Abstract
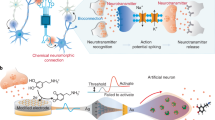
Complex organisms perceive their surroundings with sensory neurons that encode physical stimuli into spikes of electrical activities. The past decades have seen a throve of computing approaches taking inspiration from neurons, including reports of DNA-based chemical neurons that mimic artificial neural networks with chemical reactions. Yet, they lack the physical sensing and temporal coding of sensory biological neurons. Here we report a thermosensory chemical neuron based on DNA and enzymes that spikes with chemical activity when exposed to cold. Surprisingly, this chemical neuron shares deep mathematical similarities with a toy model of a cold nociceptive neuron: they follow a similar bifurcation route between rest and oscillations and avoid artefacts associated with canonical bifurcations (such as irreversibility, damping or untimely spiking). We experimentally demonstrate this robustness by encoding—digitally and analogically—thermal messages into chemical waveforms. This chemical neuron could pave the way for implementing the third generation of neural network models (spiking networks) in DNA and opens the door for associative learning.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Data supporting the findings of this study are available within the Article and its Supplementary Information. Source data are provided with this paper.
Code availability
The Mathematica notebooks used for the simulations performed in this study are available in the Supplementary Information.
References
Vriens, J., Nilius, B. & Voets, T. Peripheral thermosensation in mammals. Nat. Rev. Neurosci. 15, 573–589 (2014).
Wechselberger, M., Wright, C. L., Bishop, G. A. & Boulant, J. A. Ionic channels and conductance-based models for hypothalamic neuronal thermosensitivity. Am. J. Physiol. Regul. Integr. Comp. Physiol. 291, R518–R529 (2006).
Montell, C. Drosophila TRP channels. Pflüg. Arch. 451, 19–28 (2005).
Voets, T. et al. The principle of temperature-dependent gating in cold- and heat-sensitive TRP channels. Nature 430, 748–754 (2004).
McKemy, D. D. in TRP Ion Channel Function in Sensory Transduction and Cellular Signaling Cascades Ch. 12 (Taylor & Francis, 2007).
Roy, K., Jaiswal, A. & Panda, P. Towards spike-based machine intelligence with neuromorphic computing. Nature 575, 607–617 (2019).
Grassia, F., Levi, T., Doukkali, E. & Kohno, T. Spike pattern recognition using artificial neuron and spike-timing-dependent plasticity implemented on a multi-core embedded platform. Artif. Life Rob. 23, 200–204 (2018).
Merolla, P. A. et al. A million spiking-neuron integrated circuit with a scalable communication network and interface. Science 345, 668–673 (2014).
Davies, M. et al. Loihi: a neuromorphic manycore processor with on-chip learning. IEEE Micro 38, 82–99 (2018).
Perez-Nieves, N. & Goodman, D. Sparse spiking gradient descent. Adv. Neural Inf. Process. Syst. 34, 11795–11808 (2021).
Stromatias, E. et al. Robustness of spiking deep belief networks to noise and reduced bit precision of neuro-inspired hardware platforms. Front. Neurosci. 9, 222 (2015).
Adamatzky, A., Fullarton, C., Phillips, N., De Lacy Costello, B. & Draper, T. C. Thermal switch of oscillation frequency in Belousov–Zhabotinsky liquid marbles. R. Soc. Open Sci. 6, 190078 (2019).
Gentili, P. L., Horvath, V., Vanag, V. K. & Epstein, I. R. Belousov-Zhabotinsky ‘chemical neuron’ as a binary and fuzzy logic processor. Int. J. Unconv. Comput. 8, 177–192 (2012).
Hjelmfelt, A., Weinberger, E. D. & Ross, J. Chemical implementation of neural networks and Turing machines. Proc. Natl Acad. Sci. USA 88, 10983–10987 (1991).
Rothemund, P. W. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Genot, A. J., Bath, J. & Turberfield, A. J. Reversible logic circuits made of DNA. J. Am. Chem. Soc. 133, 20080–20083 (2011).
Qian, L. & Winfree, E. Scaling up digital circuit computation with DNA strand displacement cascades. Science 332, 1196–1201 (2011).
Seelig, G., Soloveichik, D., Zhang, D. Y. & Winfree, E. Enzyme-free nucleic acid logic circuits. Science 314, 1585–1588 (2006).
Cherry, K. M. & Qian, L. Scaling up molecular pattern recognition with DNA-based winner-take-all neural networks. Nature 559, 370–376 (2018).
Lopez, R., Wang, R. & Seelig, G. A molecular multi-gene classifier for disease diagnostics. Nat. Chem. 10, 746–754 (2018).
Okumura, S. et al. Nonlinear decision-making with enzymatic neural networks. Nature 610, 496–501 (2022).
Xiong, X. et al. Molecular convolutional neural networks with DNA regulatory circuits. Nat. Mach. Intell. 4, 625–635 (2022).
de Polavieja, G. G., Harsch, A., Kleppe, I., Robinson, H. P. & Juusola, M. Stimulus history reliably shapes action potential waveforms of cortical neurons. J. Neurosci. 25, 5657–5665 (2005).
Lankarany, M., Al-Basha, D., Ratté, S. & Prescott, S. A. Differentially synchronized spiking enables multiplexed neural coding. Proc. Natl Acad. Sci. USA 116, 10097–10102 (2019).
Caporale, N. & Dan, Y. Spike timing–dependent plasticity: a Hebbian learning rule. Annu. Rev. Neurosci. 31, 25–46 (2008).
Lobato-Dauzier, N., Cazenille, L., Fujii, T., Genot, A. & Aubert-Kato, N. Temperature-based inputs for molecular reservoir computers. In Proc. ALIFE 2020: The 2020 Conference on Artificial Life. ALIFE 2020: The 2020 Conference on Artificial Life 420–422 (ASME, 2020).
Maguire, O. R. et al. Dynamic environments as a tool to preserve desired output in a chemical reaction network. Chem. Eur. J. 26, 1676–1682 (2020).
Maguire, O. R., Wong, A. S., Westerdiep, J. H. & Huck, W. T. Early warning signals in chemical reaction networks. Chem. Commun. 56, 3725–3728 (2020).
Lemarchand, A., Berthoumieux, H., Jullien, L. & Gosse, C. Chemical mechanism identification from frequency response to small temperature modulation. J. Phys. Chem. A 116, 8455–8463 (2012).
Platkov, M. & Gruebele, M. Periodic and stochastic thermal modulation of protein folding kinetics. J. Chem. Phys. 141, 035103 (2014).
Closa, F., Gosse, C., Jullien, L. & Lemarchand, A. Identification of two-step chemical mechanisms and determination of thermokinetic parameters using frequency responses to small temperature oscillations. J. Chem. Phys. 138, 244109 (2013).
Paricio-Montesinos, R. et al. The sensory coding of warm perception. Neuron 106, 830–841 (2020).
Knowlton, W. M. et al. A sensory-labeled line for cold: TRPM8-expressing sensory neurons define the cellular basis for cold, cold pain, and cooling-mediated analgesia. J. Neurosci. 33, 2837–2848 (2013).
Bick, C., Goodfellow, M., Laing, C. R. & Martens, E. A. Understanding the dynamics of biological and neural oscillator networks through exact mean-field reductions: a review. J. Math. Neurosci. 10, 9 (2020).
Stiefel, K. M. & Ermentrout, G. B. Neurons as oscillators. J. Neurophysiol. 116, 2950–2960 (2016).
Fujii, T. & Rondelez, Y. Predator–prey molecular ecosystems. ACS Nano 7, 27–34 (2013).
Aufinger, L., Brenner, J. & Simmel, F. C. Complex dynamics in a synchronized cell-free genetic clock. Nat. Commun. 13, 2852 (2022).
Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000).
Kim, J. & Winfree, E. Synthetic in vitro transcriptional oscillators. Mol. Syst. Biol. 7, 465 (2011).
Montagne, K., Plasson, R., Sakai, Y., Fujii, T. & Rondelez, Y. Programming an in vitro DNA oscillator using a molecular networking strategy. Mol. Syst. Biol. 7, 466 (2011).
Srinivas, N., Parkin, J., Seelig, G., Winfree, E. & Soloveichik, D. Enzyme-free nucleic acid dynamical systems. Science 358, eaal2052 (2017).
Tayar, A. M., Karzbrun, E., Noireaux, V. & Bar-Ziv, R. H. Synchrony and pattern formation of coupled genetic oscillators on a chip of artificial cells. Proc. Natl Acad. Sci. USA 114, 11609–11614 (2017).
Weitz, M. et al. Diversity in the dynamical behaviour of a compartmentalized programmable biochemical oscillator. Nat. Chem. 6, 295–302 (2014).
Padirac, A. Tailoring Spatio-Temporal Dynamics with DNA Circuits. PhD thesis, Univ. Claude Bernard-Lyon I (2012).
Dehne, H., Reitenbach, A. & Bausch, A. Reversible and spatiotemporal control of colloidal structure formation. Nat. Commun. 12, 6811 (2021).
Yamagata, A., Masui, R., Kakuta, Y., Kuramitsu, S. & Fukuyama, K. Overexpression, purification and characterization of RecJ protein from Thermus thermophilus HB8 and its core domain. Nucleic Acids Res. 29, 4617–4624 (2001).
Swiler, L. P. & Wyss, G. D. A user’s guide to Sandia’s latin hypercube sampling software: LHS UNIX library/standalone version (2004).
McKemy, D. D., Neuhausser, W. M. & Julius, D. Identification of a cold receptor reveals a general role for TRP channels in thermosensation. Nature 416, 52–58 (2002).
Peier, A. M. et al. A TRP channel that senses cold stimuli and menthol. Cell 108, 705–715 (2002).
McGahan, K. & Keener, J. A mathematical model analyzing temperature threshold dependence in cold sensitive neurons. PLoS ONE 15, e0237347 (2020).
Finke, C., Freund, J. A., Rosa, E. Jr, Braun, H. A. & Feudel, U. On the role of subthreshold currents in the Huber–Braun cold receptor model. Chaos 20, 045107 (2010).
Wang, J., Liu, S., Wangy, H. & Zeng, Y. Dynamical properties of firing patterns in the Huber-Braun cold receptor model in response to external current stimuli. Neural Netw. World 25, 641–655 (2015).
Hodgkin, A. L. & Huxley, A. F. A quantitative description of membrane current and its application to conduction and excitation in nerve. J. Physiol. 117, 500 (1952).
Tsuji, S., Ueta, T., Kawakami, H., Fujii, H. & Aihara, K. Bifurcations in two-dimensional Hindmarsh–Rose type model. Int. J. Bifurc. Chaos 17, 985–998 (2007).
Hindmarsh, J. & Rose, R. A model of the nerve impulse using two first-order differential equations. Nature 296, 162–164 (1982).
Tan, E. et al. Specific versus nonspecific isothermal DNA amplification through thermophilic polymerase and nicking enzyme activities. Biochemistry 47, 9987–9999 (2008).
Bansho, Y., Furubayashi, T., Ichihashi, N. & Yomo, T. Host–parasite oscillation dynamics and evolution in a compartmentalized RNA replication system. Proc. Natl Acad. Sci. USA 113, 4045–4050 (2016).
Furubayashi, T. et al. Emergence and diversification of a host-parasite RNA ecosystem through Darwinian evolution. eLife 9, e56038 (2020).
Brette, R. Philosophy of the spike: rate-based vs. spike-based theories of the brain. Front. Syst. Neurosci. 9, 151 (2015).
Lobato-Dauzier, N. et al. Silicon chambers for enhanced incubation and imaging of microfluidic droplets. Lab Chip 23, 2854–2865 (2023).
Tanaka, G. et al. Recent advances in physical reservoir computing: a review. Neural Netw. 115, 100–123 (2019).
Naqib, F. et al. Tunable oscillations and chaotic dynamics in systems with localized synthesis. Phys. Rev. E 85, 046210 (2012).
Brette, R. Computing with neural synchrony. PLoS Comput. Biol. 8, e1002561 (2012).
Dupin, A. & Simmel, F. C. Signalling and differentiation in emulsion-based multi-compartmentalized in vitro gene circuits. Nat. Chem. 11, 32–39 (2019).
Joesaar, A. et al. DNA-based communication in populations of synthetic protocells. Nat. Nanotechnol. 14, 369–378 (2019).
Valet, M., Pontani, L.-L., Voituriez, R., Wandersman, E. & Prevost, A. M. Diffusion through nanopores in connected lipid bilayer networks. Phys. Rev. Lett. 123, 088101 (2019).
Gines, G. et al. Microscopic agents programmed by DNA circuits. Nat. Nanotechnol. 12, 351–359 (2017).
Baccouche, A. et al. Massively parallel and multiparameter titration of biochemical assays with droplet microfluidics. Nat. Protoc. 12, 1912–1932 (2017).
Genot, A. et al. High-resolution mapping of bifurcations in nonlinear biochemical circuits. Nat. Chem. 8, 760–767 (2016).
Peng, T. et al. A BaSiC tool for background and shading correction of optical microscopy images. Nat. Commun. 8, 14836 (2017).
Preibisch, S., Saalfeld, S. & Tomancak, P. Globally optimal stitching of tiled 3D microscopic image acquisitions. Bioinformatics 25, 1463–1465 (2009).
Padirac, A., Fujii, T. & Rondelez, Y. Quencher-free multiplexed monitoring of DNA reaction circuits. Nucleic Acids Res. 40, e118 (2012).
Acknowledgements
We thank K. Aihara, T. Kohno, I. Kawamata, Y. Sato, A. Estevez-Torres, J.-C. Galas and R. Brette for discussion on the manuscript. We thank FEMTO-ST (CNRS, Besancon) for the fabrication of the silicon microchambers in the frame of the RENATECH network. Neurons and ionic channels in Fig. 1 were adapted from BioRender.com. This research was supported by JSPS KAKENHI grant no. JP19KK0261 (N.A.-K.), JP20K12061 (N.A.-K. and A.J.G.), CNRS MITI Interdisciplinary Program on Biomimetism (G.G., T.L. and A.J.G.), PEPR MoleculArxiv grant no. ANR-22-PEXM-0002 (A.J.G. and Y.R.) and ERC Union Horizon 2020 grant no. 770940 (N.L.-D.).
Author information
Authors and Affiliations
Contributions
Conceptualization: N.L.-D., A.B., G.G., T.L., Y.R., N.A.-K., A.J.G. Methodology: N.L.-D., A.J.G. Investigation: N.L.-D. Visualization: N.L.-D. Funding acquisition: G.G., T.L., N.A.-K., A.J.G. Supervision: T.F., S.H.K., N.A.-K., A.J.G. Writing (original draft): N.L.-D., N.A.-K., A.J.G.
Corresponding authors
Ethics declarations
Competing interests
T.F., Y.R. and G.G. have filed a patent on the PEN DNA toolbox (patent no. WO2017141067A1, filed in Europe, Japan, Lithuania, USA, Canada).
Peer review
Peer review information
Nature Chemical Engineering thanks Lucia Marucci, Chunhai Fan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Sections 1–9, Figs. 1–28, Tables 1–3, theoretical models and discussion.
Supplementary Data 1 (download JPG )
Image of the barcoding of exonuclease concentration in encapsulated chemical neurons.
Supplementary Video 1 (download MP4 )
Chemical neuron bifurcation path.
Supplementary Video 2 (download MP4 )
Biological neuron bifurcation path.
Supplementary Video 3 (download MP4 )
Encapsulated chemical neurons in a temperature gradient.
Supplementary Code 2
Mathematica notebook for chemical neuron toy model.
Supplementary Code 2
Mathematica notebook for biological neuron toy model.
Source data
Source Data Fig. 1 (download CSV )
Statistical source data.
Source Data Fig. 4 (download CSV )
Statistical source data.
Source Data Fig. 5 (download CSV )
Statistical source data.
Source Data Fig. 6 (download CSV )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lobato-Dauzier, N., Baccouche, A., Gines, G. et al. Neural coding of temperature with a DNA-based spiking chemical neuron. Nat Chem Eng 1, 510–521 (2024). https://doi.org/10.1038/s44286-024-00087-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s44286-024-00087-5
This article is cited by
-
A recursive enzymatic competition network capable of multitask molecular information processing
Nature Chemistry (2025)
-
Oscillating chemical reaction networks stopped cold
Nature Chemical Engineering (2024)