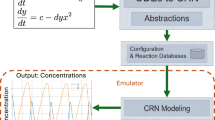
Abstract
The widespread adoption of energy-intensive computing applications has led to a growing need for energy-efficient computing approaches. Thermodynamic computing offers a promising approach for low-energy computation by leveraging the intrinsic computational capabilities of physical, chemical, or biological systems. However, the mathematical foundations of thermodynamic computing require further development to fully realize the potential energy efficiencies, as well as to assess factors like noise and operational speed. In this paper, we establish a mathematical framework for utilizing thermodynamic processes to perform fundamental operations, including addition, subtraction, multiplication, and division. We highlight the use of chemical reactions as potential computational units and explore synthetic chemical and biochemical systems as practical implementations. Additionally, we demonstrate how these principles can be applied to solving complex mathematical problems, such as ordinary differential equations (ODEs) and suggest the necessary components to implement the thermodynamic computing framework using chemical reactions based in a microfluidic device. This work enhances our understanding of thermodynamic processes for natural computing as a basis for scalable, energy-efficient computation in paradigm disruptive next-generation systems.
Similar content being viewed by others
Introduction
Understanding how thermodynamic processes can execute mathematical operations is necessary to lay the foundation for energy-efficient computing1. Toward this end, recent studies have demonstrated that energy-efficient, thermodynamic computers have the potential to achieve a linear speedup when performing fundamental linear algebra operations such as solving systems of linear equations, matrix inversion, and matrix exponentiation2,3. Natural thermodynamic systems, such as biological cells4, excel at solving optimization problems. Leveraging this ability is a grand challenge in thermodynamic computing, and indeed the development of energy-efficient gradient descent solutions based on thermodynamics is underway5.
Yet, on a fundamental level, the relationship between thermodynamics and computing has yet to be fully developed. In particular, understanding the relationship between computing and thermodynamics, and information theory and thermodynamics, is critical for developing energy-efficient and natural computing platforms.
Thermodynamic processes in chemical systems are perhaps the most studied and best understood thermodynamic systems. Previously, the time-dependent behaviors of chemical reaction networks (CRNs) have been proposed for carrying out computational operations like matrix manipulation, arithmetic functions, and solving ordinary differential equations (ODEs). Many of these proposed approaches have assumed either equilibrium, unrealistic one-way reactions, or reactions that do not conserve6. However, by leveraging thermodynamic principles, CRNs can fully execute both transient and steady-state thermodynamic computations. These capabilities offer a new avenue for scalable, energy-efficient analog computing.
Notably, biological cells serve as natural examples of unconventional computing devices, demonstrating scalability and efficiency through their inherent chemical processes. Metabolic networks perform complex mathematical functions in parallel, while subcellular compartments regulate processes like exponential decay, signal transduction, and nutrient transport. These systems link environmental perturbations to structured outputs, enabling dynamic adaptation through biochemical computation. Synthetic biochemical devices inspired by these biological mechanisms provide an opportunity to implement controlled chemical reactions for solving complex scientific problems.
However, what is lacking is a fundamental mathematical framework that relates chemical reactions to computing, which, when integrated with reservoir computing frameworks7,8,9, for example, could be used to explore the utility of CRNs in both biological and synthetic systems. Reservoir computing, based on high-dimensional dynamic systems, enables CRNs to process input signals and generate structured computational outputs efficiently10.
Our objective is to establish a framework where thermodynamic processes in general, and specifically chemical reactions, act as functional computational units, providing a foundation for composable analog computation. In the mechanical differential analyzers of V. Bush11 and others, gear ratios are used to carry out multiplications, and differential gears, similar to those found in the drive system of automobiles, are used to carry out addition. We describe the thermodynamic processes that likewise can be used to carry out multiplication, division, addition, and subtraction. We discuss both chemical and biological implementations, such as microfluidic devices, as platforms for biochemical computation, advancing scalable solutions for future scientific modeling and energy-efficient computing.
Results
Derivation of elementary operations using thermodynamics
Consider some set of n observables X = {X1, …Xn} that can take continuous values expressed in vector form x = [x1, …, xn] and can be described such that the probability density of observing the values x is given by,
Generally, in a system of n observables, we have unit vectors along a coordinate axis for each observable, \({\widehat{{\boldsymbol{X}}}}_{{\bf{1}}},\ldots ,{\widehat{{\boldsymbol{X}}}}_{{\boldsymbol{n}}}\). We define an arbitrary unit vector \({\boldsymbol{x}}=({x}_{1}{\widehat{{\boldsymbol{X}}}}_{{\bf{1}}}+\ldots +{x}_{n}{\widehat{{\boldsymbol{X}}}}_{{\boldsymbol{n}}})\), such that a distance ξ that measures the extent along x is,
Then f is such that,
and,
Defining \({\gamma }_{i}=\frac{\partial {x}_{i}}{\partial \xi }\) and,
then,
This relationship can be exploited to create an adder function. If there is an observable x3 due to a process ξ that maps x1 and x2 to x3 (ξ: x1 + x2 ↦ x3), then addition can be carried out in x-space. Specifically, to add two variables y1 + y2, first define,
where \(F({\bf{x}})=\log Pr({\bf{x}})\) for reasons that will become clear below. Then,
Moreover, since
which is obtained from Eqn (5), Eqn (12) can be given in terms of the mapped and measured variable x3,
The first term on the right-hand side carries out the mapping x1 + x2 ↦ x3 and the second term maps x3 ↦ y3. The extent of the mapping of each process is determined by the mechanical change in the thermodynamic process, Δξ. Importantly, n addition operations can be easily carried out in parallel if each respective observable xi is directly, or indirectly through other xj, mapped to a terminal observable xf,
Computationally, Eqn (15) is a general adder function for any thermodynamic process. A subtraction function,
can easily be constructed using a similar process but instead of measuring the product of the process xf, measure the amount of an intermediate variable xk needed to obtain a specific value of the product of the process xf. xf is thus the subtractant,
Multiplication and division processes similarly follow by exponentiating Eqns (15) or (17). For multiplication,
That is, multiplication and division are obtained by using the same processes as addition and subtraction but not working in \(\log\) space. Concrete examples are given next.
Application to analog systems
Experimentally, the process of mapping one object to another consists of either a deterministic or stochastic physical process ξ that relates the xi objects to xf, either directly or through other xj that may not be observable. In addition to probability densities, equations of state, defined as any function \(F\propto \log Pr\), can likewise be used for computing. If F is the equation of state, then ξ represents a process that changes the state. Consider the equation of state for an ideal gas, PV = ∑ixiRT for the case of n gas chambers each of volume fi and each containing xi moles of the gas. Suppose that ξ controls pistons that move all of the gas from the n chambers into a terminal chamber, initially empty, with volume Vf at constant temperature T. Since all of the gas from each chamber is emptied, \({\gamma }_{i}=\frac{\partial {x}_{i}}{\partial \xi }=1\). Then again using \({y}_{i}={\gamma }_{i}\log {f}_{i}\),
In this case, since the system is closed, ΔPV = 0 and ΔxfRT is simply PfVf. In short, addition can be carried out by any thermodynamic process because work is additive. Likewise, as we demonstrate below, the thermodynamic odds of one state to another, which is the exponent of the work to move from one state to another, is multiplicative in nature.
Oxidation-reduction phenomena in chemistry, as well as electrical phenomena, are represented by the thermodynamic equation of state ΔG = − zΔE where G is the free energy, z is a charge and ΔE is an electric potential. If z is a charge transferred due to chemistry (or a charge translocated in an electrical circuit), ξi is the extent of chemical reaction i (or analogously a unit length of the circuit component i), then, again representing \({y}_{i}={\gamma }_{i}\log {f}_{i}\), the relevant equations are,
where in Eqn (23) \(\frac{\partial G(z)}{\partial z}\frac{\partial z}{\partial {\xi }_{i}}=\frac{\partial V}{\partial {\xi }_{i}}\) was used. Eqns (22) and (23) describe the additivity of sequential chemical reactions i and voltages of sequential components in a circuit. The carrying out of arithmetic operations using voltages in electrical analog computing, an adder, is well known11,12.
In general for chemical reactions, the chemical free energy is G = ∑ixiμi where xi is the count or concentration of species i and μi is the chemical potential. Then from Eqns (7),
Eqn (25) is the basis for a chemical differential analyzer. For example, the differential equation,
is equivalent to,
and can be solved chemically by representing y as the system free energy G and for t the extent of the reaction ξ such that,
The first integral is simply the work done by the chemical system and the second is proportional to the extent to which the system has reacted (for example, the number of reactants consumed). Consequently,
For a mechanical differential analyzer, the integral is output onto a graph in which the position of the pen is the memory to which are added differentials of the integral. Multiplication can be carried out as the subsequent exponentiation of the integral. However, this need not be the case in thermodynamic computing, as the exponential of the free energy or entropy is the thermodynamic odds of the final state relative to the initial state. Consequently, the value of the exponential can be calculated directly by an appropriate odds ratio involving the ratio of reactant and product concentrations.
To demonstrate multiplication in chemistry using information, consider the reaction,
results in the combination of γA molecules of species A with γB molecules of species B to produce γC molecules of C. In this case,
where μi is the chemical potential of species i and β = 1/kBT is the reciprocal of the ambient energy, T is the temperature and kB is Boltzman’s constant. The chemical potential μi is related to the multinomial Boltzmann probability density,
where the probability of each species i depends on its standard free energy of formation, \({\mu }_{i}^{\circ }\),
The normalization factor is \({q}_{B}={\sum }_{i}^{M}{e}^{-{\mu }_{i}^{\circ }}\) Thus, the fi is the derivative of the probability density such that,
where the term in brackets is the normalized chemical potential, βμi.
The mapping needed for computing operations is implemented by a chemical reaction ξ where,
in which case \({\gamma }_{i}=\frac{\partial {x}_{i}}{\partial \xi }\) is the stoichiometric coefficient.
To compute a multiplication operation such as yA ⋅ yB = yC in chemical space when the number of particles of each species is large, we represent the initial values of the variable yA = fA with chemical A, yB = fB with chemical B, and yC = fC with chemical C where each is given respectively by,
To determine the value of yC after the continuous chemical operation ξ, the integral transform that is needed is the one involving the mapping ξ: xA + aB ↦ x3,
where KQ−1 is the thermodynamic odds of the reaction, K is the chemical equilibrium constant,
and Q is the chemical reaction quotient,
Substituting for yA and yB from Eqn (39),
Demonstration of chemical computation
The basic chemical operations corresponding to addition, subtraction, multiplication and division are shown in Fig. 1. In the chemical processes of addition and multiplication, the chemical reactants represent the operands and the chemical product represents the solution. These systems are open-loop control systems: the reactants are controlled and the chemical products are measured. For subtraction and division, at least one product is an operand and at least one reactant is a solution. Subtraction and division are carried out by closed-loop control: a reactant is titrated to obtain the desired chemical product.
A The relevant equation of state is the free energy, G. A and B are mapped to C through the chemical reaction ξ with signed stoichiometric coefficients \({\gamma }_{A}=\frac{\partial {n}_{A}}{\partial \xi },{\gamma }_{B}=\frac{\partial {n}_{B}}{\partial \xi }\) and \({\gamma }_{C}=\frac{\partial {n}_{C}}{\partial \xi }\). B Open-loop control allows for addition and multiplication reactions using the respective syntax below. The respective chemical solutions involve measuring reaction free energy and the product chemical potential for addition, or for multiplication, the thermodynamic odds of the reaction and the product odds (exponent of the chemical potential scaled by the thermal energy). In practice, measuring the chemical potential of the product also provides the reaction free energy since the chemical potentials of the reactants are control variables. C Closed-loop control similarly allows for subtraction and division operations.
To demonstrate the mechanics of the procedure, consider the case in which \({f}_{A}=\frac{1}{2},{f}_{B}=\frac{1}{4}\) and fC is unknown. Let the standard chemical potentials be given by \({\mu }_{A}^{\circ }={\mu }_{B}^{\circ }=-1\), \({\mu }_{C}^{\circ }=-2\), and let the system be at a non-equilibrium steady state in which the counts of the reactants are xA = 2, xB = 4. \({x}_{C}^{{\prime} }\) is measured. Consequently, the thermodynamic odds of the reaction KQ−1 are known. More precisely,
where the latter term accounts for the change in the probability as the total number of particles changes.
For the purpose of demonstration, the concentration of the product \({x}_{C}^{{\prime} }\) that is to be measured experimentally will simply be determined analytically from chemical thermodynamics (Eqs. (42) and (43)); that is, \({x}_{C}^{{\prime} }=1\) when KQ−1 = 8. Then carrying out the multiplication in chemistry,
Substituting in values,
or \(\frac{1}{2}\cdot \frac{1}{4}=\frac{1}{8}\).
A more insightful and realistic case is to carry out the operation when yA and yB are both large numbers and using parameters \({\mu }_{A}^{\circ }\), \({\mu }_{B}^{\circ }\) and \({\mu }_{C}^{\circ }\) from real chemical species. So that we know the exact solution to the problem, we take yA and yB to be such that,
Choosing a = 1.0 × 1036 gives,
First, we choose two convenient chemical species A and B to represent yA and yB such that yA = fA, yB = fB from Eqn (39). For convenience, we will use dihydroxyacetone and glyceraldehyde for A and B, respectively, and fructose for C. Their respective reference chemical potentials at pH = 7.0 and ionic strength 0.25 M are \({\mu }_{A}^{\circ }=-207.4\) kJ/mol, \({\mu }_{B}^{\circ }=-205.2\) kJ/mol, and \({\mu }_{C}^{\circ }=-426.7\) kJ/mol. At T = 298.15, the ambient energy is RT = 2.478956 kJ/mol. The respective counts of the chemicals are,
Assuming that the chemicals are in one liter (1 L) of water, division by Avogadro’s number gives concentrations in moles/liter (M),
where NAvo = 6.022141 × 1023 mol−1. Starting with no C present, only a small amount of A and B are allowed to react such that the final concentration of C is measured to be a little less than millimolar, \({n}_{C}^{{\prime} }=0.000944\) molar. At this concentration, the thermodynamic odds of the reaction are KQ−1 = 1.0 × 1018. Then the chemically computed value of yA ⋅ yB is,
The in-silico chemical calculation compares with the analytical value yA ⋅ yB = a such that
In this case, the error is due to the limited precision of the computer used to carry out the chemical calculations in-silico. In an actual chemical computer, the error will depend on the measurement precision of the starting material, nA and nB, the product, nC, and on the uncertainty in each of \({\mu }_{A}^{\circ },{\mu }_{B}^{\circ }\) and \({\mu }_{C}^{\circ }\). However, if calorimetry is used, only the chemical potential of the final product is needed, in that the value of K is the heat released during the reaction. In addition, the precision of the measurements can always be increased in practice through multiple dilutions or concentrations.
Extension to high dimensional computations
Scalar multiplication
In the examples above, the problem of finding the solution of yA ⋅ yB has been changed to the problem of finding the solution to K−1Q ⋅ fC, and we had to additionally calculate the transforms from fA → nA, fB → nB and \({n}_{C}^{{\prime} }\to {f}_{C}\). Thus, there may or may not be an advantage to using the chemical reactions to solve the multiplication problem in these simple cases.
However, consider a higher-dimensional example, in which we want to find the product fN of N + 1 random variables yi for i ∈ [0, …, N],
The chemical process for calculating fN is shown in Fig. 2. The chemical system has components {x1, …, x2N+1} which are subset into N + 1 reactants ri and N products pi. The yi = fi to be multiplied are represented by the chemical reactants ri. The pi are the reaction products with pN being the final product.
Chemical process for determining the product yN = ∏yi of N + 1 random variables yi.
The solution fN can be determined from the corresponding known reactant concentrations \({r}_{i}=\frac{{e}^{-{\mu }_{i}^{\circ }/{K}_{B}T}}{{f}_{i}}\), calculating or measuring K, and measuring pN,
The values of the intermediate products pi do not have to be stored in memory or even known. K in Eqn (70) is \(K={e}^{-\beta {\bf{S}}\cdot {\mu }^{\circ }\cdot {\bf{1}}}\), where 1 is the ones vector and \({\mu }^{\circ }={[{\mu }_{1}^{\circ },\ldots ,{\mu }_{M}^{\circ }]}^{T}\), the vector of known standard chemical potentials, and S is the stoichiometric coefficients for each of the M chemical species (reactants and products) for each of the Z reactions are captured in the stoichiometric matrix,
Combining multiplication and addition
To add two products together,
the two products are computed independently as described above, such that y = ∏yi of Ny + 1 random variables yi and z = ∏zi of Nz + 1 random variables zi, as shown in Fig. 3. The value y and z are then the input concentrations for chemicals Ay and Az. The sum of the products is then computed using the chemical reaction Ay + Az = Ayz. The chemical formulation is,
Here ri and \({r}_{i}^{{\prime} }\) are the respective reactants involved in the y-product reaction and the z-product reaction. Ky is \({K}_{y}={e}^{-\beta {{\bf{S}}}_{{\bf{y}}}\cdot {\mu }^{\circ }\cdot {\bf{1}}}\), where again 1 is the ones vector and \({\mu }^{\circ }={[{\mu }_{1}^{\circ },\ldots ,{\mu }_{M}^{\circ }]}^{T}\). Sy is the stoichiometric coefficients for each of the M chemical species (reactants and products) for each of the Zy reactions that synthesize y from the reactants ri,y. In Fig. 3, Zy = Ny. Analogous parameters are likewise defined regarding z.
Two independent products are computed chemically as described in the main text, such that yN = ∏yi of N + 1 random variables yi and zN = ∏zi of N + 1 random variables zi. The value \({y}_{{N}_{y}}+{z}_{{N}_{z}}\) are then the input concentrations for chemicals Ay and Az and are combined in the chemical reaction Ay + Az = Ayz.
Matrix multiplication
If instead the concentrations of intermediates are measured, then the reaction free energies ΔGj for each of the Z reactions can be determined. This allows for matrix-vector multiplication,
in which A is represented by S, the stoichiometric matrix, and in this case f is represented by the vector of chemical potentials, f = μ. The product b is,
This analysis is similar to that found by Aifer, et al.,2 but rather than harmonic potentials, chemical potentials are used. Notably, however, the system is easily generalized beyond equilibrium solutions since non-equilibrium boundary conditions can easily be applied. In this case, the computational solution is found at a non-equilibrium steady state. In principle, non-equilibrium boundary conditions would represent the computational problem through constraints in concentrations included in the probability density (Eqn (21)).
Moreover, given a set of V independent chemical systems k, each with its own stoichiometric matrix Sk and respective set of mapping processes (reactions) {ξj,k}, composite computations can be carried out such that,
Control surface and device design
To bridge the theoretical foundations of CRNs with practical implementations, we build on existing features and suggest the concept of a microfluidic device system designed to execute fundamental chemical computations described above. The system must support digital control interfaces to manipulate and measure signals, chemical compartmentalization strategies to enable execution of multiple operations, feedback mechanisms to chain chemical reactions in time, and consequently computations, and a mechanism to execute complex operations such as f(x) = logn(ex) enabled by the dynamics of certain species and permeable membranes13. We believe prototype systems can be implemented using current microfluidic fabrication techniques13,14, leveraging a wide range of microfluidic technologies discussed below.
Microfluidic technologies have evolved into versatile platforms supporting biological analysis, chemical synthesis, and computational modeling. Early advances focused on cell-based and droplet microfluidics, enabling single-cell manipulation15,16 and controlled droplet generation14. These systems demonstrated high precision in reagent handling and isolation, establishing foundational mechanisms for reaction control at the microscale. Further developments expanded into biomedical and environmental applications13,17, leveraging new materials and fabrication methods to integrate sensors and achieve biocompatibility and scalability.
Recent research emphasizes programmability, modularity, and adaptive control in microfluidic systems. Digital microfluidics and circuit-based network models18,19 introduced flexible droplet actuation schemes, while compiler-assisted and machine-learning-driven control frameworks20,21 have automated design and operation. Collectively, these innovations inform the proposed chemical computing architecture, combining precise micro-scale actuation, dynamic feedback control, and programmable digital interfaces to realize computationally capable biochemical systems.
Design overview
To carry out the needed operations, we propose the system presented in Fig. 4, consisting of a multilayer microfluidic chip that leverages the ideas above, incorporating:
-
Reaction chambers that host controlled CRNs, corresponding to arithmetic operations (addition, subtraction, multiplication, division) and exponential transformations15,16.
-
Microchannels and membranes engineered to modulate concentration gradients and enable transformations of the form \(\log n({e}^{[concentrations]})\), providing a proxy for nonlinear functions within the CRN framework.
-
Valves and pumps for precise flow control, enabling both open-loop and closed-loop operation as required for different classes of computation18. The valves are implemented using a crossbar network design22, borrowed from computer architecture, which allows routing of any input to any output channel or chamber.
-
Integrated sensors (optical or electrochemical) to measure reaction intermediates and final product concentrations17.
-
Digital interfaces for programmable control of fluid routing, timing, and data acquisition20,21.
Composition of readout signals can follow a reservoir computing approach to train readout signal weights.
Operational flow
The device operates in three phases:
-
1.
Initialization: Reactants corresponding to input variables (operands) are injected into designated chambers through programmable microvalves. Concentrations are set to encode input values.
-
2.
Computation: Mass-action kinetics describe the thermodynamic processes that execute CRN-based operations. For addition and multiplication, reactants are combined in open-loop chambers, and results are measured as product concentrations. For subtraction and division, closed-loop titration adjusts reactant inflow until the desired steady-state condition is reached. Exponential and logarithmic transformations are realized by leveraging semi-permeable membranes that regulate diffusion, effectively computing \(\log n({e}^{[concentrations]})\).
-
3.
Readout and Iteration: Embedded sensors capture chemical concentrations, which are digitized and fed back into the digital interface. Results can either be output externally or routed internally to subsequent chambers, enabling composition of additional CRN computations, such as a sequence of multiplications and additions.
Key features
Digital Control and Interfaces
A programmable controller synchronizes reactant injection, chamber mixing, and measurement. This enables composability of CRN operations in a manner analogous to instruction sequencing in electronic processors20.
Mechanical and Structural Aspects
The microfluidic chip incorporates elastomeric valves, micropumps, and membranes with tunable permeability. These features allow selective coupling of chemical reservoirs, ensuring robust control over concentration dynamics14,19.
Chemical Properties and Computation
Reaction free energies (G) and chemical potentials (μ) act as computational primitives. Open-loop systems leverage direct measurement of product potentials, while closed-loop control dynamically adjusts inflows to converge on the desired result. The membrane-mediated logarithmic transformation provides a mechanism to extend computation to nonlinear functions required for solving ODEs and higher-level mathematical tasks.
Computational significance
By combining precise microfluidic control with thermodynamic principles, the device acts as a biochemical analog computer. Its modularity allows chaining of operations, thereby realizing scalable reservoir computing frameworks. This approach enables experimental validation of CRN-based computation while providing a pathway toward energy-efficient, non-silicon analog computing architectures.
Discussion
Thermodynamic processes, such as chemical reactions, can be used to implement computational operations, encompassing tasks such as scalar addition, subtraction, multiplication, and division, as well as matrix manipulations, thereby enabling the solution of differential equations. This versatility highlights the potential of thermodynamic processes, and chemical reaction networks in particular, to execute complex operations typically reliant on digital systems, and to do so in a manner that minimizes the amount of energy used.
The statistical thermodynamic framework described herein elicits the question of how this framework relates to information theory, which was initially also based on statistical thermodynamic principles. In information theory, entropy is defined according to Boltzmann’s H-theorem, \({\sum }_{i}{p}_{i}\log pi\), where p(i) is the probability of object i. The term \(-\log {p}_{i}\) is referred to as the self-information of object i, I, or surprisal or a measure of uncertainty.
In statistical thermodynamics, The H-theorem \({\sum }_{i}{p}_{i}\log {p}_{i}\) is mostly of historical interest; instead, a multinomial probability density Pr is used to characterize a system, and is fully compatible with traditional statistical analyses such as Bayesian frameworks23. Entropy is defined as the logarithm of the probability density, \(\log Pr\), and is a measure of certainty of a state that includes all objects i in the system24. The greater the probability density, the larger the magnitude of the entropy. The probability of an individual object pi is simply that - the probability of i. It is proportional to the partition function of the object that includes all of the object’s internal degrees of freedom.
In information theory, probabilities are used as descriptors of a computational process. Probabilities are also descriptors of computational processes in thermodynamic computing, but in addition, computations are carried out directly using probabilities, probability ratios (odds ratios), or entropy/free energy production. Inherent in thermodynamic computing is that the computing is carried out by an explicit thermodynamic process for which the extent of the process is measure by ξ. Regardless of platform, whether it be a logic gate or the change in state in a metal-oxide semiconductor, all computational processes are thermodynamic in nature25. As such, the explicit use of thermodynamic functions in thermodynamic computing operations may provide the path to significant energy advantages over conventional computing.
The path towards thermodynamic computing with minimal energy demands may be attainable by integrating analog chemical operations with chemical reservoir computing, a concept that enables chemical systems to efficiently incorporate dynamic chemical and biochemical processes into structured outputs.
Despite their potential, implementing physical CRN-based computing devices presents significant challenges, discussed next.
Mapping computational problems onto chemical reaction networks must account for experimental constraints, including mass conservation and mitigation of reaction interference in large-scale implementations. Synthetic systems face additional hurdles in achieving robustness and repeatability, particularly in controlled environments where variations in chemical dynamics can impact computational accuracy.
The transition from a prototype hardware to an implementation-scale hardware based on off-the-shelf technologies may be economically and technically difficult at this stage of development. For instance, off-the-shelf microfluidic technologies using a digital-to-analog converter (DAC) and an analog-to-digital converter (ADC), such as that shown in Fig. 3, might consume an undesirable amount of energy. For instance, if every implemented function (addition, subtraction, multiplication, division) requires an ADC, DAC, or both, then the energy costs could be significant. However, these conversions are required only to pass the concentration output of one CRN module, representing a mathematical function, to the input of the next. This is analogous to digital systems, where modular composition also depends on encoding signals between subsystems, or to CMOS mixed-signal pipelines, where ADC/DAC stages bridge analog computation blocks.
It should be noted also that concerns regarding ADCs and DACs energy consumption apply primarily to high-speed data converters. CMOS analog or digital circuits must run at the clock rate of the target computation, and power scales with switching frequency. In contrast, CRNs in biology involving common small-molecule reactions evolve on time scales of seconds, not tens of microseconds. Therefore, high-speed conversion is not required. Commodity 10-bit ADCs span a wide energy range from the femtojoule to picojoule level per conversion, while commonly cited26,27 values of energy consumption of 10-100 mW correspond to multi-MHz samplers used in high-speed analog-digital links. Low-rate ADC/DAC operation provides sufficient temporal resolution to track the concentration trajectory and compose arithmetic modules, while staying in the ultra-low-energy regime per sample. Thus, while conversion is not free, its cost is small relative to the chemical timescales and does not likely challenge the feasibility or reproducibility of modular CRN-based ODE computation. Moreover, if each CRN representing a function were compiled into a single large CRN, and only a final product needs to be measured or controlled, then the number of DACs and ADCs could be reduced considerably. Ultimately, the feasibility of scaling up chemical computers would require careful accounting of energetic requirements and costs of advanced designs.
The rate of computation using CRNs could be inherently slow if appropriate reactions are not chosen, as the rate of chemical reactions can vary considerably. Thus, it will be important to choose reactions that have kinetics appropriate for the computational task. However, there are a number of redox reactions that have reasonable kinetics, and in fact, a single redox reaction could be used for all reactions in the CRN if each reaction is distinguishable by a distinct physical location. While a millisecond time scale for a single reaction may seem slow, chemical computing has the advantage that processes can be carried out simultaneously and in a highly parallel manner, just as with quantum computing.
On first encounter, one might think that the processes needed for a chemical computer would be too complicated to allow for large-scale or complex computations, as the number of required reactions would grow rapidly. However, the complexity of implementing these reactions may not be greater than implementing analogous electronic algorithms on CMOS devices. Certainly, implementing computations on a chemical computer would be far more feasible than implementing the same computations on a quantum computer, and a chemical computer likely has the same ability to speed up computations in that the computational processes would occur simultaneously.
The potential scalability and efficiency of chemical reaction networks for computational tasks are demonstrated by biological systems. Living systems, such as metabolic networks in biological cells, naturally perform complex mathematical functions. Future technology should allow this scalability to be extended into artificial biochemical devices, which would offer immense potential for energy-efficient, scalable solutions in scientific computing.
Thermodynamic chemical computing would require knowledge of standard thermodynamic parameters, such as equilibrium constants and standard state chemical potentials. Currently, large databases with thousands of equilibrium constants and chemical potentials are available. The accuracy of the values varies considerably, but accurate determination of equilibrium constants and chemical potentials is a mature technology area. If the accuracy of some values needs to be improved, sufficient funding only needs to be made available to do so.
To summarize, innovations in microfluidic technologies will play a critical role in scaling CRN-based computing. By fabricating miniature devices with precise control over chemical flows and interactions, researchers can create practical platforms for implementing these concepts. These advancements could bridge the gap between theoretical chemical and biochemical computation and its real-world applications, setting the stage for a paradigm shift to natural computing systems and a new era of energy-efficient, scalable computing.
Building on these findings, the development of extended mathematical models is key to broadening the scope of thermodynamic computing, and in particular, chemical and biochemical computing. Future work could focus on integrating multiscale and multiphase computations, enhancing the capacity of chemical reaction networks to simulate complex systems. Additionally, combining chemical computation with artificial intelligence frameworks may yield powerful hybrid learning systems, further expanding the applicability of this approach.
Methods
The mathematical framework is described in the Results section.
Data availability
This manuscript does not report data generation or analysis.
References
Conte, T. et al. Thermodynamic computing (2019). https://arxiv.org/abs/1911.01968 (2019).
Aifer, M. et al. Thermodynamic linear algebra. npj Unconvent. Comput. 1, 1–11 (2024).
Duffield, S., Aifer, M., Crooks, G., Ahle, T. & Coles, P. J. Thermodynamic matrix exponentials and thermodynamic parallelism. Phys. Rev. Res. 7, 013147 (2025).
Johnson, C., Delattre, H., Hayes, C. & Soyer, O. S. An Evolutionary Systems Biology View on Metabolic System Structure and Dynamics, 159–196 (Springer International Publishing, Cham, 2021). https://doi.org/10.1007/978-3-030-71737-7_8.
Donatella, K. et al. Scalable thermodynamic second-order optimization (2025). https://arxiv.org/abs/2502.08603 (2025).
Vasic, M. D. S., & Khurshid, S. Crn++: Molecular programming language. Nat Comput. 19, 391–407 (2020).
Baltussen, M. G., de Jong, T. J., Duez, Q., Robinson, W. E. & Huck, W. T. S. Chemical reservoir computation in a self-organizing reaction network. Nature 631, 549–555 (2024).
Johnson, C. G. M., Bohm Agostini, N., Cannon, W. R. & Tumeo, A. Computing with a chemical reservoir. In 2024 IEEE International Conference on Rebooting Computing (ICRC), 1–7 (2024).
Yan, M. et al. Emerging opportunities and challenges for the future of reservoir computing. Nat. Commun. 15, 1–18 (2024).
Agostini, N. B., Johnson, C. G. M., Cannon, W. R. & Tumeo, A. Chemcomp: compiling and computing with chemical reaction networks. In 2025 Design, Automation & Test in Europe Conference (DATE), 1–7 (2025).
Bush, V. The differential analyzer. a new machine for solving differential equations. J. Frankl. Inst. 212, 447–488 (1931).
Shannon, C. E. Mathematical Theory of the Differential Analyzer, 496–513 (Wiley-IEEE Press, 1993).
Niculescu, A.-G., Chircov, C., Bîrcă, A. C. & Grumezescu, A. M. Fabrication and applications of microfluidic devices: a review. Int. J. Mol. Sci. 22, 2011 (2021).
Elvira, K. S., Gielen, F., Tsai, S. S. H. & Nightingale, A. M. Materials and methods for droplet microfluidic device fabrication. Lab Chip 22, 859–875 (2022).
Wheeler, A. R. et al. Microfluidic device for single-cell analysis. Anal. Chem. 75, 3581–3586 (2003).
Tang, X. et al. Efficient single-cell mechanical measurement by integrating a cell arraying microfluidic device with magnetic tweezer. IEEE Robot. Autom. Lett. 6, 2978–2984 (2021).
Alhalaili, B., Popescu, I. N., Rusanescu, C. O. & Vidu, R. Microfluidic devices and microfluidics-integrated electrochemical and optical (bio)sensors for pollution analysis: a review. Sustainability 14, 12844 (2022).
Al-Lababidi, M. & Abdelgawad, M. Minimum movable droplet volume in digital microfluidics depends on the grounding scheme in addition to electrode size. Sens. Actuators A Phys. 355, 114299 (2023).
Rousset, N. et al. Circuit-based design of microfluidic drop networks. Micromachines 13, 1124 (2022).
Loveless, T. & Brisk, P. Compiling functions onto digital microfluidics. In Proceedings of the 21st ACM/IEEE International Symposium on Code Generation and Optimization, 136–148 (Association for Computing Machinery, New York, NY, USA, 2023). https://doi.org/10.1145/3579990.3580023.
McIntyre, D., Lashkaripour, A., Fordyce, P. & Densmore, D. Machine learning for microfluidic design and control. Lab Chip 22, 2925–2937 (2022).
Hennessy, J. L. & Patterson, D. A.Computer Architecture: A Quantitative Approach (Morgan Kaufmann, San Francisco, CA, 2019), 6th edn. See Appendix F: Interconnection Networks, discussion of the Crossbar Network.
Jaynes, E. T. Probability Theory: The Logic of Science (Cambridge University Press, Cambridge, UK, 2003).
Planck, M. The Theory of Heat Radiation (Blakiston, Philadelphia, 1914).
Bennett, C. H. The thermodynamics of computation–a review. Int. J. Theor. Phys. 21, 905–940 (1982).
Kuntz, T. G. R., Rodrigues, C. R. & Nooshabadi, S. An energy-efficient 1msps 7μw 11.9fj/conversion step 7pj/sample 10-bit SAR ADC in 90nm. In 2011 IEEE International Symposium of Circuits and Systems (ISCAS), 261–264, 2011.
Meganathan, D., Sukumaran, A., Dinesh Babu, M., Moorthi, S. & Deepalakshmi, R. A systematic design approach for low-power 10-bit 100ms/s pipelined ADC. Microelectron. J. 40, 1417–1435 (2009).
Acknowledgements
This research was supported by funding from the U.S. Department of Energy (DOE), Office of Advanced Scientific Computing Research (ASCR), as part of the ChemComp (81794) and CIS (85677) projects under the Exploratory Research for Extreme-Scale Science (EXPRESS2023 and EXPRESS2025) programs. Pacific Northwest National Laboratory (PNNL) is managed and operated by Battelle under contract DE-AC05-76RL01830 for the U.S. DOE.
Author information
Authors and Affiliations
Contributions
W.R.C. developed the mathematical fundamentals. W.R.C., C.J, N.B.A, and A.T. analyzed, discussed the results, and wrote the manuscript. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cannon, W.R., Johnson, C.G.M., Bohm Agostini, N. et al. A mathematical framework for thermodynamic computing with applications to chemical reaction networks. npj Unconv. Comput. 3, 16 (2026). https://doi.org/10.1038/s44335-026-00057-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44335-026-00057-5