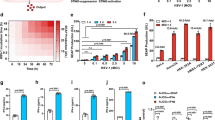
Abstract
Nanoplasmonic biosensors are powerful tools for virus research. This Perspective outlines key sensor design principles, including platform geometry, probe optimization, signal amplification, and machine learning-aided analysis, and connects them with emerging applications in virus detection, antiviral drug testing, and immune profiling. Strategic needs such as clinical validation, portable sensor development, and expanded virus coverage are also discussed to guide future efforts toward translational impact and practical implementation in real-world settings.
Similar content being viewed by others
Introduction
Nanoplasmonic biosensors are widely used to detect biological materials such as viruses, exosomes, bacteria, proteins, nucleic acids, and lipids1. While many types of biosensors exist, nanoplasmonic biosensors are highly regarded due to features such as label-free detection, rapid data acquisition, simple instrumentation, robust operation, and high surface sensitivity2,3. The coronavirus disease 2019 (COVID-19) pandemic heightened interest in developing nanoplasmonic biosensors for virus applications and research progress has accelerated over the past few years, especially with ongoing advances on the nanofabrication side that are boosting large-scale, cost-effective production possibilities for various nanostructures such as nanoholes, nanodisks, and nanospikes as well as facilitating integration into high-throughput formats such as 96-well plates.
This research progress has been stimulated by the emerging capabilities of nanoplasmonic biosensors as well as by recognizing that improved virus detection tools would be useful. Conventionally, viral infections are diagnosed by viral nucleic acid detection based on the polymerase chain reaction (PCR) assay, which has high sensitivity and specificity but requires a lab infrastructure, or by lateral flow immunoassays (LFAs) that are rapid but only qualitatively detect the presence of viral antigens such as viral surface proteins4,5. There are applications such as measuring patient responses to antiviral therapies where rapid quantification of viral antigens is needed. Likewise, quantitative measurement of virus-specific antibody levels is important to assess the host immune response and vaccine-induced protective efficacy. While such measurements can be performed using laboratory-based serological assays, less resource-intensive measurement strategies with quick readouts would be advantageous. Label-free biosensors have attractive merits to address these needs and there are promising options like electrochemical and nanoplasmonic biosensors. While electrochemical biosensors provide high sensitivity for virus detection6, nanoplasmonic biosensors stand out for their ability to interface with virus particles at the nanoscale and to enable applications that extend beyond detection. Nanoplasmonic biosensors also offer the added capability of multiplexed measurements that can be done in highly parallel formats and combined with the growing use of optical imaging-based readouts and machine learning-aided data analysis7,8.
Until recently, the primary focus of nanoplasmonic biosensors and related optical biosensors for virus applications was limited to detection9,10,11,12,13,14. While virus detection remains a critical area, we believe that nanoplasmonic sensors are competitively positioned to advance other areas of virus research as well. Examples that will be covered in this Perspective include antiviral drug evaluation and immune profiling, the latter of which has gained significant momentum due to the COVID-19 pandemic and highlights the multifaceted ways in which nanoplasmonic sensors are helping to address global health challenges. These ongoing advances also lay the groundwork to utilize next-generation nanoplasmonic biosensors in real-world virus applications.
To cover these points, we begin by introducing the measurement principles of nanoplasmonic biosensors and particular design strategies for virus applications. We then provide a detailed assessment of recent progress in utilizing nanoplasmonic biosensors across different areas of virus research. Particular emphasis is placed on sensing platform design concepts and drawing out broader themes. We conclude by critically evaluating the current state of the field, identifying key challenges and unmet needs, and proposing promising directions for future research. This analysis leads us to imagine future possibilities about how nanoplasmonic biosensors can enhance the response framework to emerging viral threats. In closing, we also briefly draw parallels to exosome studies, emphasizing the broad applicability of the biosensing issues discussed.
Benefits of nanoplasmonic biosensors for studying viruses
Nanoplasmonic biosensors have unique sensing features that are useful for studying viruses and we introduce key measurement principles underpinning these features. The sensing platforms are composed of metallic nanostructures and illumination with incident light can excite conduction-band electrons near the metal surface3. This electronic excitation creates a collective oscillation of conduction-band electrons, which amplifies the electromagnetic field intensity near the sensor surface, and controlling the nanoscale architecture of the sensing platform leads to many design possibilities.
Indeed, practical use of nanoplasmonic biosensors has been aided by ongoing advances in nanofabrication to increase engineering design possibilities as well as improve production speed, scalability, cost, and quality control15,16. For nanoparticle-type structures, there are various strategies that involve controlled deposition of solution-phase nanoparticles or bottom-up fabrication of nanostructures based on nanosphere lithography or colloidal hole-mask lithography. One advanced example is indirect nanoplasmonic sensing, which involves depositing a conformal, thin dielectric layer on top of the entire sensor surface after nanoparticle deposition17. This approach improves the physical and chemical stability of plasmonic nanoparticle transducers under the dielectric coating and permits nanoplasmonic biosensing experiments at the dielectric-solution interface. On the other hand, for nanohole-type structures, the electron beam lithography and focused ion beam techniques were pioneering top-down options but are relatively low throughput and expensive. Nanoimprinting, template-stripping, and colloidal lithography can be employed to fabricate nanoholes over larger surface areas, with more affordable cost. Depending on the application, the nanohole surface can also be covered with a dielectric coating. Together, these advances enable the affordable fabrication of plasmonic surfaces that can be useful for biosensing applications.
Depending on the type of metallic nanostructures (e.g., nanoholes or nanoparticles) as well as specific design features (e.g., shape, periodicity, ordering, height, composition), various nanoplasmonic sensing phenomena such as localized surface plasmon resonance (LSPR) and extraordinary optical transmission (EOT) can occur and involve one or more localized and/or propagating surface plasmon modes18. The spectral features will vary accordingly but they generally exhibit spectral features such as dips or peaks that are highly sensitive to changes in the local refractive index near the sensor surface, i.e., within the region of nanoplasmonic-enhanced field intensity. These features are useful to track in biosensing applications as biomacromolecules typically have higher refractive index values than solvent molecules19,20. Conventionally, nanoplasmonic biosensing experiments measure wavelength shifts in particular spectral features or changes in the intensity at a certain wavelength, which occur due to changes in the local refractive index.
A key advantage of nanoplasmonic biosensors over conventional surface plasmon resonance (SPR) biosensors is their high surface sensitivity. The enhanced field intensity is tightly confined to the sensor surface, with decay lengths as short as 5–10 nm depending on the sensor geometry21. This feature is advantageous for studying the conformational properties of proteins and lipid vesicles, including quantitatively profiling adsorption-related shape deformation events that is difficult to achieve with other widely used surface-sensitive measurement techniques such as quartz crystal microbalance-dissipation (QCM-D) due to hydrodynamic effects22,23.
These capabilities have also proven useful for investigating multivalency-induced shape deformation of virus-mimicking lipid vesicles that interact with receptor-functionalized membrane interfaces24,25. Such biophysical interactions bear a resemblance to viral infection and viral genome replication processes that occur at lipid membrane interfaces. Furthermore, like lipid vesicles and exosomes, virus particles (~50-300 nm diameter) fall within the optimal size range for detection and biophysical characterization by nanoplasmonic biosensors. While the full potential of nanoplasmonic biosensors for studying viruses remains underexplored, there has been extensive progress in certain application scopes that are linked to the sensing platform design.
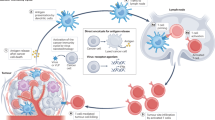
Figure 1 presents an overview of these application scopes and outlines important factors that influence sensing performance in each area. Historically, virus detection was the main topic and usually involves functionalizing the sensor surface with antibody probes that specifically recognize certain viral structural proteins (“antigens”). The viral antigens may be present in either soluble form or as part of larger virus particles and successful detection of either species is usually evidence of an ongoing viral infection. Instead of antibodies, other probes like smaller nanobodies or nucleic acid aptamers may also be used. The key sensing priority overall is to selectively capture the target antigen and prevent nonspecific adsorption of other biological materials, which is especially challenging in biological fluids. Virus detection applications have mainly utilized the high surface sensitivity of nanoplasmonic biosensors to achieve low detection limits.
A Overview of different application scopes. Virus detection was the earliest application area and involves functionalizing nanoplasmonic sensor chips with virus-capturing probes like antibodies or aptamers. The probes selectively recognize viral antigens such as viral surface proteins to capture virus particles, which can be detected by changes in the local refractive index near the sensor surface. The ability to capture virus particles in nanoplasmonic sensing platforms creates opportunities to further study the mechanisms of how antiviral drugs inhibit virus particles or characterize virus binding interactions, which are important for drug development. Analogous to how nanoplasmonic biosensors detect virus particles, these virus-drug interactions can be monitored by tracking changes in the local refractive index, which occur due to events like drug binding or virus particle disruption. In addition to capturing virus particles, nanoplasmonic sensor chips can also be functionalized with recombinant viral antigens or other suitable probes to detect circulating, virus-specific antibodies. In this case, both antibody type and concentration should be measured to determine the degree of protective immunity. This approach is also useful for measuring vaccination efficacy and testing the performance of therapeutic antibodies. Created in BioRender. Park, H. (2025) https://BioRender.com/3sblar2. B Schematic examples of virus particle capture on nanoparticle- and nanohole-based sensing platforms. Yellow shading indicates region of nanoplasmonic-enhanced electromagnetic field intensity near the sensor surface. Note that modified versions of nanoholes are sometimes referred to as nanowells or nanocups depending on the specific design. Created in BioRender. Park, H. (2025) https://BioRender.com/cn870io.
Another emerging direction of nanoplasmonic biosensors is antiviral drug evaluation where the high surface sensitivity creates unique measurement benefits compared to other biosensing options. In this case, virus particles and other experimental models mimicking viruses can be directly immobilized on the sensor surface and then the drug candidate of interest is added. The measurement emphasis is usually on tracking the interaction between the drug candidate and viral target, providing critical mechanistic insights. Appropriate selection of the nanoplasmonic sensing mode is essential for studying these interaction processes. In other cases, it may be important to indirectly assess the drug effect by measuring the disruption of a virus-cell interaction, for example.
Lastly, immune profiling is the newest and most active area of virus research for nanoplasmonic biosensors. The sensor surface is typically functionalized with a recombinant viral antigen to detect virus-specific antibodies that are circulating in blood. These capabilities are useful for diagnosing ongoing and past infections as well as for assessing protective immunity levels in communities and vaccination efficacy. An important distinction between virus detection and immune profiling lies in quantification. While virus quantification is advantageous for detection purposes, it is essential for immune profiling because antibody concentration levels must be sufficiently high for protection.
These considerations provide a broad understanding of different application scopes for virus research, which we now cover in depth to assess the latest research progress and ongoing developments.
Virus detection
Table 1 summarizes progress in utilizing nanoplasmonic biosensors for virus detection applications. In pioneering work, Yanik et al. showed that antibody-functionalized gold nanohole arrays could selectively detect small and large enveloped viruses such as vesicular stomatitis virus (VSV), pseudotyped Ebola, and vaccinia virus by utilizing EOT phenomena26. Key advantages of the measurement format included label-free readout, rapid response, high dynamic range, and compatibility with serum-containing media. Depending on the virus target, the sensor chip surface was functionalized with antibodies that recognized the viral surface proteins of a particular virus (VSV, Ebola, or vaccinia). Experimentally, it was demonstrated that viruses in the 106 to 109 plaque-forming units per mL (PFU/mL) range could be detected after ~90 min sample incubation with the sensor chip surface. Virus binding increased the local refractive index near the sensor surface, yielding wavelength shifts in the plasmonic resonance peak that ranged from ~4 to ~100 nm depending on the virus concentration. Notably, virus detection was highly sensitive to the antibody probe since negligible signal shifts (less than ~1 nm) occurred on bare sensor chips without antibody functionalization. Virus concentrations down to 106 PFU/mL were detected in serum-containing cell culture media and it was further estimated that detection sensitivity on the order of 105 PFU/mL or below could be possible based on the measured signal noise.
The clinical utility of nanoplasmonic biosensors for virus detection was further established when Inci et al. reported human immunodeficiency virus (HIV) detection in whole blood by utilizing a layer of antibody-functionalized, plasmonic gold nanoparticles27. The anti-gp120 antibody that binds to the HIV gp120 surface protein was used as the capture probe. Notably, sample preprocessing was not required and the assay had high sensitivity and reproducibility for evaluating cultured HIV subtype specimens and HIV-infected patient samples. After 60-min incubation with the virus-contaminated whole blood samples, the sensor chips were rinsed with saline and then endpoint measurements were recorded. There was minimal signal response to whole blood alone, yielding wavelength shifts in the plasmonic resonance peak of less than 1 nm. On the other hand, cultured HIV subtypes with concentrations in the range of 105–106 copies/mL caused significant ~4–9 nm peak shifts. It was further determined that the limits of detection were around ~100 and ~500 copies/mL for cultured HIV subtypes and for clinical HIV patient samples in whole blood, respectively, which are within clinically useful ranges for measuring the efficacy of antiviral drug therapies based on virus concentration levels found in patient blood. It was also discussed how the nanoplasmonic biosensing format provides a quicker readout and minimal sample processing compared to conventional PCR analysis.
The assay sensitivity was later improved by adopting a nanoparticle sandwiching strategy that involved virus particle capture by the sensing platform, followed by deposition of solution-phase, antibody-functionalized plasmonic nanoparticles on top of the virus particle layer28. The antibody recognized HBsAg, which is a hepatitis B virus (HBV) surface protein. This method successfully detected HBV in serum samples from infected patients. Experimentally, the sensor chips were incubated with serum samples for 60 min followed by buffer washing and then measurements were performed. It was possible to detect HBV specimens with concentrations ranging from 101 to 105 International Units per mL (IU/mL) and the corresponding wavelength shifts in plasmonic resonance peak ranged from 1.3 to 2.5 nm. By contrast, the diluted serum without virus yielded shifts of only 0.6 nm, which indicated selective detection of virus with statistically significant larger responses although the dependence of the measurement response on the virus concentration was rather weak. To improve the sensing performance, a sandwiching strategy was employed whereby viruses were first captured by the nanoparticle-coated sensor chip surface and then an additional layer of antibody-conjugated nanoparticles was added on top of the captured virus particles. For 104 and 105 IU/mL HBV specimens, this strategy increased the magnitude of the measurement responses by ≥50% and the larger wavelength shifts in the plasmonic resonance peak in these cases can be attributed to coupling of the LSPR-enhanced electromagnetic fields between the nanoparticle layers. Interestingly, the sensing platform was also adapted to a 96-well plate format, which can facilitate larger scale measurements in the future. Of note, the authors remarked that the observed detection limit of 101 IU/mL is also consistent with currently used PCR-based assays and such high sensitivity is required for antiviral treatment monitoring.
As reported by Hao et al., DNA aptamer-bearing dendrimers have also been attached to LSPR sensor chips in order to improve detection of the receptor binding domain (RBD) of the severe acute respiratory syndrome coronavirus 2 (SARS‑CoV‑2) spike (S) protein antigen by reducing nonspecific binding and increasing capture probe density29. A fast response time within 150 s was achieved and yielded a 6-time greater signal detecting 377 nM RBD antigen in saline compared to aptamers directly attached to gold nanoparticles without the dendrimer presentation. Importantly, the dendrimer presentation improved the limit of detection from 205 to 22 pM RBD antigen. The measurement platform was also useful for detecting pseudotyped SARS-CoV-2 in saline across the tested concentration range of 102–107 virus particles per mL (vp/mL), with a reported limit of detection of 148 vp/mL and similar detection performance was observed in an artificial saliva matrix as well. The latter example highlights the accelerating pace of nanoplasmonic biosensor developments in response to the COVID-19 pandemic and also demonstrates potential clinical utility since SARS-CoV-2 concentrations in nasopharyngeal and saliva swabbed samples are in the range of 104 to 1010 vp/mL.
Within this scope, Masterson et al. developed a multiplex assay that utilizes the LSPR properties of gold triangular nanoprisms to simultaneously detect ten SARS-CoV-2 biomarkers comprising viral nucleic acids, protein antigens, and antibodies in a single platform30. The capture probes consisted of antibodies to detect viral antigens or neutralizing antibodies and single-stranded DNA to detect RNA gene products. Initial measurements focused on incubating 10 nM RNA, antigen, or antibody targets in 10% plasma overnight and yielded peak shifts around ~10 nm after buffer washing. There were linear measurement responses over 10–7 to 10 nM analyte concentrations, indicating a high dynamic range. The corresponding limits of detection were around 90–980 aM for RNA genes, 211–246 aM for protein antigens, and 170–1000 aM for neutralizing antibodies. The measurement platform was also useful for detecting neutralizing antibodies in clinical patient samples, with ≥96% detection specificity achieved after 24 h incubation and subsequent buffer washing.
Li et al. have also fabricated a nanoplasmonic sensing platform that is composed of antibody-functionalized, vertically separated gold nanoholes and nanodisks in order to detect the SARS-CoV-2 RBD antigen31. Using this platform that had a mixture of propagating and localized surface plasmon modes, the binding affinities of 12 RBD variants were measured and showed variant-specific differences that could impact future assay design options. Experimentally, the measurements were made after 30 min incubation in saline and yielded concentration-dependent peak shifts of wild-type RBD antigen (10-2-104 ng/mL) ranging from ~0.2 to ~4 nm. Further investigation of 102 ng/mL RBD antigens showed ~2 to ~6 nm responses depending on the variant, which is related to antigen-antibody binding affinity. More detailed characterization of a target RBD variant showed a limit of detection of 27 pg/mL and a dynamic range of linear measurement response from 0.2 to 54.9 ng/mL.
Conversely, Liang et al. have explored affinity maturation strategies to engineer antibodies that bind more tightly to the SARS-CoV-2 RBD antigen for more sensitive virus detection32. The authors identified a relationship between refractive index and the optical density (OD) signal in order to conduct nanoplasmonic biosensing measurements based on OD signal intensity changes. Experimentally, the binding of a RBD-containing antigen to the antibody-functionalized sensor chip was measured over a 50 min interval, with signals reaching near saturation in ~25 min. Concentration-dependent OD signals were observed for 0.5–10 µg/mL antigen concentrations while the best-performing engineered antibody probe yielded >50% larger signals compared to the wild-type antibody. The engineered antibody profile also demonstrated stronger binding to the antigen target and the corresponding limit of detection was around 0.72 µg/mL. A similar performance improvement was observed for detecting pseudotype SARS-CoV-2 virus particles, with corresponding limits of detection of ~2500 and ~4000 vp/mL for engineered and wild-type antibody probes, respectively. In addition to OD measurements, a microscopic imaging analysis scheme was devised in tandem with the engineered antibody probe to improve limits of detection down to ~64 ng/mL and ~765 vp/mL for antigen and pseudotyped virus particles, respectively.
Instead of antibodies, Ma et al. have described the alternative use of smaller, more easily produced nanobodies as the capture probe to detect SARS-CoV-2 RBD antigen in serum based on LSPR measurements33. The antigen was incubated with the nanobody-functionalized surfaces for 30 min followed by buffer washing and then the measurement was performed. Two nanobody probes were developed and each one demonstrated highly selective binding of 0.01 µg/mL RBD antigen in saline (~2.6 nm shift) whereas other competing species such as albumins and immunoglobulins yielded negligible shifts (<0.5 nm). The probes demonstrated a wide dynamic range from 0.01 to 10,000 ng/mL antigen concentrations and maximum shift responses around 2.6–3.0 nm. Interestingly, the combination of both nanobody probes in a mixed platform yielded a larger measurement response around 3.3 nm and maintained similar detection performance in serum.
In addition, Lewis et al. have reported that an LSPR sensor chip comprising aptamer-modified gold nanoparticles can selectively detect the SARS-CoV-2 S1 protein subunit over the RBD domain and the S2 protein subunit, and further demonstrated sensing utility in spiked saliva and serum34. There was real-time monitoring of the antigen binding interactions to the aptamer-functionalized sensor chips while the response signal was recorded after ~13 min. The dynamic range of S1 antigen detection was 1–16 nM and the corresponding limit of detection was 0.26 nM. The measurement signal for 32 nM S1 antigen was a >650 response unit (RU) shift while equivalent mass concentrations of S2 or RBD antigens yielded <15 RU shifts. The measurement platform was also able to effectively detect S1 antigen in diluted saliva and serum matrices with >95% recovery.
Aside from gold nanostructures, Yang et al. demonstrated that a silver nanotriangle array could be functionalized with the human angiotensin-converting enzyme 2 (ACE2) protein, which is a natural cell membrane receptor for SARS-CoV-2 S protein binding35. The platform could detect SARS-CoV-2 spike RBD protein and human coronavirus (CoV) NL63 in buffer and saliva. Compared to other detection platforms, a unique aspect of the measurement strategy was 2 h incubation with the ACE2-functionalized sensor chip prior to aqueous rinsing and then the chip was air-dried prior to measurement. For RBD antigen, the sensing platform had a dynamic range from ~2 to ~9420 pM and the limit of detection was ~0.8 pM. As a more lab friendly whole virus substitute, human CoVs were tested in place of SARS-CoV-2. Importantly, the ACE2-binding CoV NL63 strain yielded a large, ~40 nm shift response whereas two other CoV strains that do not have ACE2 binding ability had negligible shifts (<2 nm). The limits of detection for CoV NL63 were 391 and 625 PFU/mL in buffer and saliva, respectively, demonstrating that the sensing platform had high performance even in complex matrices.
In addition to these various probe types, a recent innovation that may influence future probe design involves engineered peptides that cause plasmonic nanoparticle aggregation and colorimetric shifts that occur selectively in response to enzymatic cleavage by a norovirus or SARS-CoV-2 protease depending on the peptide sequence36,37.
Another important development has been applying machine learning techniques like neural network models to improve the detection of HBV surface antigen (HBsAg) in patient serum samples using antibody-functionalized sensing platforms38. In one study, Sun et al. reported that the neural network model improved assay limit of detection and accuracy for naked-eye detection applications and emphasized the modularity of the sensing platform. The antigen samples were incubated with the antibody-functionalized gold film for 30 min prior to transferring gold nanoparticles on top to create a sandwiched plasmonic structure. More bound antigen increased the gap thickness between the gold film and gold nanoparticles, which led to reduced reflectance.
Since antigen binding results in a weaker signal, it was beneficial to train the neural network model to improve the limit of detection for HBsAg antigen based on HBV-positive patient serum samples (1–106 IU/mL). A quasi-linear response was observed at lower antigen concentrations and the neural network model improved the limit of detection from 0.123 to 0.019 IU/mL and diagnostic accuracy from ~80% to >99% compared to manual analysis. Specifically, the neural network was trained on reflectance spectra collected for different HBsAg concentrations to overcome weak, nonlinear signal responses at low antigen concentrations, which improved prediction accuracy. Notably, the nanoplasmonic biosensor had superior results compared to LFA measurements performed in parallel, as indicated by a reduced smartphone-based error rate (1.2 vs. 2.9% for LFA) and higher sensitivity (~0.019 vs. >1 IU/mL or higher for LFAs).
Overall, judging from the reported data, several of the nanoplasmonic biosensors described above have demonstrated superior antigen detection sensitivity and dynamic range compared to LFAs. Moreover, the detected virus particle concentrations with the nanoplasmonic biosensing approach the measurement capabilities of more intensive, multi-day virus culturing assays.
From an instrumental perspective, there is also growing interest in developing low-cost, disposable nanoplasmonic biosensing chips for virus detection that are compatible with microplate readers and smartphone cameras39,40,41. Emerging ideas to improve detection performance include robot-like nanoparticles for enhanced virus particle capture42 and plasmonic nanomaterial-infused facemasks that exhibit visible color changes in response to pathogen exposure43,44,45.
Taken together, these studies highlight extensive progress in utilizing nanoplasmonic biosensors for virus detection and demonstrate how biosensor design in this context mainly focuses on (1) nanoplasmonic sensing mode, (2) capture probe type, and (3) antigen target. These design factors are essential for improving detection sensitivity, selectivity, and operability in biological fluids and continuing research in these directions is vital. At the same time, we suggest that nanoplasmonic biosensors have highly competitive merits for other virus applications, which necessitate further exploration.
Antiviral drug evaluation
A key example of these opportunities is antiviral drug testing. While most antiviral drugs inhibit viral genome replication, there is broad interest in developing new drug classes that directly target virus particles. Nanoplasmonic biosensors have proven useful for characterizing the mechanism and efficacy of antiviral drugs in this class, oftentimes providing superior measurement capabilities compared to other biosensing options. In contrast to detection-focused nanoplasmonic biosensors that mainly involve endpoint measurements, a unique feature of sensors directed at antiviral drug evaluation is real-time monitoring of virus-drug interactions to track corresponding kinetics, which help to elucidate the inhibitory mechanism. Although relatively few application examples exist at present, they clearly illustrate the analytical power of nanoplasmonic biosensing technologies within this scope and are summarized in Table 2. We begin by introducing membrane-disruptive antiviral peptides that can inhibit membrane-enveloped viruses based on curvature-selective membrane interactions46,47 and discuss the utility of different nanoplasmonic biosensing platforms.
The first nanoplasmonic biosensing platform for studying antiviral peptides was described by Goh et al. and involved a glass-supported gold nanodisk array48. A virus-mimicking lipid bilayer coating was fabricated on the sensor surface, mainly comprising lipid vesicles on top of the gold nanodisks while there were planar lipid bilayer segments on the glass surface between the nanodisks. The bare gold nanodisk platform had a bulk sensitivity of ~230 nm per refractive index unit (RIU) (nm/RIU). Transmission-mode LSPR measurements tracked the membrane-disruptive properties of an antiviral (AH) peptide towards this heterogenous platform composed of lipid vesicles and planar bilayer fragments. High AH peptide concentrations (13 µM) caused vesicle rupture, which is consistent with its viral envelope disruption mechanism. Surprisingly, however, the corresponding measurement response involved a small +0.1 nm wavelength shift in the plasmonic resonance peak due to the heterogenous lipid assembly and loss of lipid mass from the sensor surface. Conversely, low AH peptide concentrations (200 nM) were insufficient to cause vesicle rupture and only peptide binding was detected. Interestingly, the same experiment with 13 µM AH peptide on a titania-coated gold nanodisk array with a lower bulk sensitivity of ~110 nm/RIU yielded a larger measurement response. Subsequent AH peptide addition to the intact vesicle adlayer on the titania-coated gold nanodisk array caused vesicle rupture to form a planar bilayer across the entire sensor surface, not only on top of the nanodisks, which contributed to an amplified response of +1.3 nm (vs. +0.1 nm). This finding demonstrated the importance of the lipid platform architecture to sensitively detect membrane-peptide interactions because the titania-coated gold nanodisk array with lower bulk sensitivity yielded a much larger measurement response than the more sensitive, bare gold nanodisk array. This effect was due to the overall vesicle-to-bilayer transformation on the titania-coated gold nanodisk platform, which resulted in lipid mass being closer on average to the sensor surface.
Building on this work, Ferhan et al. developed a titania-coated gold nanodisk array on QCM-D sensor chips for simultaneous LSPR and QCM-D measurements49. The sensor chip had a bulk sensitivity of 150 nm/RIU and the combined measurement approach allowed more detailed tracking of the AH peptide-vesicle interaction process. The addition of 13 µM AH peptide to the intact vesicle adlayer caused a +1.2 nm shift while simultaneous QCM-D measurements permitted decoupling of lipid and solvent mass changes near the sensor surface. The vesicle rupture process to form the planar lipid bilayer on the sensor surface took ~7 min while the flow-mediated removal of excess lipid and coupled solvent took an additional 3 min.
Plasmonic nanostructures also offer potential for interfacing with virus particles and these possibilities led to exploring the use of metallic nanohole arrays to isolate individual virus-like particles for antiviral drug evaluation. This work was motivated by the demonstrated utility of AH peptide to rupture dengue virus particles and Jackman et al. developed a periodic gold nanohole array that was functionalized with a passivating polymer coating so that individual, virus-mimicking lipid vesicles could be selectively immobilized in each nanohole without requiring a capture probe50. The platform exhibited EOT sensing properties and there were multiple peaks in the transmission spectrum, which were sensitive to different regions of the nanohole geometry. The peak exhibited different bulk refractive index sensitivities (61, 221, 307 nm/RIU) and it was observed that the peaks with the highest and lowest bulk sensitivities had similarly weak surface sensitivities to detect vesicle capture in polymer-coated nanoholes (~0.3 nm wavelength shifts). By contrast, the peak with intermediate bulk sensitivity had the largest response to vesicle capture, with a three-fold larger signal (~0.9 nm shift). This finding was pivotal in establishing that spectral features can have distinct bulk and surface sensitivities, and the latter peak was thus most suitable for biosensing experiments.
According to that high-performing peak, subsequent addition of 10 µM AH peptide caused rupture of isolated lipid vesicles within the nanoholes and yielded a negative wavelength shift of –0.5 nm, which was attributed to lipid loss from within the sensor surface and confirmed by electron microscopy. Control experiments without polymer modification further verified that the sensing performance was related to the selective immobilization of vesicles within the nanoholes, as indicated by a less discernible shift (+0.2 nm) due to planar lipid bilayer formation on the nanohole sidewalls rather than vesicle rupture itself. A hallmark of this sensing platform was the ability to sequester individual viruses and virus-like particles in plasmonic nanoholes, leading to more accurate profiling of virus-drug interactions with biologically relevant readouts.
Other types of plasmonic nanostructures like silica-coated gold nanowells have proven useful for studying curvature-selective viral membrane disruption, as reported by Ferhan et al.51. The plasmonic nanowells were designed to have regions of negative and positive curvature and the entire sensor surface was conformally coated with a lipid bilayer. Thus, there were positively and negatively curved membrane regions within one sensing platform, and peptide interactions with each region could be detected separately based on distinct plasmonic signatures. Spectral peak and dip features corresponded to the positive and negative membrane curvature regions, respectively. Notably, 16 µM AH peptide only demonstrated binding to negatively curved membrane regions based on a +0.1 nm shift whereas the peptide caused extensive disruption of positively curved membrane regions, as indicated by a -0.4 nm shift. The latter shift indicates loss of lipid mass from the positively curved membrane region due to the peptide interaction. Since enveloped viruses possess positive membrane curvature, these findings provide a mechanistic foundation to rationalize the membrane-disruptive antiviral activity of the AH peptide.
In addition to the AH peptide, antimicrobial lipids such as glycerol monolaurate (GML) and lauric acid (LA) are promising antiviral molecules that disrupt microbial membranes. By utilizing LSPR measurements, Yoon et al. reported tracking the real-time membrane interactions of GML and LA addition to lipid bilayer coatings on silica-coated gold nanodisk arrays52. The measurement responses and corresponding analytical modeling revealed that GML caused membrane budding whereas LA caused tubule formation, with both interactions eventually resulting in membrane disruption. These activity profiles were consistent with their known antiviral properties while shedding light on how GML and LA distinctly disrupt membranes. Interestingly, while both compounds caused three-dimensional membrane remodeling, 2 mM LA addition caused a negative -0.4 nm maximum shift in the plasmonic resonance peak while 500 µM GML addition caused a maximum positive shift of +1.2 nm. These results demonstrate the utility of applying physics-based analytical models to interpret how label-free measurement responses are related to membrane morphological changes, and were supported by past findings obtained using different experimental methods such as fluorescence microscopy.
Additional antiviral surfactants such as cetyltrimethylammonium bromide (CTAB) and Triton X-100 (TX-100) that disrupt microbial membranes have also been explored using similar LSPR-based nanoplasmonic biosensing approaches, revealing distinct membrane-interaction profiles and varying dependencies on the critical micelle concentration53,54. For example, it was observed that cationic CTAB micelles caused extensive disruption of supported lipid bilayers and intact vesicle adlayers, with peak shifts exceeding -1 nm that indicate loss of lipid mass from the sensor surface. Conversely, CTAB monomers demonstrated binding to planar lipid membranes (+1.7 nm shift) likely through electrostatic interactions but caused extensive disruption of curved lipid vesicles that led to a -3.0 nm shift. Other compounds like TX-100 were only membrane-disruptive in the micellar state whereas TX-100 monomers demonstrated weak binding.
Another promising direction lies in testing the efficacy of small-molecule antiviral drug candidates, which has been limited to solution-phase assays so far but has excellent potential for future nanoplasmonic biosensor research. Ouyang et al. developed an assay based on functionalizing large gold nanoparticles with the SARS-CoV-2 S1 protein subunit and small gold nanoparticles with the ACE2 protein55. When the two nanoparticle types were mixed, they aggregated together due to the S1-ACE2 interaction. However, small-molecule drug candidates could disrupt this interaction, preventing nanoparticle aggregation. It was also possible to determine how clinically relevant S1 mutations affect drug efficacy. Chen et al. described a similar approach to study binding interactions between SARS-CoV-2 and SARS-CoV nucleocapsid (N) proteins and RNA strands, which are important for virus particle assembly, and to inhibit these interactions using antiviral drug candidates56.
These examples demonstrate the competitive merits of nanoplasmonic biosensors for antiviral drug development. Compared to traditional antiviral drug testing options, a unifying theme of these approaches is the emphasis on monitoring physical interactions between virus particles and drug candidates. Looking forward, future possibilities include utilizing nanoplasmonic biosensors to characterize the efficacy of antiviral drugs targeting viral enzymes such as polymerases that are involved in genome replication57.
Immune profiling
Another key opportunity relates to measuring virus-specific antibody levels, which is important for determining the protective immunity of individuals, the effectiveness of vaccination efforts, and the potential utility of therapeutic antibodies. These efforts involve functionalizing sensing surfaces with viral antigens as the capture probe to detect circulating antibody levels, highlighting the versatility of nanoplasmonic biosensing platforms to reverse the detection target from antigen (as in classical detection) to antibody. Interestingly, this topic was unexplored until the COVID-19 pandemic prompted researchers to search for new tools to measure viral immunity, as illustrated in Table 3.
In an early motivating study, Lew et al. developed antigen-functionalized gold nanoparticles for sensitive detection of SARS-CoV-2 immunoglobulin G (IgG) antibodies in patient plasma58. Several biotinylated antigens were initially tested and a short antigen from the SARS-CoV-2 S1 protein subunit yielded the largest antibody binding responses. In addition, Masterson et al. developed a high-throughput nanoplasmonic biosensing assay that was composed of gold triangular nanoprisms on a glass surface, which were functionalized with short antigens (epitopes) derived from the SARS-CoV-2 S protein sequence59. Antibody samples were incubated with the sensing platform overnight, prior to aqueous rinsing and spectroscopic measurement. The responses showed a wide dynamic range for purified SARS-CoV-2 IgG antibodies spanning ~11 orders of magnitude (10-9 to 102 nM) and reaching net shifts of up to 12 nm in the plasmonic resonance peak. An optimal ratio of two distinct epitope probes improved the limit of detection by around 10-fold and yielded an ~30 aM value. The platform showed high selectivity for SARS-CoV-2 IgG antibodies compared to other IgG antibodies and was used to screen over 100 plasma samples from COVID-19 patients and healthy controls with high specificity (90%) and sensitivity (100%). Typically, COVID-19 patients had net shifts of >8 nm whereas healthy controls were usually <4 nm according to the readout.
Various types of nanoplasmonic biosensing platforms with different geometries have also been explored following similar methodologies such as gold nanospike layers that were coated with streptavidin molecules and could bind to biotinylated antigen probes as reported by Funari et al.60. The study demonstrated the broad versatility of utilizing biotin-streptavidin chemistry because different antigens could be attached to specific sensor regions to create a multiplexed format. In one platform, it was possible to attach two types of influenza antigens and one type of SARS-CoV-2 antigen in order to simultaneously detect three monoclonal antibodies. After incubating antibody samples with the sensor chip for 2 h followed by aqueous rinsing and air drying, the SARS-CoV-2 antigen-functionalized surface detected 10 ng/mL monoclonal SARS-CoV-2 IgG with an ~4 nm shift in the plasmonic resonance peak and was insensitive to influenza A virus IgG (~0.3 nm shift). Additionally, the platform enabled antibody detection in mouse serum after immunization with a SARS-CoV-2 antigen (vaccine model). Specifically, it was possible to detect polyclonal SARS-CoV-2 IgG in mouse sera after immunization with full-length SARS-CoV-2 spike protein, as indicated by a negligible ~0.1 nm shift before immunization and an ~2.7 nm shift after immunization. Sensor chips functionalized with influenza A virus antigens demonstrated similar selectivity to influenza-specific IgG and had negligible responses to SARS-CoV-2 IgG.
Huang et al. also developed a gold nanohole array to quantify SARS-CoV-2 antibody levels based on a distinct capture strategy61. Anti-IgG secondary antibody was coated on the sensor chip walls and was used to capture SARS-CoV-2 IgG neutralizing antibodies from serum samples. Afterwards, SARS-CoV-2 RBD antigen-coated small gold nanoparticles were added to the platform and could bind to the captured antibodies, which amplified the nanoplasmonic signal based on OD measurements. The protocol was adapted to a one-step format (~15 min) and enabled rapid quantification of SARS-CoV-2 neutralizing antibody levels in clinical serum samples, achieving 96% specificity and 89% sensitivity. It was possible to improve the limit of detection to an estimated 0.2 pM antibody concentration by employing nanoporous hollow gold nanoparticles and the measured detection range was 2 to 400 pM, in which range the response showed sigmoidal behavior.
Utilizing EOT measurements, Adi et al. have demonstrated that periodic gold nanohole arrays are useful for antibody binding detection in clinical serum samples based on SARS-CoV-2 N and S protein antigens62. Antibody samples were incubated with antigen-functionalized surfaces for 1 h, followed by buffer rinsing and then endpoint imaging was performed based on measuring transmitted light intensity at two fixed wavelengths. Using clinical serum samples from patients with active (acute) or recovering (convalescent) COVID-19 infections as well as uninfected, serum control samples, a strong correlation was observed between the biosensing readouts and electrochemiluminescence (ECL) assay results. It was further noted that the protocol is quicker and simpler for the nanoplasmonic biosensing experiments compared to the ECL assay. In the nanoplasmonic biosensing experiments, it was possible to detect higher anti-S antibody signals (88% sensitivity, 100% specificity) and anti-N antibody signals (95% sensitivity, 60% specificity) in convalescent/acute sera compared to control sera.
In a follow-up study, Beisenova et al. integrated that platform with a machine learning model to estimate the SARS-CoV-2 infection and vaccination rates in a local community based on measuring antigen-specific antibody levels63. A single, fixed wavelength was used for optimal imaging after 30-min incubation and antibody levels in clinical samples were measured against a panel of six antigens. The labeled samples comprised four groups that were (1) naïve (no infection or vaccination), (2) vaccinated, (3) convalescent, and (4) convalescent and vaccinated. Clear distinctions in the antibody responses were observed in each group and a random forest-based machine learning model identified S and M antigens as highly predictive markers. The model was used to classify COVID-19 immunity status in blinded samples, and the predicted infection and vaccination rates (55% and 87%) showed excellent agreement with community-level epidemiological data (65% and 84%, respectively).
There has also been interest in naked-eye antibody detection, which has been achieved by nanoparticle-enhanced plasmonic coupling effects and is compatible with smartphone camera quantification64. In that study by Nan et al., antibody samples were incubated with an antigen-functionalized gold film for 30 min prior to transferring gold nanoparticles on top to create a sandwiched plasmonic structure as described above. Significant differences in the reflectance signal were observed between SARS-CoV-2 infected and healthy serum samples. The sensing platform was compatible with a portable, optical fiber reader (100% sensitivity, 99% specificity) and smartphone imaging (93% sensitivity, 97% specificity). Importantly, the sensing platform also demonstrated a ≥8-fold improved limit of detection compared to immunofluorescence assay (IFA) formats.
Nan et al. further extended the platform concept to support variant-specific immunity profiling65. It was identified that the SARS-CoV-2 S1 antigen and 5× diluted serum provided the best detection performance, resulting in 100% sensitivity and 100% specificity. Using the platform, it was determined that ~39% of blinded serum samples collected after the COVID-19 pandemic had high neutralizing antibody levels whereas all samples collected from before the pandemic tested negative. To improve translational prospects, the measurement readout was also adapted to a high-throughput barcode format. Technologically, these sensing capabilities demonstrate strong merits in terms of assay detection limits (compared to other assay options) and portability while streamlining of the nanoparticle transfer process would simplify the protocol for wider accessibility.
A recent study by Li et al. has also demonstrated that utilizing disorder-enhanced gold nanograin metasurfaces functionalized with SARS-CoV-2 spike protein antigen can boost nanoplasmonic sensing signals in order to monitor post-vaccination antibody levels66. It was observed that total antibody levels in convalescent serum samples produced reflectance shifts of 4–6 nm and the magnitude correlated with ELISA results. Furthermore, total antibody levels in vaccinated serum samples generated 2–6 nm shifts and showed temporal fluctuations in line with the vaccination schedule. The sensors could be regenerated with an acidic pH washing step in order to be used repeatedly, with consistent reflectance shifts.
In addition to profiling antibody levels, there is also interest in applying nanoplasmonic sensors to characterize the therapeutic activity of neutralizing antibodies that block critical virus-cell interactions. For example, Li et al. reported a nanomaterial-coupled metasurface plasmon resonance sensor to measure antibody blocking of the interaction between functionalized SARS-CoV-2 RBD antigen probes on the sensor surface and ACE2 receptors67. First, antibodies were incubated with the sensing platform, and then ACE2-functionalized gold nanoparticles were added. Successful antibody neutralization was judged by preventing nanoparticle binding and 50% inhibitory concentration (IC50) values were determined for the ACE2 binding interaction with different RBD variants. In general, neutralizing antibodies in serum were more effective against wild-type RBD while higher antibody concentrations (i.e., lower dilutions) were required to inhibit circulating mutants of global health importance. Importantly, these results were consistent with plaque reduction neutralization test results, which is a gold-standard assay for antibody evaluation. The nanoplasmonic biosensing assay also had high sensitivity (94%) and specificity (100%) and did not demonstrate cross-reactivity with IgG antibodies against other respiratory viruses.
In another antibody-related example that inspires future nanoplasmonic biosensor possibilities, Batool et al. developed a lipid membrane-coated nanoplasmonic sensing platform to detect SARS-CoV-2 virus particle binding and antibody blocking of the virus-membrane interaction68. The membrane was functionalized with ACE2 protein receptors and could detect viral antigens and whole virus binding. Using this platform, the efficacy levels of two neutralizing antibodies to block these interactions were measured for different viral strains, providing insight into the potential clinical utility of these antibodies. Similarly, Shang et al. have described utilizing SARS-CoV-2 S protein-functionalized gold nanorods and ACE2-functionalized gold nanoparticles to measure antibody neutralization69. Furthermore, SPR-type nanoplasmonic sensors have been recently used to measure neutralizing antibody levels in clinical samples from food and retail workers, providing insight into population immunity70. The diagnostic capabilities of measuring virus-specific antibody levels have also been demonstrated for veterinary applications, yielding high sensitivity and specificity for detecting SARS-CoV-2 infections in hamsters, dogs, and cats from serum samples71.
Within the nanoplasmonic biosensing context, such approaches have also been extended to nanohole-based EOT measurements for detecting antibodies that are indicative of African swine fever virus (ASFV) infection, which can be useful for agricultural biosecurity on pig farms72. Antibodies in pig serum were mixed with p30 antigen-coated gold nanoparticles, which were then added to a secondary antibody-functionalized gold nanocup surface for OD measurements. 246 serum samples, which had been validated as ASFV-positive or negative by ELISA measurements, were tested and demonstrated that the assay had an estimated high sensitivity (96%) and specificity (97%). There was a wide dynamic range spanning 1:100 to 1:16000 serum dilutions and the detection limit was comparable to that of indirect immunofluorescence while taking only 20 min in total.
Collectively, these examples demonstrate the rapid clinical exploration of nanoplasmonic biosensors for quantifying antibody levels in biological fluids. This progress highlights the accelerating pace of research in the field and emphasizes the need to strategically focus these efforts on achieving the most impactful outcomes.
Assessment of current research and future possibilities
The presented evidence from scientific reports supports the merits of utilizing nanoplasmonic biosensors in virus research. Nevertheless, like other classes of label-free biosensors such as electrochemical-based ones, existing nanoplasmonic biosensors remain largely at the research-use or precommercial stage and have not yet achieved FDA/CE approval to our knowledge. As such, there is still a gap between the current research landscape and envisioned practical applications that should be addressed by considering several issues as follows:
Challenge 1: practical utility of biosensors
While nanoplasmonic biosensing advances often target enhanced detection performance, there remains an outstanding need to focus on robustness and clinical utility. Progress in creating portable and affordable instruments is encouraging, but a critical challenge remains: the development of cost-efficient and scalable sensing platforms compatible with portable systems. Although some low-cost sensor chips are being explored, a more integrated approach combining sensor fabrication, probe functionalization, and protocol optimization is essential. Future efforts in sensor chip development should concentrate on two key directions: affordable chips that strike a balance between sensitivity and practical considerations to facilitate clinical translation, and advanced chips that leverage cutting-edge fabrication techniques to expand the scientific frontier of nanoplasmonic biosensors.
Challenge 2: more emphasis on clinical translation
It is widely accepted that nanoplasmonic biosensors possess remarkably high detection sensitivity. However, their clinical utility remains to be established and there are two strategies to address this need. First, rigorous study designs aligned with clinical regulatory guidelines should be pursued to evaluate biosensing performance in specific applications such as virus diagnostics and immune profiling. Ideally, these studies should be conducted in collaboration with clinical scientists to build credibility within the medical community and should involve large test samples. Second, the quantification merits of nanoplasmonic biosensors should be compared to gold-standard techniques. For example, while it is recognized that nanoplasmonic biosensors can detect antigens at lower concentrations than conventional diagnostics such as clinically used LFA platforms in some examples, direct comparisons are needed to prove their advantages and build a compelling business case. Such efforts can help to build motivation for seeking FDA/CE approval to use nanoplasmonic biosensors in specific diagnostic applications. As discussed above, some nanoplasmonic biosensors can demonstrate detection limits that surpass LFA sensitivity thresholds so there is strong promise in this direction.
Challenge 3: expanded focus on more virus types
The COVID-19 pandemic accelerated research on nanoplasmonic biosensors for virus applications, especially directed at SARS-CoV-2. This focus has resulted in significant benefits, including increased research activity, rapid advances to address immediate virus detection needs, and the development of new sensing platforms for immune profiling. However, it is critical to balance these positive outcomes with a broader, forward-looking approach. Expanding the scope of research to include a diverse set of viruses will provide valuable insights into optimizing capture probe designs as well as understanding antigen and antibody features of different viruses. This strategic diversification will not only deepen the understanding of biosensor performance across various viruses but also position nanoplasmonic biosensors for sustained, long-term contributions to virus research. It will also better prepare researchers to address future virus outbreaks.
Challenge 4: broaden antiviral drug testing
The development of new antiviral drug classes requires detailed mechanistic understanding to ensure clinical safety and address regulatory needs. Nanoplasmonic biosensors are uniquely suited to characterize the mechanism of antiviral drugs that directly interact with virus particles, as exemplified by the peptide and detergent cases discussed above. Various nanoplasmonic sensing platforms have shown particular strengths in characterizing drug mechanisms, especially selectivity features. These types of research should be expanded to develop more virus-specific testing platforms and to test a wider range of antiviral drug candidates. Among drug candidates tested so far, the AH peptide is the most advanced one but a more comprehensive framework should be established to investigate other antiviral peptides and therapeutic antibodies within well-defined application contexts. These testing platforms can also provide guidance to engineer more potent and selective drug candidates.
By addressing these points, we imagine the following hypothetical scenario that illustrates how suitably capable nanoplasmonic biosensors might advance healthcare responses to future viral threats:
A new virus has emerged, and sequencing analysis and protein expression are underway to generate virus-specific antibodies and viral antigens for point-of-care diagnostic development and therapeutic evaluation. Previously, the produced antibodies would be included in LFAs for virus detection using rapid test kits. Significant antibody quantities would have been required, limiting initial kit production scalability. But now with more advanced biosensors, much lower antibody quantities are needed due to microprinting and the antibodies can be printed on low-cost nanoplasmonic chips that support real-time assessment of viral presence and quantity without sample preprocessing. The chip signal is directly readable by smartphones and associated software applications can guide patients on the next steps such as isolating and seeking medical help.
Before viral antigens are produced recombinantly, other nanoplasmonic biosensor chips are already being deployed to capture virus particles from the biological fluids of infected patients. These virus platforms can be used to test the sera of recovered patients in order to identify highly effective neutralizing antibodies for therapeutic development based on binding kinetic analysis. The platforms can also be used for immune profiling, assessing pre-existing immunity, and later vaccination responses in different community populations. Rather than using recombinantly produced viral antigens in these assays, a powerful capability afforded by the sensor chips is the ability to directly use captured virus particles as the antigen. Measuring the binding kinetics of antibodies and other antiviral drug candidates against virus particles as opposed to viral protein antigens could be advantageous because effective immune responses often depend on the multivalent arrangement and other structural features influencing antigen presentation on the virus particle surface. Practically, these assays can also be initiated with captured virus particles before recombinant viral antigens are widely available.
While this discussion is speculative, it highlights future possibilities that align with the platform design features and sensor capabilities being pursued in current research areas.
New research directions
In addition to strengthening capabilities in ongoing research areas and realizing possibilities like the discussed scenario, the following topics represent excellent opportunities to harness the most compelling features of nanoplasmonic biosensors. These features include the high surface sensitivity of the measurement response and the proven capability to analyze the conformational properties of biomacromolecules (e.g., proteins) and biological nanoparticles (e.g., vesicles, virus particles).
Opportunity 1: virus-cell binding interactions
Virus binding to cell membrane receptors is a critical step in viral infection pathways, often involving shape changes in the virus particle and/or cell membrane73. While other surface-sensitive measurement techniques have been used to study these binding processes74, there is an outstanding need to explore the multivalent ligand-receptor interactions in greater detail. Nanoplasmonic biosensors have already demonstrated exceptional capabilities for studying multivalent binding of lipid vesicles and extending these approaches to study viruses represents a promising frontier. Potential applications include measuring strain-specific virus binding affinities to membrane-associated receptors, detecting viral fusion events involving membrane-protein interactions, and evaluating therapeutic performance of competitive binders and neutralizing antibodies.
Opportunity 2: antigen presentation
Nanoplasmonic biosensors have proven useful for studying the conformational properties of adsorbed proteins on alumina surfaces, which are related to widely used adjuvants in vaccine applications75. Greater effort is needed to bridge these measurement capabilities with vaccine-related protein engineering. For example, recent protein engineering efforts have focused on controlling the orientation of bound antigens on aluminum-based adjuvant materials to modulate humoral immune responses76. Developing nanoplasmonic sensing surfaces that mimic the composition of adjuvant materials could provide an innovative platform for studying antigen binding and adsorption-related orientation. The high surface sensitivity of nanoplasmonic biosensors is ideally suited for this application and could enable correlations between antigen engineering and conformational properties.
Exosome possibilities
While our focus is to discuss emerging opportunities to use nanoplasmonic biosensors for virus applications, we also want to briefly comment on sensor utility for many different types of exosomes. For example, cancer exosomes are widely regarded as important modulators of the immune system and impact the effectiveness of cancer immunotherapies. They also share some physicochemical similarities with membrane-enveloped viruses in terms of physical shape, lipid bilayer coating, and size range (~50-300 nm diameter). As such, there have been extensive efforts to develop nanoplasmonic biosensors for detecting cancer exosomes and exosomal proteins in clinical samples77,78,79,80,81,82, largely mirroring the sensor geometries and antibody-based capture schemes discussed here. These nanoplasmonic biosensing capabilities have also been applied to evaluate the efficacy of exosome-related anticancer therapies83 and therapeutic antibodies84. In the latter respect, there has also been exploration of antibody-functionalized supported lipid bilayer coatings to capture cancer exosomes in other contexts85, which could be useful for future nanoplasmonic biosensing applications like isolating exosomes from biological fluids as well as testing the efficacy of anti-exosome-therapeutics. We foresee that areas like antiviral drug evaluation and immune profiling that are discussed in the virus context can also be applied more actively to the cancer exosome field through concepts such as anti-exosome drug evaluation (e.g., peptides and antibodies that target exosomes) and immune profiling (e.g., antibody responses to cancer exosome-based vaccines).
In addition to cancer exosomes, other types of exosomes are also promising detection targets and are related to various translational medicine applications such as diagnosing cardiovascular, neurodegenerative, and infectious diseases as well as particular conditions like diabetes and organ injuries81,86,87. Exosomes exist in a wide range of biological fluids such as plasma, urine, and cerebrospinal fluid, and careful consideration of isolation strategies and possible matrix effects on nanoplasmonic biosensing signals is warranted. In addition to exosomes, related biological nanoparticles such as bacterial membranes can also be studied, providing insights into human gut health and environmental analysis88.
Conclusion and outlook
In closing, nanoplasmonic biosensors have demonstrated excellent potential for virus research and growing measurement capabilities have broad implications for translational medicine applications. We foresee tremendous growth opportunities that will come from developing clinically useful instruments, validating diagnostic and immune profiling capabilities against established technologies, and strategically targeting high-priority viruses where nanoplasmonic biosensors can provide a competitive edge while maintaining scientific breadth. These biosensing tools will also be useful for diagnostic testing and immune profiling in resource-limited settings as well as for pandemic preparedness. Furthermore, expanding fundamental research on virus-related interactions such as virus-cell binding dynamics and controlling antigen presentation will be critical to leverage the most advanced measurement capabilities of these biosensors to help realize next-generation diagnostic, therapeutic, and vaccine technologies.
Regarding future possibilities, we have discussed many experimental opportunities above but wish to briefly comment more on data analysis opportunities, especially given recent advances on the machine learning side. Nanoplasmonic biosensors generate information-rich optical datasets and there are various areas where data interpretation can be improved by machine learning techniques. Several examples covered here included random forest classifiers to rank the relative importance of different antigen probes and neural networks to improve reflectance-based detection limits. Looking forward, it may also be possible to recognize patterns in how spectral signatures change over time in order to identify specific viral targets or quantify their levels, which can be especially important in multiplexed systems. These techniques could also support automatic correction of background noise or signal drift to improve detection limits and help convert raw sensor data into clinically meaningful results in real-time. We foresee that next-generation nanoplasmonic biosensor systems will combine the latest advances in nanofabrication, biointerfacial science, and optimal imaging with these emerging capabilities to create instruments with highly integrated hardware and software packages for not only data acquisition but also for clinically relevant data interpretation.
Data availability
No datasets were generated or analysed during the current study.
References
Li, R. et al. Application of nanoplasmonic biosensors based on nanoarrays in biological and chemical detection. Opt. Expr. 31, 21586–21613 (2023).
Willets, K. A. & Van Duyne, R. P. Localized surface plasmon resonance spectroscopy and sensing. Annu. Rev. Phys. Chem. 58, 267–297 (2007).
Špačková, B., Wrobel, P., Bocková, M. & Homola, J. Optical biosensors based on plasmonic nanostructures: A review. Proc. IEEE 104, 2380–2408 (2016).
Heskin, J. et al. Evaluating the performance characteristics of five lateral flow assays for the detection of the SARS-CoV-2 nucleocapsid antigen. Sci. Rep. 12, 8811 (2022).
Kakkar, S. et al. Lateral flow assays: Progress and evolution of recent trends in point-of-care applications. Mater. Today Bio 28, 101188 (2024).
Brazaca, L. C. et al. Biosensing strategies for the electrochemical detection of viruses and viral diseases – A review. Anal. Chim. Acta 1159, 338384 (2021).
Mauriz, E., Dey, P. & Lechuga, L. M. Advances in nanoplasmonic biosensors for clinical applications. Analyst 144, 7105–7129 (2019).
Masson, J.-F., Biggins, J. S. & Ringe, E. Machine learning for nanoplasmonics. Nat. Nanotechnol. 18, 111–123 (2023).
Li, Z. et al. Plasmonic-based platforms for diagnosis of infectious diseases at the point-of-care. Biotechnol. Adv. 37, 107440 (2019).
Mauriz, E. Recent progress in plasmonic biosensing schemes for virus detection. Sensors 20, 4745 (2020).
Soler, M. et al. How nanophotonic label-free biosensors can contribute to rapid and massive diagnostics of respiratory virus infections: COVID-19 case. ACS Sens. 5, 2663–2678 (2020).
Hassan, M. M., Sium, F. S., Islam, F. & Choudhury, S. M. A review on plasmonic and metamaterial based biosensing platforms for virus detection. Sens. Bio-Sens. Res. 33, 100429 (2021).
Wang, J., Xu, Z. & Kotsifaki, D. G. Plasmonic and metamaterial biosensors: A game-changer for virus detection. Sens. Diagn. 2, 600–619 (2023).
Tsalsabila, A. et al. Progress and outlooks in designing photonic biosensor for virus detection. Adv. Opt. Mater. 12, 2400849 (2024).
Masson, J.-F., Murray-Méthot, M.-P. & Live, L. S. Nanohole arrays in chemical analysis: manufacturing methods and applications. Analyst 135, 1483–1489 (2010).
Kurt, H. et al. Nanoplasmonic biosensors: Theory, structure, design, and review of recent applications. Anal. Chim. Acta 1185, 338842 (2021).
Larsson, E. M., Syrenova, S. & Langhammer, C. Nanoplasmonic sensing for nanomaterials science. Nanophotonics 1, 249–266 (2012).
Jackman, J. A., Rahim Ferhan, A. & Cho, N.-J. Nanoplasmonic sensors for biointerfacial science. Chem. Soc. Rev. 46, 3615–3660 (2017).
Jackman, J. A., Ferhan, A. R. & Cho, N.-J. Surface-based nanoplasmonic sensors for biointerfacial science applications. Bull. Chem. Soc. Jpn. 92, 1404–1412 (2019).
Jaikishan, S., Lavainne, M. & Wiedmer, S. K. Nanoplasmonic sensing for studies on liposomes and extracellular vesicles. Sens. Actuators Rep. 7, 100192 (2024).
Zhdanov, V. P. Basics of the LSPR sensors for soft matter at interfaces. Plasmonics 18, 971–982 (2023).
Jackman, J. A. et al. Nanoplasmonic ruler to measure lipid vesicle deformation. Chem. Commun. 52, 76–79 (2016).
Jackman, J. A. et al. Quantitative profiling of nanoscale liposome deformation by a localized surface plasmon resonance sensor. Anal. Chem. 89, 1102–1109 (2017).
Park, H. et al. Unraveling how multivalency triggers shape deformation of sub-100 nm lipid vesicles. J. Phys. Chem. Lett. 12, 6722–6729 (2021).
Park, H. et al. Multivalency-induced shape deformation of nanoscale lipid vesicles: Size-dependent membrane bending effects. J. Phys. Chem. Lett. 13, 1480–1488 (2022).
Yanik, A. A. et al. An optofluidic nanoplasmonic biosensor for direct detection of live viruses from biological media. Nano Lett. 10, 4962–4969 (2010).
Inci, F. et al. Nanoplasmonic quantitative detection of intact viruses from unprocessed whole blood. ACS Nano 7, 4733–4745 (2013).
Inci, F. et al. Enhancing the nanoplasmonic signal by a nanoparticle sandwiching strategy to detect viruses. Appl. Mater. Today 20, 100709 (2020).
Hao, X., St-Pierre, J.-P., Zou, S. & Cao, X. Localized surface plasmon resonance biosensor chip surface modification and signal amplifications toward rapid and sensitive detection of COVID-19 infections. Biosens. Bioelectron. 236, 115421 (2023).
Masterson, A. N. et al. Multiplexed and high-throughput label-free detection of RNA/spike protein/IgG/IgM biomarkers of SARS-CoV-2 infection utilizing nanoplasmonic biosensors. Anal. Chem. 93, 8754–8763 (2021).
Li, F. et al. Affinity exploration of SARS-CoV-2 RBD variants to mAb-functionalized plasmonic metasurfaces for label-free immunoassay boosting. ACS Nano 17, 3383–3393 (2023).
Liang, J. et al. Nanoplasmonic sensor optimization via digital imaging analysis and antibody evolution. Sens. Actuators, B 418, 136287 (2024).
Ma, Z. et al. Sensitivity-enhanced nanoplasmonic biosensor using direct immobilization of two engineered nanobodies for SARS-CoV-2 spike receptor-binding domain detection. Sens. Actuators, B 383, 133575 (2023).
Lewis, T., Giroux, E., Jovic, M. & Martic-Milne, S. Localized surface plasmon resonance aptasensor for selective detection of SARS-CoV-2 S1 protein. Analyst 146, 7207–7217 (2021).
Yang, Y. et al. Silver nanotriangle array based LSPR sensor for rapid coronavirus detection. Sens. Actuators, B 359, 131604 (2022).
Ling, C. et al. Valence-driven colorimetric detection of norovirus protease via peptide-AuNP interactions. Chem. Commun. 59, 12459–12462 (2023).
Jin, Z. et al. Peptide amphiphile mediated co-assembly for nanoplasmonic sensing. Angew. Chem., Int. Ed. 62, e202214394 (2023).
Sun, W. et al. Neural network enables high accuracy for hepatitis B surface antigen detection with a plasmonic platform. Nano Lett. 24, 8784–8792 (2024).
Huang, L. et al. One-step rapid quantification of SARS-CoV-2 virus particles via low-cost nanoplasmonic sensors in generic microplate reader and point-of-care device. Biosens. Bioelectron. 171, 112685 (2021).
Wallace, S. et al. Multiplexed biosensing of proteins and virions with disposable plasmonic assays. ACS Sens. 8, 3338–3348 (2023).
Bido, A. T. et al. Detection of SARS-CoV-2 in saliva by a low-cost LSPR-based sensor. Anal. Methods 15, 3955–3966 (2023).
Li, R. et al. Versatile nanorobot hand biosensor for specific capture and ultrasensitive quantification of viral nanoparticles. Mater. Today Bio 16, 100444 (2022).
Zaccagnini, F. et al. Multifunctional FFP2 face mask for white light disinfection and pathogens detection using hybrid nanostructures and optical metasurfaces. Small 20, 2400531 (2024).
Ryu, J. S. et al. Advancing SARS-CoV-2 variant detection with high affinity monoclonal antibodies and plasmonic nanostructure. Adv. Funct. Mater. 34, 2405340 (2024).
Hwang, Y. et al. Nanopot plasmonic sensor platform for broad spectrum virus detection. Chem. Eng. J. 505, 159484 (2025).
Jackman, J. A. et al. Deciphering how pore formation causes strain-induced membrane lysis of lipid vesicles. J. Am. Chem. Soc. 138, 1406–1413 (2016).
Jackman, J. A. et al. Therapeutic treatment of Zika virus infection using a brain-penetrating antiviral peptide. Nat. Mater. 17, 971–977 (2018).
Zan, G. H., Jackman, J. A., Kim, S.-O. & Cho, N.-J. Controlling lipid membrane architecture for tunable nanoplasmonic biosensing. Small 10, 4828–4832 (2014).
Ferhan, A. R., Jackman, J. A. & Cho, N.-J. Integration of quartz crystal microbalance-dissipation and reflection-mode localized surface plasmon resonance sensors for biomacromolecular interaction analysis. Anal. Chem. 88, 12524–12531 (2016).
Jackman, J. A. et al. Plasmonic nanohole sensor for capturing single virus-like particles toward virucidal drug evaluation. Small 12, 1159–1166 (2015).
Ferhan, A. R. et al. Nanoplasmonic sensing architectures for decoding membrane curvature-dependent biomacromolecular interactions. Anal. Chem. 90, 7458–7466 (2018).
Yoon, B. K. et al. Real-time nanoplasmonic sensing of three-dimensional morphological changes in a supported lipid bilayer and antimicrobial testing applications. Biosens. Bioelectron. 174, 112768 (2021).
Duša, F., Chen, W., Witos, J. & Wiedmer, S. K. Nanoplasmonic sensing and capillary electrophoresis for fast screening of interactions between phosphatidylcholine biomembranes and surfactants. Langmuir 34, 5889–5900 (2018).
Jaikishan, S. et al. Fragment-based approach to study fungicide-biomimetic membrane interactions. Soft Matter 20, 5954–5968 (2024).
Ouyang, Y. et al. Virus-like plasmonic nanoprobes for quick analysis of antiviral efficacy and mutation-induced drug resistance. Anal. Chem. 95, 5009–5017 (2023).
Chen, Y. et al. Plasmonic imaging of multivalent NTD–nucleic acid interactions for broad-spectrum antiviral drug analysis. Anal. Chem. 96, 9551–9560 (2024).
Cho, N.-J. et al. Reconstitution and functional analysis of a full-length hepatitis C virus NS5B polymerase on a supported lipid bilayer. ACS Cent. Sci. 2, 456–466 (2016).
Lew, T. T. S. et al. Epitope-functionalized gold nanoparticles for rapid and selective detection of SARS-CoV-2 IgG antibodies. ACS Nano 15, 12286–12297 (2021).
Masterson, A. N. & Sardar, R. Selective detection and ultrasensitive quantification of SARS-CoV-2 IgG antibodies in clinical plasma samples using epitope-modified nanoplasmonic biosensing platforms. ACS Appl. Mater. Interfaces 14, 26517–26527 (2022).
Funari, R., Fukuyama, H. & Shen, A. Q. Nanoplasmonic multiplex biosensing for COVID-19 vaccines. Biosens. Bioelectron. 208, 114193 (2022).
Huang, L. et al. Novel nanostructure-coupled biosensor platform for one-step high-throughput quantification of serum neutralizing antibody after COVID-19 vaccination. Biosens. Bioelectron. 199, 113868 (2022).
Adi, W., Biswas, D., Shelef, M. A. & Yesilkoy, F. Multiplexed COVID-19 antibody quantification from human sera using label-free nanoplasmonic biosensors. Biomed. Opt. Express 13, 2130–2143 (2022).
Beisenova, A. et al. Machine-learning-aided multiplexed nanoplasmonic biosensor for COVID-19 population immunity profiling. Sens. Diagn. 2, 1186–1198 (2023).
Nan, J. et al. Thickness-sensing sandwiched plasmonic biosensors enable label-free naked-eye antibody quantification. Nano Lett. 22, 9596–9605 (2022).
Nan, J. et al. Naked-eye readable microarray for rapid profiling of antibodies against multiple SARS-CoV-2 variants. Nano Lett. 23, 10892–10900 (2023).
Li, F. et al. Plasmonic nanograin metasurface with disorder-enhanced biosensing for SARS-CoV-2 variant and antibodies. Adv. Funct. Mater. 34, 2401983 (2024).
Li, R. et al. Metasurface-driven and nanomaterial-coupled plasmonic biosensor for the rapid and quantitative clinical identification of neutralizing antibodies against SARS-CoV-2 variants. Adv. Funct. Mater. 33, 2306145 (2023).
Batool, R. et al. Biomimetic nanoplasmonic sensor for rapid evaluation of neutralizing SARS-CoV-2 monoclonal antibodies as antiviral therapy. Biosens. Bioelectron. 226, 115137 (2023).
Shang, J. et al. Single-particle rotational sensing for analyzing the neutralization activity of antiviral antibodies. Talanta 279, 126606 (2024).
Coutu, J. et al. Large-scale validation of a plasmonic sensor for SARS-CoV-2 pseudo-neutralization with a cohort of food and retail workers. Sens. Diagn. 3, 850–862 (2024).
Giarola, J. F. et al. Validation of a plasmonic-based serology biosensor for veterinary diagnosis of COVID-19 in domestic animals. Talanta 271, 125685 (2024).
Zhao, Y. et al. One-step rapid and sensitive ASFV p30 antibody detection via nanoplasmonic biosensors. Microbiol. Spectr. 10, e02343–02322 (2022).
Overeem, N. J., van der Vries, E. & Huskens, J. A dynamic, supramolecular view on the multivalent interaction between influenza virus and host cell. Small 17, 2007214 (2021).
Rydell, G. E., Dahlin, A. B., Höök, F. & Larson, G. QCM-D studies of human norovirus vlps binding to glycosphingolipids in supported lipid bilayers reveal strain-specific characteristics. Glycobiology 19, 1176–1184 (2009).
Park, H. et al. Comparing protein adsorption onto alumina and silica nanomaterial surfaces: Clues for vaccine adjuvant development. Langmuir 37, 1306–1314 (2021).
Xu, D. et al. Vaccine design via antigen reorientation. Nat. Chem. Biol. 20, 1012–1021 (2024).
Im, H. et al. Label-free detection and molecular profiling of exosomes with a nano-plasmonic sensor. Nat. Biotechnol. 32, 490–495 (2014).
Park, J. et al. Analyses of intravesicular exosomal proteins using a nano-plasmonic system. ACS Photonics 5, 487–494 (2018).
Raghu, D. et al. Nanoplasmonic pillars engineered for single exosome detection. PLoS One 13, e0202773 (2018).
Liang, K. et al. Nanoplasmonic quantification of tumour-derived extracellular vesicles in plasma microsamples for diagnosis and treatment monitoring. Nat. Biomed. Eng. 1, 0021 (2017).
Chin, L. K. et al. Plasmonic sensors for extracellular vesicle analysis: From scientific development to translational research. ACS Nano 14, 14528–14548 (2020).
Hu, R. et al. Exploring aptamer-based metasurfaces for label-free plasmonic biosensing of breast tumor-derived exosomes. Adv. Opt. Mater. 12, 2401180 (2024).
Pan, S. et al. Extracellular vesicle drug occupancy enables real-time monitoring of targeted cancer therapy. Nat. Nanotechnol. 16, 734–742 (2021).
Batool, R., Soler, M., Singh, R. & Lechuga, L. M. A novel biomimetic nanoplasmonic sensor for rapid and accurate evaluation of checkpoint inhibitor immunotherapy. Anal. Bioanal. Chem. 416, 7295–7304 (2024).
Shin, S. et al. Curvature-sensing peptide inhibits tumour-derived exosomes for enhanced cancer immunotherapy. Nat. Mater. 22, 656–665 (2023).
Neettiyath, A., Chung, K., Liu, W. & Lee, L. P. Nanoplasmonic sensors for extracellular vesicles and bacterial membrane vesicles. Nano Converg. 11, 23 (2024).
Liu, W., Chung, K., Yu, S. & Lee, L. P. Nanoplasmonic biosensors for environmental sustainability and human health. Chem. Soc. Rev. 53, 10491–10522 (2024).
Amrollahi, P., Zheng, W., Monk, C., Li, C.-Z. & Hu, T. Y. Nanoplasmonic sensor approaches for sensitive detection of disease-associated exosomes. ACS Appl. Bio Mater. 4, 6589–6603 (2021).
Acknowledgements
We acknowledge support from the National Research Foundation of Korea (NRF) grants funded by the Korean government (MSIT) (RS-2024-00405818, RS-2024-00460012, RS-2022-NR066735). This work was also partially supported by the SKKU Global Research Platform Research Fund, Sungkyunkwan University, 2024, and by the National Research Foundation, Prime Minister’s Office, Singapore, under its Campus for Research Excellence and Technological Enterprise (CREATE) program. Schematic illustrations were created with BioRender.com under an academic lab subscription.
Author information
Authors and Affiliations
Contributions
H.P. and J.A.J. contributed to study conceptualization and literature review. H.P. was responsible for figure creation. J.A.J. contributed to table compilation. J.A.J. prepared the original draft. H.P. and J.A.J. critically reviewed and edited the manuscript. Both authors checked the final version of the manuscript and approved submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Park, H., Jackman, J.A. Nanoplasmonic biosensors for detecting viruses and combating viral infections. npj Biosensing 2, 22 (2025). https://doi.org/10.1038/s44328-025-00043-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44328-025-00043-0
This article is cited by
-
Disposable Point of Care multiplexed plasmonic biosensor for rapid and specific identification of respiratory viruses
npj Biosensing (2026)
-
Detection Strategies for Begomoviruses: A Focus Review of Methods
Molecular Biotechnology (2026)