MoleculeNet: a benchmark for molecular machine learning. Chem. Sci. 9, 513–530 (2018).
Gebauer, N. W. A., Gastegger, M., Hessmann, S. S. P., Müller, K.-R. & Schütt, K. T. Inverse design of 3D molecular structures with conditional generative neural networks. Nat. Commun. 13, 973 (2022).
Jing, B., Eismann, S., Suriana, P., Townshend, R. J. L. & Dror, R. Learning from protein structure with geometric vector perceptrons. In International Conference on Learning Representations (ICLR, 2021); https://openreview.net/forum?id=1YLJDvSx6J4
Bao, F. et al. Equivariant energy-guided SDE for inverse molecular design. In Eleventh International Conference on Learning Representations (ICLR, 2023); https://openreview.net/forum?id=r0otLtOwYW
Huang, H., Sun, L., Du, B. & Lv, W. Learning joint 2-D and 3-D graph diffusion models for complete molecule generation. IEEE Trans. Neural Netw. Learn. Syst. 35, 11857–11871 (2024).
Landrum, G. et al. RDKit: Open-Source Cheminformatics Software (RDKit, 2016).
Buttenschoen, M., Morris, G. M. & Deane, C. M. PoseBusters: AI-based docking methods fail to generate physically valid poses or generalise to novel sequences. Chem. Sci. 15, 3130–3139 (2024).
Morgan, H. L. The generation of a unique machine description for chemical structures—a technique developed at Chemical Abstracts Service. J. Chem. Doc. 5, 107–113 (1965).
Kim, S. et al. PubChem in 2021: new data content and improved web interfaces. Nucleic Acids Res. 49, D1388–D1395 (2021).
Ho, J. & Salimans, T. Classifier-free diffusion guidance. In NeurIPS 2021 Workshop on Deep Generative Models and Downstream Applications (2021); https://openreview.net/forum?id=qw8AKxfYbI
Karras, T. et al. Guiding a diffusion model with a bad version of itself. Adv. Neural Inf. Process. Syst. 37, 52996–53021 (2024).
Boffi, N. M., Albergo, M. S. & Vanden-Eijnden, E. How to build a consistency model: learning flow maps via self-distillation. In Thirty-ninth Annual Conference on Neural Information Processing Systems (2025); https://openreview.net/forum?id=Di5apl8HSH
Liu, G.-H., Choi, J., Chen, Y., Miller, B. K. & Chen, R. T. Q. Adjoint Schrödinger bridge sampler. In Thirty-ninth Annual Conference on Neural Information Processing Systems (2025); https://openreview.net/forum?id=rMhQBlhh4c
Dunn, I. & Koes, D. R. FlowMol3: flow matching for 3D de novo small-molecule generation. Preprint at https://arxiv.org/abs/2508.12629 (2025).
Dunn, I. & Koes, D. R. Mixed continuous and categorical flow matching for 3D de novo molecule generation. Preprint at https://arxiv.org/abs/2404.19739 (2024).
Gat, I. et al. Discrete flow matching. Adv. Neural Inf. Process. Syst. 37, 133345–133385 (2024).
Le, T., Cremer, J., Noe, F., Clevert, D.-A. & Schütt, K. T. Navigating the design space of equivariant diffusion-based generative models for de novo 3D molecule generation. In Twelfth International Conference on Learning Representations (ICLR, 2024); https://openreview.net/forum?id=kzGuiRXZrQ
Axelrod, S. & Gómez-Bombarelli, R. GEOM, energy-annotated molecular conformations for property prediction and molecular generation. Sci. Data 9, 185 (2022).
Becke, A. D. Density-functional thermochemistry. III. The role of exact exchange. J. Chem. Phys. 98, 5648–5652 (1993).
Lee, C., Yang, W. & Parr, R. G. Development of the Colle–Salvetti correlation-energy formula into a functional of the electron density. Phys. Rev. B 37, 785–789 (1988).
Zeng, C., Jin, J. & Liu, M. PropMolFlow: property-guided molecule generation with geometry-complete flow matching. Zenodo https://doi.org/10.5281/zenodo.17726328 (2025).
Moret, M. et al. Leveraging molecular structure and bioactivity with chemical language models for de novo drug design. Nat. Commun. 14, 114 (2023).
O’Boyle, N. M. et al. Open Babel: an open chemical toolbox. J. Cheminform. 3, 33 (2011).
Frisch, M. J. et al. Gaussian 16 revision C.01 (2016).
Zeng, C., Jin, J. & Mingjie, L. PropMolFlow code. Zenodo https://doi.org/10.5281/zenodo.17702415 (2025).
Acknowledgements
We acknowledge funding from NSF Grant OAC-2311632 and from the AI and Complex Computational Research Award of the University of Florida. S.M. also acknowledges support from the Simons Center for Computational Physical Chemistry (Simons Foundation grant 839534). We also acknowledge UFIT Research Computing for computational resources and consultation, as well as the Nvidia AI Technology Center at UF. We would like to thank A. Morehead (University of Missouri) for providing the checkpoint models for the conditional generation of GeoLDM.
Author information
Authors and Affiliations
Contributions
C.Z., J.J. and M.L. conceptualized the study. C.Z. and J.J. developed the methodology, implemented the code and ran the computational experiments under the guidance of M.L. C.A. developed the PropMolFlow demo web interface under the guidance of C.Z. and M.L. M.T., G.K., E.B.T., R.G.H., A.R. and S.M. provided guidance for various parts of this project. C.Z., M.L. and S.M. wrote the original draft, and all authors participated in the discussion and review of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Computational Science thanks Andreas Luttens and Arne Schneuing for their contribution to the peer review of this work. Primary Handling Editor: Kaitlin McCardle, in collaboration with the Nature Computational Science team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Chemical validity and sampling efficiency of PropMolFlow against five baseline models.
a, Molecule stability. b, RDKit validity. c, Uniqueness. d, Closed-shell ratio. e, PoseBusters validity. f, Sampling time. The y-axis for sampling time uses a broken scale to expand 0–150 min and compress 150–360 min by a ratio of 5 for visual clarity. Each box plot summarizes the metric values computed for six molecular properties (n = 6) and 10000 sampled molecules. The median is shown as a solid line. The edges of the box correspond to the first and third quartiles, and the whiskers extend to values within 1.5 × interquartile ranges. All individual data points are overlaid as black dots. PropMolFlow results use the top-performing models of each property in the ID tasks for property alignment.
Extended Data Fig. 2 Performance of GVP property predictors without and with DFT relaxation.
a, Comparison between Target, DFT and GVP shows the reliability of GVP to evaluate the MAE metrics commonly used for property-guided generation. Comparison between GVP and GVP-R, and between DFT and DFT-R shows the structural dependence of GVP-predicted and DFT-predicted properties, respectively. Comparison between GVP and DFT with and without relaxation shows the reliability of GVP in capturing the ground-truth DFT values over raw and relaxed structures. ‘Target’, ‘DFT’ and ‘GVP’ are respective input, DFT-calculated and GVP-predicted property values on raw molecules. ‘DFT-R’ and ‘GVP-R’ refer to values evaluated on DFT-relaxed molecules. The ‘d’ indicates the MAE distances between two property value vectors. b, Pairwise comparison between Target, DFT and GVP on raw molecules. c, GVP versus DFT for both raw and DFT-relaxed molecules. d, Root mean squared distances (RMSDs) and normalized property sensitivity due to DFT relaxation. Molecules with the highest RMSDs for each property and their DFT and GVP values are shown. In molecule representation, gray, red, blue, and white colors indicate C, O, N, and H, respectively. Property values for α, Δϵ, and Cv are in units of Bohr3, eV, and cal/(mol ⋅ K), respectively.
Extended Data Fig. 3 Interpolation study by varying property values.
The minimum and maximum target properties in red and corresponding DFT-calculated properties in blue are provided below the configurations. All molecules chosen here pass the filtering criteria and have DFT values closest to the target properties. C, H, O, and N are in gray, white, red, and blue colors, respectively. Property units are provided in the square brackets under each property symbol.
Extended Data Fig. 4 Toward out-of-distribution generation.
a, Distribution of DFT-calculated and GVP-predicted values for PropMolFlow generated molecules, and the property distribution of the QM9 training data is also included. The vertical black dashed line in histograms represents the target property value q0.99, which corresponds to the 99th quatile of training data distributions. Curves on top of histograms are fitted with a kernel density estimation. b, Three example molecules that do not exist in QM9 but are found in a larger PubChem dataset. Numbers below the configurations are DFT calculated property values on raw molecules generated by PropMolFlow models. C, H, O, N, and F are in gray, white, red, blue, and yellow colors, respectively. Property values for α, Δϵ, and Cv are in units of Bohr3, eV, and cal/(mol ⋅ K), respectively. c, Maximum Tanimoto similarity of generated and filtered molecules compared to the training data using a Morgan fingerprint. Dashed lines indicate the 0.8 similarity cutoff to define novel molecules.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–14, Tables 1–14, proofs, extended results and discussions, and additional references.
Source data
Source Data Fig. 1 (download ZIP )
Statistical source data.
Source Data Extended Data Fig. 1 (download CSV )
Statistical source data.
Source Data Extended Data Fig. 2 (download ZIP )
Statistical source data.
Source Data Extended Data Fig. 3 (download CSV )
Statistical source data.
Source Data Extended Data Fig. 4 (download ZIP )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zeng, C., Jin, J., Ambrose, C. et al. PropMolFlow: property-guided molecule generation with geometry-complete flow matching. Nat Comput Sci 6, 233–242 (2026). https://doi.org/10.1038/s43588-025-00946-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43588-025-00946-y