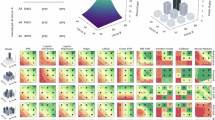
Abstract
Machine learning (ML) models are often based on complex black-box architectures that are difficult to interpret. This interpretability problem can hinder the use of ML in fields like medicine, ecology, and insurance, and has boosted research in interpretable machine learning (IML). Here, we propose a novel approach for the functional decomposition of black-box predictions, which is a core concept of IML. This approach replaces the prediction function with a surrogate model consisting of simpler subfunctions, providing insights into the direction and strength of the main feature contributions and their interactions. Our method is based on a concept termed “stacked orthogonality”, which ensures that the main effects capture as much functional behavior as possible. To compute the subfunctions, we combine neural additive modeling with an efficient post-hoc orthogonalization procedure. Our method yielded plausible results in an analysis of stream biological condition in the Chesapeake Bay watershed (United States).
Similar content being viewed by others
Introduction
Machine learning (ML) has increased greatly in both popularity and significance, driven by an increase in methods, computing power and data availability1, making it an useful tool in the development of artificial intelligence (AI). On May 29, 2025, a search on Web of Science for publications including the term “machine learning” yielded more than 474,000 results, corresponding to an average annual increase by more than 22% since 2006. ML models are often characterized by their high generalizability, making them particularly successful when used for supervised learning tasks like classification and risk prediction. In recent years, ML models based on deep artificial neural networks (ANNs) have led to groundbreaking results in the development of high-performing prediction models2.
The high prediction accuracy of modern ML models is usually achieved by optimizing complex “black-box” architectures with thousands of parameters. As a consequence, they often result in predictions that are difficult, if not impossible, to interpret. This interpretability problem can hinder the use of ML in fields like medicine, ecology, and insurance, where an understanding of the model and its inner workings is paramount to ensure user acceptance and fairness3. In a recent environmental study, for example, we explored the use of ML to derive predictions of stream biological condition in the Chesapeake Bay watershed (CBW) of the mid-Atlantic coast of North America4. Clearly, if these predictions are intended to inform future management policies (projecting, e.g., changes in land use, climate, and watershed characteristics), they need to be interpretable in terms of relevant features as well as the directions and strengths of the feature effects. We will return to this example below in order to illustrate our proposed methodology.
Interpretable machine learning
In recent years, the need for understanding ML models has boosted research in the field of interpretable machine learning (IML3,5,6). In this field, interpretability is commonly defined as “the degree to which a human can understand the cause of a decision”7. A related concept considered separately in some works is explainability, which describes “the internal logic and mechanics that are inside a ML system”8. Because the methodology presented in this work applies to both concepts, we will not distinguish between the two.
The focus of this paper is on IML for supervised learning tasks, which involve a set of features X = {X1, …, Xd} to derive predictions of a qualitative or quantitative outcome variable Y. Denoting the model (i.e., the prediction function) by \(F(X)\in {\mathbb{R}}\), interpretability can generally be achieved in two ways: The first approach is to impose an interpretable structure on F during the learning process (“model-based” or “by-design” interpretability5,6). An example of this approach is the least absolute shrinkage and selection operator9, which, in its basic form, assumes F to be linear in the features. Consequently, each feature effect is interpretable in terms of a real-valued coefficient. The second approach, which is particularly applicable to black-box models, aims to achieve interpretability by post-processing an already learned prediction model (“post hoc” interpretability6,10). Here we will consider model-agnostic post-processing methods, which can be applied to a broad range of prediction functions regardless of the ML method applied to the training data3. Popular examples of model-agnostic methods include partial dependence plots (PDP) and accumulated local effects (ALE) plots. The underlying principle of these methods is to measure the variability of the prediction function F with respect to changes in subsets of the features X (an approach that is closely linked to the concept of sensitivity analysis in numerical and nonlinear regression modeling11,12).
While PDP and ALE plots have become established methods in IML, they are not without limitations. For example, PDP have been criticized for ignoring the correlations between the feature of interest and the other features, relying on data points with a very low probability of being observed. This “extrapolation” issue may result in misleading effect estimates when the features are correlated3. Similarly, PDP may hide possible interaction effects of the features, a problem that can be alleviated by individual conditional expectation plots in some cases3,13. While ALE plots avoid extrapolation of the data14, Grömping15 observed that these plots do not generally identify the linear shapes of the main effects in a linear prediction model. As a consequence, the feature effects depicted by ALE plots may show systematic deviations from the respective effects in the model formula (for which an explanation is sought). The method proposed in this paper is not affected by these issues: It avoids hiding feature interactions by explicitly including these terms in the estimation procedure and is solely based on the multivariate feature distribution to avoid extrapolation. Furthermore, it does not alter the shapes of the main effects in a linear model.
Functional decomposition
The basic idea of our method is to achieve interpretability by decomposing the prediction function F (depending on all features X) into a set of simpler ("better interpretable”) functions depending on subsets of the features only. More specifically, let ϒ = {1, …, d} the set of feature indices and \({\mathcal{P}}(\Upsilon )\) the power set (i.e., the set of all subsets) of ϒ. Then F can be decomposed into a sum of functions
where \(\mu \in {\mathbb{R}}\) is an intercept term and, for any \(\theta \in {\mathcal{P}}(\Upsilon )\backslash {{\emptyset}},{X}_{\theta }\) denotes the subset of features with indices θ. For example, if d = 3 and θ = {1, 3}, then Xθ is given by {X1, X3}. Accordingly, the intercept term can be defined as \(\mu ={f}_{{{\emptyset}}}\). Note that the last sum in Eq. (1) consists of only one summand.
In IML, the main focus is usually on the subset of functions fθ with ∣θ∣ = 1 ("main effects”, first sum in Eq. (1)) and ∣θ∣ = 2 ("two-way interactions”, second sum in Eq. (1)). For main effects, fθ depends on only one feature Xj, j ∈ ϒ, allowing for a simple graphical analysis that plots the values of fθ(Xj) against the values of Xj. For example, in the aforementioned study on stream biological condition, the main effect of 30-year mean annual precipitation shows a positive association between the amount of precipitation and the predicted values of stream condition (see Fig. 3 below). Two-way interactions, on the other hand, can be visualized using heatmaps or contour plots. For example, Fig. 4 below presents a plot of the interaction between the elevation of the sample sites and the percentage of upstream catchment area as developed, showing elevations at which land use for development leads to low biotic integrity. As demonstrated by these examples, both main effects and two-way interactions allow for simple graphical interpretations of the respective feature effects, whereas the functions fθ with ∣θ∣ > 2 (termed “multivariate feature interactions”) constitute the less interpretable parts of F.
This paper presents a novel approach to specify and compute the functions fθ, given a fixed (possibly black-box) prediction function F. The proposed method also allows the measurement of the “degree of interpretability” by quantifying the importance of the main and two-way interaction effects in Eq. (1). We emphasize that our methodology is designed to decompose the prediction function F but not to learn it from a set of data. Accordingly, we assume that F is not subject to sampling variability but has been derived previously by the application of some ML method. Our method is based on regularity conditions that are similar to those described by Hooker16; however, we consider a different type of functional decomposition and also employ a different computational methodology.
Conditions on the features and the prediction function
It is clear from Eq. (1) that the functions fθ are not uniquely defined. For example, let d = 2, μ = 0, and F(X1, X2) = X1 + X1 ⋅ X2. Then the sets of functions {f1(X1) = X1, f2(X2) = 0, f12(X1, X2) = X1 ⋅ X2} and {f1(X1) = 0.5 ⋅ X1, f2(X2) = 0, f12(X1, X2) = 0.5 ⋅ X1 + X1 ⋅ X2} both satisfy Eq. (1). As a consequence, further assumptions are needed to derive a unique representation of Eq. (1).
Our first set of assumptions is on the features X = {X1, …, Xd}. In line with Hooker16, we consider the features as real-valued random variables, assuming that X1, …, Xd are defined on a joint probability space with probability function PX. We further assume that each Xj, j ∈ ϒ, has bounded support. Note that these are rather weak assumptions in practice, allowing X to include both continuous and categorical features (the latter encoded by sets of dummy variables).
Regarding the functions in Eq. (1), we assume that each fθ, and also F, is square integrable with respect to PX. Again, this is a rather weak assumption, as square integrable functions emerge from many popular ML methods. They include, for instance, the piecewise prediction functions obtained from random forests and tree boosting, and also many ANN predictors after transformation by a sigmoid activation function. Following Hooker16, we define the variance of \({f}_{\theta },\theta \in {\mathcal{P}}(\Upsilon )\backslash {{\emptyset}}\), by \({\sigma }_{\theta }^{2}=\int{f}_{\theta }^{2}({X}_{\theta })d{P}_{X}\), the variance of F by \({\sigma }_{F}^{2}=\int{(F(X)-\mu )}^{2}d{P}_{X}\), and the covariance of fθ and \({f}_{{\theta }^{{\prime} }},\theta ,{\theta }^{{\prime} }\in {\mathcal{P}}(\Upsilon )\backslash {{\emptyset}}\), by \({\sigma }_{\theta {\theta }^{{\prime} }}=\int{f}_{\theta }({X}_{\theta }){f}_{{\theta }^{{\prime} }}({X}_{{\theta }^{{\prime} }})d{P}_{X}\). Without loss of generality, we assume that each \({f}_{\theta },\theta \in {\mathcal{P}}(\Upsilon )\backslash {{\emptyset}}\), is centered around zero, i.e., ∫ fθ(Xθ) dPX = 016 (p. 714). Finally, we assume that the functions \({f}_{\theta },\theta \in {\mathcal{P}}(\Upsilon )\backslash {{\emptyset}}\), are linearly independent. This assumption means that each fθ forms a closed subspace of the Hilbert space of square integrable functions. In practice, it implies that each fθ carries unique information about F and that all functions fθ are non-zero.
Generalized functional analysis of variance (ANOVA)
Next, we define a set of requirements to describe the relations between the functions fθ. Our main requirement is that the summands in Eq. (1) are well separated, meaning that higher-order effects (i.e., functions with large ∣θ∣) do not contain any components of lower-order effects with small ∣θ∣ (see below for a mathematical treatment). In particular, we require that predictive information explained by a main effect is not contained in the higher-order effects that include the corresponding feature (purity criterion, Molnar3, Section 8.4). A related requirement is optimality, meaning that lower-order functions should capture as much functional behavior as possible16.
To implement the above requirements, Hooker16 proposed a decomposition termed generalized functional ANOVA. With this approach, the functions in Eq. (1) are required to be hierarchically orthogonal, satisfying the constraints
Hierarchical orthogonality implies that for any given \({\theta }^{{\prime} }\), the effect \({f}_{{\theta }^{{\prime} }}({X}_{{\theta }^{{\prime} }})\) is orthogonal to all higher-order effects fθ(Xθ) with \({X}_{\theta }\supseteq {X}_{{\theta }^{{\prime} }}\)16,17. It thus provides an implementation of the purity criterion, ensuring that higher-order effects are uncorrelated with lower-order effects. Furthermore, the constraints in Eq. (2) provide an implementation of optimality because, according to Eq. (2), all lower-order effects \({f}_{{\theta }^{{\prime} }}\) are orthogonal projections of the combined effects \({f}_{{\rm{comb}}({\theta }^{{\prime} },\theta )}:= {f}_{{\theta }^{{\prime} }}+{f}_{\theta }\) onto the respective lower-order subspaces. It follows from the Hilbert projection theorem18 that the lower-order effects \({f}_{{\theta }^{{\prime} }}\) capture as much of the variance of \({f}_{{\rm{comb}}({\theta }^{{\prime} },\theta )}\) (i.e., as much functional behavior of \({f}_{{\rm{comb}}({\theta }^{{\prime} },\theta )}\)) as possible.
In his original work on generalized functional ANOVA, Hooker16 specified conditions for the uniqueness of the functions fθ. He considered a more general definition of the integral in Eq. (2), allowing for weight functions other than the probability density function of X. Based on the same decomposition, Chastaing et al.19 studied further assumptions on the feature distribution PX. The authors also introduced a coefficient to measure the importance of individual feature combinations. For each \(\theta \in {\mathcal{P}}(\Upsilon )\backslash {{\emptyset}}\), this coefficient is defined as \({S}_{\theta }=({\sigma }_{\theta }^{2}+{\sum }_{{\theta }^{{\prime} }\ne \theta }{\sigma }_{\theta {\theta }^{{\prime} }})/{\sigma }_{F}^{2}\) (generalized Sobol sensitivity index, Chastaing et al.19, p. 2427).
Computational challenges
In recent years, functional decomposition has been acknowledged as a key concept in making ML models explainable3. In practice, however, the application of functional decomposition methods remains challenging. This is mainly due to the computational and numerical issues associated with the estimation of the feature effects fθ. In fact, despite the availability of algorithms to achieve hierarchical orthogonality16,19,20, state-of-the-art methods still involve systems of equations that are, even for a moderate feature count, “complex and computationally intensive”3. Here we introduce stacked orthogonality, an alternative approach to implement purity and optimality. Based on the conditions of stacked orthogonality, we will present an algorithm to estimate the functions fθ in a computationally efficient manner.
Functional decomposition with stacked orthogonality
Analogous to generalized functional ANOVA, our method is based on the functional decomposition in Eq. (1). However, instead of the hierarchical orthogonality constraints in Eq. (2), we require the functions fθ to meet the stacked orthogonality constraints
where k ∈ ϒ denotes the effect level. Throughout the paper, we will use the terms “order” and “level” interchangeably. Unlike hierarchical orthogonality, which requires the effect of each individual feature combination θ to be uncorrelated with higher-order effects, the conditions in Eq. (3) provide a level-wise implementation of the purity criterion: for each level k, the sum of all level-k effects is required to be uncorrelated with the sum of all lower-level effects (including the intercept with \(| {\theta }^{{\prime} }| =0\))—hence the term “stacked orthogonality”. In addition to implementing purity, the constraints in Eq. (3) also provide a level-wise implementation of optimality. This is because, according to Eq. (3), the sum of the “lower-order” effects (with levels <k) is an orthogonal projection of the sum of the “current-order” effects (with levels ≤k) onto the lower-order subspace. It follows from the Hilbert projection theorem that the sum of lower-order effects captures as much of the variance of the sum of the current-order effects (i.e., as much functional behavior at the current level) as possible.
A convenient feature of stacked orthogonality is that the variance of F can be decomposed in a level-wise fashion, giving rise to the calculation of level-wise coefficients of explained variation. More specifically, for each k ∈ ϒ, we define the fraction of \({\sigma }_{F}^{2}\) explained by the k-th level as
By definition, it holds that \(\mathop{\sum }\nolimits_{k = 1}^{d}{I}_{k}=1\) if the stacked orthogonality constraints in (3) are met. Consequently, by calculating I1 (fraction of \({\sigma }_{F}^{2}\) explained by the main effects) and I2 (fraction of \({\sigma }_{F}^{2}\) explained by the two-way interaction effects), it is possible to quantify the degree of interpretability of the prediction model F. We emphasize that the definition in Eq. (4) is different from the generalized Sobol sensitivity indices in ref. 19, as the latter refer to contributions of individual feature combinations \(\theta ,\theta \in {\mathcal{P}}(\Upsilon )\), whereas Ik, k ∈ ϒ, measures the level-wise contributions of all features. For example, in the aforementioned study on stream biological condition, the contribution of the main effects (as measured by I1) was 80.6%. The contribution of the interaction effects (I2) was 2.5%, leading to a total of 83.1% model interpretability.
Estimation by neural additive models and post-hoc orthogonalization
As stated above, the application of functional decomposition methods strongly depends on the availability of a user-friendly algorithm to compute the functions fθ. To arrive at the decomposition in Eq. (1) satisfying the stacked orthogonality constraints, we propose the following three-step procedure:
In the first step, we generate a sample of n data points \({\mathcal{S}}={\{{F}_{i},{X}_{i1},\ldots ,{X}_{id}\}}_{i = 1,\ldots ,n}\), where Xij, j ∈ ϒ, and Fi = F({Xi1, …, Xid}) denote the j-th feature value and the value of the prediction function, respectively, of the i-th data point. For instance, the data could be sampled from an available set of training data that were used previously for the learning of F. In this case, the probability measure PX is given by the distribution of the feature values in the training data. Alternatively, one could use a grid of feature values to generate \({\mathcal{S}}\) (corresponding to uniformly distributed features) or some other reference distribution for which an explanation is sought.
In the second step, we use the data generated in Step 1 to obtain initial estimates \({f}_{\theta }^{0}\) of the functions \({f}_{\theta },\theta \in {\mathcal{P}}(\Upsilon )\backslash {{\emptyset}}\). This is done by fitting a neural additive model (NAM21) of the form
where Xiθ are the values of Xθ corresponding to the i-th data point. Model fitting is performed using a backpropagation procedure, with each function \({f}_{\theta }^{0}\) represented by an ANN depending on the respective feature subset Xθ (see Fig. 1 for an illustration). As demonstrated by Agarwal et al.21, NAMs allow for modeling a wide range of functional shapes, exploiting the property of ANNs to approximate general classes of functions arbitrarily well22,23,24,25,26,27,28,29,30. Compared to Agarwal et al.21, our only additional requirement (needed for Step 3 below) is that all ANNs in Eq. (5) are linear in their output layers. More specifically, we require each vector \({{\bf{f}}}_{\theta }^{0}={({f}_{\theta }^{0}({X}_{1\theta }),\ldots ,{f}_{\theta }^{0}({X}_{n\theta }))}^{\top }\in {{\mathbb{R}}}^{n},\theta \in {\mathcal{P}}(\Upsilon )\backslash {{\emptyset}}\), to be of the form
where \({{\bf{U}}}_{\theta }\in {{\mathbb{R}}}^{n\times {b}_{\theta }}\) and \({b}_{\theta }\in {\mathbb{N}}\) are the outputs and the number of units, respectively, of the penultimate layer, and \({{\bf{w}}}_{\theta }^{0}\in {{\mathbb{R}}}^{{b}_{\theta }}\) is a vector of weights. Note that Eq. (5) does not contain an intercept term. Accordingly, the initial estimate of μ is given by \({\mu }^{0}={f}_{{{\emptyset}}}^{0}=0\), and we define \({b}_{{{\emptyset}}}=1,{{\bf{U}}}_{{{\emptyset}}}={(1,\ldots ,1)}^{\top }\in {{\mathbb{R}}}^{n\times 1}\), and \({{\bf{w}}}_{{{\emptyset}}}^{0}=0\). Updates of the initial intercept vector \({{\bf{f}}}_{{{\emptyset}}}^{0}={{\bf{U}}}_{{{\emptyset}}}{{\bf{w}}}_{{{\emptyset}}}^{0}\in {{\mathbb{R}}}^{n}\) will be computed during the post-hoc orthogonalization procedure described below. Also note that it is possible to extend the NAM by an additional nonlinear activation function in the output layer. This approach may, for example, be convenient when the prediction space is constrained to an interval (e.g., when predictions are given by a set of probabilities Fi ∈ [0, 1]). In this case, our method would require linearity only for the functions \({f}_{\theta }^{0}\) but not necessarily for the output of the NAM. The latter would then be given by Fi = g(ηi), where g denotes the activation function and ηi is the right-hand side of (5). Accordingly, the stacked orthogonality constraints would not apply to Fi but to ηi, which is analogous to the interpretation of generalized additive models (GAMs31, where g takes the role of an inverse “link function”). For notational convenience, we will only consider linear NAM outputs in the remainder of this paper. For details on the specification of the ANNs in Eq. (5), we refer to the “Methods” section.
In the example considered here, there are two features X1 and X2. Accordingly, the set of functions \({f}_{\theta }^{0},\theta \in {\mathcal{P}}(\Upsilon )\backslash {{\emptyset}}\), is given by the two main effects \({f}_{1}^{0}({X}_{1}),{f}_{2}^{0}({X}_{2})\) and the two-way interaction \({f}_{12}^{0}({X}_{1},{X}_{2})\). Each function is represented by a fully connected artificial neural network (ANN). The units in the penultimate layers of the ANNs are denoted by \({U}_{1}\in {{\mathbb{R}}}^{{b}_{1}},{U}_{2}\in {{\mathbb{R}}}^{{b}_{2}}\) and \({U}_{12}\in {{\mathbb{R}}}^{{b}_{12}}\), where b1, b2, and b12 are the widths of the layers. The outputs of the ANNs are given by the dot products \({U}_{1}^{\top }{{\bf{w}}}_{1}^{0},{U}_{2}^{\top }{{\bf{w}}}_{2}^{0}\) and \({U}_{12}^{\top }{{\bf{w}}}_{12}^{0}\), where \({{\bf{w}}}_{1}^{0},{{\bf{w}}}_{2}^{0}\) and \({{\bf{w}}}_{12}^{0}\) are vectors of weights. The prediction function F(X1, X2) is given by the sum of the three dot products (hence the term neural additive model). The parameters of the ANNs are estimated jointly by backpropagation. Details on model fitting and the specification of the ANN architectures are given in the “Methods” section.
We emphasize that we do not use NAMs for supervised learning, i.e., to derive the relationship between an outcome variable Y and a set of features X1, …, Xd. Instead, we consider the predicted values Fi as the outcome of the NAM. Given Xi, these values are deterministic, and hence the right-hand side of Eq. (5) does not include a residual error term. Put differently, the right-hand side of Eq. (5) defines a “surrogate model” for the prediction model F. Importantly, because we want to arrive at an exact decomposition of the form Eq. (1), we do not aim to avoid overfitting the data. Instead, we run the backpropagation procedure until it achieves an (almost) perfect correlation between the left-hand side and the right-hand side of Eq. (5). This is possible due to the approximation properties of ANNs (see the section on experiments with synthetic data). We further note that model fitting can be done very conveniently using established ANN implementations in Python32 and R33 (see the attached code at https://github.com/Koehlibert/ONAM).
In the third step, we apply a post-hoc orthogonalization procedure to the initial estimates \({f}_{\theta }^{0}\). This is necessary to ensure that the final estimates satisfy the stacked orthogonality conditions in Eq. (3). The post-hoc orthogonalization procedure considered here is an extension of the method by Rügamer34; it proceeds in an iterative manner, starting at the highest interaction level and descending down to the main effects. We describe the first two iterations of the procedure in a non-technical way. A formal definition of the algorithm is given in the “Methods” section.
In the first iteration of the post-hoc orthogonalization procedure, the idea is to achieve orthogonality between the d-way interaction effect and the sum of all lower-order effects (∣θ∣ < d). To this end, the vector of d-way interactions (given by \({{\bf{f}}}_{\Upsilon }^{0}\)) is projected onto the column space spanned by the “lower-order” matrices Uθ, ∣θ∣ < d (including \({{\bf{U}}}_{{{\emptyset}}}\), which is a vector of ones). Next, \({{\bf{f}}}_{\Upsilon }^{0}\) is replaced by the vector orthogonal to this space, giving the new vector of d-way interactions \({{\bf{f}}}_{\Upsilon }^{1}\). Note that \({{\bf{f}}}_{\Upsilon }^{1}\) has zero mean, as the lower-order column space contains a column of ones, and as \({{\bf{f}}}_{\Upsilon }^{1}\) is orthogonal to this space. The lower-order functions are updated by adding the projected values of \({{\bf{f}}}_{\Upsilon }^{0}\) to the initial lower-order functions, giving new functions \({{\bf{f}}}_{\theta }^{1},| \theta | < d\) (including a new intercept \({{\bf{f}}}_{{{\emptyset}}}^{1}\)).
In the second iteration, the idea is to achieve orthogonality between the sum of the effects of order d − 1 and the sum of all effects with ∣θ∣ < d − 1. Analogous to the first iteration, the effects of order d − 1 are summed up and projected onto the column space spanned by the matrices Uθ, ∣θ∣ < d − 1 (again including \({{\bf{U}}}_{{{\emptyset}}}\)). Next, each \({{\bf{f}}}_{\theta }^{1}\) with ∣θ∣ = d − 1 is replaced by its respective vector orthogonal to this column space, giving new estimates \({{\bf{f}}}_{\theta }^{2}\) of the effects of order d − 1. The functions with θ < d − 1 are updated in the same way as in the first iteration, resulting in new estimates \({{\bf{f}}}_{\theta }^{2}\), ∣θ∣ < d − 1, whereas the “higher-order” vector \({{\bf{f}}}_{\Upsilon }^{1}\) is left unchanged (\({{\bf{f}}}_{\Upsilon }^{1}\equiv {{\bf{f}}}_{\Upsilon }^{2}\)).
Iterating the above procedure (i.e., establishing orthogonality between the sums of the current-order and the lower-order effects while leaving higher-order effects unchanged) ensures stacked orthogonality of the final estimates \({{\bf{f}}}_{\theta }^{d-1}\). As a result, one obtains the desired decomposition of the prediction function F. We emphasize that post-hoc orthogonalization does not require re-fitting the NAM in Eq. (5) but can be performed rather efficiently by multiplying a set of matrices and vectors. In case of a high(er)-dimensional feature set, the number of summands in Eq. (5) can easily be reduced to a subset of “relevant” effects, see Remark 2 below.
Remark 1
NAM fitting is based on ANN layers with prespecified numbers of hidden units. We note that these numbers may not always be sufficient to closely approximate the true underlying functions, especially when the latter are highly non-linear. To address this issue and to “further improve accuracy and reduce the high-variance that can result from encouraging the model to learn highly nonlinear functions”, Agarwal et al.21 proposed to compute the final function estimates by an average of multiple NAM fits ("ensemble approach”). In line with this strategy, we stabilize our function estimates by fitting an ensemble of NAMs with different weight initializations and by applying the post-hoc orthogonalization procedure to each member of the ensemble. Afterwards, the orthogonalized estimates are averaged, giving vectors of the form \({\bar{{\bf{f}}}}_{\theta }^{d-1}=\mathop{\sum }\nolimits_{r = 1}^{R}{{\bf{f}}}_{\theta }^{d-1,r}/R\), where R is the size of the ensemble and \({{\bf{f}}}_{\theta }^{d-1,r}\) refers to the post-hoc-orthogonalized estimate of the r-th ensemble member. Note that this procedure does not substantially increase the run time of the algorithm, as NAM fitting with different weight initializations can be parallelized. We further note that the averaged estimates are no longer guaranteed to satisfy the stacked orthogonality constraints in Eq. (3). To overcome this problem, we add a final post-hoc orthogonalization step to our algorithm, replacing the outputs Uθ by the averaged vectors \({\bar{{\bf{f}}}}_{\theta }^{d-1}\) and applying the above procedure to the averaged estimates.
Remark 2
In settings with a large number of features, the number of interaction terms in Eq. (5) is very high (\(\mathop{\sum }\nolimits_{l = 2}^{d}(\begin{array}{c}d\\ l\end{array})\), growing exponentially in d). In these cases, one may be interested in the interpretation of only a small subset of effects. For example, in the aforementioned study on stream biological condition, we analyzed all main effects and three two-way interaction effects (instead of all possible 524,268 interaction terms defined by the 19 features, see below). The stacked orthogonality approach can easily be adapted to these settings; all one has to do is to redefine the NAM in Step 2. To this end, let \(\Theta \subset {\mathcal{P}}(\Upsilon )\backslash {{\emptyset}}\) represent the effects of interest, and let \({\mathcal{P}}(\Upsilon )\backslash (\Theta \cup {{\emptyset}})\) be the corresponding set of “non-interesting” effects. Then \({\mathcal{P}}(\Upsilon )\backslash (\Theta \cup {{\emptyset}})\) can be removed from the lower-order sums in Eq. (5) and absorbed into the last summand \({f}_{\Upsilon }^{0}\). Post-hoc orthogonalization can be applied to the resulting NAM fit as before.
Remark 3
The coefficient Ik can be computed by replacing the variance terms in Eq. (4) with their respective sample variances obtained from the post-hoc-orthogonalized ensemble average.
A schematic overview of the procedure is given in Algorithm 1. Pseudocode for the post-hoc orthogonalization procedure is presented in Algorithm 2 in the “Methods” section. Our method is implemented in Python and R; all source code and data are publicly available at https://github.com/Koehlibert/ONAM.
Algorithm 1
Schematic overview of the procedure
Input Prediction model F, n, R, Θ
1: Step 1 – Generate \({\mathcal{S}}={\{{F}_{i},{X}_{i1},\ldots ,{X}_{id}\}}_{i = 1,\ldots ,n}\)
2: for r in 1, …, R do
3: Step 2 – Fit NAM to give initial estimates \({{\bf{f}}}_{\theta }^{0,r}\)
4: Step 3 – Apply post-hoc orthogonalization
5: (Algorithm 2):
6: for m in 1, …, d − 1 do
7: o ← d − m + 1
8: Project current-order effects \({{\bf{f}}}_{\theta }^{m-1,r},| \theta | =o\),
9: onto lower-order effects \({{\bf{f}}}_{\theta }^{m-1,r},| \theta | < o\)
10: Update current-order effects \({{\bf{f}}}_{\theta }^{m,r},| \theta | =o\),
11: by vectors orthogonal to projections
12: Update lower-order effects \({{\bf{f}}}_{\theta }^{m,r},| \theta | < o\),
13: by adding projections to \({{\bf{f}}}_{\theta }^{m-1,r},| \theta | < o\)
14: Update higher-order effects \({{\bf{f}}}_{\theta }^{m,r},| \theta | > o\),
15: by \({{\bf{f}}}_{\theta }^{m-1,r},| \theta | > o\)
16: end for
17: end for
18: \({\bar{{\bf{f}}}}_{\theta }^{d-1}=\mathop{\sum }\nolimits_{i = 1}^{R}{{\bf{f}}}_{\theta }^{d-1,r}/R,\theta \in \Theta\)
19: Apply post-hoc orthogonalization (Algorithm 2)
20: to \({\bar{{\bf{f}}}}_{\theta }^{d-1},\theta \in \Theta\)
21: Update \({\bar{{\bf{f}}}}_{\theta }^{d-1},\theta \in \Theta\) by post-hoc-orthogonalized
22: estimates
Output\({\bar{{\bf{f}}}}_{\theta }^{d-1},\theta \in \Theta\)
Results
Illustration: predictive modeling of stream biological condition
To illustrate our methodology, we analyzed a set of environmental data collected by Maloney et al.4. The aim of this study was to analyze the condition of small, non-tidal streams (upstream area ≤200 km2) in the CBW on the mid-Atlantic coast of North America. For this purpose, the authors modeled the relationship between stream biological condition (outcome variable) and a set of 19 landscape measures (feature variables) using data from 4605 sites in the CBW (see Fig. 2). Details on the study design and the collection of samples have been provided in Section 2 of Maloney et al.4. Stream biological condition was assessed by the Chesapeake Basin-wide Index of Biotic Integrity (Chessie BIBI), which is a multi-metric index derived from stream benthic macroinvertebrate samples. The Chessie BIBI measures the biological quality of streams and wadeable rivers on a scale ranging from 0 to 10035. The list of features, which includes information on land use, climate, and natural watershed characteristics, is presented in SI Appendix S1.
Points are colored by Chesapeake Bay Index of Biotic Integrity (Chessie BIBI) score, with 0 indicating very poor biological condition and 100 indicating excellent biological condition. Inset shows the study area in relation to the United States. Basemap data from U.S. Census Bureau, 2022 (https://www.census.gov/geographies/mapping-files/timeseries/geo/tiger-line-file.html), accessed using the tigris package in R: Walker K (2024), tigris: Load Census TIGER/Line Shapefiles. R package version 2.1, https://CRAN.Rproject.org/package=tigris. Geographic coordinate system North American Datum of 1983. Watershed boundary from U.S. Geological Survey, 2022, National Hydrography Dataset (ver. USGS National Hydrography Dataset Best Resolution (NHD) for Hydrologic Unit (HU) 8 – 2022), https://prdtnm.s3.amazonaws.com/index.html?prefix=StagedProducts/Hydrography/NHD/HU8.
Deriving accurate predictions of the Chessie BIBI supports the restoration and conservation of streams in the watershed because this index is used by a key management group as a measure of stream health in meeting its goal to improve stream health of 10% of stream miles above a 2008 baseline36. Accurate predictions are particularly important for streams located at unsurveyed sites, for which the Chessie BIBI cannot be measured directly due to the sheer length of stream kilometers in the watershed (estimated to be over 220,000 km of streams with upstream areas ≤200 km2). At the same time, predictions of the Chessie BIBI need to be interpretable if they are intended to inform future management policies (e.g., projecting changes in land use, climate and watershed characteristics). Maloney et al.4 addressed these issues by fitting a random forest model to a training data set of size 3684 and by applying IML techniques (partial dependence and ALE plots, Friedman’s H-statistic, permutation importance, and Shapley values4) to the resulting black-box predictions. A descriptive summary of the training data and the random forest predictions is given in SI Appendix S2. Details on model fitting and the evaluation of prediction accuracy have been given in Sections 2.3 and 3, respectively, of Maloney et al.4.
Here, we investigate whether our decomposition method is able to yield plausible predictor-response relationships that are in line with Maloney et al.4. To this end, we applied our three-step algorithm to the aforementioned random forest model. The effects of interest (\(\Theta \subset {\mathcal{P}}(\Upsilon )\backslash {{\emptyset}}\)) included the 19 main effects and three two-way interactions (forest × development, forest × elevation, and development × elevation). We analyzed these interactions because the respective features were found to have the highest overall interaction strengths (Maloney et al.4, p. 7).
To reduce the impact of outliers on visualization, we excluded data points that exceeded at least one of the 99-th percentiles of the continuous variables. The reduced training data (n = 3114) were used for NAM fitting. The NAM ensemble consisted of 50 models with random weight initializations.
Figure 3 presents the main effects of the percentages of upstream catchment area as forest, developed, and barren, and the main effect of 30-year annual precipitation, along with the respective PDP and ALE plots adapted from Maloney et al.4. The contributions of the other features are presented in SI Appendix S3. The value of the summary measure I1 was 0.806, suggesting that 80.6% of the random forest prediction could be explained by the 19 main effects.
The first column of the figure presents the main effects of the percentages of upstream catchment area as forest (A1), developed (B1), and barren (C1), and the main effect of 30-year annual precipitation (D1) on the random forest prediction of the Chessie BIBI. The main effects were obtained by applying the proposed three-step algorithm to the training data of Maloney et al.4. Partial dependence plots (PDP) and accumulated local effect (ALE) plots are shown in the second (A2, B2, C2, D2) and third (A3, B3, C3, D3) columns, respectively. The values on the y-axes correspond to deviations of the predicted Chessie BIBI from the mean prediction. Note that PDP were mean-centered to ensure comparability with the main effects and ALE plots.
The main effects obtained from our method suggest that predicted Chessie BIBI scores tend to decrease with increasing development and barren land cover and increase with increasing forest cover and precipitation in upstream catchments, agreeing with the functional relationships in Maloney et al.4 and supporting relationships that have been consistently identified in previous studies37,38,39. Panel A1 of Fig. 3 suggests an almost linear positive trend with upstream catchment area as forest (reflecting lower anthropogenic disturbance), whereas the PDP and ALE plots (Panels A2 and A3) are characterized by (essentially) positive but nonlinear associations. The main effect of upstream catchment area as developed is similar to the respective PDP and ALE plots (Panels B1, B2, B3 of Fig. 3). For the effect of upstream catchment area as barren (Panels C1, C2 and C3), PDP and ALE plots both show nonlinear negative effects on the prediction of the Chessie BIBI. For this feature, the main effect obtained from our method has a different shape, suggesting a slight increase in the predicted Chessie BIBI at about 1.5% of upstream catchment area as barren. The effect of upstream total precipitation shown in panels D1, D2, and D3 of Fig. 3 suggests a positive association of this feature with the prediction of the Chessie BIBI. In contrast to the PDP and ALE plots, which reach a plateau at around 1300 mm, our method indicates an almost linear positive effect across the whole range of upstream total precipitation. Taken together, our method shows similar general patterns as the PDP and ALE plots in Maloney et al.4. However, there are also differences in the shapes of the curves, which could be the result of our method shifting predictive information to the main effects and/or due to extrapolation issues affecting the PDPs.
The two-way interaction between development land cover and site elevation is visualized in Fig. 4. It shows how the sum of the two main effects of development land cover and site elevation is altered by the addition of the two-way interaction term, indicating a less negative effect of development land cover on the prediction of biotic integrity at sites with a low elevation. This interaction was also reported in Maloney et al.4, who attributed it to elevation being a possible surrogate for stream slope, which has been shown to affect the development and stream biotic integrity relationship40; elevation gradient has also been shown to affect species distributional patterns41, including benthic macroinvertebrates42. Overall, we found a rather small interaction effect of development land cover and site elevation. The other two-way interactions are visualized in SI Appendix S3. The value of the summary measure I2 was 0.025, suggesting that another 2.5% of the random forest prediction could be explained by the three two-way interaction effects (in addition to the 80.6% contributed by the main effects).
The left panel (A) depicts the sum of the two main effects of development land cover and site elevation, visualizing, in particular, the negative effect of development land cover on the prediction of the Chessie BIBI. The right panel (B) was obtained by adding the two-way interaction between development land cover and site elevation to the sum of the two main effects. It suggests a less negative effect of development land cover on the prediction of biotic integrity at sites with a low elevation.
Analysis of data sets from other research domains
We further applied our method to data from the Salt River Pima-Maricopa Indian Community of the Salt River Reservation, Arizona, diabetes study and the Boston Housing study. For the diabetes data43, we considered a gradient boosting machine44 that yielded probability predictions for the binary outcome “diabetes” (yes/no). The decomposition of this model involved a logistic activation function in the output layer. For the Boston Housing data43, we considered an XGBoost model45 to predict housing prices (measured in USD 1000’s). The results of our analysis, which are presented in SI Appendix S4, demonstrate that the proposed algorithm also works well when combined with other ML methods than random forests. They also show that our algorithm yields plausible associations in other fields than ecology.
Experiments with synthetic data
In addition to analyzing real-world data, we investigated whether our method is able to extract the subfunctions fθ from a synthetic additive prediction function. To this end, we constructed predictions defined by
where X1, …, X10 followed a multivariate uniform distribution on [−3, 3]10. In our experiments, we considered three scenarios with different sets of functional forms for the main and two-way interaction effects (for details, see SI Appendix S5). In order to define the true decomposition that our method should recover, we orthogonalized these functions in a large data set of size n = 100,000 (see the attached code on GitHub). Using the obtained orthogonal functions, we generated 10 independent samples \({\{{F}_{i},{X}_{i1},\ldots ,{X}_{i10}\}}_{i = 1,\ldots ,n}\) of size n ∈ {2000, 5000} to which we applied our method. The feature values were generated by sampling data points \({\{{Z}_{i1},\ldots ,{Z}_{i10}\}}_{i = 1,\ldots ,n}\) from a multivariate normal distribution with zero mean, unit variance, and equicorrelation 0.5, and by applying the univariate standard normal cumulative distribution function Φ(⋅) to give Xij = 6 ⋅ (Φ(Zij) − 0.5), j = 1, …, 10. We used the ANN architecture described in the “Methods” section, setting the number of ensemble members to 10.
Figure 5 presents the estimated main effects obtained in the three scenarios with n = 2000. Despite some variation, which is likely due to differences in the empirical distribution functions of the features, and some tendency to oversmooth the effects in highly nonlinear regions (which could be addressed by increasing the complexity of the NAM architecture), our method performed well in approximating the true main effects. Similar results were obtained for the two-way interaction terms and in the scenarios with n = 5000 (SI Appendix S6). The average values of the summary measures I1 and I2 were 0.370, 0.918, 0.983, and 0.605, 0.079, 0.017, respectively, in the scenarios with n = 2000, and 0.389, 0.922, 0.984, and 0.586, 0.075, 0.015, respectively, in the scenarios with n = 5000.
The blue lines visualize the main effects f1(X1), f2(X2), f3(X3), as obtained by applying the proposed three-step algorithm to samples of size n = 2000 each. The black lines correspond to the true post-hoc-orthogonalized main effects defined in SI Appendix S5 (A1–A3: scenario 1, B1–B3: scenario 2, C1–C3: scenario 3).
Run-time of the algorithm
To analyze the run-time of the proposed algorithm, we extended the experimental setup by additional sets of features and interaction effects. The results of this analysis, which are presented in SI Appendix S7, demonstrate that the run-time of the algorithm is approximately linear in the size of Θ. More than 99% of the computational effort was due to the fitting of the NAMs in Step 2 of the procedure, which is based on established implementations in Python. Further details on the run-time of the algorithm are given in SI Appendix S7.
Discussion
In recent years, techniques to improve the interpretability of black-box models have become a key component of ML methodology. As part of this methodology, functional decomposition is considered a “core concept of ML interpretability”3.
In this paper, we provided support for a novel concept for the decomposition of black-box prediction functions into explainable feature effects. In line with earlier approaches by Hooker16, the idea of our method is to replace the original prediction function with a surrogate model consisting of simpler, “better interpretable” subfunctions. The latter allows for a graphical representation of the main feature contributions and their interactions, providing insights into the direction and strength of the effects.
Our concept of stacked orthogonality is designed to achieve purity of the subfunctions; it implies that predictive information explained by the main effects is not contained in the higher-order effects. At the same time, stacked orthogonality implies that lower-order functions (offering a high degree of interpretability) capture as much functional behavior as possible. Another contribution of this work is the development of a user-friendly algorithm to estimate the subfunctions from data. It is based on the fitting of a NAM, which allows the approximation of feature effects using ANN architectures, and an efficient post-hoc orthogonalization method to achieve stacked orthogonality. The proposed algorithm yielded plausible feature effects in our application examples. Furthermore, it was able to approximate the true underlying subfunctions in our numerical experiments.
A key requirement for establishing interpretability is that the (black-box) model F does not generate any non-admissible predictions (i.e., predictions “out of range”). For instance, in our application on stream biological condition, all predicted Chessie BIBI values were admissible in the sense that they were included in the support of Y (i.e., in the interval [0, 100], which was guaranteed by the design of the random forest model). Under this assumption, and given that our algorithm achieves perfect correlation between the black-box sample predictions F1, …, Fn and the decomposed NAM outputs, it is guaranteed that the latter are admissible as well. Importantly, since users of our method are essentially free to decide about the number and locations of the sample points, any sample point of interest could be included in F1, …, Fn (and thus “forced” to be admissible). Beyond that, it is, of course, possible that out-of-sample points Fnew, Xnew,1, …, Xnew,d (not contained in \({\mathcal{S}}={\{{F}_{i},{X}_{i1},\ldots ,{X}_{id}\}}_{i = 1,\ldots ,n}\) and thus not used for NAM fitting) may result in dissimilarities between the original black-box prediction Fnew and the fitted NAM value \({\hat{F}}_{{\rm{new}}}\) (obtained by feeding Xnew,1, …, Xnew,d into the trained NAM). In this case, a non-admissible value \({\hat{F}}_{{\rm{new}}}\) (e.g., a BIBI value larger than 100 or a probability larger than 1) may be obtained. Thus, if users want to be sure that the algorithm will always produce admissible outputs (regardless of whether the inputs are contained in the sample \({\mathcal{S}}\) or not), they should include an appropriate activation function in the output layer of the NAM, as discussed in the section on Estimation by NAM and post-hoc orthogonalization. For example, in SI Appendix S4 we used a logistic activation function to allow for out-of-sample probability decompositions contained in [0, 1]. Analogously, one may extend the NAM for the Chessie BIBI by an activation function of the form \(g({\eta }_{i})=100\cdot {(1+\exp (-{\eta }_{i}))}^{-1}\) (to ensure that out-of-sample decompositions are contained in [0, 100]).
Despite the aforementioned advantages, our method is not without limitations. First, NAM fitting (and thus estimation of the subfunctions) is limited to rather low-dimensional feature sets. It should be emphasized, however, that our method allows users to specify subsets of “effects of interest” and to shift all “uninteresting” effects to the highest-order interaction level. This strategy preserves the practicability of the proposed method even when the overall number of higher-order interactions is prohibitively large. It also contributes to preserving the interpretability of the decomposition as a whole. A second limitation is that our concept of stacked orthogonality is not primarily designed for quantifying the overall contributions of single features. Instead, our summary measures Ik quantify the contributions of the effect levels (e.g., all main effects or all interaction effects considered together), or more generally, the contributions of the aforementioned “effects of interest” to the overall black-box prediction. On the other hand, our method does not preclude users from calculating generalized Sobol sensitivity indices (as defined by Chastaing et al.19) to summarize the overall contributions of single features.
In addition to the graphical comparisons presented here, the proposed algorithm could be compared to other IML methods in a more quantitative way. To date, however, there is still a lack of consensus on how to best conduct such larger-scale benchmark experiments. This methodological gap, which has been acknowledged in several overview works3,46,47, is partly due to the fact that IML methods may involve very different objective functions and/or may focus on very different aspects of explainability/interpretability. Consequently, recent benchmark experiments have mainly dealt with specific subclasses of IML methods (like feature importance48, GAMs49, and counterfactual interpretability50), whereas a commonly accepted performance metric applying to more general classes of IML methods is lacking (cf. Kadir et al.51). Further research could increase understanding of how the absence of such a metric can hinder the conduct of larger-scale IML method comparisons.
The applications considered in this paper are mainly based on black-box predictions derived from tabular data. Since our algorithm is based on ANNs, it could, in principle, be applied to text or image data as well. Although the post-hoc orthogonalization step of our algorithm is largely independent of the structure of the feature data, we note that visualizations of text- or image-based feature effects may require more sophisticated techniques than the 2D plots presented in this work2,52. In addition to exploring other types of data structures, analyses could be conducted on how sensitive NAM fitting is to changes in the target population (i.e., to changes in the probability measure PX). Regarding the latter issue, we note that the sampling procedure in Step 1 of our algorithm can be adapted to match the desired distributional characteristics. In particular, the sampling procedure can be adapted to include additional samples from specific feature subspaces (thereby refining the estimates in these regions) or to estimate the subfunctions in “counterfactual” regions not contained in the training data.
Finally, we emphasize that our method is designed to explain the inner workings of a black-box model. It can not be used to evaluate the features’ ability to predict the outcome variable Y. This is a general aspect of post-hoc functional decomposition3,16,19 and can be deduced from the basic equation in Eq. (1). In fact, since the left-hand side of Eq. (1) is entirely dependent on the prediction function F but not on Y, the decomposition in Eq. (1), and thus also the NAM in Eq. (5), do not incorporate any information on how well Y can be predicted by F and its subfunctions fθ. Put differently, the subfunctions obtained from our method will only have a meaningful interpretation if the underlying black-box model is useful in predicting the outcome of interest.
Methods
Details on NAM fitting
As stated above, each function \({f}_{\theta }^{0}\) in Eq. (5) is represented by a separate ANN. This representation is generally not restricted to a specific network architecture but can be adapted to the learning task(s) as needed. For our numerical experiments, we used fully connected ANNs with five hidden layers each. The numbers of units were 256, 128, 64, 32, and 8, starting with the first hidden layer. Rectified linear unit activation functions were used in the first four hidden layers; a linear activation function was used in the last hidden layer with bθ = 8. The NAM was fitted using backpropagation with the mean squared error loss and the Adam optimizer (ref. 53, see the attached code on GitHub). The backpropagation procedure was run until convergence.
Details on post-hoc orthogonalization
In Step 3 of our method, we apply the following algorithm to process the initial intercept estimate \({\mu }^{0}={f}_{{{\emptyset}}}^{0}=0\) and the NAM estimates \({f}_{\theta }^{0},\theta \in {\mathcal{P}}(\Upsilon )\backslash {{\emptyset}}\). The superscript r has been omitted for ease of notation.
Input: Vectors of initial estimates \({{\bf{f}}}_{\theta }^{0}={{\bf{U}}}_{\theta }{{\bf{w}}}_{\theta }^{0}\in {{\mathbb{R}}}^{n},\theta \in {\mathcal{P}}(\Upsilon )\).
For m = 1 to d − 1:
-
11.
Define the actual interaction order by d − m + 1.
-
12.
Define the actual set of effects by \({\mathcal{A}}=\{\theta \in {\mathcal{P}}(\Upsilon ):| \theta | =d-m+1\}\). Let \({f}_{{\mathcal{A}}}^{m-1}={\{{{\bf{f}}}_{\theta }^{m-1}\}}_{\theta \in {\mathcal{A}}}\) be the set of function estimates of order d − m + 1.
-
13.
Define the set of lower-order effects by \({\mathcal{L}}=\{\theta \in {\mathcal{P}}(\Upsilon ):| \theta | < d-m+1\}\). Let \({f}_{{\mathcal{L}}}^{m-1}={\{{{\bf{f}}}_{\theta }^{m-1}\}}_{\theta \in {\mathcal{L}}}\) be the set of function estimates of order lower than d − m + 1.
-
14.
Define the set of higher-order effects by \({\mathcal{H}}=\{\theta \in {\mathcal{P}}(\Upsilon ):| \theta | > d-m+1\}\). Let \({f}_{{\mathcal{H}}}^{m-1}={\{{{\bf{f}}}_{\theta }^{m-1}\}}_{\theta \in {\mathcal{H}}}\) be the set of function estimates of order higher than d − m + 1.
-
15.
Define the matrix \({\bf{U}}={[{{\bf{U}}}_{\theta }]}_{\theta \in {\mathcal{L}}}\) by concatenating the output matrices \({{\bf{U}}}_{\theta },\theta \in {\mathcal{L}}\) (including the single-column matrix \({{\bf{U}}}_{{{\emptyset}}}={(1,\ldots ,1)}^{\top }\) for the intercept). By definition, U is of dimension n × B, where \(B={\sum }_{\theta \in {\mathcal{L}}}{b}_{\theta }\). We assume that the architectures of the ANN terms in Eq. (5) have been specified such that n ≥ B.
-
16.
Compute the matrix \({\bf{P}}={\bf{U}}{({{\bf{U}}}^{\top }{\bf{U}})}^{-1}{{\bf{U}}}^{\top }\) (assuming U is of full rank). By definition, multiplication of a vector \({\bf{x}}\in {{\mathbb{R}}}^{n}\) with P is equivalent to projecting x onto the column space spanned by U. In case U is not of full rank, we adapt the algorithm as described below.
-
17.
Compute the sum of the actual function estimates by \({{\bf{z}}}_{{\mathcal{A}}}^{m-1}={\sum }_{\theta \in {\mathcal{A}}}{{\bf{f}}}_{\theta }^{m-1}\).
-
18.
Update the actual effects \({f}_{{\mathcal{A}}}^{m}\) by projecting \({{\bf{z}}}_{{\mathcal{A}}}^{m-1}\) onto the column space of U and by setting \({f}_{{\mathcal{A}}}^{m}\) equal to the vectors that are orthogonal to this projection. This gives \({f}_{{\mathcal{A}}}^{m}={\{({\bf{I}}-{\bf{P}}){{\bf{f}}}_{\theta }^{m-1}\}}_{\theta \in {\mathcal{A}}}\), where I is the identity matrix of size n.
-
19.
Update the lower-order effects \({f}_{{\mathcal{L}}}^{m}\) by adding the projections of \({{\bf{z}}}_{{\mathcal{A}}}^{m-1}\) to \({f}_{{\mathcal{L}}}^{m-1}\). This gives \({f}_{{\mathcal{L}}}^{m}={\{{{\bf{f}}}_{\theta }^{m-1}+{{\bf{U}}}_{\theta }{[{({{\bf{U}}}^{\top }{\bf{U}})}^{-1}{{\bf{U}}}^{\top }{{\bf{z}}}_{{\mathcal{A}}}^{m-1}]}_{\theta }\}}_{\theta \in {\mathcal{L}}}\), where \({[{({{\bf{U}}}^{\top }{\bf{U}})}^{-1}{{\bf{U}}}^{\top }{{\bf{z}}}_{{\mathcal{A}}}^{m-1}]}_{\theta }\) is a vector of length bθ. It contains those elements of the vector \({({{\bf{U}}}^{\top }{\bf{U}})}^{-1}{{\bf{U}}}^{\top }{{\bf{z}}}_{{\mathcal{A}}}^{m-1}\) that match the positions of the columns of Uθ in U.
-
20.
The higher-order effects are not updated, i.e., \({f}_{{\mathcal{H}}}^{m}={f}_{{\mathcal{H}}}^{m-1}={\{{{\bf{f}}}_{\theta }^{m-1}\}}_{\theta \in {\mathcal{H}}}\).
Algorithm 2
Post-hoc orthogonalization
Input Initial estimates \({{\bf{f}}}_{\theta }^{0}={{\bf{U}}}_{\theta }{{\bf{w}}}_{\theta }^{0}\in {{\mathbb{R}}}^{n},\theta \in {\mathcal{P}}(\Upsilon )\)
1: for m in 1, …, d − 1 do
2: 1.1 o ← d − m + 1
3: 1.2 \({\mathcal{A}}\leftarrow \{\theta \in {\mathcal{P}}(\Upsilon ):| \theta | =o\},{f}_{{\mathcal{A}}}^{m-1}\leftarrow {\{{{\bf{f}}}_{\theta }^{m-1}\}}_{\theta \in {\mathcal{A}}}\)
4: 1.3 \({\mathcal{L}}\leftarrow \{\theta \in {\mathcal{P}}(\Upsilon ):| \theta | < o\},{f}_{{\mathcal{L}}}^{m-1}\leftarrow {\{{{\bf{f}}}_{\theta }^{m-1}\}}_{\theta \in {\mathcal{L}}}\)
5: 1.4 \({\mathcal{H}}\leftarrow \{\theta \in {\mathcal{P}}(\Upsilon ):| \theta | > o\},{f}_{{\mathcal{H}}}^{m-1}\leftarrow {\{{{\bf{f}}}_{\theta }^{m-1}\}}_{\theta \in {\mathcal{H}}}\)
6: 1.5 \({\bf{U}}\leftarrow {[{{\bf{U}}}_{\theta }]}_{\theta \in {\mathcal{L}}}\)
7: 2.1 Compute \({\bf{P}}={\bf{U}}{({{\bf{U}}}^{\top }{\bf{U}})}^{-1}{{\bf{U}}}^{\top }\)
8: 2.2 Compute \({{\bf{z}}}_{{\mathcal{A}}}^{m-1}={\sum }_{\theta \in {\mathcal{A}}}{{\bf{f}}}_{\theta }^{m-1}\)
9: 3.1 Update
10: \({f}_{{\mathcal{A}}}^{m}={\{({\bf{I}}-{\bf{P}}){{\bf{f}}}_{\theta }^{m-1}\}}_{\theta \in {\mathcal{A}}}\)
11: 3.2 Update
12: \({f}_{{\mathcal{L}}}^{m}={\{{{\bf{f}}}_{\theta }^{m-1}+{{\bf{U}}}_{\theta }{[{({{\bf{U}}}^{\top }{\bf{U}})}^{-1}{{\bf{U}}}^{\top }{{\bf{z}}}_{{\mathcal{A}}}^{m-1}]}_{\theta }\}}_{\theta \in {\mathcal{L}}}\)
13: 3.3 Update
14: \({f}_{{\mathcal{H}}}^{m}={f}_{{\mathcal{H}}}^{m-1}={\{{{\bf{f}}}_{\theta }^{m-1}\}}_{\theta \in {\mathcal{H}}}\)
15: end for
Update \({\{{{\bf{f}}}_{\theta }^{d-1}\}}_{\theta \in {\mathcal{P}}(\Upsilon )\backslash {{\emptyset}}}\) by mean-centered vectors
Output\({\{{{\bf{f}}}_{\theta }^{d-1}\}}_{\theta \in {\mathcal{P}}(\Upsilon )}\)
A schematic overview of the post-hoc orthogonalization procedure is given in Algorithm 2. The updates in Step 3.2 imply that each \({{\bf{f}}}_{\theta }^{m}\) can be written in the form \({{\bf{U}}}_{\theta }{{\mathbf{\beta }}}_{\theta }^{m}\), where \({{\mathbf{\beta }}}_{\theta }^{m}\) is a vector of coefficients of length bθ. Consequently, one obtains
where \({[{{\mathbf{\beta }}}_{\theta }^{m-1}]}_{\theta \in {\mathcal{L}}}\) denotes the concatenation of the coefficient vectors \({{\mathbf{\beta }}}_{\theta }^{m-1}\) (i.e., a vector of length B). According to Eq. (8), the sum of the lower-order effects is orthogonal to the sum of actual effects, and the final result of the algorithm satisfies the stacked orthogonality constraints in Eq. (3).
In the final step, we center the vectors \({{\bf{f}}}_{\theta }^{d-1},\theta \in {\mathcal{P}}(\Upsilon )\backslash {{\emptyset}}\), by subtracting their respective means. This ensures that all functions are centered around zero, as assumed in the Subsection “Conditions on the features and the prediction function”. Note that the centering does not affect the above orthogonality proof, as the actual effects \({{\bf{f}}}_{\theta }^{m},\theta \in {\mathcal{A}}\), are left unchanged in later iterations (implying \({{\bf{f}}}_{\theta }^{m}={{\bf{f}}}_{\theta }^{d-1}\) for these effects), and as the sum of the mean-centered actual effects is equal to the sum of the uncentered actual effects \({\sum }_{\theta \in {\mathcal{A}}}{{\bf{f}}}_{\theta }^{m}\) in the first line of Eq. (8). The latter result is due to the fact that the sum \({\sum }_{\theta \in {\mathcal{A}}}{{\bf{f}}}_{\theta }^{m}\) has zero mean, being orthogonal to \({{\bf{U}}}_{{{\emptyset}}}={(1,\ldots ,1)}^{\top }\). By the same argument, the centering does not affect the value of the intercept term.
In case U is not of full rank, we project \({{\bf{z}}}_{{\mathcal{A}}}^{m-1}\) onto a full-rank subspace of the column space of U. More specifically, we consider the pivoted QR decomposition
where \(\tilde{{\bf{Q}}}\in {{\mathbb{R}}}^{n\times n}\) is a unitary matrix, \(\tilde{{\bf{R}}}\in {{\mathbb{R}}}^{n\times B}\) is an upper triangular matrix with diagonal elements r11, …, rBB, and \(\tilde{{\bf{P}}}\in {{\mathbb{R}}}^{B\times B}\) is a permutation matrix arranging the columns of U such that ∣r11∣≥…≥∣rBB∣. Denoting the rank (i.e., the number of non-zero singular values) of U by rU, we define \(\tilde{{\bf{U}}}\in {{\mathbb{R}}}^{n\times {r}_{{\bf{U}}}}\) by those columns of U corresponding to first rU diagonal elements of \(\tilde{{\bf{R}}}\). The positions of these columns are indicated by the entries of the permutation matrix \(\tilde{{\bf{P}}}\). Accordingly, we define the matrices \({\tilde{{\bf{U}}}}_{\theta },\theta \in {\mathcal{L}}\), by those columns of Uθ contained in \(\tilde{{\bf{U}}}\), and we perform Steps 2 and 3 of the above algorithm with U and Uθ replaced by \({\tilde{{\bf{U}}}}_{\theta }\) and \({\tilde{{\bf{U}}}}_{\theta }\), respectively.
Data availability
Chessie BIBI data for the Predictive Modeling of Biological Stream Health are available at https://datahub.chesapeakebay.net/LivingResourcesand for predictors data are available at https://www.usgs.gov/apps/ecosheds/#/, https://doi.org/10.5066/P944W2WT, and https://doi.org/10.5066/P9EUVT4E. All code used for figure generation and analyzed data is furthermore available on github: https://github.com/Koehlibert/ONAM.
Code availability
An implementation of the algorithm, along with installation instructions, is available on GitHub. The repository also contains all code necessary to replicate simulation results.
References
Sarker, I. H. Machine learning: algorithms, real-world applications and research directions. SN Comput. Sci. 2, 160 (2021).
Alber, M. et al. iNNvestigate neural networks! J. Mach. Learn. Res. 20, 93 (2019).
Molnar, C. Interpretable Machine Learning–a Guide for Making Black Box Models Explainable, 2 edn (Independently published, 2022).
Maloney, K. O. et al. Explainable machine learning improves interpretability in the predictive modeling of biological stream conditions in the Chesapeake Bay Watershed, USA. J. Environ. Manag. 322, 116068 (2022).
Chen, V. et al. Applying interpretable machine learning in computational biology – pitfalls, recommendations and opportunities for new developments. Nat. Methods 21, 1454–1461 (2024).
Murdoch, W. J., Singh, C., Kumbier, K., Abbasi-Asl, R. & Yu, B. Definitions, methods, and applications in interpretable machine learning. Proc. Natl. Acad. Sci. USA 116, 22071–22080 (2019).
Miller, T. Explanation in artificial intelligence: insights from the social sciences. Artif. Intell. 267, 1–38 (2019).
Linardatos, P., Papastefanopoulos, V. & Kotsiantis, S. Explainable AI: a review of machine learning interpretability methods. Entropy 23, 18 (2021).
Tibshirani, R. Regression shrinkage and selection via the lasso. J. R. Stat. Soc. Ser. B 58, 267–288 (1996).
Turbé, H., Bjelogrlic, M., Lovis, C. & Mengaldo, G. Evaluation of post-hoc interpretability methods in time-series classification. Nat. Mach. Intell. 5, 250–260 (2023).
Da Veiga, S., Gamboa, F., Iooss, B. & Prieur, C. Basics and trends in sensitivity analysis: Theory and practice in R (Society for Industrial and Applied Mathematics, 2021).
Lepore, A., Palumbo, B. & Poggi, J.-M. (eds) Interpretability for Industry 4.0: Statistical and Machine Learning Approaches (Springer, 2022).
Welchowski, T., Maloney, K. O., Mitchell, R. & Schmid, M. Techniques to improve ecological interpretability of black-box machine learning models–case study on biological health of streams in the United States with gradient boosted trees. J. Agric. Biol. Environ. Stat. 27, 175–197 (2022).
Apley, D. W. & Zhu, J. Visualizing the effects of predictor variables in black box supervised learning models. J. R. Stat. Soc. Ser. B 82, 1059–1086 (2020).
Grömping, U. Model-agnostic effects plots for interpreting machine learning models. Tech. Rep., Reports in Mathematics, Physics and Chemistry: Department II, Beuth University of Applied Sciences Berlin, 1/2020.
Hooker, G. Generalized functional ANOVA diagnostics for high-dimensional functions of dependent variables. J. Comput. Graph. Stat. 16, 709–732 (2007).
Stone, C. J. The use of polynomial splines and their tensor products in multivariate function estimation. Ann. Stat. 22, 118–171 (1994).
Luenberger, D. G. Optimization by Vector Space Methods (Wiley, 1969).
Chastaing, G., Gamboa, F. & Prieur, C. Generalized Hoeffding-Sobol decomposition for dependent variables–application to sensitivity analysis. Electron. J. Stat. 6, 2420–2448 (2012).
Lengerich, B., Tan, S., Chang, C.-H., Hooker, G. & Caruana, R. Purifying interaction effects with the functional ANOVA: an efficient algorithm for recovering identifiable additive models. In: Chiappa, S. & Calandra, R. (eds) Proc. Twenty Third International Conference on Artificial Intelligence and Statistics, vol. 108, 2402–2412 (Proceedings of Machine Learning Research, 2020).
Agarwal, R. et al. Neural additive models: interpretable machine learning with neural nets. In: Ranzato, M., Beygelzimer, A., Dauphin, Y., Liang, P. S. & Wortman Vaughan, J. (eds) Proc. 35th Conference on Neural Information Processing Systems (NeurIPS 2021), vol. 34, 4699–4711 (Advances in Neural Information Processing Systems, 2021).
Cai, Y. Achieve the minimum width of neural networks for universal approximation. Preprint at: https://doi.org/10.48550/arXiv.2209.11395 (2023).
Cybenko, G. Approximation by superpositions of a sigmoidal function. Math. Control Signals Syst. 2, 303–314 (1989).
Hornik, K. Approximation capabilities of multilayer feedforward networks. Neural Netw. 4, 251–257 (1991).
Hornik, K., Stinchcombe, M. & White, H. Multilayer feedforward networks are universal approximators. Neural Netw. 2, 359–366 (1989).
Ismailov, V. A three layer neural network can represent any multivariate function. J. Math. Anal. Appl. 523, 127096 (2023).
Ismayilova, A. & Ismailov, V. On the Kolmogorov neural networks. Neural Netw. 176, 106333 (2024).
Kidger, P. & Lyons, T. Universal approximation with deep narrow networks. In Abernethy, J. & Agarwal, S. (eds) Proc. Thirty Third Conference on Learning Theory, vol. 125, 2306–2327 (Proceedings of Machine Learning Research, 2020).
Park, S., Yun, C., Lee, J. & Shin, J. Minimum width for universal approximation. In Proc. Ninth International Conference on Learning Representations. https://openreview.net/forum?id=O-XJwyoIF-k (OpenReview.net, 2021)
Shen, Z., Yang, H. & Zhang, S. Optimal approximation rate of ReLU networks in terms of width and depth. J. de. Math. ématiques Pures et. Appliquées 157, 101–135 (2022).
Wood, S. N. Generalized Additive Models–an Introduction with R, 2 edn (Chapman and Hall/CRC, 2017).
Van Rossum, G. & Drake, F. l. Jr. The Python language reference. Python Software Foundation: Wilmington, DE, USA. Version 3.15, accessed on Apr 26, 2025. https://docs.python.org/3/reference/index.html, (2025).
R Core Team, R: A Language and Environment for Statistical Computing Version 4.2.2, accessed on Dec 31, 2022. R Foundation for Statistical Computing (Vienna, Austria, 2022). https://www.R-project.org.
Rügamer, D. A new PHO-rmula for improved performance of semi-structured networks. In Krause, A. et al. (eds) Proc. 40th International Conference on Machine Learning, vol. 202, 29291–29305 (Proceedings of Machine Learning Research, 2023).
Smith, Z. M., Buchanan, C. & Nagel, A. Refinement of the Basin-wide Index of Biotic Integrity for non-tidal streams and wadeable rivers in the Chesapeake Bay Watershed. Interstate Commission on the Potomac River Basin Report 17-2 (2017).
Chesapeake Bay Program. 2015-2025 stream health management strategy–v.3 (2020). https://d18lev1ok5leia.cloudfront.net/chesapeakebay/documents/2015-2025-Stream-Health-Management-Strategy.pdf, accessed August 15, 2024.
Allan, J. D. Landscapes and riverscapes: the influence of land use on stream ecosystems. Annu. Rev. Ecol. Evol. Syst. 35, 257–284 (2004).
Carlisle, D. M., Falcone, J. & Meador, M. R. Predicting the biological condition of streams: use of geospatial indicators of natural and anthropogenic characteristics of watersheds. Environ. Monit. Assess. 151, 143–160 (2009).
Hill, R. A. et al. Predictive mapping of the biotic condition of conterminous U.S. rivers and streams. Ecol. Appl. 27, 2397–2415 (2017).
Snyder, C. D., Young, J. A., Villella, R. & Lemarié, D. P. Influences of upland and riparian land use patterns on stream biotic integrity. Landsc. Ecol. 18, 647–664 (2003).
Gaston, K. J. Global patterns in biodiversity. Nature 405, 220–227 (2000).
Namayandeh, A., Beresford, D. V., Somers, K. M. & Dillon, P. J. Difference in benthic invertebrate communities of headwater streams can be detected using a short elevation gradient. Int. Aquat. Res. 10, 153–164 (2018).
Leisch, F., Dimitriadou, E. & Hornik, K. mlbench: Machine Learning Benchmark Problems Accessed on Dec 30, 2024. R package version 2.1-6 (2024). https://cran.r-project.org/web/packages/mlbench.
Friedman, J. H. Greedy function approximation: a gradient boosting machine. Ann. Stat. 29, 1189–1232 (2001).
Chen, T. & Guestrin, C. XGBoost: a scalable tree boosting system. In Krishnapuram, B. et al. (eds) Proc. 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining (KDD ’16), 785–794 (Association for Computing Machinery, 2016).
Bodria, F. et al. Benchmarking and survey of explanation methods for black box models. Data Min. Knowl. Discov. 37, 1719–1778 (2023).
Carvalho, D. V., Pereira, E. M. & Cardoso, J. S. Machine learning interpretability: a survey on methods and metrics. Electronics 8, 832 (2019).
Hooker, S., Erhan, D., Kindermans, P.-J. & Kim, B. A benchmark for interpretability methods in deep neural networks. Preprint at: https://doi.org/10.48550/arXiv.1806.10758 (2019).
Kruschel, S. et al. Challenging the performance-interpretability trade-off: an evaluation of interpretable machine learning models. Business & Information Systems Engineering. In press, https://doi.org/10.1007/s12599-024-00922-2 (2025).
Kan, Z., Rezaei, S. & Liu, X. Benchmarking counterfactual interpretability in deep learning models for time series classification. Preprint at: https://doi.org/10.48550/arXiv.2408.12666 (2024).
Kadir, M. A., Mosavi, A. & Sonntag, D. Evaluation metrics for XAI: a review, taxonomy, and practical applications. In IEEE 27th International Conference on Intelligent Engineering Systems (INES), 000111–000124 (IEEE, 2023).
Zeiler, M. D. & Fergus, R. Visualizing and understanding convolutional networks. In Fleet, D., Pajdla, T., Schiele, B. & Tuytelaars, T. (eds.) Proc. 13th European Conference on Computer Vision (ECCV 2014), 8689 of Lecture Notes in Computer Science, 818–833 (Springer International Publishing, Cham, 2014).
Kingma, D. P. & Ba, J. Adam: A method for stochastic optimization. Tech. Rep. arXiv: https://doi.org/10.48550/arXiv.1412.6980 (2017).
Acknowledgements
We thank Giles Hooker for providing valuable advice on functional decomposition. We used a data set from Maloney et al.4 who merged data of the Chessie BIBI (available at https://datahub.chesapeakebay.net) with the 1:24,000 map scale Spatial Hydro-Ecological Decision System (SHEDS, https://www.usgs.gov/apps/ecosheds) predictor data set. What is analyzed here is a subset of both data sets; future users should refer back to these original sources for metadata and original files. We deeply thank all the data providers. Support for KOM and LJB was provided by the U.S. Geological Survey Ecosystems Mission Area and the U.S. Geological Survey Chesapeake Bay Activities. Any use of trade, firm, or product names (including use in the supplementary information to this article) does not imply endorsement by the U.S. Government. We thank Thomas Prince for proofreading the manuscript.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
D.K. and M.S. conceived the methodology. D.K. developed and implemented the algorithm. The manuscript was written by D.K. and M.S. with input from all authors. D.R. contributed to the theoretical development of the algorithm. K.O.M. and L.J.B. provided data and contributed to the empirical analyses. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Köhler, D., Rügamer, D., Boyle, L.J. et al. Achieving interpretable machine learning by functional decomposition of black-box models into explainable predictor effects. npj Artif. Intell. 1, 34 (2025). https://doi.org/10.1038/s44387-025-00033-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44387-025-00033-7