Abstract
Freshwater ecosystems in densely populated areas are susceptible to plastic pollution, often reaching concentrations higher than those in marine systems. The ‘plastisphere,’ a specialized microbial community on plastic surfaces, plays a crucial role in determining macroplastic fate and ecological impact. Here, in a 12-week mesocosm experiment simulating urban freshwater systems exposed to macroplastic, we investigated factors affecting polymer-specific biofilm formation, microbial community dynamics and macroplastic settling behaviour. Biofilm growth was significantly influenced by time and plastic type, with rough surfaces (for example, PS (polysytrene) lids, HDPE (high-density polyethylene) bags) supporting higher cell densities. Biofilm development altered buoyancy, increasing sinking rates of dense (PS lids, PLA (polylactic acid (plant-based biopolymer)) cups) and those with thick biofilms (HDPE and PE (polyethylene) bags). Algae accounted for ~99% of the estimated biofilm thickness. Microbial communities in the biofilm exhibited clear temporal succession with peak abundances of bacteria, algae and cyanobacteria over time. However, no differences were detected between biodegradable and conventional plastics. Early dominance of known biofilm-forming and potential plastic-degrading bacteria declined over time, indicating limited biodegradation potential. Overall, water quality primarily shaped microbial community composition, while plastic properties governed biofilm development and settling. Most tested plastics (six of eight) sank during the experiment, highlighting freshwater systems’ vulnerability to plastic accumulation and associated water quality degradation.
Similar content being viewed by others
Main
Plastics are a major pollutant of aquatic ecosystems in the Anthropocene. The amount of plastic flowing into the ocean is expected to triple by 20401. About 80% of plastic waste is generated in terrestrial ecosystems, which is then transported into rivers, lakes and ponds before reaching the ocean, making it essential to understand the fate of plastic waste in inland waters. Inland waters not only transport plastic into the ocean but also serve as potential sinks for its accumulation2. A cross-national survey of inland waters2 showed that especially lakes and reservoirs in densely populated areas are vulnerable to plastic contamination, with concentrations even exceeding those reported in hotspot marine areas. The presence of macroplastics in aquatic ecosystems poses potential risks to the environment and human health3,4,5. Macroplastics—plastic debris larger than 5 mm—are increasingly recognized as important pollutants in freshwater systems, although the majority of research on freshwater plastics (76%) focuses on microplastics. Macroplastics are commonly household items, such as cups and straws, that are typically used and disposed in their entirety, rather than fragmented or broken-down plastic pieces6.
Formation of a specialized microbial community on plastic surfaces, known as the ‘plastisphere’, has generated considerable attention7,8,9. The plastisphere encompasses a diverse range of microorganisms, including bacteria, fungi and algae, which interact with the plastic substrate10. Microbial colonization on plastic surfaces forms biofilm matrices that support the attachment and growth of microorganisms. These biofilms have been shown to modify the physical and chemical properties of the plastic debris11 and have a potential for plastic degradation. On macroplastics, biofilm growth and biofouling modify surface characteristics and buoyancy, influencing transport pathways and bioavailability across habitats12. Over time, biofouling and weathering processes can fragment macroplastics into microplastics, thereby extending their persistence and ecological impacts within aquatic environments13. Biofilms play a major role in increasing plastic ingestion rates and can also serve as a nutritious food source for zooplankton14,15. While ingestion has often been studied in the context of microplastics, it is also a major concern for macroplastics, with numerous reports of ingestion across freshwater16,17 and marine ecosystems18. Furthermore, plastics move over long distances, facilitating microbial dispersal of invasive species and even transporting pathogenic species19,20,21. Understanding the factors that drive biofilm growth on plastic surfaces in aquatic ecosystems is therefore critical for comprehending the fate and impact of plastic debris in these ecosystems.
Multiple factors determine the microbial community composition on macroplastics. Geographic location seems to play a key role in the uniqueness of the plastisphere22,23. The microbial community on plastics has been observed to differ from the surrounding water and when compared to microbial communities on other non-plastic substrates artificial substrates such as glass21 or natural substrates such as wood24. Plastics have distinct surface characteristics, chemical nature and leach organic matter and additives, which can influence the colonizing microbial communities. Biofilm formation on microplastics has been shown to be polymer specific, with surface roughness playing an important role25. In addition, biodegradable plastics have been observed to facilitate greater colonization rates by bacteria exhibiting higher levels of cell-specific activity compared to non-biodegradable plastics26. There is growing evidence that plastic colonization can also be shaped by environmental factors, for example, temperature, dissolved oxygen and nutrients27.
Numerous studies have described the composition and diversity of the plastisphere either focusing on snapshot sampling or short-term studies, and often not focusing on environmentally realistic plastic items22,28. Furthermore, research focusing on the quantification of biofilm formation, its abundance, diversity and its implications for the fate of macroplastics, particularly in freshwater environments, is completely lacking with previous research primarily focusing on plastic colonization in marine ecosystems26,28,29,30. Freshwater plastisphere studies are, however, limited10,31,32,33, and the insights found in marine environments are not directly translatable to freshwater ecosystems9.
To address these research gaps, we aimed to investigate how commonly used intact macroplastic objects often found in aquatic ecosystems—such as cups, lids, bags and straws6,34—and their characteristics—influence microbial colonization, temporal changes in bacterial community composition and their fate in freshwater ecosystems. More specifically, the objectives of this study were: (1) to determine the formation and growth of biofilms on different commonly used macroplastic items in freshwater ecosystems with increasing exposure time; (2) to determine the temporal changes in bacterial succession and community composition on the different plastic items; (3) to investigate the influence of characteristics such as plastic item morphology, size, surface roughness, polymer identity, degradability, density and surface area on microbial community composition and diversity and (4) to estimate the implications of biofilm formation on the fate of macroplastic items.
To this end, we conducted a mesocosm experiment for 12 weeks using controlled indoor mesocosms that allowed us to mimic an urban freshwater pelagic ecosystem. The use of mesocosms is motivated by the fact that they offer a high degree of ecological realism and thus provide an intermediate step between field conditions and small-scale lab experiments. Despite this potential, the literature reveals a lack of mesocosm studies addressing plastic fate and biofilm growth at comparable temporal and spatial scales35,36. Each mesocosm was exposed to one type of commonly used intact plastic object commonly found in aquatic ecosystems, such as cups, lids, bags and straws6,34. The state of these objects, microbial colonization and water quality variables were monitored over time.
Biofilm development and the fate of plastic items
The biofilm formation showed different temporal dynamics for bacteria, algae and cyanobacteria (Fig. 1). Bacterial cell counts per cm2 exhibited a distinct triphasic pattern consisting of initial colonization, growth and maturation phases across all plastic items (Fig. 1a). Temporal dynamics had a significant effect on bacterial biofilm growth (ANOVA, effect of time, p = 2.2 × 10−4). After eight weeks, bacterial cell counts peaked, with significant differences between W2 and W8 (p = 2.56 × 10−4) and between W4 and W8 (p = 5.53 × 10−5), followed by a significant reduction at W12 (t-test, p = 0.0017). Plastic substrates had a statistically significant role on bacterial biofilm formation (ANOVA, effect of plastic item, p = 0.004). Post hoc pairwise tests of the main effect of plastic revealed that the HDPE (high-density polyethylene)-bag-1 treatment, in particular, displayed significantly higher cell counts compared to the treatment with PLA (polyactic acid (plant-based biopolymer)-cup and PP (polyethylene)-straw (post hoc t-test, p = 0.048 and p = 0.042, respectively). The interaction between plastic substrate and time was not statistically significant (ANOVA, Plastic × Time effect, p = 0.111), indicating that the type of plastic did not play a significant role in determining the changes over time.
a–d, Barplot of biofilm growth expressed as density (cell cm−2) of heterotrophic bacteria (a), algae (b) and cyanobacteria (c) and the estimated thickness of the biofilm layer (d) for different plastic items and time periods. Each bar represents the mean of three plastic replicates and error bars indicate standard deviation, represented only in the positive direction for visual clarity. W2, W4, W8 and W12 indicate the weeks at which sampling was conducted (that is, week 2, week 4, week 8 and week 12).
Algal growth followed an almost identical pattern to bacterial growth in the biofilm across all plastic items (Fig. 1b). Both time and plastic type had a significant effect on algal biofilm growth (ANOVA, effect of time, p = 5.2 × 10−9; effect of plastic type, p = 2.8 × 10−4). The interaction between plastic type and time was statistically significant (ANOVA, Plastic × Time effect, p = 0.002), indicating that plastic identity played a significant role in the temporal development of algae. Post hoc tests on the effect of plastics at each time point indicated that algal biofilm growth at W8 (p = 6.8 × 10−4) and W12 (p = 0.042) was significantly influenced by plastic substrates. At W8, HDPE-bag-1 and PS-lid had significantly higher algal biofilm density compared to all plastic items (Extended Data Table 1 for p-values). Post hoc tests at W12 showed significant differences only between PS-lid compared to PE-bag-1, PE-bag-2 and PLA-cup (Extended Data Table 2 provides p-values).
Unlike bacteria and algae, cyanobacteria growth only reached the highest cell density towards the end of the experiment at W12 (Fig. 1c). These dynamics in the biofilm were significantly influenced by time and marginally by plastic type (ANOVA, effect of time, p = 1.57 × 10−5; effect of plastic type, p = 0.039). The interaction between plastic type and time was statistically significant (ANOVA, Plastic × Time effect, p = 0.002), indicating that similar to the algal contribution to the biofilm, plastic type had a significant effect on the patterns of cyanobacterial biofilm formation over time. Post hoc tests on the effect of plastic type at each time point indicated that W8 (p = 0.022) and W12 (p = 0.033) were significantly different. Pairwise post hoc analyses at W8 revealed HDPE-bag-2 had significantly higher cyanobacterial biofilm growth compared to PE-bag-2, PP-cup, PP-straw and PLA-cup (Extended Data Table 3 provides p-values).
Both initial density and biofilm formation had a visible impact on the fate of the plastic items. High-density plastics (PS lids and PLA cups) settled right after immersion. Plastics such as HDPE-bag-1 and HDPE-bag-2, which experienced a steep increase in biofilm thickness during the early weeks of the experiment, settled mostly by week 6 (Supplementary Fig. 1). The estimated biofilm thickness varied between 0.8 µm and 22 µm across different plastic items (Fig. 1d), with algae accounting for over 99% of the biofilm composition, whereas bacteria and cyanobacteria contributed less than 1% of the biofilm thickness (Extended Data Fig. 1). Similar to the algal dynamics, time and plastic type had a significant effect on biofilm thickness (ANOVA, effect of time, p = 5.4 × 10−9; effect of plastic type, p = 2.8 × 10−4). Remarkably, the interaction between plastic type and time was statistically significant (ANOVA, Plastic × Time effect, p = 0.003), indicating that plastic identity played a role in the temporal development of biofilm thickness. HDPE-bag-1 and PS-lid had significantly higher biofilm thickness compared to the other plastic items at W8, peaking at around 20 µm and 22 µm respectively (Extended Data Table 4 provides p-values). PS-lid also showed significantly higher biofilm thickness at W12, compared to PE-bag-1, PE-bag-2 and PLA-cup (Extended Data Table 5 provides p-values). When cell counts were normalized by plastic weight (as a proxy for biofilm-to-plastic ratios), HDPE bags showed the highest relative biofilm mass, followed by PE bags. In contrast, the PS-lid and PLA-cup exhibited the lowest ratios, corresponding to the lowest cell counts per gram of plastic (Extended Data Fig. 2).
Microbial community composition dynamics across plastic items
The composition of the bacterial communities colonizing the plastic items changed over time, showing a clear succession of different taxa (Fig. 2). After two weeks (W2), the community was mainly dominated by Proteobacteria (over 75% in most plastics). Firmicutes, Planctomycetes, Bacteroidota, Actinobacteriota and Cyanobacteria (together comprising 20%) were the other phyla present in the early colonization of plastics. From week eight (W8) to week 12 (W12), however, Verrucomicrobiota and Cyanobacteria started to prevail at the expense of Proteobacteria’s abundance. The presence of Firmicutes declined in almost all plastic items over time. While Bacteroidota and Actinobacteriota abundance remained constant during the different time intervals, Patescubacteria started to grow after eight weeks. Desulfobacterota showed persistence in the mesocosms containing PE-bag-1 and HDPE-bag-2 starting from week four (W4) and peaked at week eight (W8) with relative abundances of 14% and 18%, respectively. The relative abundance of Desulfobacterota continued to increase in the PE-bag-1 mesocosm, reaching 23% by week 12 (W12). In contrast, Desulfobacterota were also detected in the mesocosm with the PLA-cup at W8, although it constituted only about 2% of the microbial community.
Each bar represents the mean of three replicates. Each bar shows taxa grouped to different phyla. The category ‘others’ represent all taxa below 1% relative abundance.
Subsequent taxonomic analysis at the genus level revealed that the initial colonization of plastic items was predominantly comprised of five genera: Sphingobium, Pseudomonas, Exiguobacterium, Gemmobacter and Hydrogenophaga, which collectively accounted for over 25% of the microbial community (Extended Data Fig. 3). Other genera, such as Sphingorhabdus, Blastomonas, Bosea and Flavobacterium, exhibited relatively stable abundances throughout the different biofilm formation stages (initial, growth and maturation). In contrast, the maturation phase (W12) was characterized by a diverse array of species with lower abundances. Notably, the community compositions associated with biodegradable plastics (for example, HDPE bags and PLA-cup) and non-biodegradable (for example, PP-cup and PE bags) plastics showed similarities.
The ordination of the samples through principal coordinate analysis (PCoA) (a measure of beta diversity) indicated the overlap in microbial community based on time periods (Extended Data Fig. 4). Particularly, the PCoA analysis showed no overlap between microbial communities in week two (W2) and week 12 (W12), whereas considerable overlap of W2–W4 and W8–W12 samples was revealed, indicating the microbial communities between these time intervals to be more similar.
Microbial alpha diversity across plastic items
We used the Shannon diversity index, which accounts for both species richness and evenness, to assess temporal changes in the alpha diversity of microbial communities on the macroplastic items. Wilcoxon rank-sum tests revealed that HDPE-bag-1 exhibited a significantly higher Shannon diversity of microbial communities on most plastic items, except PE-bag-2 and HDPE-bag-2 (Fig. 3a). Additionally, the Shannon diversity index of the microbial communities present on PE-bag-2 and HDPE-bag-2 was significantly different from the Shannon diversity index on PP straws. The biofilm of other plastic items did not show significant differences in Shannon diversity indices, indicating that the microbial community of these plastic items had similar diversity.
a,b, Alpha diversity for different plastic items (a) and time periods (b) expressed as Shannon diversity index. The line inside the box represents the median, whereas the whiskers represent the lowest and highest values within the interquartile range. Individual sample values are shown as dots: in a, 12 dots represent four time points with three replicates each, and in b, 24 dots represent eight plastic items with three replicates each. Statistical differences were assessed using a two-sided Wilcoxon rank-sum test for non-parametric data. P values were adjusted for multiple comparisons using the Benjamini–Hochberg method.
Further analyses of alpha diversity focusing on time periods revealed an increase of microbial diversity over time (Fig. 3b). Particularly, significant differences were observed between W2 and W12 (p = 0.00043) and between W4 and W12 (p = 0.00039).
Factors influencing plastisphere microbial community composition
Microbial community composition among plastic items was influenced by multiple factors: the environment that is, dissolved oxygen (DO), light availability, pH, total inorganic phosphorus (TIP) and total inorganic nitrogen (TIN), plastic traits (that is, plastic item, plastic degradability, plastic morphology, polymer identity, density) and time (Fig. 4). The variance partitioning analysis (VPA) allowed us to separate the total variance in microbial communal composition into fractions explained by different predictor variables, that is, environmental factors, plastic traits and temporal dynamics. The VPA showed that environmental factors explained most of the variation in microbial community composition (31%), followed by temporal dynamics (26%) and plastic traits (24%). Interestingly, the microbial community was significantly influenced by temporal changes in the environment (21%). Because plastic traits remained stable over time, the variability (7%) explained by the interaction between plastic and environment can be attributed to changes in the environment due to the presence of the plastics.
The circles represent the variance explained by environmental factors (environment), plastic traits (plastic) and exposure time (time), overlapping areas indicate shared explained variance between variables. Numbers inside the circles indicate the proportion of total variation attributed to each factor and their interaction. The pink circle represents environmental factors, the green circle represents plastic items and the blue circle represents time.
We further examined the influence of individual environmental factors (Extended Data Fig. 5) in shaping the microbial community over the different time periods using redundancy analysis (RDA) (Table 1). The amount of variance explained by the sum of environmental factors increased from 14.99 % (W2) to 29.44% (W12), indicating that the environment plays a larger effect in shaping the microbial community composition over time. These changes in the environment were predominantly governed by variation in DO and light availability (for example, variance explained 14,86%, p = 0.001 at week 12) (Table 1 and Extended Data Fig. 5). Nutrients (that is, TIN and TIP) also became more influential over time, significantly increasing the variance explained from 3.15% at week 2 to 10.02% at week 12 (p = 0.001) (Table 1 and Extended Data Fig. 5).
Discussion
Factors driving biofilm development on plastics
In our study, biofilm growth was observed on all plastic items, but certain plastic substrates exhibited significantly higher biofilm growth compared to others. Our results emphasize the significant impact of both temporal dynamics and the identity of plastic substrates on the growth of biofilms. The maturation phase of bacteria and algae was achieved after eight weeks, followed by a significant decrease at week 12. The peak in microbial abundance observed at week 8 (based on flow cytometry data), along with the notable increase in community diversity at week 8 (comparable to that at week 12) suggests that the biofilm may have reached the maturation phase during this period. Environmental changes occurring during the experiment, such as decreasing light levels and water temperatures, mimic transition into winter. The initial trend observed in our study was similar to that of Li et al.27, who observed an increase in biomass growth on various plastics (PVC, PP, PE, PS and PU) in brackish waters over six weeks. Similarly, Dussud et al.26 reported a maturation phase in biofilm growth on PE and OXO-biodegradable plastics after 22 days in marine conditions. Compared to heterotrophic bacteria and algae, cyanobacterial biofilm followed a distinct temporal trend, with the highest growth reached at week 12. This suggests that antecedent biofilm formation and environmental factors such as light availability and nutrient fluctuations may be crucial in shaping cyanobacterial biofilm communities8,37. Specifically, higher nutrient levels positively correlated with biofilm growth27. Our results suggest that under autumnal conditions in the Netherlands (and possibly at similar latitudes) biofilm development on plastic surfaces may follow comparable patterns. It is important to recognize, however, that the maturation phase observed in this specific experimental set-up may differ under other environmental contexts.
Besides environmental factors, biofilm development has been shown to be influenced by several plastic traits such as plastic morphology, polymer identity, degradability, buoyancy, chemical nature38 and roughness of the substrate39. We observed that the biodegradable bags (HDPE-bag-1) and the non-biodegradable coffee cup lids (PS-lid), exhibited significantly higher cell density compared to other plastic items. Previous research on biofilm growth found significantly higher biomass on PS27, PVC40 compared to PP, PE or PET. PVC had a higher surface roughness than PET and PP, which explains the higher biomass growth, as substrates with higher surface roughness provide more favourable habitats for biofilm growth and development41. Dussud et al.26 found that at similar surface roughness, colonization increased with increasing polarity (polarity of OXO-biodegradable substrates > PE plastics), indicating that besides surface roughness, polarity may also play in determining the colonization process. Higher surface polarity enhances the adsorption of conditioning films, creating a more favourable environment for bacterial attachment26. The higher cell densities observed on HDPE-bag-1 and PS-lid suggest that their surface properties (for example, roughness and polarity) might be similar and more conducive to microbial colonization26,42. It is worth noting that surface roughness is influenced not only by polymer type but also by the characteristics of the product itself.
Changes in buoyancy and environmental fate
Buoyancy of plastic is influenced by several factors, such as, polymer density, biofilm growth43, the leaching of organic matter and fragmentation resulting from biodegradation and photodegradation44. In our study, the development of biofilms specially changed the buoyancy of plastic bags (that is, HDPE-bag-1 and HDPE-bag-2) contributing to higher sinking rates. Nevertheless, we observed that PP cups developed a biofilm thickness similar to that of PE bags but did not sink, whereas the PE bags did. Our observation was similar to the finding of Miao et. al.36 who observed a decreased density in PP sheets (5 mm × 5 mm) in riverine systems. However, our findings differ with those of Chen et al.37, who reported that biofilm formation led PP sheets in a freshwater lake to lose buoyancy. This discrepancy could be explained by a density loss of the cups due to leaching and biodegradation36, which could counteract the weight gained by biofilms. In addition to the combined effects of polymer and biofilm densities, the morphology of plastic items seems to have a key role determining the buoyancy45.
The non-buoyant plastic items, the PLA-cup and PS-lid, sank immediately at the start of the experiment, whereas others, such as the PP-cup and PP-straw, remained afloat throughout. This highlights the importance of the initial polymer density in determining the vertical transport of plastics in aquatic ecosystems. The influence of biofilm development was particularly evident for plastic bags. Thin, low-mass items, such as HDPE bags, showed the highest biofilm-to-plastic ratios and settled first, followed by PE bags (Extended Data Fig. 2), indicating their higher susceptibility to biofilm-driven buoyancy changes. In contrast, although PS lids showed high biofilm colonization (reflected by biofilm thickness and cell density per surface area) they had among the lowest biofilm-to-plastic mass ratios. This highlights the role of plastic item morphology, as thicker and denser plastic types such as PLA and PS accumulated relatively little biofilm in proportion to their mass, resulting in a lower likelihood of buoyancy alteration. These findings are consistent with ref. 25, who reported that biofilm development differentially affects sinking behaviour depending on polymer type, size and density, suggesting that rivers may retain larger PET particles, while lighter plastics are more likely to be transported downstream. In our study, among the eight plastic items tested, two were initially negatively buoyant, while four became negatively buoyant due to biofilm growth. This suggests that the majority of such macroplastic items are likely to remain in inland waters, particularly in lentic systems. It is important to note, as emphasized by both our study and previous work, that biofilm formation is substrate specific25 and also site specific46. Therefore, direct extrapolation of our findings should be approached with caution; generalization may be possible for similar plastic items exposed to comparable environmental conditions or seasonal contexts. Understanding these intricate settling dynamics between biofilms and real plastic objects is crucial to develop more reliable predictive models of plastic transport and fate in freshwater ecosystems12.
Importantly, this settling process determines not only the fate of plastics in aquatic ecosystems but also the spatial distribution and potential accumulation zones, which directly affects the mass flow of plastics into the oceans47. Moreover, the build-up of plastics in sediments could have long-term ecological effects causing the disruption of benthic habitats48,49. Our study emphasizes the vital necessity to consider biofilm–plastic interactions in the context of global plastic pollution management strategies by integrating these multifaceted factors.
Factors driving microbial community composition and diversity on plastics
We showed that changes in the environmental conditions over time and differences among plastic substrates shape the microbial community composition and diversity of the plastisphere. Specifically, dissolved oxygen and light availability were the main drivers shaping the colonizing community, with nutrient dynamics becoming significant over time. These findings align with previous studies highlighting the greater influence of environmental variables such as DO and nutrients on community composition on plastics37,50. However, we found alpha diversity to be significantly higher on oxo-biodegradable bags (that is, HDPE-bag-1) compared to most plastic items (except HDPE-bag-2 and PE-bag-2). These results confirm the variance partitioning analysis, indicating that while the surrounding environment is the primary driving factor in plastics community composition, this process can be influenced by the characteristics of the specific plastic substrate. Oxo-biodegradable bags (HDPE-bag-1) exhibited higher biofilm thickness and alpha diversity, suggesting that biofilm productivity may enhance biofilm diversity. This is similar to the pattern observed in terrestrial ecosystems where higher productivity can increase trait and phylogenetic diversity under certain conditions51. However, PP straws showed significantly lower alpha diversity compared to PE-bag-2 despite having similar biofilm thickness, indicating that higher colonization rates do not necessarily equate to higher diversity. This is analogous to the transient competitive exclusion observed in phytoplankton at high productivity levels52.
The microbial communities found in our study were dominated by Proteobacteria along with Planctomycetes, Bacteroidota, Actinobacteriota and Cyanobacteria. This is similar to those reported in other studies in freshwater ecosystems31,32,33. Proteobacteria and Bacteroidota phyla represent pioneer colonizers and are recognized for their production of extracellular polymeric substances (EPS)53. The secreted EPS forms a complex matrix that supports microbial attachment and aggregation, enhances nutrient cycling and acts as a nutrient-rich substrate for other species54. In contrast, Cyanobacteria seem to be late colonizers that take advantage of the already mature biofilm, as confirmed by the patterns observed in our microbial community composition and flow cytometry data. The colonization trends were similar across plastic items, with the majority of the founder taxa not maintaining high abundance over time and gradually changing, reflecting the community succession taking place during the colonization process55. Importantly, we did not find differences in the community composition between biodegradable and non-biodegradable plastics. For example, both types exhibited a high relative abundance of Desulfobacteriota, which are sulfide-reducing, strictly anaerobic bacteria56. The presence of these bacteria was associated with treatments with large plastic bags (that is, non-biodegradable PE-bag-1 and OXO-biodegradable bag HDPE-bag-2), which correlated with low DO levels in the mesocosm (Table 1 and Extended Data Fig. 5). This observation aligns with our finding that environmental factors and other plastic traits (that is, roughness and buoyancy) rather than biodegradability or polymer identity, are the key drivers of biofilm microbial communities. At the genus level, species composition of the plastisphere did not substantially differ between the different plastic items, but there was a clear link with exposure time. Genera belonging to the phylum Proteobacteria (that is, Sphingobium, Pseudomonas, Gemmobacter, Hydrogenophaga and Firmicutes (Exiguobacterium) dominated the initial colonization. Importantly, Sphingorhabdus is a known biofilm former, commonly found in freshwater ecosystems31. In our study, we found potentially plastic-degrading bacteria such as Pseudomonas27, Sphingobium57 and Hydrogenophaga with relatively high abundance in the first month. Roager and Sonnenschein58 found that Hydrogenophaga species are prevalent colonizers on polymers such as PE, PP and PS. Additionally, it has been found that the genera Hydrogenophaga can decompose aliphatic biodegradables such as polyhydroxyalkanoate and polyhydroxybutyrate59.
Conclusion
First, biofilm thickness contributed to buoyancy changes and increased sinking rates of plastic bags, primarily due to algae, highlighting the importance of understanding the role of algae in governing the plastisphere. Second, our results demonstrate that plastic traits play an important role in biofilm growth. For instance, plastics items with similar surface properties, such as HDPE bags and PS lids, fostered significantly higher biofilm growth compared to PLA cups and PP straws.
Third, focusing on the role of bacteria in macroplastic colonization, we observed a reduced presence of potential plastic-degrading bacteria in the biofilm, suggesting its limited ability to biodegrade plastics under environmentally relevant conditions. The microbial community was to a large extent influenced by temporal changes in water quality, with several water quality parameters, in turn, being significantly changed by the presence of the plastics. Water quality parameters, such as dissolved oxygen and light availability were key environmental drivers of this community composition. We demonstrate here how plastic types influence microbial composition while that same composition and growth affect the fate of plastic items in the ecosystem. Because macroplastics are the source of micro- and nanoplastics, our results also have implications for the risks posed by these particles, which are much more bioavailable. These more complex interactions can best be studied at the ecosystem scale under controlled conditions. Therefore, it is important to continue using mesocosm studies in future research.
Methods
Experimental design and sample collection
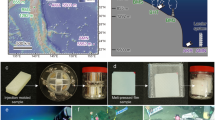
We conducted a 12-week mesocosm experiment to evaluate the process of microbial colonization of different macroplastic items and to identify the possible driving factors. To this end, we used a gradient design, where we exposed eight types of intact plastic object (details in Supplementary Table 1) to natural aquatic communities in indoor mesocosms. The experiment was conducted in controlled indoor mesocosms referred to as Limnotrons (0.97 m diameter and 1.37 m deep; ref. 60).
Initially, most macroplastics are positively buoyant upon entry in the aquatic system and stay in the water column61, thus, we were interested in the pelagic system. Limnotrons were filled with 988 l of water from a nearby mesotrophic shallow reference pond at the Netherlands Institute of Ecology, Wageningen, the Netherlands (coordinates in decimal degrees: 51.986759, 5.671079). All mesocosms were started with similar environmental conditions by homogenizing the water before inoculating the mesocosm. For all mesocosms, regardless of treatment, light and temperature conditions were changed weekly following the ambient seasonality from September to December similar to the reference pond (Supplementary Fig. 2). The indoor experiment was conducted from 12 September to 5 December 2019. We chose this period because it is expected that environmental conditions prevailing through this timeframe both enhance degradation and biofilm formation processes, with the system experiencing both warmer and cooler temperatures. This timeframe effectively mimics the transition through the seasons, providing a balance of conditions that promote biofilm growth while avoiding algal blooms seen in warmer months. To facilitate gas exchange between air and water we deployed one axial fan (AC axial compact fan 4850 Z; EBM-papst St. Georgen GmbH and Co. KG) and placed an aquarium pump (EHEIM compact 300; EHEIM GmbH and Co. KG) at 80 cm below the water surface in each mesocosm. Before the onset of the experimental treatments, the Limnotrons were left to stabilize for one week. After, the macroplastic items (described in the section below) were introduced into the Limnotrons, with an additional Limnotron serving as control (without addition of macroplastics). A total of 36 plastic items, previously cleaned with 70% ethanol, were introduced to each Limnotron with one type of plastic per Limnotron. Every two weeks, the Limnotrons were topped up with aerated ground water to compensate for water losses through evaporation and due to sampling.
Plastic samples were collected every two weeks (W2, W4, W6, W8, W10, W12). Using sterilized gloves (with 70% ethanol), six plastic items were gently removed from each Limnotron (48 in total; six items per eight Limnotrons) and placed in individual sterilized trays. Samples were then transferred to a sterilized surface for cutting, inside a thoroughly cleaned laminar flow cabinet prepared for microbiological work, equipped with flame and positive pressure. Using autoclaved scissors, forceps and a ruler, one square (2 × 2 cm) was cut from each plastic item (a total of six squares per Limnotron). Three squares were used for sequencing of the biofilm and three for flow cytometric analysis. The squares were consistently taken from the same area of each item across all time points. Each plastic square was stored in an individually labelled, autoclaved universal glass vial (30 ml) and frozen at −20 °C until further analysis.
Only samples from week two (W2), week four (W4), week eight (W8) and week 12 (W12) were used for sequencing and flow cytometry, resulting in a total of 192 samples: 96 samples (8 plastic items × triplicate × 4 time points) for DNA extraction and microbial community sequencing, and 96 samples (8 plastic items × triplicate × 4 time points) for flow cytometric enumeration (bacteria, cyanobacteria and algae).
Macroplastic items
All plastic objects used were made of commercially available non-biodegradable plastics62 plastics or biodegradable plastics58. We used four intact non-biodegradable plastic items, that is: (1) polypropylene cups (referred to as PP-cup; Asda groceries); (2) polystyrene lids (referred to as PS-lid; Asda groceries); (3) bin liner polyethylene bags, (referred to as PE-bag-1; Asda groceries); and (4) polyethylene bags, (referred to as PE-bag-2; Asda groceries). The four remaining Limnotrons were exposed to oxo-biodegradable plastics that is: (5) high-density polyethylene bags (referred to as HDPE-bag-1; Eco Hound/EHVR240); (6) high-density polyethylene bags (referred to as HDPE-bag-2; Sabco Packaging Ltd.); (7) polyethylene straws (referred to as PE-straw; Asda groceries) and (8) polylactic acid (plant-based biopolymer63 cups (referred to as PLA-cup; Vegware R360Y). For each plastic item, a picture with descriptions, the manufacturer’s name, initial dimensions and polymer ID confirmed by ATR-FTIR (attenuated total reflectance-Fourier transform infrared) spectroscopy (Cary 630 FTIR Spectrometer, Agilent Technologies Inc.) is provided as Supplementary Information (Supplementary Table 1).
Water quality monitoring and nutrients sampling/analysis
Water quality was monitored weekly using multichannel probes for simultaneous measurements of pH, dissolved oxygen (DO) and conductivity (EC) (IntelliCAL Probes, Hach). Light and temperature were recorded constantly by loggers (HOBO Pendant MX Temperature/Light Data Logger) located inside the Limnotrons at approximately 60 cm below water surface. A custom-made climate control system automatically recorded and regulated the water temperature (SpecView 32/859, SpecView Ltd.). For the analysis of dissolved inorganic nutrients, water samples (1 l) were collected weekly from the three Limnotron’s sampling ports. Briefly, 15 ml sub-samples were filtered through a pre-washed 0.2-μm glass microfibre filter (Whatman GF/F, Maidstone) and the filtrate was immediately stored at −20 °C. Just before analysis, samples were slowly thawed at 4 °C and the dissolved inorganic nutrients \({\mathrm{NO}}_{2}^{\,\,-}\), \({\mathrm{NO}}_{3}^{\,\,-}\), \({\mathrm{NH}}_{4}^{\,\,+}\) and \({\mathrm{PO}}_{4}^{\,\,3-}\) were measured using a QuAAtro39 Auto-Analyzer (SEAL Analytical Ltd.).
Flow cytometry
Bacteria, algae and cyanobacteria were analysed in triplicate using flow cytometry (BD influx cell sorter, Biosciences). To dislodge the biofilm from plastic samples 10 ml of sterile phosphate buffered saline was added into universal glass tubes, vortexed for 1 min and sonicated for another minute at high power (probe sonicator). Plastic samples were then removed and vortexed for an additional 1 min. Detached cell samples were fixed with glutaraldehyde (25%, grade I; Sigma) to a final concentration of 0.5% (v/v), left for 30 min at 4 °C in the dark and preserved at −80 °C until further analysis. Following a protocol adapted from Marie et al.64, detached cell samples were thawed in a water bath at 37 °C, removing the detached cell samples just before complete thawing. For bacterial cell enumeration, samples were stained with nucleic acid-specific dye SYBR Green I and incubated for 15 min in the dark before analysis. For photosynthetic microbes, that is, algae and cyanobacteria, cell count samples were directly analysed without any prior stain.
DNA extraction and amplicon sequencing
The taxon composition of the communities colonizing the macroplastics was identified using DNA extraction and amplicon sequencing. In total, there were 96 samples (8 plastic items × triplicate × 4 time points). Genomic DNA was extracted using the QIAGEN DNeasy PowerWater Kits (Qiagen Benelux B.V.), following the manufacturer’s protocol. DNA samples were checked for quality and quantity using Nanodrop ND-2000 (Thermo Fisher Scientific). The 96 DNA samples were submitted for sequencing to Génome Québec (McGill University). To obtain the microbial profiles the hypervariable region of the conserved subunit 16S rRNA gene, V3-V4 region was amplified using the primer set 341-F (CCTACGGGNGGCWGCAG) and 805-R (GACTACHVGGGTA TCTAATCC). Diluted and undiluted samples were used to optimize PCR conditions for amplifying the 314F–805-R fragment. A clear band was observed on agarose gels in tested samples. The PCR conditions that produced the best results were applied to all samples in the project. All samples were successfully amplified. Sequencing was performed on a single lane of Illumina MiSeq platform at Génome Québec (https://genomequebec.com/).
Data analysis
Analysis of DNA sequences
Raw sequencing FASTQ files were analysed using the DADA2 pipeline65 in R. Sequences were filtered to remove primers, adaptors and barcodes. Sequences with low quality ends were trimmed. Sequences were denoised using a parametric error model and merged, and chimaeras were removed. Taxonomy of resulting amplicon sequence variants (ASVs)65 was assigned using the SILVA reference database (v138). For downstream analyses, we used the phyloseq R package to generate a phyloseq object, which combines assigned ASVs and associated sample data.
Analysis of biodiversity metrics
The Shannon diversity index66 was used to determine the temporal changes in alpha diversity of the microbial communities present on the various macroplastic items. This index measures species diversity in a community (alpha diversity) accounting for both species richness (the number of species) and evenness (how evenly individuals are distributed among those species). A higher Shannon index indicates a more diverse and evenly distributed community. We employed the ‘estimate_richness’ function available in the ‘phyloseq’ package (version 1.36.0). Statistical significance of the differences in Shannon diversity was tested with the non-parametric Wilcoxon rank-sum test. Beta diversity was visualized using principal coordinate analysis (PCoA) with Bray–Curtis distance.
Variance partitioning analysis (VPA) was used to decompose the total variance of the response community by three explanatory matrices, that is, environment (containing the measured water quality parameters), plastic (containing the five plastic traits: plastic item, plastic degradability, plastic morphology, polymer identity, density) and time (containing the time periods in which sampling took place). Redundancy analysis (RDA) was used to explore correlations between plastic microbial composition and environmental factors (that is, DO, Light, pH, total inorganic phosphorus (TIP) and total inorganic nitrogen (TIN)) per time period. Species data was Hellinger transformed and environmental data scaled. The output of the RDA model was tested for significance using the ‘anova.cca’ function and 999 permutations. The analysis and visualization of PCoA, VPA and RDA was conducted using the ‘vegan’ package (version 2.6-4) in R.
Drivers of biofilm formation
Biofilm thickness was estimated by calculating the weighted average biovolume of algae and cyanobacteria based on community composition and relative abundance from Frenken et al.67. For bacteria, we used the average biovolume values from Bratbak68 and Frølund et al.69 (Supplementary Information provides details). To examine the effects of plastic treatments and time on biofilm growth (that is, flow cytometry bacteria, algae and cyanobacteria counts per cm−2) a two-way repeated-measures ANOVA test was used. Repeated measurements ANOVAs take into account the effect of time, thus accounting for temporal autocorrelation. The normality of the data was assessed using the Shapiro–Wilk test, and the homogeneity of variances was examined through Levene’s test. These assumptions were met for all variables. The assumption of sphericity was tested using Mauchly’s Test. In cases where sphericity was not met, Greenhouse–Geisser correction was applied. Post hoc analysis was followed by pairwise t-tests to compare the differences between pairs of plastic items. The Benjamini and Hochberg70 method was used to adjust the obtained p-values and correct for multiple testing. Significance level was set at 0.05. These analyses were performed using the ‘rstatix’ package in R (version 0.7.2).
Data availability
All datasets generated during this study that support the findings are available via Zenodo at https://doi.org/10.5281/zenodo.18750270 (ref. 71). The 16S rRNA gene amplicon sequencing reads are available in the NCBI Sequence Read Archive (BioProject, PRJNA1427709).
Code availability
All R scripts used to generate the results and figures are available via Zenodo at https://doi.org/10.5281/zenodo.18750270 (ref. 71).
References
Lau, W. W. Y. et al. Evaluating scenarios toward zero plastic pollution. Science 369, 1455–1461 (2020).
Nava, V. et al. Plastic debris in lakes and reservoirs. Nature 619, 317–322 (2023).
Noventa, S. et al. Paradigms to assess the human health risks of nano- and microplastics. Microplast. Nanoplast. 1, 9 (2021).
Coffin, S. et al. Development and application of a health-based framework for informing regulatory action in relation to exposure of microplastic particles in California drinking water. Microplast. Nanoplast. 2, 12 (2022).
World Health Organization Microplastics in Drinking-Water (World Health Organization, 2019); https://iris.who.int/handle/10665/326499
Winton, D. J., Anderson, L. G., Rocliffe, S. & Loiselle, S. Macroplastic pollution in freshwater environments: focusing public and policy action. Sci. Total Environ. 704, 135242 (2020).
Zettler, E. R., Mincer, T. J. & Amaral-Zettler, L. A. Life in the ‘plastisphere’: microbial communities on plastic marine debris. Environ. Sci. Technol. 47, 7137–7146 (2013).
Wright, R. J., Erni-Cassola, G., Zadjelovic, V., Latva, M. & Christie-Oleza, J. A. Marine plastic debris: a new surface for microbial colonization. Environ. Sci. Technol. 54, 11657–11672 (2020).
Barros, J. & Seena, S. Plastisphere in freshwaters: an emerging concern. Environ. Pollut. 290, 118123 (2021).
Amaral-Zettler, L. A., Zettler, E. R. & Mincer, T. J. Ecology of the plastisphere. Nat. Rev. Microbiol. 18, 139–151 (2020).
Oberbeckmann, S., Osborn, A. M. & Duhaime, M. B. Microbes on a bottle: substrate, season and geography influence community composition of microbes colonizing marine plastic debris. PLoS ONE 11, e0159289 (2016).
Lofty, J., Wilson, C. & Ouro, P. Biofouling changes the settling dynamics of macroplastic plates. Commun. Earth Environ. 5, 750 (2024).
Andrady, A. L. Weathering and fragmentation of plastic debris in the ocean environment. Mar. Pollut. Bull. 180, 113761 (2022).
Vroom, R. J. E., Koelmans, A. A., Besseling, E. & Halsband, C. Aging of microplastics promotes their ingestion by marine zooplankton. Environ. Pollut. 231, 987–996 (2017).
Amariei, G., Rosal, R., Fernández-Piñas, F. & Koelmans, A. A. Negative food dilution and positive biofilm carrier effects of microplastic ingestion by D. magna cause tipping points at the population level. Environ. Pollut. 294, 118622 (2022).
Blettler, M. C. M. & Mitchell, C. Dangerous traps: macroplastic encounters affecting freshwater and terrestrial wildlife. Sci. Total Environ. 798, 149317 (2021).
Merga, L. B., Redondo-Hasselerharm, P. E., Van Den Brink, P. J. & Koelmans, A. A. Distribution of microplastic and small macroplastic particles across four fish species and sediment in an African lake. Sci. Total Environ. 741, 140527 (2020).
Yaghmour, F. et al. Junk food: polymer composition of macroplastic marine debris ingested by green and loggerhead sea turtles from the Gulf of Oman. Sci. Total Environ. 828, 154373 (2022).
Masó, M., Garcés, E., Pagès, F. & Camp, J. Drifting plastic debris as a potential vector for dispersing Harmful Algal Bloom (HAB) species. Sci. Mar. 67, 107–111 (2003).
Quilliam, R. S., Jamieson, J. & Oliver, D. M. Seaweeds and plastic debris can influence the survival of faecal indicator organisms in beach environments. Mar. Pollut. Bull. 84, 201–207 (2014).
Kirstein, I. V., Wichels, A., Krohne, G. & Gerdts, G. Mature biofilm communities on synthetic polymers in seawater - specific or general?. Mar. Environ. Res. 142, 147–154 (2018).
Amaral-Zettler, L. A. et al. The biogeography of the plastisphere: implications for policy. Front. Ecol. Environ. 13, 541–546 (2015).
Coons, A. K., Busch, K., Lenz, M., Hentschel, U. & Borchert, E. Biogeography rather than substrate type determines bacterial colonization dynamics of marine plastics. PeerJ https://doi.org/10.7717/peerj12135 (2021).
Oberbeckmann, S., Kreikemeyer, B. & Labrenz, M. Environmental factors support the formation of specific bacterial assemblages on microplastics. Front. Microbiol. https://doi.org/10.3389/fmicb.2017.02709 (2018).
Vercauteren, M., Lambert, S., Hoogerwerf, E., Janssen, C. R. & Asselman, J. Microplastic-specific biofilm growth determines the vertical transport of plastics in freshwater. Sci. Total Environ. 910, 168399 (2024).
Dussud, C. et al. Colonization of non-biodegradable and biodegradable plastics by marine microorganisms. Front. Microbiol. https://doi.org/10.3389/fmicb.2018.01571 (2018).
Li, W. et al. Colonization characteristics of bacterial communities on plastic debris influenced by environmental factors and polymer types in the haihe estuary of bohai bay, China. Environ. Sci. Technol. 53, 10763–10773 (2019).
Debroas, D., Mone, A. & Ter Halle, A. Plastics in the North Atlantic garbage patch: a boat-microbe for hitchhikers and plastic degraders. Sci. Total Environ. 599–600, 1222–1232 (2017).
Oberbeckmann, S., Loeder, M. G. J., Gerdts, G. & Osborn, A. M. Spatial and seasonal variation in diversity and structure of microbial biofilms on marine plastics in Northern European waters. FEMS Microbiol. Ecol. 90, 478–492 (2014).
Ogonowski, M. et al. Evidence for selective bacterial community structuring on microplastics: selective bacterial community structuring. Environ. Microbiol. 20, 2796–2808 (2018).
Di Pippo, F. et al. Microplastic-associated biofilms in lentic Italian ecosystems. Water Res. 187, 116429 (2020).
González-Pleiter, M. et al. Microbial colonizers of microplastics in an Arctic freshwater lake. Sci. Total Environ. 795, 148640 (2021).
Wright, R. J., Langille, M. G. I. & Walker, T. R. Food or just a free ride? A meta-analysis reveals the global diversity of the Plastisphere. ISME J. 15, 789–806 (2021).
de Lange, S. I. et al. Sample size requirements for riverbank macrolitter characterization. Front. Water https://doi.org/10.3389/frwa.2022.1085285 (2023).
Amaral-Zettler, L. A., Zettler, E. R., Mincer, T. J., Klaassen, M. A. & Gallager, S. M. Biofouling impacts on polyethylene density and sinking in coastal waters: a macro/micro tipping point?. Water Res. 201, 117289 (2021).
Miao, L. et al. Effects of biofilm colonization on the sinking of microplastics in three freshwater environments. J. Hazard. Mater. 413, 125370 (2021).
Chen, X. et al. Effects of microplastic biofilms on nutrient cycling in simulated freshwater systems. Sci. Total Environ. 719, 137276 (2020).
Lorite, G. S. et al. The role of conditioning film formation and surface chemical changes on Xylella fastidiosa adhesion and biofilm evolution. J. Colloid Interface Sci. 359, 289–295 (2011).
Riedewald, F. Bacterial adhesion to surfaces: the influence of surface roughness. PDA J. Pharm. Sci. Technol. 60, 164–171 (2005).
Miao, L. et al. Distinct microbial metabolic activities of biofilms colonizing microplastics in three freshwater ecosystems. J. Hazard. Mater. 403, 123577 (2021).
McGivney, E. et al. Rapid physicochemical changes in microplastic induced by biofilm formation. Front. Bioeng. Biotechnol. https://doi.org/10.3389/fbioe.2020.00205 (2020).
Reisser, J. et al. Millimeter-sized marine plastics: a new pelagic habitat for microorganisms and invertebrates. PLoS ONE 9, e100289 (2014).
Kooi, M., van Nes, E. H., Scheffer, M. & Koelmans, A. A. Ups and downs in the ocean: effects of biofouling on vertical transport of microplastics. Environ. Sci. Technol. https://doi.org/10.1021/acs.est.6b04702 (2017).
Browne, M. A., Galloway, T. & Thompson, R. Microplastic—an emerging contaminant of potential concern?. Integr. Environ. Assess. Manage. 3, 559–561 (2007).
Kooi, M. et al. The effect of particle properties on the depth profile of buoyant plastics in the ocean. Sci. Rep. 6, 33882 (2016).
Vincent, A. E. S., Chaudhary, A., Kelly, J. J. & Hoellein, T. J. Biofilm assemblage and activity on plastic in urban streams at a continental scale: site characteristics are more important than substrate type. Sci. Total Environ. 835, 155398 (2022).
Jambeck, J. R. et al. Plastic waste inputs from land into the ocean. Science 347, 768–771 (2015).
Redondo-Hasselerharm, P. E., Gort, G., Peeters, E. T. H. M. & Koelmans, A. A. Nano- and microplastics affect the composition of freshwater benthic communities in the long term. Sci. Adv. 6, eaay4054 (2020).
Redondo-Hasselerharm, P. E., Rico, A. & Koelmans, A. A. Risk assessment of microplastics in freshwater sediments guided by strict quality criteria and data alignment methods. J. Hazard. Mater. 441, 129814 (2023).
Rogers, K. L., Carreres-Calabuig, J. A., Gorokhova, E. & Posth, N. R. Micro-by-micro interactions: how microorganisms influence the fate of marine microplastics. Limnol. Oceanogr. Lett. 5, 18–36 (2020).
Brun, P. et al. The productivity-biodiversity relationship varies across diversity dimensions. Nat. Commun. 10, 5691 (2019).
Vallina, S. M. et al. Global relationship between phytoplankton diversity and productivity in the ocean. Nat. Commun. 5, 4299 (2014).
Dang, H. & Lovell, C. R. Bacterial primary colonization and early succession on surfaces in marine waters as determined by amplified rRNA gene restriction analysis and sequence analysis of 16S rRNA genes. Appl. Environ. Microbiol. 66, 467–475 (2000).
Lipsman, V., Shlakhter, O., Rocha, J. & Segev, E. Bacteria contribute exopolysaccharides to an algal-bacterial joint extracellular matrix. npj Biofilms Microbiomes 10, 1–16 (2024).
Brislawn, C. J. et al. Forfeiting the priority effect: turnover defines biofilm community succession. ISME J. 13, 1865–1877 (2019).
Rosenberg, E. The Prokaryotes: Deltaproteobacteria and Epsilonproteobacteria (Springer Berlin Heidelberg, 2014).
Song, D., Chen, X. & Xu, M. Characteristics and functional analysis of the secondary chromosome and plasmids in sphingomonad. Int. Biodeterior. Biodegrad. 171, 105402 (2022).
Roager, L. & Sonnenschein, E. C. Bacterial candidates for colonization and degradation of marine plastic debris. Environ. Sci. Technol. 53, 11636–11643 (2019).
Miao, L. et al. Spatio-temporal succession of microbial communities in plastisphere and their potentials for plastic degradation in freshwater ecosystems. Water Res. 229, 119406 (2023).
Vasantha Raman, N. et al. Effect of an antidepressant on aquatic ecosystems in the presence of microplastics: a mesocosm study. Environ. Pollut. 357, 124439 (2024).
Kuizenga, B., van Emmerik, T., Waldschläger, K. & Kooi, M. Will it float? Rising and settling velocities of common macroplastic foils. ACS ES&T Water 2, 975–981 (2022).
Barnes, D. K. A., Galgani, F., Thompson, R. C. & Barlaz, M. Accumulation and fragmentation of plastic debris in global environments. Philos. Trans. R. Soc. B: Biol. Sci. 364, 1985–1998 (2009).
Abdelmoez, W., Dahab, I., Ragab, E. M., Abdelsalam, O. A. & Mustafa, A. Bio- and oxo-degradable plastics: insights on facts and challenges. Polym. Adv. Technol. 32, 1981–1996 (2021).
Marie, D., Brussaard, C. P. D., Thyrhaug, R., Bratbak, G. & Vaulot, D. Enumeration of marine viruses in culture and natural samples by flow cytometry. Appl. Environ. Microbiol. 65, 45–52 (1999).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Shannon, C. & Weaver, W. The Mathematical Theory of Communication (Univ. of Illinois Press, 1949).
Frenken, T. et al. Warming accelerates termination of a phytoplankton spring bloom by fungal parasites. Glob. Change Biol. 22, 299–309 (2016).
Bratbak, G. Bacterial biovolume and biomass estimations. Appl. Environ. Microbiol. 49, 1488–1493 (1985).
Frølund, B., Palmgren, R., Keiding, K. & Nielsen, P. H. Extraction of extracellular polymers from activated sludge using a cation exchange resin. Water Res. 30, 1749–1758 (1996).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Gebreyohanes Belay, B. M., de Senerpont Domis, L. N. & Koelmans, A. A. The role of biofouling and microbial colonization in shaping macroplastic fate in freshwaters. Zenodo https://doi.org/10.5281/zenodo.18750270 (2026).
Acknowledgements
We thank S. Hesari and D. Waasdorp for their valuable assistance monitoring and sampling during the experiment. We also thank S. Wiezer for running the flow cytometer. We would like to extend our thanks to E. Harrison, R. Snow, A. Kruszelnicki and C. Gwinnett from Staffordshire University for the fruitful collaboration. B.M.G.B. acknowledges funding from the Gieskes Strijbis Fund (Microplastics in the foodweb) and the Royal Dutch Academy of Sciences (KNAW). L.N.d.S.D. acknowledges funding from H2020-INFRAIA-project number 731065 ‘AQUACOSM: Network of Leading European AQUAtic MesoCOSM Facilities Connecting Mountains to Oceans from the Arctic to the Mediterranean’.
Author information
Authors and Affiliations
Contributions
B.M.G.B.: conceptualization, data curation, formal analysis, investigation, methodology, project administration, validation, visualization, writing–original draft and writing–review and editing. A.A.K.: conceptualization, formal analysis, supervision, validation and writing–review and editing. L.N.d.S.D.: conceptualization, formal analysis, methodology, resources, supervision, validation and writing–review and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Water thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Total estimated thickness of the biofilm layer for different plastic items and time periods.
Stacked bar chart of total estimated biofilm thickness (µm) at sampling weeks 2, 4, 8, and 12, showing the relative contributions of bacteria, cyanobacteria, and algae. Note y-axis is on a log₁₀ scale.
Extended Data Fig. 2 Growth of the biofilm layer for different plastic items and time periods.
Bar plot of biofilm growth expressed as density (cells g−1), with flow cytometry cell counts normalized by each plastic item’s mass, serving as a proxy for plastic–biofilm mass ratio.
Extended Data Fig. 3 Relative abundance of top-15 bacterial genera for different plastic items and time periods.
Relative abundance profiles of bacterial genera (top 15) per plastic item and time (W2, W4, W8, W12). Each bar represents the mean of three biological replicates. Each bar shows taxa grouped to different genera and others represent all remaining taxa.
Extended Data Fig. 4 Principal coordinates analysis of plastisphere microbial communities for different plastic items and time periods.
Principal coordinates analysis (PCoA) of plastisphere microbial communities on plastic substrates based on Bray–Curtis dissimilarities calculated from amplicon sequence variant (ASV) relative abundances. Symbols represent samples, and distances between symbols represent similarities between samples. Ellipses indicate the 95% confidence intervals for each time period.
Extended Data Fig. 5 Time series of water quality parameters.
Time series of water quality measurements over weeks W1–W13 for (a) light availability (lux), (b) dissolved oxygen (mg L−1), (c) electrical conductivity (µS cm−1), (d) pH, (e) total dissolved inorganic phosphorus (mg L−1), and (f) total dissolved inorganic nitrogen (mg L−1).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1 and 2, Table 1 and Estimated biofilm thickness calculation.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gebreyohanes Belay, B.M., Koelmans, A.A. & de Senerpont Domis, L.N. The role of biofouling and microbial colonization in shaping macroplastic fate in freshwaters. Nat Water (2026). https://doi.org/10.1038/s44221-026-00629-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44221-026-00629-6