Abstract
Background
Ovarian cancer is a major female reproductive health issue with heterogeneous biological features on its subtypes, which may require different therapeutic strategies. Glucagon-like peptide-1 receptor (GLP-1R) agonists were reported to be beneficial for ovarian cancer, but the causal effects and mechanisms on its heterogeneous subtypes remain unclear.
Methods
We used genetic variants robustly associated with gene expression, protein level, splicing event, and DNA methylation of GLP-1R in six endocrine-related tissues (N ≤ 35,431) as genetic instruments to proxy the effect of GLP-1R agonism. To increase power, we conducted a meta-analysis of genome-wide association studies of ovarian cancer (29,066 cases, 461,542 controls), and identified 12 genome-wide associated variants, including two previously unreported variants: rs77247401 (MIR1208) and rs56159231 (PLEKHM1).
Results
Here we show that gene expression of GLP-1R in pancreas is associated with a reduced risk of overall ovarian cancer risk odds ratio ([OR] = 0.94, 95% confidence interval [CI] 0.89-1.00) and endometrioid ovarian cancer (ENOC; OR = 0.83, 95% CI = 0.72-0.95), which the finding is validated using splicing event of GLP-1R in pancreas (OR = 0.13, 95% CI = 0.02-0.86). However, null association is found for GLP-1R expression in pancreas with other ovarian cancer subtypes. The phenome-wide MR followed by mediation MR identifies six body composition and metabolic factors as mediators, including 18:2 linoleic acid.
Conclusions
The protective effect of GLP-1R agonists on ovarian cancer, especially ENOC, needs further validation in large-scale and well-conducted clinical trials.
Plain language summary
The class of drugs known as GLP-1 receptor (GLP-1R) agonists are known to have a range of health benefits. However, their effect on ovarian cancer, which is a significant health concern for women worldwide, has been unclear. GLP-1R agonists act on a protein expressed in the outside of cells, called the GLP-1 receptor. In our study, we used human genetic data to predict activity of the GLP-1 receptor in over 490,000 individuals. We found that GLP-1R activity in the pancreas was associated with a lower risk of a specific subtype of ovarian cancer called endometrioid ovarian cancer. This protective effect appeared to be partly influenced by changes in body composition and molecules in the blood, such as linoleic acid. Our results suggest that GLP-1R agonists could help prevent certain forms of ovarian cancer. Further clinical studies are needed to confirm this possibility.
Similar content being viewed by others
Introduction
Ovarian cancer is the most lethal gynecologic malignancy, with a dismal 50% 5-year survival rate, primarily due to late-stage diagnosis and aggressive tumor biology1,2. It is highly heterogeneous, with epithelial ovarian cancer (EOC) representing approximately 90% of cases2,3. EOC is further classified into five major histological subtypes—high-grade serous (HGSOC), low-grade serous (LGSOC), mucinous (MOC), endometrioid (ENOC), and clear cell (CCOC) carcinoma—each exhibiting distinct risk factors, molecular profiles, and clinical behaviors3,4,5. Understanding the unique biology of these subtypes is critical for developing targeted therapies and improving patient outcomes.
Glucagon-like peptide-1 receptor (GLP-1R) agonists have emerged as effective therapies for multiple metabolic disorders, demonstrating significant benefits in weight management and cardiovascular risk reduction in individuals with obesity or type 2 diabetes (T2D)6,7,8.The 2025 American Diabetes Association (ADA) Standards of Care have broadened the use of GLP-1R agonists beyond glycemic control, recommending them for patients with T2D and chronic kidney disease, as well as those with T2D, obesity, or prediabetic individuals with nonalcoholic fatty liver disease and heart failure with preserved ejection fraction9. Beyond diabetes and obesity, GLP-1R agonists exhibit a multifaceted impact on cancer progression across various malignancies10. Concerns regarding their potential association with cancer risk, particularly thyroid cancer, have prompted regulatory warnings and contraindications for patients with a history of medullary thyroid carcinoma or multiple endocrine neoplasia type 211. While preclinical and clinical studies have explored the role of GLP-1R signaling in various cancers, its impact on ovarian cancer and its histological subtypes remains poorly understood12. This calls for more extensive research to unravel the intricate relationships between the GLP-1R agonists and ovarian cancer and subtypes, providing valuable insights for clinicians and researchers alike.
Existing epidemiology studies provided inconclusive evidence for the association between GLP-1R agonists and ovarian cancer. A multi-center retrospective cohort study in individuals with T2D suggested that usage of GLP-1R agonists may reduce ovarian cancer risk compared to insulin or metformin usage13. However, subsequent analyses using global pharmacovigilance data failed to replicate these findings14. Randomized controlled trials (RCTs), which is the gold-standard approach to extend indication of marketed drugs, have been limited by short follow-up periods and insufficient number of cases to detect the effect of GLP-1R agonists on ovarian cancer15,16,17. Alternative approaches are needed to evaluate the causal relationship between the two.
Application of Mendelian randomization (MR) principles is a parallel approach to estimate the effect of GLP-1R expression on ovarian cancer, which may proxy the effects of GLP-1R agonists. By leveraging germline genetic variants as instruments, MR minimizes confounding from sociodemographic, behavioral, and health-related factors, providing more reliable causal estimates18,19. Till now, one MR study has explored the effect of GLP-1R agonism on ovarian cancer risk, which suggested no association. However, its methodological limitations undermine the validity of its conclusions20. Moreover, no study has yet investigated the effects of GLP-1R expression on the distinct histological subtypes of ovarian cancer, nor has the potential mediating role of weight loss or other metabolic pathways5,21,22.
In this study, we investigate the association of GLP-1R expression on ovarian cancer and five major histological subtypes with a GWAS meta-analysis followed by MR. Here we show that pancreatic GLP-1R expression is associated with a reduced risk of endometrioid ovarian cancer. We further demonstrate that this protective effect is partly mediated by specific body composition and metabolic factors, including linoleic acid using phenome-wide MR and mediation MR23. These findings suggest that GLP-1R agonists may represent a promising targeted therapeutic strategy for endometrioid ovarian cancer, warranting further clinical investigation.
Methods
Study overview and design
The overall study design was summarized in Fig. 1. We began by leveraging data from genetic association studies of omics traits to select comprehensive genetic instruments for GLP-1R expression24,25,26,27,28,29,30,31,32,33,34,35,36,37,38. Then, a large-scale genome-wide association study (GWAS) meta-analysis was performed for ovarian cancer using summary statistics from the OCAC4, UK Biobank39, and FinnGen40. The GWAS meta-analysis results for overall ovarian cancer, as well as subtype-specific GWAS results for EOC from the Ovarian Cancer Association Consortium (OCAC)4, were used as outcomes in the MR analyses (Supplementary Data 1). To explore potential mediators linking GLP-1R expression to ovarian cancer subtypes, we conducted phenome-wide analysis followed by two-step MR analyses23. In addition, we performed validation analyses using the MELODI Presto platform41, to externally validate findings from prior studies and confirm the identified mediators. Our reporting adheres to the STROBE-MR (Strengthening the Reporting of MR Studies) guidelines42,43.
Flowchart depicting the study design to explore causal relationships between GLP-1R expression and ovarian cancer (created with biorender.com). QTLs quantitative trait loci, GWAS genome-wide association study, OCAC Ovarian Cancer Association Consortium, LD linkage disequilibrium.
Genome-wide association study meta-analysis of ovarian cancer
To enhance statistical power, we conducted a GWAS meta-analysis of ovarian cancer using summary statistics from three large-scale cohorts: the OCAC(N cases = 25,509, N controls = 40,941), the UK Biobank (N cases = 1218, N controls = 198,523), and FinnGen (Release 12; N cases = 2339, N controls = 222,078)4,39,40. Meta-analysis was conducted using the inverse variance-weighted fixed-effect method implemented in METAL44 and using easyQC45 for quality control. Chromosome positions were lifted over to Genome Build 38 using the UCSC Lift Genome Annotations tool (https://genome.ucsc.edu/cgi-bin/hgLiftOver). The linkage disequilibrium score (LDSC)46 regression was implemented to evaluate the heritability of the trait, the genomic inflation factor (λ), and the intercept. The conditional analysis of top association variants (P-value < 5 × 10-8) was conducted using GCTA-COJO47. Functional annotation of the top association variants was conducted using RegulomeDB v248 and HaploReg49,50. The RegulomeDB score evaluates the strength of functional genomic evidence, with a lower score indicating stronger supporting evidence (details provided in Supplementary Data 2).
We also used the FUMA for functional mapping and annotation of the genome-wide association Study51 (Supplementary Note 1). The putative causal variants were prioritized by combined annotation dependent depletion (CADD) score52. The CADD score > 12.37 indicates a potentially deleterious effect. Phenotypes associated with identified loci were collected by searching the GWAS Catalog (https://www.ebi.ac.uk/gwas/home). Furthermore, candidate SNPs were mapped to protein-coding genes in SNP2GENE and then the mapped genes were annotated in biological contexts with the GENE2FUNC module of FUMA. This included a tissue specificity test against differentially expressed genes (DEGs) across all 54 GTEx v8 tissues, with significance defined by a Bonferroni-corrected P-value < 0.05; and a gene set enrichment analysis against the MSigDB, for which significance was defined by a Benjamini-Hochberg false discovery rate (FDR) threshold of q < 0.05.
Assessment of druggability and drug repurposing potential
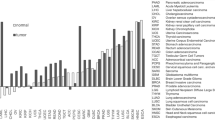
To systematically evaluate the translational potential of the potential effector genes, we searched the Citeline PharmaProjects database (accessed October 2025), a comprehensive repository tracking the development history of over 100,000 drug candidates worldwide. We cross-referenced the genes of interest against this database to identify those encoding targets of approved or investigational therapeutic agents. To characterize the therapeutic potential of associated genes, we curated detailed drug information—including name, indication, mechanism of action, and development status—for all genes with existing drug development programs.
Instrument selection of GLP-1R expression
Fifteen studies estimating the genetic associations of molecular traits of GLP-1R were included in the initial instrument search24,25,26,27,28,29,30,31,32,33,34,35,36,37,38. The effect sizes of these QTLs were reported in standard deviation (SD) units in the original studies.
Source of instruments
Only QTLs associated with molecular traits (e.g., DNA methylation, gene expression, splicing events, and protein levels) of GLP-1R in endocrine tissues were selected as candidate instruments. The tissues included the hypothalamus, pituitary, thyroid, small intestine, and pancreas. For QTLs lacking tissue-specific data, those in whole blood or plasma were used to proxy the effects of GLP-1R expression in the circulation system.
Selection of instruments
Tissue-specific eQTL data for GLP-1R were extracted from GTEx v8, with the majority of donors in the GTEx consortium representing participants from the general population. Cis-acting DNA methylation, gene expression, and splicing QTLs were prioritized as primary instruments because these variants are more likely to exert direct effects on GLP-1R and reduce the likelihood of pleiotropy. Given the importance of proteins in drug target identification, we utilized cis-acting protein QTLs (pQTLs) as our core set of genetic instruments. To complement this primary analysis and assess robustness, we also employed a set of trans-pQTLs as supplementary instruments53. Linkage disequilibrium (LD) clumping (r² <0.001) was performed to identify independent instruments, utilizing LD reference panels from the European participants in the 1000 Genomes Project. A stringent clumping window of 10,000 kb was applied, and only bi-allelic SNPs with a minor allele frequency (MAF) > 0.01 were included. QTLs with F-statistics exceeding 10 were selected to mitigate the risk of weak instrument bias53.
To evaluate potential sex-specific bias of the genetic instruments for GLP-1R expression, we utilized sex-biased eQTLs (sb-eQTLs) from the GTEx v8 database54. Specifically, we extracted the summary statistics for all cis-eQTLs associated with expressions of GLP-1R. The presence of a significant sb-eQTL was determined based on a genotype-by-sex (G × Sex) interaction test, as defined in the original GTEx v8 analysis. We applied the same statistical threshold, considering an sb-eQTL to be non-significant if the Storey q-value (FDR) was greater than 0.25. As supportive evidence, we also confirmed that the directions of effects of the eQTLs were consistent between males and females in the sex-stratified analyses.
Outcome selection
The primary outcomes were overall ovarian cancer and its five major histological subtypes: high-grade serous (HGSOC), low-grade serous (LGSOC), endometrioid (ENOC), mucinous (MOC), and clear cell (CCOC) carcinoma. Body mass index (BMI; N = 532,396) and Type 2 diabetes(T2D; N = 933,970) were included as positive control outcomes55,56. For histological subtype analyses, GWAS summary statistics were obtained from the OCAC, which included 40,941 controls and cases stratified by histotype: 13,037 HGSOC, 1012 LGSOC, 2810 ENOC, 1417 invasive MOC, and 1366 CCOC4.
MR analyses between GLP-1R expression and ovarian cancer
We performed two-sample MR using the TwoSampleMR package in ref. 57. In the primary MR analysis, we estimated the effect of GLP-1R expression on ovarian cancer and its subtypes. The genetic variants robustly associated with molecular traits of GLP-1R were used to proxy GLP-1R expression (each molecular trait was treated as one exposure). In the positive control analysis, effect of GLP-1R expression on T2D and BMI were estimated. The IVW method was used as discovery method when two or more instruments were included in the analysis, whereas the Wald ratio method was used when only one instrument was available. When an instrument was missed from the outcome GWAS, a proxy genetic variant with square of pair-wise correlation (LD r2) > 0.8 was selected. A Bonferroni-corrected P-value threshold was estimated by multiplying the four types of GLP-1R expression (DNA methylation, gene expression, splicing events, and protein levels as exposures) with genetic correlation adjusted outcomes. We used the P-value threshold (P-value < 0.05/4, i.e., 0.0125) as heuristic to identify a feasible number of MR findings.
Candidate mediators filtering using EpiGraphDB platform
To identify potential mediators linking GLP-1R expression to ovarian cancer, we utilized the EpiGraphDB platform, a biomedical knowledge graph database containing 2204 genome-wide summary datasets from the IEU OpenGWAS project58. Candidate mediators were screened in two processes: (1) Pre-computed MR results for endometrioid ovarian cancer (GWAS ID “ieu-a-1125”) were queried, and phenotypes with MR P-value < 0.05 were selected; (2) Phenotypes were further excluded given the study population, sex, ancestry, heritability, and data quality. Only those phenotypes passed both processes were selected as candidate mediators, and subsequently included in the mediation analysis. Detailed information was shown in Supplementary Fig. 1.
Mediation analysis using two-step MR framework
Mediation analysis was performed using a two-step MR approach23. In the first step, univariate MR was used to estimate the effect of GLP-1R expression on candidate mediators (denoted as “a”). In the second step, genetic instruments for the mediators were selected (P-value < 5 × 10-8; LD clumping r2 < 0.001), and the effects of mediators on ovarian cancer subtypes were estimated using univariate MR (denoted as “b”). The indirect effect of GLP-1R expression on ovarian cancer via the mediator was calculated as the product of the two effects (a × b). The proportion of mediation was estimated as (a × b) / c, where “c” represents the total effect of GLP-1R expression on the outcome. Mediators were considered valid if they showed evidence of being influenced by GLP-1R expression and had a causal effect on ENOC in the expected direction, with both associations requiring P-value < 0.05 with an FDR correction at q < 0.05.
MR sensitivity analyses
MR relies on three key assumptions: (1) relevance, where genetic instruments are robustly associated with the exposure of interest; (2) exchangeability, which means that there are no unmeasured common causes (confounders) between the instruments and the outcome; and (3) exclusion restriction, where the instruments affect the outcome solely through the exposure under investigation59.
Verify the relevance assumption
Instrument strength was evaluated using the mean Cragg–Donald F-statistics calculated as \((\frac{n-k-1}{k})(\frac{{R}^{2}}{1-{R}^{2}})\), where R² represents the variance in exposure explained by instruments, n is the sample size for the exposure, and k is the number of instruments. A mean F-statistic > 10 indicates sufficient instrument strength for MR analysis.
Falsify the exchangeability assumption
Genetic colocalization analysis was performed for MR findings with a P-value < 0.05 (Supplementary Note 2). Colocalization, a Bayesian model, estimates the probability that a single variant affects both the GLP-1R expression and an ovarian cancer outcome. Both conventional colocalization methods60,61 and a recently developed approach, PWCoCo, which accounts for multiple signals in one region, were used to estimate colocalization probability. For exposures lacking sufficient genetic variants coverage or missing key information (for example, allele frequency or effect size), we conducted the “LD check” analysis53. For the tested MR estimates, we considered findings with colocalization probability ≥ 80% as “colocalized”, and LD checked findings with r2 ≥ 0.8 as “colocalized”.
Falsify the exclusion restriction assumption
Sensitivity analyses were conducted using MR-Egger, weighted median and weighted mode methods to assess the robustness of the IVW estimates19,62. Heterogeneity was assessed using Cochran’s Q statistic, with high heterogeneity suggesting invalid instruments59.
Sensitivity and Validation Analyses for Mediation Findings
Assessment of sex-based heterogeneity in the two-step MR analysis
To address potential concerns that sex-specific effects might confound our mediation analysis, we employed two approaches to rigorously evaluate the impact of sex on our findings. First, we repeated the two-step MR mediation analysis using exclusively female-specific genetic data, including female cis-eQTLs for GLP-1R expression in the pancreas (from GTEx v8) and female-specific GWAS summary statistics for both the body composition mediators and endometrioid ovarian cancer. Second, we applied the MRSamePopTest63 package to formally test for heterogeneity in the genetic associations (from GLP-1R expression to body composition mediators) between female-specific and sex-combined GWAS data. The non-significant results from this test provide evidence against substantial sex-based confounding in the exposure-mediator relationship.
Multivariable MR analysis with weak instruments
To strengthen causal inference and account for potential pleiotropy in the mediation pathway, we implemented a cis-Multivariable MR (cis-MVMR) framework64. We utilized all available cis-eQTLs for GLP-1R in pancreatic tissue under relaxed instrument selection criteria (P-value < 0.05) followed by LD pruning (r² <0.95). The analysis from exposure to mediators and outcomes employed robust principal components generalized method of moments (PC-GMM), and analysis from mediators to outcome by robust principal components multivariate generalized method of moments (PC-MVGMM) to model effects across multiple mediators. We systematically assessed the robustness of these analyses by varying the number of principal components.
Given that weak instrument bias and high correlation among mediators can render standard multivariable MR estimates unstable and imprecise, we applied MR-Mediator Bayesian Averaging (MR-MBA)65 to complement the cis-MVMR analysis. This method identifies the most important mediator by comparing all possible combinations of mediators, providing a more robust ranking under conditions of weak instrument strength.
Associations for these six mediators were considered significant based on the FDR threshold of q < 0.05 used in the validation analysis stage.
External validation of potential mediators
To investigate potential mediators between GLP-1R expression and ENOC, a comprehensive literature search was conducted, with the search period ending on October 12, 2024. To identify relevant literature, keywords “Endometrioid Ovarian Cancer”, “Ovarian Cancer”, “Genetic Factors”, “Risk Factors”, “Environmental Factors”, and “Biomarkers” were used to query major biomedical databases, including PubMed, Web of Science, and Embase. In addition, MELODI Presto41, a platform designed to explore semantic triples derived from biomedical literature, was employed to systematically assess potential mediating pathways41. Specific queries, including “GLP-1R agonists” and “endometrioid ovarian cancer”, were used to identify enriched triples from individual searches and overlapping triples across query lists. We further explored the impact of mediators on other EOC subtypes using MR analyses, to evaluate the potential heterogeneity across all EOC subtypes.
Statistics and Reproducibility
The complete and reproducible analysis code is available (https://zenodo.org/records/17994065)66 and as described in the Code Availability section below. Primary causal estimates were derived using the IVW method for MR analyses. Sensitivity analyses were conducted using MR-Egger, weighted median, weighted mode methods. A Bonferroni-corrected P-value threshold (P < 0.0125) was applied to account for multiple testing across the four molecular trait types (gene expression, protein, splicing, and methylation) used as exposures. For GWAS meta-analysis, the inverse variance-weighted fixed-effect method was implemented in METAL. Genetic colocalization was assessed using both conventional Bayesian methods and the PWCoCo method, with a posterior probability ≥80% considered evidence of colocalization. In mediation analyses, the FDR of q < 0.05 was used to determine significant mediators.
The sample sizes for all GWAS used in this study were predetermined by the respective contributing consortia. No additional sample size calculation was performed, as the analyses utilized publicly available summary-level data. Replicates in this genetic study are defined as: (1) independent genetic instruments for the same exposure, (2) consistent results across different MR methods, and (3) validation of primary findings in independent datasets or using alternative data modalities. All analyses were conducted once on the available summary statistics. The reporting of this study follows the STROBE-MR guidelines to ensure transparency and reproducibility.
Results
Genetic associations and functional annotation of ovarian cancer
We performed a GWAS meta-analysis of ovarian cancer by combining the previously published effort from ovarian cancer consortium (OCAC), UK Biobank and FinnGen (Supplementary Data 1). Our meta-analysis constitutes 25.5 million SNPs tested in 29,066 ovarian cancer cases and 461,542 controls. This analysis identified 1434 genetic signals associated with ovarian cancer (P-value < 5×10-8; MAF > 1%;HetPVal > 0.05). The quantile-quantile (Q-Q) plot showed a substantial deviation of the test statistics from the null expectation, consistent with a polygenic architecture for the disease (Supplementary Fig. 2). We evaluated potential confounding and SNP-based heritability using linkage disequilibrium score (LDSC) regression46. The LDSC analysis indicated minimal confounding bias, as evidenced by a genomic inflation factor (λGC) of 1.07, an intercept of 1.02 (SE = 0.003) close to unity, and a low inflation ratio (0.22, SE = 0.03). The SNP-based heritability estimated on the observed scale was 0.78% (SE = 0.04%), which was highly significant (Z = 21.09, P-value = 9.41 × 10⁻⁹⁹), further supporting a polygenic basis for ovarian cancer.
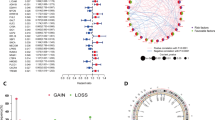
The conditional analysis using GCTA-COJO uncovered a total of 12 independent genome-wide significant signals, two of which have not been previously reported47 (Fig. 2a, Table 1). Functional annotation using RegulomeDB v248 revealed the previously unidentified variant rs77247401 with moderate likelihood to be regulatory (rank = 4, score = 0.609), and the variant rs56159231 with high likelihood to be a functional variant (rank=1 f, score=0.554). A RegulomeDB rank score of 1 f, reflecting strong functional genomic support. Rs77247401 (C > CTG) resides in the regulatory region of the lncRNA RP11-89M16.1 and disrupts binding sites for the glucocorticoid receptor and Nkx2 transcription factors, implicating altered hormonal signaling and developmental pathways in ovarian carcinogenesis annotated by HaploReg49,50. Besides, it has high linkage disequilibrium (LD r² = 0.95, D = 0.98) with rs10088218, a SNP reported to be associated with ovarian cancer in GWAS. The intronic variant rs56159231 may influence ovarian cancer risk by disrupting Erα and TCF11::MafG binding sites, modulating PLEKHM1-mediated autophagy.
a Manhattan plot of the GWAS meta-analysis for ovarian cancer. The analysis included 490,608 individuals (29,066 cases and 461,542 controls). The x-axis shows genomic positions, and the y-axis displays −log₁₀ P values. The red horizontal line indicates the genome-wide significance threshold (P-value < 5 × 10⁻⁸). Lead variants surpassing this threshold are annotated with their nearest genes. b Forest plot of MR associations between multi-omics GLP-1R instruments and overall ovarian cancer risk. Genetic instruments were derived from molecular quantitative trait loci across multiple layers: DNA methylation (meQTL, N = 32,851), gene expression (eQTL, N = 174–29,621), alternative splicing (sQTL, N = 305), and protein abundance (pQTL, N = 35,329–35,431). Ovarian cancer summary statistics were obtained from a GWAS meta-analysis (N = 490,608). Causal estimates were generated using the Wald ratio or inverse-variance weighted (IVW) method. Points represent the causal estimate for the odds ratio (OR), and error bars indicate 95% confidence intervals. All P-values were derived from two-sided Wald ratio or inverse-variance weighted tests. No associations in this panel met the Bonferroni correction threshold (P-value < 0.0125). See Supplementary Data 8 for detailed results. c Forest plot of MR associations between GLP-1R expression in pancreas and ovarian cancer subtypes. Type 2 diabetes (T2D; 80,154 cases and 853,816 controls) and body mass index (BMI; N = 532,396) were included as positive controls. Causal estimates were derived using the Wald ratio method. Points represent the causal estimate for the OR for ovarian cancer subtypes and T2D and the causal estimate for the beta coefficient for BMI; error bars represent 95% confidence intervals. All P-values were derived from two-sided Wald ratio tests. Sample sizes for ovarian cancer subtypes were as follows: high-grade serous (N = 53,978), low-grade serous (N = 41,953), endometrioid (N = 43,751), mucinous (N = 42,358), and clear cell (N = 42,307). Associations surviving Bonferroni correction (P-value < 0.0125) are indicated in blue. See Supplementary Data 8 and 9 for full cohort details.
The SNP2GENE module of Functional Mapping and Annotation (FUMA)51 provided extensive functional annotation for SNPs in genomic areas identified by lead SNPs. The variant rs6755777 (locus 2q31.1) was predicted to have a potentially deleterious effect (CADD score > 12.37)52. Phenome-wide association analysis with GWAS catalog revealed extensive pleiotropy between risk loci and various traits. Notably, several loci demonstrated significant genetic correlations with metabolic traits relevant to GLP-1R agonist response (Supplementary Data 3). Locus 9q34.2, which encompasses the ABO gene previously established in both ovarian cancer and diabetes, exhibited particularly broad pleiotropy, associating with ten GLP-1R agonist-related traits including type 2 diabetes, lipid parameters, and blood pressure. Similarly, loci implicated in body composition (locus 10p12.31) and glycemic traits (locus 15q26.1) demonstrated specific associations with relevant metabolic phenotypes. These findings provide convergent genetic evidence for shared biological pathways between metabolic regulation and ovarian cancer susceptibility.
Comprehensive enrichment analyses across tissue-specific expression and biological pathways with GENE2FUNC51 module of FUMA further delineated the functional landscape of the mapped genes of ovarian cancer in SNP2GENE. Tissue-specific expression analysis revealed significant enrichment of associated genes in metabolically active tissues, most prominently in the kidney cortex (Padj-value = 6.30×10⁻⁵), with nominally significant enrichment in the gastrointestinal tract (transverse colon, Padj-value = 0.03). Although visceral omental adipose tissue showed only nominal association (Padj-value = 0.23), this collective pattern suggests a broader metabolic context for genetic susceptibility (Supplementary Fig. 3). Pathway analysis further uncovered strong enrichment in fundamental developmental processes, with the most significant signal in embryonic skeletal system development (Padj-value = 2.11 × 10⁻⁷)—a pathway involving multiple HOX gene family members (HOXD10, HOXD9, HOXD4, HOXD1, and the HOXB1-B6 cluster). Parallel investigation of regulatory mechanisms revealed significant involvement of epigenetic pathways, including targets of the PRC2 (Padj-value = 0.0078) and genomic regions overlapping breast cancer amplicons (17q21/q25; Padj-value = 1.11×10⁻⁹; Supplementary Data 4). These findings establish that ovarian cancer genetic architecture is shaped by core developmental programs and conserved epigenetic mechanisms, pointing to fundamental processes of cell fate determination as key modifiers of disease risk.
Druggable genome and drug repurposing potential
Leveraging the Citeline PharmaProjects database (accessed October 2025), we identified two genes among the 12 candidate effector genes for ovarian cancer that are targeted by existing drug development programs. These programs encompassed 79 unique pharmacological agents, 61 of which have anti-cancer indications (Supplementary Data 5). Among these, TERT emerged as the most prominent target, supported by multiple clinical-stage candidates, such as the immuno-oncology vaccine UV-1 (Phase II) and the direct telomerase inhibitor ateganosine (Phase II). The microtubule-associated protein tau gene (MAPT), although primarily linked to neurological disorders, is also under investigation in oncology through the tau aggregation inhibitor LAM-226, currently in active preclinical development, suggesting an alternative mechanistic direction for therapeutic exploration. This structured druggability assessment thus defines a therapeutically actionable gene set originating from ovarian cancer genetic risk loci, providing a rational foundation for targeted drug development and repurposing strategies.
Summary of instrument selection of GLP-1R expression
Genetic instruments for GLP-1R expression were selected from 15 studies that estimated associations between genetic variants and molecular phenotypes related to GLP-1R in individuals of European ancestry. These phenotypes included DNA methylation (N = 32,851)24, gene expression (N ranging from 174 to 29,621)25,26,27, alternative splicing events (N = 305)28 and protein levels (N ranging from 35,329 to 35,431)29,30,31,32,33,34,35,36,37,38, which these variants were noted as molecular quantitative trait loci (QTLs). Given the endocrine-related functions of GLP-1R, we focused on molecular QTLs in tissues, such as the hypothalamus, pituitary, thyroid, small intestine, pancreas, whole blood, and plasma. We selected multiple molecular phenotypes here since the association across multiple independent datasets can strengthen the robustness of the findings.
Instruments were selected based on minor allele frequency (MAF) > 1%, F-statistic > 10, and being robustly associated with molecular phenotypes of GLP-1R (P-value < 5×10−8). Linkage disequilibrium (LD) based clumping (LD r2 < 0.001 from 1000 Genome Europeans) was applied to retain independent instruments. After selection, eight cis-acting QTLs and two trans-acting pQTLs passed the stringent selection process and were designated as primary instruments (Supplementary Data 6). In addition, a clear trend was observed where QTLs from the deCODE, GoDMC, and eQTLGen consortia demonstrated higher F-statistics (sample sizes: 23,482–35,431; F-statistics: 54–162) compared to QTLs from GTEx v8 (sample sizes: 237–574; F-statistics: 14–28). The sex-specific analysis using GTEx v8 cis-eQTL data found no evidence to support sex heterogeneity in the genetic regulation of GLP-1R expression in endocrine tissues54 (Supplementary Fig. 4, Supplementary Data 7). None of the tested genetic instruments reached the significance threshold for a G × Sex interaction (qval ≤ 0.25). For instance, in key tissues, such as the pancreas (rs880347; G × Sex qval = 1.00) and thyroid (rs10305420; G × Sex qval = 1.00) the genetic effects on GLP-1R expression were not significantly different between females and males. Furthermore, all instruments in these tissues exhibited consistent direction of effects in males and females. The stability of these genetic effects across sexes provides robust evidence that the selected genetic instruments are not subject to substantial sex-specific bias.
Association of GLP-1R expression on positive control and ovarian cancer outcomes
For the positive control analyses, MR showed protective effects of GLP-1R expression on both T2D and BMI when using eQTL and sQTL in pancreas and eQTL in thyroid as instruments (Supplementary Data 8).
The MR results assessing the effect of GLP-1R expression on ovarian cancer and its subtypes are presented in Fig. 2b,c. Utilizing meta-analyzed ovarian cancer GWAS, we found a marginal protective effect of GLP-1R expression on overall ovarian cancer risk (odds ratio [OR] of 0.94 (95% confidence interval [CI]: 0.89–1.00) per standard deviation increase in GLP-1R expression in pancreas). In histotype-specific analyses, a protective effect of GLP-1R expression on ENOC was identified (OR = 0.83, 95% CI: 0.72–0.95) per standard deviation increase in GLP-1R expression in pancreas). In addition, an OR of 0.13 (95% CI: 0.02–0.86) was observed for ENOC per standard deviation increase in intron excision ratios of GLP-1R, a proxy for alternative splicing in pancreas. For CCOC, protective effects were observed for DNA methylation levels of CpG sites cg05924652 (OR = 0.48, 95% CI: 0.23–0.97) and cg25014318 (OR = 0.33, 95% CI: 0.12–0.96; Supplementary Data 9). Only the association of GLP-1R expression, proxied by eQTL in the pancreas, with reduced ENOC risk remained robust after applying a Bonferroni-corrected P-value threshold of 0.0125. The colocalization analysis between GLP-1R expression and ovarian cancer signals in the cis-acting region of GLP-1R showed a colocalization probability of 93.5% under the condition of a robust signal in the exposure data (Supplementary Data 9, Supplementary Fig. 5).
Mediation effect of GLP-1R expression, mediators and ovarian cancer subtypes
For those ovarian cancer subtypes that showed robust evidence in the main MR analysis, we next performed mediation MR analysis to identify putative mediators underlying this subtype-specific relationship. An initial phenome-wide screening of 2,204 candidate mediators was conducted using the “MR-EVE” function implemented in the EpiGraphDB platform, which identified 347 phenotypes at least marginally associated with the ovarian cancer subtypes (MR P-value < 0.05)58. We further applied a set of inclusion criteria, including (1) the GWAS being conducted in participants of European ancestry; (2) with female participants; (3) excluding phenotypes with overlapped samples from OCAC, and (4) without MR estimates. A total of 256 phenotypes were selected as potential mediators and included in the follow-up two-step MR analyses (Fig. 3a, Supplementary Fig. 1, Supplementary Data 10).
a Phenome-wide association scan for endometrioid ovarian cancer. The Manhattan plot displays associations between genetic instruments for GLP-1R expression and 2204 phenotypes, categorized by domain. The y-axis shows the strength of association (−log₁₀ P value) based on Mendelian randomization analysis results from EpiGraphDB. Red and blue horizontal lines indicate false discovery rate (FDR) thresholds of P-value < 0.05 and the genome-wide significance level (P-value < 5 × 10⁻⁸), respectively. b Schematic of significant mediators in the GLP-1R–endometrioid ovarian cancer pathway. The diagram summarizes mediators identified through two-step MR analysis, showing the proportion mediated for each. All mediated proportions were statistically significant after FDR correction (P-value < 0.05). c Association of candidate mediators with ovarian cancer histotypes. Forest plot displays MR estimates for six mediators—body mass index, waist circumference, hip circumference, arm fat mass, arm predicted mass, and 18:2 linoleic acid—across five ovarian cancer subtypes: high-grade serous (HGSOC), low-grade serous (LGSOC), mucinous (MOC), endometrioid (ENOC), and clear cell (CCOC). Points represent the causal estimate for the odds ratio (OR) derived from inverse-variance weighted MR analysis; error bars indicate 95% confidence intervals. All P-values were derived from two-sided inverse-variance weighted tests. Sample sizes for mediator–outcome analyses ranged from 41,953 to 462,117 individuals across phenotype combinations. Associations with an FDR < 0.05 were considered statistically significant after correction for multiple comparisons. Associations with the ENOC subtype that were significant after FDR correction for multiple testing are highlighted in blue.
In step one of the Mediation MR analysis, we estimated the association of GLP-1R expression (as exposure), proxied by eQTL in the pancreas, with the 256 candidate mediators (as outcomes). Of these, GLP-1R expression was associated with 32 phenotypes (MR P-value < 0.05). These phenotypes could be classified into five categories: anthropometric or bioimpedance measurements, metabolic and biochemical measurements, bone and density measurements, vascular and circulatory system indicators, and other factors. In step two, 24 of the 32 candidate mediators (treated as exposures) were found to be associated with ENOC (as outcome), where the exposure-mediator, mediator-outcome and exposure-outcome MR estimates showed consistent direction of effects. Full results of the two-step MR analyses were shown in Supplementary Data 11.
Among the 24 potential mediators, BMI, waist circumference, hip circumference, arm fat mass and arm predicted mass,18:2 linoleic acid (LA) showed additional evidence to support their mediation role linking GLP-1R expression with ENOC (Table 2, Supplementary Data 12–13). The causal inference for the six identified mediators was robust to multiple testing, as associations in the two-step MR analysis remained significant after false discovery rate (FDR) correction for both the exposure-mediator and mediator-outcome stages. The additional evidence was obtained from a set of MR sensitivity analyses methods, where at least one of these methods showed same direction of effect as the main MR estimate (using inverse variance weighted [IVW]) (Supplementary Data 12). We found no evidence to support the presence of horizontal pleiotropy, as indicated by the MR-Egger intercept test (P-value > 0.05), or between-instrument heterogeneity (Cochran’s Q P-values > 0.05). The mediation analysis revealed that the six identified mediators collectively explained a substantial proportion of the total effect of GLP-1R expression on endometrioid ovarian cancer. Linoleic acid (18:2 LA) was the most influential mediator, accounting for 7.7% of the effect, followed by arm predicted mass (2.8%), hip circumference (2.3%), arm fat mass (2.0%), waist circumference (1.8%), and BMI (1.5%) (Fig. 3b).
Sensitivity analysis and external validation of mediation pathway
Additional sensitivity analyses confirmed that the identified mediation pathways related to body composition were not substantially biased by sex-specific effects (Supplementary Figs. 6–8, Supplementary Data 14–17). The female-specific two-step MR analysis corroborated the direction and nature of our primary findings, despite reduced statistical power due to smaller sample sizes. While the associations between GLP-1R expression and the female-specific body composition mediators were directionally consistent with the primary analysis. Crucially, the effects of these female-specific mediators on ENOC risk were robust, with BMI, hip circumference, waist circumference, and arm fat mass all showing positive associations following FDR correction (e.g. BMI IVW OR = 1.36, P-value = 0.002; Supplementary Data 15). Second, the MRSamePopTest63 demonstrated no significant sex-based heterogeneity for any of the five body composition mediators (All P-value > 0.05; Supplementary Data 17). The minimal differences in genetic effect estimates (Beta difference range: 0.001-0.003) between female and sex-combined data for the exposure-mediator relationships indicate that sex is not a major effect modifier at this step, validating the use of sex-combined data for these mediators in our primary analysis.
Mediation analysis with weak instruments established 18:2 LA as the primary mediator through which GLP-1R expression influences endometrioid ovarian cancer risk (Supplementary Data 18–19). To address pleiotropy and formally test mediation, we performed cis-MVMR using 181 pancreatic GLP-1R cis-eQTLs (P-value < 0.05)64 (Supplementary Fig. 9, Supplementary Data 18). Robust PC-GMM analysis confirmed GLP-1R expression significantly reduces ENOC risk (OR = 0.86, 95% CI: 0.82–0.91; P-value < 0.001) and negatively associated with all six mediators (all P-value < 0.001, remaining significant after FDR correction at q < 0.05). However, robust MVPC-GMM analysis showed little evidence to support the independent mediator effects, with conditional F statistics <1 indicating weak instrument bias. The significance for the mediator effect may have been masked by the complexity of the model. The MR-Bayesian Model Averaging (MR-BMA)65 analysis was applied to overcome this analytical challenge. The results, robustly ranked 18:2 LA as the top mediator (posterior probability = 0.38), followed by arm fat mass (posterior Probability = 0.20) and body mass index (posterior probability = 0.19) (Supplementary Data 19). This finding is consistent with our initial two-step MR results and provides strong evidence that 18:2 LA is the most likely primary mediator through which GLP-1R expression influences endometrioid ovarian cancer risk.
Evidence from literature search validated the association of the five body composition traits with ENOC67,68,69. In contrast, 18:2 LA was identified as a previously unreported mediator of ovarian cancer. Furthermore, MELODI Presto41 highlighted a similar mediation pathway by mining literature evidence, wherein GLP-1R agonists modulated obesity-related factors that contributed to the risk of ENOC (Supplementary Data 20).
We further validated whether the six ENOC related mediators were associated with other ovarian cancer subtypes, and found little evidence to support, such association (Fig. 3c, Supplementary Data 21). Only few exceptions were observed, including hip circumference and arm fat mass marginally associated with HGSOC (OR for hip circumference = 1.19, 95% CI 1.07–1.34, P-value = 0.01; OR for arm fat mass = 1.13, 95% CI 1.00–1.27, P-value = 0.04), and 18:2 LA marginally associated with invasive MOC (OR = 1.23, 95% CI 1.01–1.50, P-value = 0.04).
Discussion
The heterogeneity of ovarian cancer subtypes poses a significant challenge for therapeutic treatment. In this study, we demonstrated that GLP-1R expression in pancreas has a potential protective effect on overall ovarian cancer risk. Subtype analysis showed robust evidence that GLP-1R expression in pancreas is associated with a reduced risk of ENOC. Mediation analyses further identified six anthropometric and metabolic traits—such as BMI, waist circumference, and 18:2 LA—as key mediators of this protective effect. These findings highlight the potential of GLP-1R as a modulator of ENOC risk, suggesting that GLP-1R agonists may offer a therapeutic strategy for ENOC through their effects on body weight and lipid metabolism.
Beyond our focus on the effect of GLP-1R expression on ovarian cancer, our GWAS meta-analysis provides a broader architectural context for ovarian cancer susceptibility. The polygenic risk is significantly enriched in fundamental developmental programs and conserved epigenetic mechanisms, illustrating a complex genetic landscape where pathways of cell fate determination contribute to disease risk. This finding is consistent with the established concept that cancer cells recapitulate developmental processes, such as the epithelial-to-mesenchymal transition, which is essential for embryonic morphogenesis and is reinstated in carcinomas to facilitate invasion and metastasis70. Importantly, this genetic landscape also exhibits extensive pleiotropy with metabolic traits. This aligns with large-scale epidemiological evidence demonstrating a substantial global cancer burden attributable to metabolic syndrome71,72. Together, these insights create a convergent biological backdrop that reinforces the specific, protective role of GLP-1R-mediated metabolic homeostasis identified in our MR analyses. This integration of discovery-based GWAS and causal inference frameworks offers a more comprehensive etiological model for ovarian cancer.
Previous studies provided inconclusive evidence on the effect of GLP-1R agonists on ovarian cancer, largely due to the absence of long-term RCTs with enough number of ovarian cancer cases14,15,16,17. The association with ovarian cancer we identified aligns with the study showing a favorable association among women with T2D13. Recently, Sun et al. used a two-sample MR approach to investigate the association between GLP-1R agonism and cancer risk, concluding that GLP-1R agonism does not increase ovarian cancer risk20. However, a few technique issues existed in this study. First, the inclusion of 22 highly correlated genetic instruments with almost perfect LD violates the assumption of instrument variable analysis, which the instrument needs to be independent. This leads to over-precise MR estimates. Second, the instruments were only extracted from eQTLs in blood samples, neglecting tissue-specific effects of GLP-1R. Third, no positive control analysis for body weight was performed. Fourth, discrepancies in outcome data—such as mismatched sample sizes and overlapping datasets between discovery and replication phases due to FinnGen—compromised the validity of their conclusions. Fifth, issues in the two-sample MR harmonization process, particularly the incorrect assumption that all alleles are on the forward strand. Using the right strand information, we were not able to replicate the positive-control effect of GLP-1R on T2D using their instruments. In contrast, our study selected robust genetic instruments and identified a protective effect of GLP-1R expression on ovarian cancer and its subtypes.
Ovarian cancer subtypes are characterized by distinct pathogenesis, molecular features, and responses to standard chemotherapy5,21,73,74. The heterogeneity of EOC poses a significant challenge for treatment, as adjuvant therapy is typically guided by tumor grade and stage rather than histological subtype74. Previous studies have linked ENOC to endometriosis, suggesting that reduced GLP-1 levels in peritoneal fluid may contribute to endometriosis development75,76,77. In consistent with these studies, we showed that GLP-1R expression exerts distinct effects on ovarian cancer subtypes, with a particularly protective role in ENOC. In addition, most risk factors exhibit considerable heterogeneity across histologic subtypes of EOC4. Consistently, our MR analyses showed that mediators associated with ENOC were unlikely to influence other invasive EOC subtypes.
Our study identified several key mediators underlying the relationship between GLP-1R expression and ENOC. First, the use of GLP-1R agonists for weight management has been expanding rapidly78. Our results underscore the central role of weight loss in mediating the protective effect of GLP-1R expression on ENOC. Second, 18:2 LA emerged as an alternative mediator that was not reported by any existing study. LA, an essential polyunsaturated fatty acid obtained primarily from diet, plays a critical role in lipid metabolism and obesity79. Excessive LA intake can contribute to obesity-related conditions, such as fat accumulation, inflammation, and fatty liver disease80,81,82. The recent discovery that dietary LA is directly sensed through an FABP5-mTORC1 signaling axis to drive cancer progression provides a compelling mechanistic context for our finding83. This evidence establishes LA not merely as a nutrient but as a potent bioactive molecule capable of directly activating a key oncogenic pathway. Therefore, we hypothesize that the protective effect of GLP-1R signaling may operate by counteracting this tumor-promoting pathway identified in this study. GLP-1R activation may either reduce the bioavailability of LA for FABP5 binding or modulate downstream metabolic processes that ultimately suppress mTORC1 hyperactivation. Our genetic data thus position GLP-1R signaling as a critical upstream regulator of a pathogenic lipid-sensing pathway, shifting the focus from LA intake to its downstream signaling consequences.
Our study has several limitations. First, the estimated effects should be interpreted cautiously. The MR estimates reflect the long-term effects of GLP-1R expression on outcome risk. They do not capture the short-term effects of GLP-1R agonists in individuals with T2D or obesity. Second, as the primary datasets for both the exposure and mediators included male participants, our analysis necessarily assumed homogeneous effects across sexes. Although a prior study reported generally minimal sex differences in eQTLs, which supports the reliability of our instruments54. We also directly evaluated this assumption through sensitivity analyses. These analyses confirmed the absence of significant sex-specific bias in either our genetic instruments or the resulting causal estimates. Nevertheless, future studies with access to larger, sex-stratified datasets would be valuable to further investigate this aspect. Third, our study estimated the effects of GLP-1R expression only in a European population. Future studies should investigate the effects of GLP-1R agonists in populations with diverse ancestries and disease statuses. While we employed Steiger filtering to test the direction of causation as a sensitivity analysis, we note that its statistical power can be suboptimal when binary outcomes are involved. Therefore, the primary evidence for directionality in our cis-MR framework rests not on this statistical test, but on the strong prior biological knowledge that GLP-1R signaling influences downstream metabolic traits and cancer risk, and not vice versa. Finally, the statistical power to detect effects was limited for certain outcomes, particularly those with insufficient numbers of cases (e.g. LGSOC and CCOC), and thus we observed some wide confidence intervals for some of the MR estimates.
In conclusion, our findings provide evidence that GLP-1R expression is associated with a reduced risk of ENOC, partially mediated through its effects on body weight and lipid metabolism. The identification of this fatty acid pathway, robust to multiple sensitivity checks, offers an alternative mechanistic hypothesis beyond weight regulation alone. Further validation through clinical trials and exploration of the underlying mechanisms will be essential to fully understand the therapeutic potential of GLP-1R agonists in preventing ovarian cancer. Elucidating these modifiable factors could inform clinical prevention strategies and therapeutics for ovarian cancer, while also explaining the heterogeneity observed across ovarian cancer subtypes.
Data availability
The data of molecular traits used to proxy GLP-1R expression were available via deCODE genetics website (https://www.decode.com/summarydata/), the eQTL Gen website (https://www.eqtlgen.org), the GoDMC website (http://mqtldb.godmc.org.uk), the MetaBrain website (https://metabrain.nl/) and the Genotype-Tissue Expression (GTEx) Portal website (https://www.gtexportal.org). The summary statistics of ovarian cancer for GWAS meta-analyses used in this study are deposited in the IEU OpenGWAS database (GWAS IDs ieu-a-1120 and ieu-b-4963) and FinnGen r12 defined as II Neoplasms, from cancer register (ICD-O-3) (https://r12.finngen.fi/pheno/C3_OVARY_EXALLC). The GWAS summary statistics for the ovarian cancer subtypes are available at https://gwas.mrcieu.ac.uk with their GWAS IDs listed in the Supplementary Data. The GWAS summary statistics for positive control outcomes are downloaded from the GWAS Catalog (https://www.ebi.ac.uk/gwas/) and the accession codes are as follows: body mass index (GCST90029007) and type 2 diabetes (GCST90018926). The GWAS summary statistics for mediators were derived from EpiGraphDB database (https://www.epigraphdb.org/). No additional permissions or applications are required to access these datasets. The drug information utilized in this study was obtained from the Citeline database (https://clinicalintelligence.citeline.com/). Full access to the Citeline platform is commercially licensed and not freely available to all researchers. The source data for Figs. 1, 2 and 3 are available in Supplementary Data.
Code availability
The MR pipeline assessing the effect of GLP-1R expression on ovarian cancer is publicly available on the Omics Harbor GitHub repository (https://github.com/geneinmylife/GLP1R-OC-project) and has been permanently archived on Zenodo (https://doi.org/10.5281/zenodo.17994065).
References
Ajayi, A. F., Oyovwi, M. O., Akano, O. P., Akanbi, G. B. & Adisa, F. B. Molecular pathways in reproductive cancers: a focus on prostate and ovarian cancer. Cancer Cell Int. 25, 33 (2025).
Webb, P. M. & Jordan, S. J. Global epidemiology of epithelial ovarian cancer. Nat. Rev. Clin. Oncol. 21, 389–400 (2024).
Cho, K. R. & Shih, I.-M. Ovarian cancer. Annu. Rev. Pathol. 4, 287–313 (2009).
Phelan, C. M. et al. Identification of 12 new susceptibility loci for different histotypes of epithelial ovarian cancer. Nat. Genet. 49, 680–691 (2017).
Wentzensen, N. et al. Ovarian cancer risk factors by histologic subtype: an analysis from the ovarian cancer cohort consortium. J. Clin. Oncol. 34, 2888–2898 (2016).
Lincoff, A. M. et al. Semaglutide and cardiovascular outcomes in obesity without diabetes. N. Engl. J. Med. 389, 2221–2232 (2023).
Wharton, S. et al. Daily Oral GLP-1 receptor agonist orforglipron for adults with obesity. N. Engl. J. Med. 389, 877–888 (2023).
Srivastava, G. & Kumar, R. B. Once-weekly semaglutide in adults with overweight or obesity. N. Engl. J. Med. 385, e4 (2021).
American Diabetes Association Primary Care Advisory Group Standards of Care in Diabetes—2025 Abridged for Primary Care. Clin. Diab. 43, 182 (2025).
Ibrahim, S. S. et al. The effect of GLP-1R agonists on the medical triad of obesity, diabetes, and cancer. Cancer Metastasis Rev. 43, 1297–1314 (2024).
Pasternak, B. et al. Glucagon-like peptide 1 receptor agonist use and risk of thyroid cancer: scandinavian cohort study. BMJ 385, e078225 (2024).
Zheng, Z. et al. Glucagon-like peptide-1 receptor: mechanisms and advances in therapy. Signal Transduct. Target Ther. 9, 234 (2024).
Wang, L., Xu, R., Kaelber, D. C. & Berger, N. A. Glucagon-like peptide 1 receptor agonists and 13 obesity-associated cancers in patients with type 2 diabetes. JAMA Netw. Open 7, e2421305 (2024).
Montastruc, J.-L. GLP 1 receptor agonists and obesity-associated cancers: a disproportionality analysis in Vigibase®. Eur. J. Clin. Pharm. 80, 1999–2001 (2024).
Pratley, R. et al. Oral semaglutide versus subcutaneous liraglutide and placebo in type 2 diabetes (PIONEER 4): a randomised, double-blind, phase 3a trial. Lancet 394, 39–50 (2019).
Marso, S. P. et al. Liraglutide and cardiovascular outcomes in type 2 diabetes. N. Engl. J. Med. 375, 311–322 (2016).
O’Neil, P. M. et al. Efficacy and safety of semaglutide compared with liraglutide and placebo for weight loss in patients with obesity: a randomised, double-blind, placebo and active controlled, dose-ranging, phase 2 trial. Lancet 392, 637–649 (2018).
Smith, G. D. & Ebrahim, S. Mendelian randomization’: can genetic epidemiology contribute to understanding environmental determinants of disease? Int. J. Epidemiol. 32, 1–22 (2003).
Zheng, J. et al. Recent developments in mendelian randomization studies. Curr. Epidemiol. Rep. 4, 330–345 (2017).
Sun, Y. et al. Association of glucagon-like peptide-1 receptor agonists with risk of cancers-evidence from a drug target Mendelian randomization and clinical trials. Int. J. Surg. 110, 4688–4694 (2024).
Yarmolinsky, J. et al. Genetically proxied glucose-lowering drug target perturbation and risk of cancer: a Mendelian randomisation analysis. Diabetologia 66, 1481–1500 (2023).
Riedinger, C. J. et al. Glucagon-like peptide-1 (GLP-1) receptor agonists for weight management: a review for the gynecologic oncologist. Gynecol. Oncol. 190, 1–10 (2024).
Carter, A. R. et al. Mendelian randomisation for mediation analysis: current methods and challenges for implementation. Eur. J. Epidemiol. 36, 465–478 (2021).
Garrido-Martín, D., Borsari, B., Calvo, M., Reverter, F. & Guigó, R. Identification and analysis of splicing quantitative trait loci across multiple tissues in the human genome. Nat. Commun. 12, 727 (2021).
GTEx Consortium The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Võsa, U. et al. Large-scale cis- and trans-eQTL analyses identify thousands of genetic loci and polygenic scores that regulate blood gene expression. Nat. Genet. 53, 1300–1310 (2021).
de Klein, N. et al. Brain expression quantitative trait locus and network analyses reveal downstream effects and putative drivers for brain-related diseases. Nat. Genet. 55, 377–388 (2023).
Min, J. L. et al. Genomic and phenotypic insights from an atlas of genetic effects on DNA methylation. Nat. Genet. 53, 1311–1321 (2021).
Robins, C. et al. Genetic control of the human brain proteome. Am. J. Hum. Genet. 108, 400–410 (2021).
Wingo, T. S. et al. Brain proteome-wide association study implicates novel proteins in depression pathogenesis. Nat. Neurosci. 24, 810–817 (2021).
Sun, B. B. et al. Genomic atlas of the human plasma proteome. Nature 558, 73–79 (2018).
Hansson, O. et al. The genetic regulation of protein expression in cerebrospinal fluid. EMBO Mol. Med. 15, e16359 (2022).
Sun, B. B. et al. Plasma proteomic associations with genetics and health in the UK Biobank. Nature 622, 329–338 (2023).
Pietzner, M. et al. Mapping the proteo-genomic convergence of human diseases. Science 374, eabj1541 (2021).
Ferkingstad, E. et al. Large-scale integration of the plasma proteome with genetics and disease. Nat. Genet. 53, 1712–1721 (2021).
Zhang, J. et al. Plasma proteome analyses in individuals of European and African ancestry identify cis-pQTLs and models for proteome-wide association studies. Nat. Genet. 54, 593–602 (2022).
Yang, C. et al. Genomic atlas of the proteome from brain, CSF and plasma prioritizes proteins implicated in neurological disorders. Nat. Neurosci. 24, 1302–1312 (2021).
Western, D. et al. Proteogenomic analysis of human cerebrospinal fluid identifies neurologically relevant regulation and implicates causal proteins for Alzheimer’s disease. Nat. Genet. 56, 2672–2684 (2024).
Mitchell, R. et al. UK Biobank Genetic Data: MRC-IEU Quality Control, version 2. University of Bristol https://doi.org/10.5523/BRIS.1OVAAU5SXUNP2CV8RCY88688V (2019).
Kurki, M. I. et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 613, 508–518 (2023).
Elsworth, B. & Gaunt, T. R. MELODI Presto: a fast and agile tool to explore semantic triples derived from biomedical literature. Bioinformatics 37, 583–585 (2021).
Skrivankova, V. W. et al. Strengthening the reporting of observational studies in epidemiology using mendelian randomisation (STROBE-MR): explanation and elaboration. BMJ 375, n2233 (2021).
Skrivankova, V. W. et al. Strengthening the reporting of observational studies in epidemiology using mendelian randomization: the STROBE-MR statement. JAMA 326, 1614–1621 (2021).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Winkler, T. W. et al. Quality control and conduct of genome-wide association meta-analyses. Nat. Protoc. 9, 1192–1212 (2014).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Dong, S. et al. Annotating and prioritizing human non-coding variants with RegulomeDB v.2. Nat. Genet. 55, 724–726 (2023).
Ward, L. D. & Kellis, M. HaploReg v4: systematic mining of putative causal variants, cell types, regulators and target genes for human complex traits and disease. Nucleic Acids Res. 44, D877–D881 (2016).
Ward, L. D. & Kellis, M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 40, D930–D934 (2012).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
Rentzsch, P., Witten, D., Cooper, G. M., Shendure, J. & Kircher, M. CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res. 47, D886–D894 (2019).
Zheng, J. et al. Phenome-wide Mendelian randomization mapping the influence of the plasma proteome on complex diseases. Nat. Genet. 52, 1122–1131 (2020).
Oliva, M. et al. The impact of sex on gene expression across human tissues. Science 369, eaba3066 (2020).
Loh, P.-R., Kichaev, G., Gazal, S., Schoech, A. P. & Price, A. L. Mixed-model association for biobank-scale datasets. Nat. Genet. 50, 906–908 (2018).
Sakaue, S. et al. A cross-population atlas of genetic associations for 220 human phenotypes. Nat. Genet. 53, 1415–1424 (2021).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. Elife 7, e34408 (2018).
Liu, Y. et al. EpiGraphDB: a database and data mining platform for health data science. Bioinformatics 37, 1304–1311 (2021).
Davies, N. M., Holmes, M. V. & Davey Smith, G. Reading Mendelian randomisation studies: a guide, glossary, and checklist for clinicians. BMJ 362, k601 (2018).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet 10, e1004383 (2014).
An efficient and robust tool for colocalisation: Pair-wise Conditional and Colocalisation (PWCoCo) | bioRxiv. https://doi.org/10.1101/2022.08.08.503158v1.
Davey Smith, G. & Hemani, G. Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum. Mol. Genet. 23, R89–R98 (2014).
Woolf, B. et al. MRSamePopTest: introducing a simple falsification test for the two-sample mendelian randomisation ‘same population’ assumption. BMC Res Notes 17, 27 (2024).
Patel, A. et al. Robust use of phenotypic heterogeneity at drug target genes for mechanistic insights: Application of cis-multivariable Mendelian randomization to GLP1R gene region. Genet. Epidemiol. 48, 151–163 (2024).
Zuber, V., Colijn, J. M., Klaver, C. & Burgess, S. Selecting likely causal risk factors from high-throughput experiments using multivariable Mendelian randomization. Nat. Commun. 11, 29 (2020).
Zheng, J. & Liu, J. Mendelian randomization study of GLP-1R effects on ovarian cancer subtypes mediated by metabolic factors. Zenodo https://doi.org/10.5281/zenodo.17994065 (2025).
Chen, Y. et al. Plasma metabolites and risk of seven cancers: a two-sample Mendelian randomization study among European descendants. BMC Med. 22, 90 (2024).
Feng, Y. et al. Causal effects of genetically determined metabolites on cancers included lung, breast, ovarian cancer, and glioma: a Mendelian randomization study. Transl. Lung Cancer Res. 11, 1302–1314 (2022).
Freuer, D. et al. Body fat distribution and risk of breast, endometrial, and ovarian cancer: a two-sample mendelian randomization study. Cancers 13, 5053 (2021).
Nieto, M. A. Epithelial plasticity: a common theme in embryonic and cancer cells. Science 342, 1234850 (2013).
Xing, Q.-Q. et al. Global burden of common cancers attributable to metabolic risks from 1990 to 2019. Medicine 4, 168–181 (2023).
Wang, H. et al. DLAT is involved in ovarian cancer progression by modulating lipid metabolism through the JAK2/STAT5A/SREBP1 signaling pathway. Cancer Cell Int. 25, 25 (2025).
Kuroki, L. & Guntupalli, S. R. Treatment of epithelial ovarian cancer. BMJ 371, m3773 (2020).
Chen, S. et al. A review of the clinical characteristics and novel molecular subtypes of endometrioid ovarian cancer. Front. Oncol. 11, 668151 (2021).
Guidozzi, F. Endometriosis-associated cancer. Climacteric 24, 587–592 (2021).
Wang, L. et al. Endometriosis and epithelial ovarian cancer: a two-sample Mendelian randomization analysis. Sci. Rep. 13, 21992 (2023).
Krasnyi, A. M. et al. The levels of ghrelin, glucagon, visfatin and Glp-1 are decreased in the peritoneal fluid of women with endometriosis along with the increased expression of the CD10 protease by the macrophages. Int. J. Mol. Sci. 23, 10361 (2022).
Zheng, Z. et al. Glucagon-like peptide-1 receptor: mechanisms and advances in therapy. Sig Transduct. Target Ther. 9, 1–29 (2024).
Sanders, T. A. B. Omega-6 fatty acids and cardiovascular disease. Circulation 139, 2437–2439 (2019).
Zhu, T., Lu, X.-T., Liu, Z.-Y. & Zhu, H.-L. Dietary linoleic acid and the ratio of unsaturated to saturated fatty acids are inversely associated with significant liver fibrosis risk: a nationwide survey. Front. Nutr. 9, 938645 (2022).
Jeyapal, S. et al. Substitution of linoleic acid with α-linolenic acid or long chain n-3 polyunsaturated fatty acid prevents Western diet induced nonalcoholic steatohepatitis. Sci. Rep. 8, 10953 (2018).
Juárez-Hernández, E., Chávez-Tapia, N. C., Uribe, M. & Barbero-Becerra, V. J. Role of bioactive fatty acids in nonalcoholic fatty liver disease. Nutr. J. 15, 72 (2016).
Koundouros, N. et al. Direct sensing of dietary ω-6 linoleic acid through FABP5-mTORC1 signaling. Science 387, eadm9805 (2025).
Acknowledgements
This work was supported by grants from the Noncommunicable Chronic Diseases–National Science and Technology Major Project (2024ZD0531500, 2024ZD0531502) and the National Key Research and Development Program of China (2022YFC2505203). J.L.L., M.X., W.Q.W., Y.F.B. and G.N. are supported by the National Natural Science Foundation of China (82088102, 81970728 and 81941017) and the Shanghai Municipal Education Commission–Gaofeng Clinical Medicine Grant Support (20161307 and 20152508 Round 2). J.L.L., M.X., W.Q.W., Y.F.B. and G.N. are members of the Innovative Research Team of High-level Local Universities in Shanghai.
Author information
Authors and Affiliations
Contributions
Conceptualization: J.Z.; Formal analysis: J.J.L. and Z.H.C.; Methodology: J.Z.; Investigation: J.J.L., Z.H.C., Q.Y., H.L., S.Y.W., M.L., T.G.W., Z.Y.Z., M.X., Y.H.C., Y.X., J.L.L., and Q.H.G.; Visualization: J.J.L. and Z.H.C.; Funding acquisition: L.M.W., M.X., J.Z., J.L.L., W.Q.W., Y.F.B., and G.N.; Project administration: G.N. and W.Q.W.; Supervision: L.M.W., W.Q.W., Y.F.B., and J.Z.; Writing—original draft: J.J.L.; Writing—review and editing: Q.Y. and J.Z.; L.M.W., W.Q.W., Y.F.B., and J.Z. are the guarantors of this work and shall take responsibility for the full access and integrity of the data. All authors have approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests. The funding sources had no role in the design of the study, collection and analysis of data, interpretation of results, and decision to publish.
Peer review
Peer review information
Communications Medicine thanks Yiwen Liang, Benjamin Woolf and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, J., Chen, Z., Yang, Q. et al. Mendelian randomization study of GLP-1R effects on ovarian cancer subtypes mediated by metabolic factors. Commun Med 6, 113 (2026). https://doi.org/10.1038/s43856-026-01379-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43856-026-01379-y