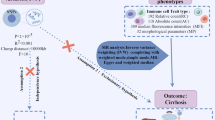
Abstract
Patients with advanced chronic liver disease who have underlying cirrhosis are highly susceptible to bacterial infections, which significantly increase the risk of complications and mortality, compounded by escalating antimicrobial resistance. The current gold standard for infection detection and antimicrobial resistance (AMR) profiling remains dependant on traditional microbiological methods. These conventional approaches are slow, labour-intensive, and often fail to deliver timely and accurate results, delaying critical antimicrobial treatment decisions. Clinical metagenomics (CMg) is emerging as a transformative molecular-based tool in infection diagnostics. By enabling the direct sequencing of pathogens from patient-derived samples, CMg offers rapid and comprehensive identification of pathogens and their resistance profiles. Incorporating this technology into the clinical management of patients with cirrhosis has potential to address diagnostic challenges, reduce reliance on broad-spectrum antibiotics and improve outcomes. To effectively incorporate CMg into infection diagnostics, it will be essential to embed of point-of-care sequencing, standardisation of AMR databases, and accessibility to bioinformatics workflows.

Similar content being viewed by others
The increasing burden of infections and antimicrobial resistance in advanced chronic liver disease and ongoing unmet needs of rapid infection diagnostics
Cirrhosis represents an advanced stage of chronic liver disease (ACLD) and is marked by diminished liver synthetic function and clinical complications, resulting from sustained parenchymal injury and scarring. In the UK, it affects around 60,000 people, leading to recurrent hospitalisations and high mortality, contributing to over 4000 deaths annually1. Cirrhosis progresses from stable (‘compensated’) to decompensated stages, with the latter marked by complications that often require hospitalisation and can be life-threatening, including portal hypertensive bleeding, ascites, spontaneous bacterial peritonitis and hepatic encephalopathy. Due to cirrhosis-associated immune dysfunction2 patients are particularly vulnerable to bacterial infections3, which can precipitate acute-on-chronic liver failure (ACLF), characterised by multi-organ failure and extremely high short term mortality rates4.
A delay in the administration of antibiotics in ACLD is associated with an increase of 7.6% in mortality per hour in the first 6 h of presentation5. In a nested cohort study of 635 patients with ACLD and septic shock who developed ACLF, in-hospital mortality was 75.6%6. The lack of rapid and accurate infection diagnostics has led to the frequent, poorly targeted and often inappropriate prescription of broad-spectrum antimicrobials, contributing to the increasing burden of antimicrobial resistance (AMR)7,8 further exacerbating infectious complications in patients with ACLD9.
The current gold standard for diagnosing bacterial infection in biofluid samples (ascitic fluid, blood, urine) obtained from patients with ACLD involves traditional culture-based microbiological methods, which are labour-intensive and often require long turnaround times. Samples undergo incubation for 2-5 days, followed by Gram staining, species identification using matrix-assisted laser desorption/ionisation time-of-flight (MALDI-TOF) mass spectrometry, and antimicrobial susceptibility testing (AST), most commonly by disk diffusion. In the case of spontaneous bacterial peritonitis (SBP), which is the infection of ascitic fluid, the most commonly suspected organisms are Escherichia coli and Klebsiella pneumoniae in community-acquired cases, while nosocomial SBP is increasingly associated with multidrug-resistant organisms such as Enterococcus spp.10. Current guidelines for the diagnosis of SBP have not evolved in over 20 years11. Diagnosis still relies on basic microscopy including Gram staining and assessment of cell morphology, alongside polymorphonuclear leukocyte (PMN) count threshold12, and leads to negative culture results in up to 60% of cases, likely due to the low bacterial concentration and relatively sterile nature of ascitic fluid. Similarly, diagnostics of bacterial pathogens in blood samples requires up to five days of incubation, with an additional 48 h for antibacterial testing to produce a complete antibiogram and confirm antibiotic sensitivities13. Delays in identifying pathogens contributes to worsening outcomes and higher mortality rates in cirrhosis, with disturbances of the ‘gut-liver axis’ causing microbiome dysbiosis, intestinal barrier failure, cirrhosis-associated immune dysfunction and systemic inflammation all adversely synergising to further exacerbate infection-related complications3. This further underscores the necessity for advanced point-of-care diagnostics to rapidly and accurately identify infecting organisms, while simultaneously characterising the resistance profiles of increasingly prevalent multi-drug resistant (MDR) pathogens.
Infection diagnostic approaches in clinical practice have undergone limited innovation in the past few decades, underscoring the urgent need to evolve these methods; harnessing next-generation sequencing techniques is one approach to improve patient outcomes and enhance antimicrobial stewardship14. In response, novel point-of-care testing technologies have emerged, utilising automated infection identification methods to address the unmet need for rapid detection and AMR profiling of clinical pathogens15. For example, nucleic acid amplification technologies (NAAT) have become a powerful tool for detecting clinically significant pathogens16. NAAT-based platforms frequently use multiplexed diagnostic panels that can identify multiple target genes, thus providing both the identity and AMR profile of the pathogen(s) of interest17.
Beyond conventional microbiological culture: clinical metagenomics in infection diagnostics
While NAAT have advanced pathogen identification primarily by reducing turnaround times to ~6–12 h and enabling the specific detection of AMR genes, they remain limited by their targeted nature16. Untargeted technologies such as long-read clinical metagenomics (CMg) hold even greater promise due to its ability to provide a more comprehensive analysis without pre-selected target genes, enabling the identification of a broader range of pathogens and resistance mechanisms in a single test18. Building on these advancements, CMg has recently emerged as a promising tool in the field of rapid infection diagnostics19. Long-read CMg allows for comprehensive, untargeted detection of bacterial pathogens within a single clinical sample, collected in a manner that reflects the likely site of infection. This method provides a more complete picture of the microbial landscape, which is particularly beneficial in cases of polymicrobial infections - common amongst ACLD patients who undergo invasive procedures20 and are repeatedly exposed to healthcare environment-related nosocomial risks - thus enabling more informed and effective clinical decisions.
Long read CMg has demonstrated success in rapidly diagnosing lower respiratory infections by accurately detecting AMR genes, with a turnaround time of six hours from sample to result21. Additionally, it can be optimised to generate AMR profiles for pathogens in clinical blood samples within 12 hours22 and has shown effectiveness in diagnosing orthopaedic prosthesis-associated infections23. The sensitivity, high resolution, and rapid turnaround of CMg can transform infection diagnostics and revolutionise treatment options for patients, facilitating timely, targeted, and life-saving interventions.
The untargeted nature of CMg sequencing, especially with long-read sequencing technologies, brings further benefit in allowing for the assembly of Metagenome-Assembled Genomes (MAGs)24. This provides detailed insights into the genetic architecture of the pathogens present, revealing the genomic location and context of AMR genes. Understanding the exact positioning of these genes within the genome can shed light on their mechanisms of dissemination, such as horizontal gene transfer, which is critical for devising strategies to combat the spread of resistance. However, these methods may not always fully resolve AMR-gene contextualisation, particularly for plasmids with broad host ranges, and complementary approaches such as Hi-C sequencing can further strengthen plasmid-host linkage and genomic context25. Additionally, CMg can uncover virulence factors and other gene functions that may play a role in pathogenicity and infection mechanisms. While not all virulence factors are immediately clinically actionable, this information can enhance risk stratification, support infection control strategies, and contribute to longer-term improvements in patient management and antimicrobial stewardship26,27. Unlike isolate genomics, which requires culturing a single organism, CMg can detect virulence genes directly from clinical samples without the need for culture, an important advantage since some pathogens cannot be cultured using standard diagnostic methods28. Furthermore, CMg enables detection of virulence factors in mixed microbial populations, providing context on virulence in polymicrobial infections and revealing determinants in organisms or genetic settings that might be missed by isolate-based approaches29. This wealth of genetic information not only aids in the immediate clinical management of infections but also contributes to a deeper understanding of the molecular mechanisms driving antimicrobial resistance and virulence. While this area is still largely exploratory, such insights have the potential to inform the development of new therapeutic and preventative strategies in the future. CMg can also provide crucial epidemiological surveillance capabilities, enabling healthcare institutions to rapidly identify emerging pathogens30, track the spread of infections, and implement timely infection control measures, which are essential for preventing and managing hospital-acquired infections. The value of metagenomic epidemiology has already been demonstrated in other contexts, such as the genomic surveillance of SARS-CoV-2 during the COVID-19 pandemic31, and these approaches hold clear potential for application in ACLD to monitor infection dynamics and emerging resistance patterns. Incorporating CMg-based surveillance in this setting could enable earlier detection of resistant pathogens and transmission events, guide antimicrobial stewardship, and inform timely infection control measures. Such an approach has the potential to reduce infection-related morbidity and mortality in ACLD while also contributing to broader efforts to combat the spread of AMR.
Future direction and overcoming challenges for clinical implementation
Blood and ascites derived from patients with ACLD are usually naturally sterile and even when infected tend to contain relatively low bacterial concentrations, rendering molecular-based pathogen detection particularly challenging32,33. Despite the promising applications of CMg, further optimisation adjusted to these specific sample types is needed to begin to harness the potential of CMg in routine clinical practice28, particularly in cirrhosis. Additionally, the untargeted nature of CMg can detect microbial DNA that may not reflect active infection, including signals from translocating or colonising organisms. This raises challenges in clinical interpretation, as it becomes difficult to distinguish true pathogens from background commensal organisms or contaminants28, particularly in polymicrobial results, which are common in hospitalised patients with ACLD34. Without strong clinical correlation and robust, standardised laboratory methodologies, such findings risk prompting unnecessary treatments. While CMg offers powerful surveillance and stewardship benefits, it may inadvertently lead to indiscriminate antimicrobial use if results are not carefully contextualised35.
As with all metagenomics-based approaches, there is a pressing need for the standardisation of lab-based workflows and standard operating procedures, to ensure reproducibility of the data36. The implementation of contamination controls alongside host DNA depletion approaches are critical steps and require careful attention to ensure accurate results. Without standardised controls, the reliability of metagenomic data can significantly vary37,38, hindering its progression into the clinical diagnostic setting.
Besides the limitations in laboratory-based experimental approaches, significant challenges also lie in connecting genomic data with phenotypic resistance profiles. Many AMR databases lack consistent accuracy and are not regularly updated, leading to discrepancies in results39,40. Selecting the most reliable database and tools remains complex, as there is limited consensus on which resources offer the most accurate representation of AMR genes. A consistent approach to AMR profiling through standardised workflows, bioinformatics pipelines and databases will improve the comparability of results across different laboratories, facilitating more effective tracking of resistance patterns41. These considerations need to be addressed while aiming to create a simplified bench-to-bedside workflow for diagnostic laboratory staff and clinicians, as the current process requires significant expertise. Making this technology more accessible and cost-effective will be key to its integration into everyday clinical use, enabling broader application of this powerful diagnostic tool in combating infectious diseases.
One of the major limitations of CMg is its perceived high cost, although actual expenses can vary significantly depending on the platform used, sequencing depth, sample batching, and whether the workflow is carried out onsite or outsourced. For instance, portable devices such as the Oxford Nanopore MinION can process multiple samples at relatively low per-sample costs (approximately £80 depending on workflow and batching) when run efficiently in-house42,43. However, when factoring in the full diagnostic pipeline, including specialised reagents, infrastructure, computational analysis, and clinical interpretation, the overall cost remains a consideration, particularly for routine diagnostics in resource-limited settings42. However, with continued technological advancements, a gradual reduction in costs is anticipated in line with the evolution of sequencing-based technologies. Emerging strategies such as sample multiplexing, which allows multiple samples to be processed simultaneously, are already being implemented in UK clinical diagnostic settings36. These innovations not only improve efficiency but also pave the way for broader, more cost-effective clinical adoption in the near future. The substantial economic burden associated with prolonged hospital stays due to delayed or ineffective treatment of infections are important to highlight as an ‘opportunity cost’ of not evolving diagnostic approaches. A recent multicentre interventional study demonstrated that CMg significantly improved pathogen and resistance detection in patients with sepsis, reduced unnecessary antimicrobial use, and showed potential to shorten hospital stays, translating into meaningful cost savings for healthcare systems44.
CMg has the potential to revolutionise infection diagnostics in ACLD by enabling rapid, accurate pathogen identification and antimicrobial resistance (AMR) profiling. Unlike traditional methods, which are slow and often ineffective, CMg offers a cutting-edge, untargeted approach that provides comprehensive insights into the microbial landscape. To integrate CMg into clinical practice, its current limitations must be systematically addressed, and relevant clinical trials need to be well-designed and delivered, in collaboration with key stakeholders, including diagnostic pathologists, microbiologists and infectious diseases experts, within a broader antimicrobial stewardship strategy for tackling AMR in cirrhosis45.
This breakthrough technology with correct implementation should enhance ACLD patient outcomes by facilitating timely, targeted treatments, reducing unnecessary antibiotic use, and combating the rising tide of AMR. Given that infections and antimicrobial resistance are major drivers of mortality in ACLD where 30% of patients die within 1 month after infection and another 30% die by 1 year46, the introduction of rapid and accurate diagnostic strategies such as CMg holds immense potential to transform healthcare, improving survival rates and quality of life for patients with ACLD. As these approaches are operationalised in the context of cirrhosis-related infections, there are much wider translational benefits to be realised in other conditions where infections have a detrimental role, including but not limited to critical care environments47, immunosuppressive states48 and organ transplantation49. As advancements continue, the integration of CMg into routine clinical practice will drive a new era of precision medicine, ensuring faster, more effective, and life-saving interventions.
Data availability
No datasets were generated or analysed during the current study.
References
Williams, R. & Horton, R. Liver disease in the UK: a Lancet Commission. Lancet 382, 1537–1538 (2013).
Albillos, A. et al. Cirrhosis-associated immune dysfunction. Nat. Rev. Gastroenterol. Hepatol. 19, 112–134 (2022).
Van der Merwe, S. et al. The multifactorial mechanisms of bacterial infection in decompensated cirrhosis. J. Hepatol. 75, S82–S100 (2021).
Arroyo, V. et al. Acute-on-chronic liver failure: a new syndrome that will re-classify cirrhosis. J. Hepatol. 62, S131–S143 (2015).
Ferrer, R. et al. Empiric antibiotic treatment reduces mortality in severe sepsis and septic shock from the first hour: results from a guideline-based performance improvement program. Crit. Care Med. 42, 1749–1755 (2014).
Arabi, Y. M. et al. Antimicrobial therapeutic determinants of outcomes from septic shock among patients with cirrhosis. Hepatology 56, 2305–2315 (2012).
Fernandez, J. et al. Multidrug-resistant bacterial infections in patients with decompensated cirrhosis and with acute-on-chronic liver failure in Europe. J. Hepatol. 70, 398–411 (2019).
Piano, S. et al. Epidemiology and effects of bacterial infections in patients with cirrhosis worldwide. Gastroenterology 156, 1368–1380 e10 (2019).
Jalan, R. et al. Bacterial infections in cirrhosis: a position statement based on the EASL Special Conference 2013. J. Hepatol. 60, 1310–1324 (2014).
Koulaouzidis, A. et al. Spontaneous bacterial peritonitis. Postgrad. Med. J. 83, 379–383 (2007).
Rimola, A. et al. Diagnosis, treatment and prophylaxis of spontaneous bacterial peritonitis: a consensus document. Int. Ascites Club. J. Hepatol. 32, 142–153 (2000).
Bobadilla, M., Sifuentes, J. & Garcia-Tsao, G. Improved method for bacteriological diagnosis of spontaneous bacterial peritonitis. J. Clin. Microbiol. 27, 2145–2147 (1989).
Cohen, J. et al. Sepsis: a roadmap for future research. Lancet Infect. Dis. 15, 581–614 (2015).
Shallcross, L. & O’Brien, A. Antimicrobial resistance in liver disease: better diagnostics are needed. Lancet Gastroenterol. Hepatol. 2, 151–153 (2017).
Land, K. J. et al. REASSURED diagnostics to inform disease control strategies, strengthen health systems and improve patient outcomes. Nat. Microbiol. 4, 46–54 (2019).
Vasala, A., Hytonen, V. P. & Laitinen, O. H. Modern tools for rapid diagnostics of antimicrobial resistance. Front. Cell Infect. Microbiol. 10, 308 (2020).
Candel, F. J. et al. The role of rapid multiplex molecular syndromic panels in the clinical management of infections in critically ill patients: an experts-opinion document. Crit. Care 28, 440 (2024).
Burnham, C. D. et al. Diagnosing antimicrobial resistance. Nat. Rev. Microbiol 15, 697–703 (2017).
Wheeler, N. E. et al. Innovations in genomic antimicrobial resistance surveillance. Lancet Microbe 4, e1063–e1070 (2023).
Wu, H. X. et al. Clinical evaluation of metagenomic next-generation sequencing method for the diagnosis of suspected ascitic infection in patients with liver cirrhosis in a clinical laboratory. Microbiol Spectr. 11, e0294622 (2023).
Themoula Charalampous, G. L. K. & O’Grady, Justin Applying clinical metagenomics for the detection and characterisation of respiratory infections. in Eur Respir Monogr, (2019).
Moragues-Solanas, L. et al. Development and proof-of-concept demonstration of a clinical metagenomics method for the rapid detection of bloodstream infection. BMC Med Genomics 17, 71 (2024).
Street, T. L. et al. Clinical metagenomic sequencing for species identification and antimicrobial resistance prediction in orthopedic device infection. J. Clin. Microbiol. 60, e0215621 (2022).
Ko, K. K. K., Chng, K. R. & Nagarajan, N. Metagenomics-enabled microbial surveillance. Nat. Microbiol 7, 486–496 (2022).
Stalder, T. et al. Linking the resistome and plasmidome to the microbiome. ISME J. 13, 2437–2446 (2019).
Valik, J. K. et al. Genomic virulence markers are associated with severe outcomes in patients with Pseudomonas aeruginosa bloodstream infection. Commun. Med. 4, 264 (2024).
de Nies, L. et al. PathoFact: a pipeline for the prediction of virulence factors and antimicrobial resistance genes in metagenomic data. Microbiome 9, 49 (2021).
Chiu, C. Y. & Miller, S. A. Clinical metagenomics. Nat. Rev. Genet. 20, 341–355 (2019).
Napit, R. et al. Metagenomic analysis of human, animal, and environmental samples identifies potential emerging pathogens, profiles antibiotic resistance genes, and reveals horizontal gene transfer dynamics. Sci. Rep. 15, 12156 (2025).
Charalampous, T. et al. Evaluating the potential for respiratory metagenomics to improve treatment of secondary infection and detection of nosocomial transmission on expanded COVID-19 intensive care units. Genome Med 13, 182 (2021).
von Fricken, M. E. et al. Editorial: metagenomics for epidemiological surveillance in One Health. Front. Microbiol. 14, 1191946 (2023).
Selway, C. A., Eisenhofer, R. & Weyrich, L. S. Microbiome applications for pathology: challenges of low microbial biomass samples during diagnostic testing. J. Pathol. Clin. Res. 6, 97–106 (2020).
Yang, C.J. et al. 16S rRNA next-generation sequencing may not be useful for examining suspected cases of spontaneous bacterial peritonitis. Medicina 60, (20240.
Campbell, K. A., Trivedi, H. D. & Chopra, S. Infections in cirrhosis: a guide for the clinician. Am. J. Med 134, 727–734 (2021).
Chen, H. et al. Early application of metagenomics next-generation sequencing may significantly reduce unnecessary consumption of antibiotics in patients with fever of unknown origin. BMC Infect. Dis. 23, 478 (2023).
Butler, I. et al. Standardization of 16S rRNA gene sequencing using nanopore long read sequencing technology for clinical diagnosis of culture negative infections. Front Cell Infect. Microbiol 15, 1517208 (2025).
AWHO collaborative study to evaluate the candidate WHO International Reference Reagent for DNA extraction of the Gut Microbiome. Available from: https://www.who.int/publications/m/item/who-bs-2023.2455 (2023).
A WHO collaborative study to evaluate the candidate 1st WHO International Reference Reagents for Gut Microbiome analysis by Next-Generation Sequencing. Available from: https://www.who.int/publications/m/item/who-bs-2022.2416 (2023).
Feldgarden, M. et al. Validating the AMRFinder tool and resistance gene database by using antimicrobial resistance genotype-phenotype correlations in a collection of isolates. Antimicrob Agents Chemother. 63, (2019).
Frost, I. et al. Status, challenges and gaps in antimicrobial resistance surveillance around the world. J. Glob. Antimicrob. Resist. 25, 222–226 (2021).
Hassall, J. et al. Limitations of current techniques in clinical antimicrobial resistance diagnosis: examples and future prospects. NPJ Antimicrob. Resist. 2, 16 (2024).
Govender, K. N. et al. Metagenomic sequencing as a pathogen-agnostic clinical diagnostic tool for infectious diseases: a systematic review and meta-analysis of diagnostic test accuracy studies. J. Clin. Microbiol 59, e0291620 (2021).
Loit, K. et al. Relative Performance of MinION (Oxford Nanopore Technologies) versus Sequel (Pacific Biosciences) Third-Generation Sequencing Instruments in Identification of Agricultural and Forest Fungal Pathogens. Appl. Environ. Microbiol. 85 (2019).
Xu, F. et al. Impact of metagenomics next-generation sequencing on etiological diagnosis and early outcomes in sepsis. J. Transl. Med. 23, 394 (2025).
Patel, V. C. & Williams, R. Antimicrobial resistance in chronic liver disease. Hepatol. Int. 14, 24–34 (2020).
Fernandez, J. et al. Management of bacterial and fungal infections in cirrhosis: The MDRO challenge. J. Hepatol. 75, S101–S117 (2021).
Charalampous, T. et al. Routine metagenomics service for ICU patients with respiratory infection. Am. J. Respir. Crit. Care Med. 209, 164–174 (2024).
Deng, Q. et al. Nanopore-based metagenomic sequencing for the rapid and precise detection of pathogens among immunocompromised cancer patients with suspected infections. Front. Cell Infect. Microbiol. 12, 943859 (2022).
Wu, H. et al. The role of metagenomic next-generation sequencing in diagnosing and managing post-kidney transplantation infections. Front. Cell Infect. Microbiol. 14, 1473068 (2024).
Acknowledgements
V.C.P. has received funding support by King’s College Hospital Charity (Registered charity number: 1165593) and the Foundation for Liver Research (Registered charity number: 268211/1134579).
Author information
Authors and Affiliations
Contributions
Conceptualisation: A.L.L. Formal analysis: M.M., C.S. Investigation: A.L.L. Project administration: M.M., V.C.P. Supervision: C.S., V.C.P. Visualization: M.M. Writing—original draft: M.M., C.S. Writing—review and editing: A.L.L.
Corresponding author
Ethics declarations
Competing interests
V.C.P. declares consultancy roles with Resolution Therapeutics, Emles Bioventures, AlfaSigma S.p.A., AstraZeneca, Norgine Pharmaceuticals Ltd, and Menarini Diagnostics Ltd.
Transparency declarations
VCP has delivered paid lectures for Norgine Pharmaceuticals Ltd, Alfasigma S.p.A. and Menarini Diagnostics Ltd. The views expressed in the publication are those of the author(s) and not necessarily those of the NHS, DHSC, ‘arms’ length bodies, DSIT or other government departments.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mohamad, M., Sergaki, C. & Patel, V.C. Enhancing infection diagnostics in advanced chronic liver disease: harnessing clinical metagenomics for rapid pathogen and antimicrobial resistance detection. npj Antimicrob Resist 4, 3 (2026). https://doi.org/10.1038/s44259-025-00171-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44259-025-00171-7