Abstract
Neuroinflammation is a key process associated with Alzheimer’s disease (AD). There is interest in developing New Approach Methodologies (NAMs) by using human in-vitro complex systems such as brain organoids, combined with machine learning and computational approaches, to reproducibly and robustly evaluate monoclonal antibodies and other therapeutic modalities on these human-derived systems. Herpesviruses such as herpes simplex virus 1 (HSV-1) had been shown to be associated with AD risk and molecular pathology. Building on top of previously reported work, we used herpes simplex virus 1 (HSV-1) infection in 2D dissociated cells from human cerebral organoids (dcOrgs) to recapitulate AD-associated molecular readouts, such as high co-abundance of intracellular beta amyloid (Aβ) and phosphorylated tau (pTau) with HSV-1. Secreted Aβ42/40 ratios in conditioned media were lower from HSV-1-infected dcOrgs, compared to mock dcOrgs. Differentially expressed transcripts from bulk and single-cell RNA sequence data in HSV-1-infected dcOrgs were enriched for AD-associated GWAS genes. Our high-throughput, quantitative framework represents a comprehensive approach to harness on the strengths of 2D dcOrgs for high-throughput applications such as therapeutic screens and can complement the 3D brain organoids and animal models for neuroinflammation in AD.
Similar content being viewed by others
Introduction
There is great interest in developing high-throughput combinatorial New Approach Methodology (NAM) through the integration of human donor-derived microphysiological systems, such as complex human brain organoids, combined with computational machine learning (ML) approaches, to evaluate monoclonal antibodies and therapeutics for rare and common human diseases such as Alzheimer’s disease (AD) and related dementias (ADRDs). These NAMs can complement findings from animal models to increase the success rates for monoclonal antibodies and therapeutic compounds in human diseases.
Neuroinflammation plays a central role in Alzheimer’s disease (AD) and several recent studies on human post-mortem brains have uncovered neuroinflammation-induced molecular signatures associated with AD using single-cell RNA sequencing (scRNA-seq)1,2,3,4. Human genetics genome-wide association studies (GWAS) have identified microglia as a key cell type that is involved in innate immune pathways associated with AD5,6,7. In parallel, previous studies reported that herpes simplex virus 1 (HSV-1) is found in adult post-mortem brain samples and is associated with amyloid plaques of AD patients8,9,10,11. Subsequently, additional studies reported associations between herpesviruses and AD, as well as associations between HSV-1 and a major genetic risk factor for AD, apolipoprotein E (APOE) ε4 allele12,13,14,15,16,17,18,19. Epidemiological studies on large-scale population cohorts provided further evidence through the discovery that anti-herpetic medication reduced dementia risk in patients with HSV-1 infections and that increased risk for AD is associated with herpes simplex encephalitis (HSE)20,21,22,23,24,25,26,27,28.
Risk factors for AD and dementia include recurrent viral infections, such as by HSV-1 and human cytomegalovirus (HCMV)8,10,26,29,30,31,32,33. The anti-viral drug acyclovir (ACV) is highly effective in reducing HSV-1 replication and reducing clinical symptoms34. As such, clinical trials using the oral anti-herpetic drug valacyclovir (VCV) as a potential treatment for AD-associated neuroinflammation are currently underway and VCV may prove to be a promising AD therapeutic35,36,37,38. In addition, immunity against another herpesvirus, varicella zoster virus (VZV), was associated with decreased risk of dementia39,40,41.
Several recent studies reported successful recapitulation of AD-associated phenotypes and mechanistic insights into neuroinflammation-induced AD molecular signatures, by using HSV-1-infected 3D human brain organoids or reactivation of HSV-1 in 3D brain organoids42,43. These organoids contain significant proportions of cell types that are more commonly found in human fetal brains than adult brains, such as neural progenitor cells44,45,46,47,48,49,50, and therefore, are imperfect models for neurodegenerative diseases. However, despite being imperfect, the use of human brain organoids had already led to novel, exciting molecular insights into AD neuropathology27,51,52,53,54. Researchers discovered that HSV-1 catalyzed beta amyloid (Aβ) aggregation and elevated tau phosphorylation (pTau) in human brain organoids, leading to aberrations associated with encephalitis and AD27,28,43,53,54,55,56,57,58,59.
The use of 2D dissociated cells from human cerebral organoids (that we termed as “dcOrgs”) may have some differences compared to the use of 3D cerebral organoids for viral infections or drug treatment, such as the uniform accessibility of cells to viruses or drug treatment in 2D dcOrgs, versus the accessibility of cells on the surface of 3D cerebral organoids to viruses or drug treatment28 and attenuated innate immune activation observed in 3D cerebral organoids compared to 2D dcOrgs57,60. The 2D dcOrgs comprise of a heterogeneous population of cell types, including neurons, glia cells, stem cells and progenitor cells, that may more faithfully recapitulate the heterogeneous cell type populations found in human brains, over the use of co-culture systems (such as astrocyte and neuron co-cultures).
Given these exciting prior discoveries, we evaluated the use of HSV-1 infection in 2D dcOrgs to recapitulate AD-associated neuroinflammatory molecular and multi-transcriptomic signatures, and to enable a high-throughput, rapid and quantitative NAM for neuroinflammation in AD. In combination with computational analytical methods development, we sought to identify on-target and off-target molecular and transcriptomic effects of the anti-herpetic drug acyclovir (ACV)61. We discovered that HSV-1-induced neuroinflammation in dcOrgs led to molecular and transcriptomic signatures associated with AD through complex cell autonomous effects, including increased intracellular co-abundance of aggregated Aβ and HSV-1 and differential expression of transcripts associated with AD in HSV-1-infected dcOrgs compared to mock dcOrgs. Our framework can be used to generate novel, exciting biological hypotheses as well as high-throughput therapeutic target screens.
Results
HSV-1 infection in 2D dissociated cells from cerebral organoids
An ideal in-vitro complex system for neuroinflammation in AD is comprised of heterogeneous cell types, including neurons and glia cells, such as astrocytes and microglia (Fig. 1a). The resident innate immune cells in the central nervous system (microglia) are differentiated from the mesodermal lineage in-vitro62,63. As such, it was widely thought that cOrgs did not contain microglia. However, earlier studies discovered and characterized CD11b+ functional microglia that differentiated innately within these spontaneously differentiated cerebral organoids64,65,66,67,68. We differentiated 3D cOrgs for 2-4 months, followed by dissociation and culturing of the 2D dcOrgs for another month. We evaluated the percentages of cell types in 6-month dcOrgs by using flow cytometry and found that 7.1% of dcOrgs were positive for P2RY12, which is a marker for microglia (Fig. S1), and our observed fraction was consistent with reported fractions found in human and mouse brains69.
a Schematic of using dcOrgs for high-throughput assays. b Microscope images (20X magnification) of the mock, infected, treated or UV-HSV-1 dcOrgs, as well as mock, HSV-1+ or treated hiPSCs. c PCA plot using the human transcripts from the samples, using RNA-seq, color-coded by batches. d PCA plot of the HSV-1 viral transcripts from the samples, using RNA-seq, color-coded by batches.
We conducted HSV-1 infections in dcOrgs and consistently observed that >50% of cells were GFP+ 23 h post-inoculation (Fig. 1b and Fig. S2). ACV treatment on HSV-1-infected dcOrgs (that we termed as “Treated dcOrgs”) resulted in effective rescue and few GFP+ cells were observed under the microscope. As a negative control experiment, we infected dcOrgs with HSV-1 that had been UV-inactivated (that we termed as “UV-HSV-1 dcOrgs”) and observed no detectable GFP+ cells or plaques (Fig. S3). In parallel, we used influenza A (IAV) infected dcOrgs as another negative control. HSV-1 is a neurotropic DNA virus and IAV is a respiratory RNA virus. Unlike HSV-1, influenza was not associated with increased risk of AD70.
High inter-sample correlations observed in dcOrgs and hiPSCs using bulk RNA sequencing
To further evaluate the system with regard to the variability across biological and technical replicates of dcOrgs, we conducted bulk RNA-seq on dcOrgs and human induced pluripotent stem cells (hiPSCs) under different conditions (Fig. 1c, Data S1). Each set of conditions was performed in triplicate, followed by total RNA extraction, ribosomal RNA depletion and sequencing. We observed high correlations between replicate mock dcOrgs or HSV-1-infected dcOrgs (Pearson’s \(r\)>0.99; Fig. S4, Data S1). The correlations between mock versus HSV-1-infected dcOrg pairs were lower (\(r\)=0.75–0.82), indicating that there were many more transcriptome-wide perturbations than expected from the baseline technical variability. The correlations between HSV-1-infected versus treated dcOrg pairs ranged from \(r\)=0.81–0.89, indicating that ACV treatment had partially rescued some of the transcriptomic perturbations due to HSV-1 infection.
The correlations for biological replicates were high (\(r\)≥0.91 for mock or treated dcOrgs and \(r\)≥0.73 for HSV-1-infected dcOrgs, Data S1). The correlations between mock versus HSV-1-infected or treated hiPSC pairs were higher (\(r\)=0.95–1), because fewer human transcripts were differentially expressed due to HSV-1 infection in hiPSCs compared to dcOrgs. Previously, we found that RNA-seq data on cerebral organoids differentiated from 25 donors71 had similarly high inter-individual correlations \({r}^{2}\) > 0.7, with a mean \({r}^{2}\) of 0.94, indicating that dcOrgs differentiated from many individual donors can be a highly reproducible platform for HSV-1-transduced neuroinflammation.
Investigation of the top three principal components showed that most of the variation (57.2%) in the human transcripts was captured by the first principal component (PC1), as shown in the principal components analysis (PCA) plot in Fig. 1c and Fig. S5. The dcOrgs were also distinctively separated by their conditions into well-defined clusters on PC1. dcOrgs with UV-inactivated HSV-1 (UV-HSV-1) were clustered close to mock dcOrgs on PC1, whereas treated dcOrgs formed their own cluster between mock and HSV-1-infected dcOrgs on PC1. All hiPSCs clustered together into a single group regardless of the condition (mock, HSV-1-infected or treated). A fraction (0.96–1.4%) of all transcripts were viral transcripts in infected hiPSCs, and 0.028–0.035% of all transcripts were viral transcripts in treated hiPSCs. There were lower viral transcript counts in the hiPSCs than dcOrgs, despite using a higher multiplicity of infection (MOI) for the hiPSCs versus dcOrgs (MOI of 4 and 2, respectively), consistent with reports that stem cells are resistant to viral infection, unlike differentiated cells72,73.
To elucidate the viral transcriptome-wide expression patterns of HSV-1, we performed PCA using detected HSV-1 transcripts and found that most of the variation (62.4%) was captured by PC1 (Fig. 1d). All the HSV-1-infected dcOrgs clustered together on PC1 and PC2. However, both sets of treated dcOrgs showed distinctive clusters, primarily in PC1 and PC2. We found that 0.15-0.17% of all transcripts were viral transcripts in the first set of treated dcOrgs (dcOrgs1), 2.4-3% of total transcripts were viral transcripts in the second set of treated dcOrgs (dcOrgs2), whereas 60-81% of all transcripts were viral transcripts in HSV-1-infected dcOrgs, similar to our observation from scRNA-seq data from HSV-1-infected dcOrgs. There were 14-20 times more HSV-1 transcripts in the second set of treated dcOrgs compared to the first set of treated dcOrgs, suggesting that ACV treatment in dcOrgs2 was not as effective as in dcOrgs1 on suppressing viral transcript expression.
High co-abundance correlations in intracellular Aβ and HSV-1 were predominantly driven by Aβ aggregates versus Aβ monomers
Several prior studies have shown that accumulation of intracellular Aβ (aggregated Aβ42 in particular) precedes the accumulation of extracellular amyloid plaques and may be an early event in the pathogenesis of AD74,75,76. In the mammalian brain, amyloid plaques are primarily produced by neurons and are also produced by other cell types such as astrocytes and oligodendrocytes77,78,79,80,81. We next evaluated the impact of viral infections in dcOrgs on intracellular AD-associated molecular readouts, such as the abundance of intracellular Aβ or pTau. We performed 3-channel flow cytometry to quantify intracellular protein abundance from 20,000-30,000 single cells for each condition, followed by machine learning based gating (Fig. 2a). A Zombie Violet dye was used to label dead cells, Alexa Fluor-647 or Allophycocyanin (APC) fluorophore-conjugated antibodies were used to detect the abundance of Aβ, pTau or cell-type specific markers, and the HSV-1 virus was tagged with GFP on VP26.
a Schematic of the flow cytometry experiments and computational analyses. Replicate experiments are biological replicate experiments that were conducted at different times. The results from the conditions (mock, infected, treated) within a replicate experiment were conducted at the same time on the same batch of dcOrgs and viral/ACV stocks to minimize technical differences across the conditions. b UMAP plots showing the normalized fluorophore intensities for Aβ aggregates using aducanumab (red), HSV-1 (green) and Zombie dye (blue) across mock, HSV-1-infected and ACV-treated samples. c UMAP plots showing the normalized fluorophore intensities for Aβ aggregates using solanezumab (red), HSV-1 (green) and Zombie dye (blue) across mock, HSV-1-infected and ACV-treated samples. d Pairwise correlation heatmaps using the intensities for HSV-1, Zombie dye and aducanumab (Aβ aggregates) or solanezumab (Aβ monomers) across mock, HSV-1-infected and ACV-treated samples, calculated by taking the differences in correlations between infected dcOrgs and mock dcOrgs, or treated dcOrgs and mock dcOrgs. e Boxen plots showing the aducanumab or solanezumab fluorophore intensities for GFP- cells (without HSV-1) versus GFP+ cells (with HSV-1) within the same HSV-1-infected sample, as well as GFP- cells versus GFP+ cells within the same ACV-treated sample.
We evaluated the protein abundance of intracellular Aβ species, as well as co-detection of Aβ species and HSV-1, by using monoclonal antibodies for Aβ. The aducanumab antibody, which is selective for aggregated Aβ, was the first AD treatment to have received approval from the U.S. Food and Drug Administration (FDA)82,83,84. In parallel, we used the solanezumab antibody that selectively binds to monomeric Aβ and had failed to slow cognitive decline in people with preclinical AD or mild dementia85,86.
Uniform Manifold Approximation and Projection (UMAP) plots and co-detection analyses on the normalized intensities from the flow cytometry results revealed that there were higher intracellular aggregated Aβ in HSV-1-infected dcOrgs compared to mock dcOrgs or ACV-treated dcOrgs (Fig. 2b-c). High co-detection of HSV-1 and intracellular Aβ42 abundance was found primarily observed in dead cells within dcOrgs (Figure S6), similar to our prior observations from HCMV-infected dcOrgs33. We found strong correlations in the co-detection of intracellular Aβ aggregates and HSV-1 using aducanumab (Δ\(r\)=0.77 and 0.78 for two replicates; Fig. 2d). There was a positive but weaker correlation in the co-abundance of intracellular Aβ monomers and HSV-1 using solanezumab (Δ\(r\)=0.65 and 0.64 across two replicates; Fig. 2d). ACV treatment on HSV-1-infected dcOrgs reduced the correlations in co-abundance between Aβ aggregates and monomers with HSV-1 (Δ\(r\)=0.26-0.35; Fig. 2d). These results indicated that HSV-1 abundance in dcOrgs is highly correlated with the intracellular abundance of aggregated Aβ in dead cells, and that the abundance of aggregated Aβ could be reduced by ACV treatment.
Direct comparisons in HSV-1- cells versus HSV-1+ cells within the same samples indicate cell intrinsic effects on increased abundance of Aβ aggregates and monomers
Because not all cells within the HSV-1-infected dcOrgs or ACV-treated dcOrgs were positive for HSV-1 abundance, we sought to understand if the increased abundance of Aβ aggregates and monomers were driven by cell intrinsic or extrinsic factors. If the increased abundance in these Aβ readouts were driven by cell intrinsic factors, we would expect to observe differences in the intensity distributions of Aβ from cells that were positive for HSV-1 protein abundance (HSV-1+ cells) versus cells that did not have HSV-1 protein abundance (HSV-1- cells) within the same dcOrg sample. Alternatively, if the increased abundance in the Aβ was driven by cell extrinsic factors, we would not expect to observe differences in the intensity distributions from HSV-1- cells versus HSV-1+ cells. We found that the aducanumab intensities in live HSV-1+ cells were significantly higher than the intensities in live HSV-1- cells, indicating that increased intracellular aggregated Aβ abundance was driven by cell intrinsic factors (Fig. 2d). Similar results were obtained for the solanezumab intensities (Fig. 2d).
Co-abundance of HSV-1 and aducanumab were higher in dead cells versus live cells in HSV-1-infected dcOrgs (Δ\(r\)=0.75 and 0.84 in dead cells versus Δ\(r\)=0.38 and 0.62 in live cells; Figure S6a-d). Similarly, co-abundance of HSV-1 and solanezumab were higher in dead cells versus live cells in HSV-1-infected dcOrgs (Δ\(r\)=0.81 and 0.79 in dead cells versus Δ\(r\)=0.63 and 0.67 in live cells; Figure S6e-h). We evaluated IAV-infected dcOrgs and observed that IAV and aducanumab were more highly co-detected in live cells in IAV-infected dcOrgs (Δ\(r\)=0.44 and 0.42; Fig. 3a-d, Figure S7a-d), versus dead cells in IAV-infected dcOrgs (Δ\(r\)=0.12 and 0.16). Similarly, co-abundance of IAV and solanezumab were higher in live cells in IAV-infected dcOrgs (Δ\(r\)=0.22 and 0.25; Figure S7e-h), compared to dead cells in IAV-infected dcOrgs (Δ\(r\)=-0.01 and -0.02).
a UMAP plots showing the normalized fluorophore intensities for Aβ aggregates using aducanumab (red), IAV (green) and Zombie dye (blue) across mock and IAV-infected samples. b UMAP plots showing the normalized fluorophore intensities for Aβ aggregates using solanezumab (red), IAV (green) and Zombie dye (blue) across mock and IAV-infected samples. A cell with a darker blue color had a higher intensity from the Zombie dye, compared to a cell with a lighter blue, white or black color; however, gating for live/dead cells was conducted by using the raw intensities and not by using the normalized intensities (that are represented in the UMAP plots). c Pairwise correlation heatmaps using the intensities for IAV, Zombie dye and aducanumab (Aβ aggregates) or solanezumab (Aβ monomers) across mock and IAV-infected samples. The change in correlations (Δcorrelations) were calculated by taking the differences in correlations between infected dcOrgs and mock dcOrgs. d Boxen plots showing the aducanumab or solanezumab fluorophore intensities for GFP- cells (without IAV) versus GFP+ cells (with HSV-1) within the same IAV-infected sample in live-gated cells.
These results supported a hypothesis that cell intrinsic factors may be contributing to HSV-1 infection on aducanumab intensities and support the observations that HSV-1 infection led to increased intracellular abundance of Aβ aggregates and monomers, but preferentially of aggregated Aβ.
Intracellular abundance of HSV-1 in dcOrgs is most strongly correlated with aggregated Aβ, total tau, pTau-181 and pTau-217
To further define the effects of HSV-1-induced neuroinflammation on Aβ aggregation and tau phosphorylation in dcOrgs, we compared the co-abundance of HSV-1 in dcOrgs with several antibodies targeting the precursor APP, Aβ, Aβ1-40, total tau, pTau-181, pTau-217 and pTau-202,205. In terms of the amyloid antibodies, we observed strong correlation in intensities of HSV-1 with Aβ using the 6E10 antibody (Δ\(r\)=0.62 and 0.51 across 2 replicates; Fig. 4a).
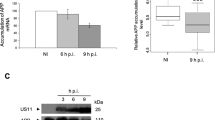
a Pairwise Δ\(r\) heatmaps using the intensities for HSV-1, Zombie dye and APP, Aβ, tau and different pTau residues across HSV-1-infected dcOrgs and ACV-treated dcOrgs. The Δ\(r\) values were calculated by taking the difference in correlations from infected dcOrgs and mock dcOrgs or treated dcOrgs and mock dcOrgs. b Top panel shows the concentrations of Aβ42/40/38 in pg/mL, Aβ42/40 ratios and Aβ42/38 ratios detected from conditioned media that had undergone heat inactivation for mock, HSV-1 + , or treated dcOrgs, and dcOrgs with UV-inactivated HSV-1 (UV-HSV-1). Bottom panel shows the concentrations of Aβ42/40/38 in pg/mL, Aβ42/40 ratios and Aβ42/38 ratios detected from conditioned media that had undergone heat inactivation for mock or IAV-infected (IAV + ) dcOrgs. P-values shown were calculated using 2-sided t-tests with comparison to mock dcOrgs.
Total tau and hyperphosphorylated tau are additional hallmark biomarkers of AD. Site-specific tau phosphorylation changes had been associated with clinical biomarkers87,88,89. pTau-Thr217 (pTau-217) and pTau-Thr181 (pTau-181) had been reported be closely linked to amyloid and tau pathology90; pTau-Thr205 (pTau-205) was associated with a decline in synaptic homeostasis and later stages of disease progression; whereas there were no significant changes in pTau-Ser202 (pTau-202) throughout disease progression87,88,89,91,92. We found strong correlations in intensities of HSV-1 with total tau using the HT7 antibody (Δ\(r\)=0.77 and 0.63; Fig. 4a), pTau-181 (Δ\(r\)=0.85 and 0.76), pTau-217 (Δ\(r\)=0.73 and 0.72), pTau Ser202 and Thr205 (AT8 antibody; Δ\(r\)=0.38 and 0.72).
Conditioned media from HSV-1-infected dcOrgs or HSV-1-infected and ACV-treated dcOrgs had consistently lower concentrations of secreted Aβ42, Aβ40 and Aβ38 compared to conditioned media from mock dcOrgs (2-tailed t-test P≤3.2×10-5; Fig. 4b, Data S2). There were significantly lower secreted Aβ42/40 or Aβ42/38 ratios from conditioned media with HSV-1-infected dcOrgs (2-tailed t-test P = 0.0012; Fig. 4b), that were primarily driven by lower levels of secreted Aβ42. ACV treatment restored secreted Aβ42/40 ratios to similar ratios as detected in conditioned media with mock dcOrgs but did not completely restore secreted Aβ42/38 ratios.
IAV infection in dcOrgs similarly led to decreased concentrations of secreted Aβ42, Aβ40 and Aβ38 detected in the conditioned media, compared to conditioned media with mock dcOrgs (2-tailed t-test P≤0.0098; Fig. 4b). However, secreted Aβ42/40 and Aβ42/38 ratios in conditioned media with IAV-infected dcOrgs did not differ significantly from ratios detected in conditioned media with mock dcOrgs (1-tailed Wilcoxon P = 0.72 and 0.095 respectively; Fig. 4b). We made similar observations using conditioned media that were UV-irradiated to inactivate residual viruses (Figure S8).
These results showed that HSV-1 infection in dcOrgs resulted in significantly lower ratios of secreted Aβ42/40, but IAV infection in dcOrgs did not affect the ratios of secreted Aβ42/40. In addition, ACV treatment of HSV-1-infected dcOrgs rescued the ratios of secreted Aβ42/40 to similar ratios as detected from mock dcOrgs. The results indicated that HSV-1 infection in dcOrgs led to a lower proportion of Aβ42 peptides that were secreted into the media, consistent with prior reports from AD patients’ CSF samples93,94.
Cell type enrichment analyses indicate a depletion of neurons and enrichment of glia in HSV-1-infected dcOrgs
Glia-neuronal morphological and functional changes that are important in AD include astrogliosis (marked by increased proportions of GFAP+ reactive astrocytes) and neuronal loss95,96. We sought to evaluate differences in the HSV-1-infected dcOrgs by performing flow cytometry with 15 cell-type markers to evaluate the enrichment or depletion of cell types. Using the cell counts from the flow cytometry results, we found that there were decreased proportions of NeuN+ neurons, VMAT2+ dopaminergic neurons, TuJ1+ immature neurons and EOMES+ intermediate progenitor cells in HSV-1-infected dcOrgs compared to mock dcOrgs, which indicated that there was likely to be neuronal loss in dcOrgs due to HSV-1 infection (Fig. 5, Figure S9-11). Live glia was mostly increased in proportions, such as GFAP+ astrocytes, GLAST+ astrocytes, Iba1+ microglia, P2RY12+ microglia, and O4+ oligodendrocytes. ACV treatment of HSV-1-infected dcOrgs reduced the depletion in proportions of neurons and neural progenitor cells and reduced the increased proportions of reactive glia cells (Fig. 5). CD4+ or CD45+ monocytes/macrophages did not show a huge increase in proportions among infected live cells and the proportions were similar after ACV treatment, which showed that the increased proportions of Iba1+ or P2RY12+ cells were likely to be driven by microglia and not monocytes nor macrophages. Our results were consistent with reactive astrogliosis, activation of microglia and neuronal death previously observed in AD patients4,97,98,99.
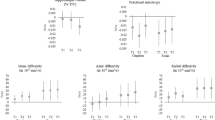
Odds ratios calculated for HSV-1-infected live dcOrgs versus mock live dcOrgs, or ACV-treated live dcOrgs versus mock live dcOrgs, using 15 cell-type markers. Multiple sets of replicate experiments were performed for each marker (depicted by the colored points). Shaded points represent significant odds ratios with P≤0.05 and open points represent non-significant odds ratios with P > 0.05. The tan bars represent average odds ratios<1 across both replicates and the blue bars represent average odds ratios>1 across both replicates.
Abundance of Iba1+ microglia and GFAP+ astrocytes were highly positively correlated with HSV-1
We next asked about the correlations in co-abundance of HSV-1 with cell-type marker abundance using our flow cytometry approach and conducted two sets of independent experiments for each marker (Figs. S12-14). We observed positive correlations in the co-abundance of HSV-1 and NeuN+ neurons (Δ\(r\)=0.26 and 0.24; Figure S12-14), as well as positive correlations in the co-abundance of HSV-1 and EOMES+ intermediate progenitor cells (Δ\(r\)=0.3 and 0.44). Correlation analyses of glia cell-type marker abundance with HSV-1 protein abundance pointed to Iba1+ microglia with the highest correlations (Δ\(r\)=0.66 and 0.39). There were negative correlations observed in the abundance of HSV-1 and P2RY12+ microglia (Δ\(r\)=-0.28). We also observed positive correlations in the abundance of GFAP+ astrocytes with HSV-1 (Δ\(r\)=0.32 and 0.49; Figure S13), and GLAST+ astrocytes with HSV-1 (Δ\(r\)=0.49 and 0.19). To ensure the reproducibility of the Δ\(r\) values, we conducted extensive replication experiments to establish the distributions of Δ\(r\) for the antibody markers used in our study and found that this approach can enable us to detect experimental failures that led to outlier Δ\(r\) values in an unbiased manner (Figure S15-19). These results showed that there were high correlations in the co-abundance of HSV-1 with neuronal and glia markers (NeuN, EOMES, Iba1, GFAP and GLAST), indicating that there were complex interactions between HSV-1 with neuronal cells, microglia and astrocytes to recapitulate the observed neuroinflammatory molecular and transcriptomic signatures.
Gene set enrichment analyses identified that differentially expressed transcripts due to HSV-1-infection in dcOrgs were enriched for AD-associated genes
After characterizing HSV-1-infected 2D dcOrgs and observing that many of the previously reported molecular signatures associated with AD that were observed in HSV-1-infected 3D brain organoids27,55,56,58, could be recapitulated in HSV-1-infected 2D dcOrgs, we further dissected the transcriptomic signatures of HSV-1-infected 2D dcOrgs. The identification of AD-associated transcriptomic readouts will enable us to use the massive RNA-seq and scRNA-seq datasets available from AD human post-mortem brains to identify if there may be subsets of individuals whose AD pathology may have arisen from chronic neuroinflammation due to HSV-1 reactivation, or more broadly, neurotropic pathogens. Identifying AD-associated transcriptomic readouts will also enable us to use emerging scRNA-seq technologies to identify cell types and cell type specific networks associated with neuroinflammation-induced AD molecular pathology such as aggregated Aβ, pTau, secreted Aβ42/40 ratios and cell-type proportional changes that we had observed.
We performed four sets of analyses to identify differentially expressed genes (DEGs) from HSV-1-infected dcOrgs versus mock dcOrgs (HSV-1-Inf-vs-Mock dcOrgs), HSV-1-infected dcOrgs versus HSV-1-infected and ACV-treated dcOrgs (HSV-1-Inf-vs-Treated dcOrgs), and HSV-1-infected dcOrgs versus dcOrgs with UV-inactivated HSV-1 (HSV-1-Inf-vs-UV-HSV-1 dcOrgs), shown in Data S3 and volcano plots in Figure S20. There were no viral transcripts detected in RNA-seq data from UV-HSV-1 dcOrgs, confirming the efficiency of viral inactivation (Data S4, Figure S3). The first two sets of experiments (HSV-1-Inf-vs-Mock dcOrgs and HSV-1-Inf-vs-Treated dcOrgs) had two sets of replication experiments (dcOrgs1 and dcOrgs2). In addition, we performed DEG analyses to compare HSV-1-infected hiPSCs from the same donor versus mock hiPSCs (Inf-vs-Mock hiPSCs) and HSV-1-infected hiPSCs versus HSV-1-infected and ACV-treated hiPSCs (Inf-vs-ACV hiPSCs).
Gene ontology analyses showed that biological processes such as regulation of apoptosis, autophagy and mitochondrion organization were enriched among DEGs in both sets of Inf-vs-Mock dcOrgs and Inf-vs-ACV dcOrgs1 but not in Inf-vs-ACV dcOrgs2 (Data S5). Pathways such as integrin signaling pathway, gonadotropin-releasing hormone receptor pathway and angiogenesis were enriched among DEGs in both sets of Inf-vs-Mock dcOrgs and Inf-vs-ACV dcOrgs1 but angiogenesis was not an enriched pathway in Inf-vs-ACV dcOrgs2 (Data S5).
Next, we obtained lists of genes that were associated to 21 common neurodegenerative, neuropsychiatric or autoimmune diseases, as well as human traits and related diseases identified from GWAS studies and reported in the GWAS Catalog100. To evaluate if the DEGs in infected versus mock dcOrgs were enriched for genes associated to any of the 21 common diseases or traits, we conducted gene set enrichment analyses (GSEA)101,102 with the gene lists in Data S6.
We observed that DEGs in both sets of HSV-1-infected versus mock dcOrgs (Inf-vs-Mock dcOrgs1 and Inf-vs-Mock dcOrgs2) were enriched for AD-associated genes (P = 0.039 and P = 5.1×10-3 respectively; Fig. 6a, Data S6). Similarly, the DEGs in HSV-1-infected dcOrgs versus dcOrgs with UV-inactivated HSV-1 (Inf-vs-UV dcOrgs2) were enriched for AD-associated genes (P = 0.015), providing further support that the transcriptomic profiles of dcOrgs with UV-inactivated HSV-1 were similar to mock dcOrgs. As control experiments, we evaluated whether the DEGs in HSV-1-infected versus mock hiPSCs might be enriched for AD-associated genes, but we did not observe any enrichment (P = 0.67; Fig. 6a). We conducted GSEA using a published list of DEGs identified from HSV-1-infected versus mock glioblastoma-derived (SH-SY5Y) neurons103, but we did not observe any enrichment for the neuronal DEGs with AD-associated genes (P = 0.38). We wondered whether inflammation induced by an RNA virus such as IAV, might similarly perturb AD-associated genes and therefore, we performed GSEA on DEGs identified from IAV-infected versus mock dcOrgs. However, we did not observe an enrichment for AD-associated genes (P = 0.79; Fig. 6a). Collectively, these results indicate that HSV-1-induced neuroinflammation in dcOrgs led to transcriptomic perturbations in AD-associated genes that may be driven by cell types other than neurons within the dcOrgs.
a Figure showing the GSEA results for HSV-1-infected cells or IAV-infected cells versus other conditions (mock, ACV or UV). -log10(Nominal P-value) from GSEA (y-axis) of the differentially expressed genes from our datasets with GWAS-associated genes for common diseases and traits (x-axis): Alzheimer’s disease (AD), amyotrophic lateral sclerosis (ALS), multiple sclerosis (MS), Parkinson’s disease (PD), Attention-deficit hyperactivity disorder (ADHD), autism spectrum disorders (ASD), bipolar disorder (BD), major depressive disorder (MDD), obsessive-compulsive disorder (OCD), schizophrenia (SCZ), Tourette syndrome (TS), Crohn’s disease (CD), inflammatory bowel disease (IBD), psoriasis (PSO), rheumatoid arthritis (RA), type 1 diabetes (T1D), type 2 diabetes (T2D), body mass index (BMI), brain volume (BRAIN), height (HEIGHT), waist-to-hip ratio (WHR). b Figure showing the GSEA results with a focus on the ACV-treated dcOrgs. -log10 (Nominal P-value) from GSEA (y-axis) of the differentially expressed genes from our datasets with GWAS-associated genes for common diseases and traits (x-axis). c Schematic figure of the scRNA-seq experiments where the cells from the mock samples were defined as “true uninfected” (TU) in tan colors, and the cells from the HSV-1-infected dcOrgs that did not contain many viral transcripts were defined as “abortively infected” (AI) in tan colors, and the cells from the HSV-1-infected dcOrgs that contained viral transcripts were defined as “true infected” (TI) in green colors. The TI and AI cells are collectively defined as “infected” (I). d -log10(Nominal P-value) from GSEA (y-axis) of the pseudobulk differential expression of various comparisons (TI versus TU, PU versus TU, I versus TU or TI versus PU). e Scatter plot for our discovery dataset showing the log2 fold change in infected versus ACV-treated dcOrgs, Inf-vs-ACV dcOrgs1 (y-axis) and infected versus mock dcOrgs, Inf-vs-Mock dcOrgs1 (x-axis), with the Pearson’s \(r\) correlations of the log2 fold changes of transcripts in each category shown on the bottom right. f Scatter plot for our replication dataset showing the log2 fold change in infected versus ACV-treated dcOrgs, Inf-vs-ACV dcOrgs2 (y-axis) and infected versus mock dcOrgs, Inf-vs-Mock dcOrgs2 (x-axis), with the Pearson’s correlations (\(r\)) of the log2 fold changes of transcripts in each category shown on the bottom right. g Venn diagram showing the numbers and percentages of genes that were rescued or not rescued in both our discovery and replication datasets (dcOrgs1 and dcOrgs2). h Venn diagram showing the numbers and percentages of genes that were in the exacerbated 1 or exacerbated 2 categories in both our discovery and replication datasets (dcOrgs1 and dcOrgs2).
HSV-1 and IAV infections in dcOrgs perturbed the expression of transcripts in several innate immune pathways
The expression of transcripts in several human immune pathways (cGAS/STING, IFN, IRF3, JAK/STAT, RIG-I and TLR) were significantly differentially perturbed in both sets of HSV-1-infected versus mock dcOrgs (Data S3-4, Figure S21a). ACV treatment rescued the expression for some of these transcripts (Figure S21b). As expected, HSV-1-infected dcOrgs versus dcOrgs with UV-inactivated HSV-1 showed similar differential expression as HSV-1-infected versus mock dcOrg samples (Figure S21c). IAV-infected versus mock dcOrgs did not result in much significant differential expression of these human immune transcripts, except for transcripts in the RIG-I and TLR pathways (Figure S21d). Similarly, the expressions of these transcripts were not significantly perturbed in hiPSCs (Figure S22). These results from HSV-1-infected dcOrgs are consistent with reports that the cGAS/STING pathway is a major pathway associated with aging-related inflammation in neurodegenerative diseases such as AD58,104.
A strong transcriptional activation of the innate immune response was observed in our RNA-seq datasets from HSV-1 infection in dcOrgs and reported previously from HSV-1 infection in 2D dcOrgs and 3D cerebral organoids28,57. This innate immune induction is most likely produced by microglia or astrocytes within the cerebral organoids, although a small fraction of neurons had been reported to produce type I interferon response105. Neurons had also been reported to be activated directly by microbes to release neuropeptides and modulate the innate immune response106,107. As such, the interferon induction observed in our data is likely to be primarily driven by glia cells and to a lesser extent, by neurons within these dcOrgs.
ACV treatment in HSV-1-infected dcOrgs rescued the expression in AD-associated genes
To determine the effect of ACV, we evaluated if ACV treatment of HSV-1-infected dcOrgs could prevent the transcriptomic perturbations in AD-associated genes that we had observed. GSEA analyses of our discovery dataset (Inf-vs-ACV dcOrgs1) showed an enrichment for AD-associated transcripts (P = 0.023; Fig. 6b), indicating that ACV treatment could rescue HSV-1-induced transcriptomic perturbations in AD-associated genes. However, our replication dataset (Inf-vs-ACV dcOrgs2) did not show an enrichment for AD-associated transcripts, indicating that ACV treatment in our replication experiment did not rescue HSV-1-induced transcriptomic perturbations in AD-associated genes (P = 0.53; Fig. 6b). As there were 14-20 times more HSV-1 viral transcripts detected in our replication dataset compared to our discovery dataset (Fig. 1d), we hypothesized that there might be different extents of inhibition of HSV-1 DNA synthesis in the two experiments.
Using a previously reported annotation of viral transcripts108, we observed that there was a high correlation in the P-value ranked differential expression for the leaky late viral transcripts (γ1) across both datasets (Spearman’s ρ = 0.63, P = 1.6×10-3), but not for the true late viral transcripts (γ2) across both datasets (Spearman’s ρ = 0.2, P = 0.47; Figure S23, Data S7). These results were consistent with prior reports that ACV treatment can perturb the expression of true late (γ2) viral genes, but not the expression of immediate early, early or leaky late (γ1) viral genes108,109. The results also indicated that the observed enrichment of AD-associated genes was due to dosage-dependent expression of γ2 viral genes, indicating that the expression of one or more γ2 genes led to differential expression in many of the AD-associated genes.
We performed GSEA using the DEGs from ACV-vs-Mock dcOrgs2 and observed an enrichment for AD-associated genes (P = 0.028; Fig. 6b), which showed that the transcriptomic profile of ACV-treated dcOrgs2 was similar to HSV-1-infected dcOrgs; thus, the results from ACV-vs-Mock dcOrgs2 were similar to Inf-vs-Mock dcOrgs2 and showed a similar enrichment for AD-associated genes. These results indicated that ACV treatment can rescue transcriptomic perturbations in AD-associated DEGs due to HSV-1 infection by a mechanism that is dependent on inhibition of viral DNA synthesis. The results are different from our prior results that ACV treatment cannot rescue transcriptomic perturbations in type 1 diabetes (T1D)-associated DEGs due to HSV-1 infection in stem cell derived pancreatic islets (sc-islets)110.
Cell type enrichment from bulk RNA-seq data aligns with results from flow cytometry
We ran our previously published Orgo-Seq method71 to predict for cell type enrichment or depletion by using our bulk RNA-seq datasets and a previously published scRNA-seq reference dataset111. We observed that astrocytes were increased in proportions across all our datasets, whereas excitatory neurons and inhibitory neurons were decreased in proportions across most of our datasets (Figure S24), which were consistent with our results from the flow cytometry experiments. However, microglia, oligodendrocytes and oligodendrocyte progenitor cells were predicted to be depleted across most of our RNA-seq datasets. To validate the gene expression for some of the markers, we conducted quantitative real-time PCR (qRT-PCR) and found that the expression of astrocytes (GFAP) and oligodendrocytes (SOX10) were increased in HSV-1-infected dcOrgs versus mock dcOrgs, whereas the expression of neural progenitor cells (Nestin) and neurons (RBFOX3 and MAP2) were decreased (Figure S25).
Single-cell RNA-seq analyses indicated that AD-associated DEGs from HSV-1-infected dcOrgs were driven by cells that were exposed to HSV-1 but did not contain viral transcripts
To identify transcriptomic changes at individual single cells, we generated scRNA-seq data using Parse Evercode WT Mini V2 with 2D HSV-1-infected versus mock dcOrgs (Figs. S26-29). We conducted analyses using pseudobulk data from subsets of the cells from our scRNA-seq data. Individual cells within the mock dcOrgs were defined as “true uninfected” (TU) cells. Individual cells in HSV-1-infected dcOrgs but did not contain many viral transcripts were collectively defined as “pseudo-uninfected” (PU) cells that may be comprised of cells that were uninfected, or “abortively infected” (AI) cells (Fig. 6c). Individual cells in HSV-1-infected dcOrgs that contained viral transcripts were defined as “true infected” (TI) cells. We could also combine the PU and TI cells to collectively analyze these cells as “infected” (I) cells.
There were multiple cell types that had preferentially higher levels of viral transcripts (in the TI cells compared to the TU cells), as shown in Data S9, such as cortical neurons, glia progenitor cells, interneurons, mesoderm-derived cells, oligodendrocytes, BMP-responsible cells, cilia-bearing cells, proteoglycan-expressing cells and unfolded protein responsible cells (mapped to the Tanaka reference dataset). By mapping the cell types to the Allen reference dataset, there were similar cell types that had preferentially higher levels of viral transcripts in the TI cells compared to the TU cells, such as microglia-PVM, endothelial cells, oligodendrocytes, oligodendrocyte progenitor cells (OPCs), and neuronal subtypes such as chandelier, L2/3 IT, L4 IT, L5 ET, L5 IT, L6 CT, L6 IT, L6 IT Car3, L6b, Lamp5, Lamp5 Lhx6, Pvalb, Sncg, Sst, Sst Chodl and VIP neurons.
Unlike the GSEA results from the bulk RNA-seq data, we found that there was no enrichment for AD-associated genes in the I/TU analyses from the 2D scRNA-seq data (P = 0.14; Fig. 6d; Data S6). Instead, we did observe an enrichment for AD-associated genes in the PU/TU analyses (P = 3.7×10-3). These PU cells did not harbor high abundance of viral transcripts but had been exposed to other cells that had high abundance of viral transcripts (TI cells). These PU cells may be comprised of abortively infected (AI) cells and prior literature reported that astrocytes were the main AI cells that produce interferon beta in HSV-1-infected mammalian brains112,113,114. Using the scRNA-seq data, we identified changes in cell type proportions among the TU, TI, PU and I cells and found that there were multiple cell types that increased or decreased in proportions (Data S9). However, these results remain to be replicated with additional scRNA-seq experiments.
ACV treatment exacerbated the expression for 19-23% of human transcripts that were expressed in dcOrgs
To explore the contribution of unexpected transcriptomic perturbations due to ACV treatment, we defined two categories of unexpectedly perturbed transcripts. First, we defined “Exacerbated 1” group as genes whose dysregulation were exacerbated by ACV treatment. For instance, if HSV-1 infection in dcOrgs down-regulated the human gene expression compared to mock dcOrgs, but ACV treatment on HSV-1-infected dcOrgs further down-regulated the gene expression compared to HSV-1-infected dcOrgs, or the reciprocal scenario with up-regulation, the gene would be classified into the Exacerbated 1 group. Next, we defined “Exacerbated 2” group as genes where HSV-1 infection in dcOrgs did not perturb the human gene expression but ACV treatment in HSV-1-infected dcOrgs significantly perturbed the expression of the gene.
Collectively, both groups of exacerbated genes comprised of 19% or 23% among all genes that were expressed in dcOrgs (Fig. 6e-f). We observed that exacerbated genes comprised of similarly high percentages of 22% or 23% among AD-associated GWAS genes (Data S8). On the other hand, the expression for most of the genes (22% or 40%) were rescued by ACV treatment, and similarly 26% or 41% of AD-associated genes were rescued by ACV treatment. These results reaffirmed that ACV treatment can rescue most AD-associated transcriptomic perturbations due to HSV-1 infection, despite resulting in high percentages of exacerbated gene expression.
ACV treatment rescued AD-associated gene expression in dcOrgs
We further explored the overlaps between transcripts that were rescued or not rescued across the dcOrg replicates. The highest overlap was among transcripts that were rescued in dcOrgs1 but were not rescued in dcOrgs2 (Fig. 6g), which further supported our observations that ACV treatment was more effective in the discovery experiment (Inf-vs-ACV dcOrgs1) than the replication experiment (Inf-vs-ACV dcOrgs2). As expected, the second highest overlap in transcripts were rescued in both dcOrgs1 and dcOrgs2, demonstrating that despite the dosage-dependent differences of ACV treatment in both sets of experiments, there were several transcripts in common that were rescued across both datasets.
The highest overlap in AD-associated transcripts was driven by transcripts that were rescued in both dcOrgs1 and dcOrgs2 (Figure S30). There was an enrichment in the ratios of AD-associated transcripts that were rescued by ACV treatment, compared to the overall ratios of all transcripts that were rescued by ACV treatment (OR = 1.33, P = 0.062), indicating that ACV treatment may have a preferential rescue for AD-associated genes in HSV-1-infected dcOrgs. These results are consistent with our prior results that ACV treatment did not rescue the expression of T1D-associated genes in HSV-1-infected sc-islets110.
Similarly, there were modest enrichment of RA-associated or T1D-associated transcripts that were rescued in both dcOrgs1 and dcOrgs2, compared to the overall set of all transcripts (OR = 1.33 and 1.47, P = 0.068 and 0.074 respectively, Figure S30). There were 7 other diseases or traits with modest enrichments of transcripts that were rescued in both dcOrgs1 and dcOrgs2, such as Parkinson’s disease (PD; OR = 1.55, P = 0.032), ADHD (OR = 1.24, P = 0.071), obsessive compulsive disorder (OCD; OR = 1.96, P = 0.0043), schizophrenia (SCZ; OR = 1.21, P = 0.069), inflammatory bowel disease (IBD; OR = 1.29, P = 0.088), height (OR = 1.14, P = 0.03) and waist-hip-ratio (WHR; OR = 1.34, P = 7.3×10-4). These results indicate that ACV treatment may rescue the expression of AD-associated genes and genes associated with several other common diseases or traits.
On the other hand, the 4-way Venn diagrams depicting both groups of exacerbated genes did not show high overlaps in any subsets that were common between both dcOrg replicates (Fig. 6h, Figure S31). This observation indicated that different sets of genes were transcriptionally exacerbated by each ACV treatment, unlike the rescued genes.
Analyses on human post-mortem brain RNA-seq revealed that 25-31% of patients with late-onset AD have transcriptomic signatures similar to HSV-1 infected dcOrgs
A previous study used RNA-seq data generated using human post-mortem brain samples from patients with late-onset AD (LOAD) and performed molecular subtyping of the LOAD patients into 5 subtypes (A, B1, B2, C1 and C2)115. Globally, there were weaker fold change differences observed in RNA-seq data from human post-mortem brains versus dcOrgs (Figure S32a). For better visualization, we rescaled the fold change differences observed in human post-mortem brains to illustrate that there were some pathways with high degrees of similarities in fold changes between the human post-mortem brains and dcOrgs, such as Amyloid, Blalock, Immune, Synapse/Myelin and Tau (Figure S32b).
We conducted a modified gene set enrichment analysis test using the DEGs from the dcOrgs with the most informative DEGs from the post-mortem brains and identified a significant enrichment of the DEGs in dcOrgs with subtype A (Figure S32c). Inf-vs-Mock dcOrgs1, Inf-vs-Mock dcOrgs2, Inf-vs-ACV dcOrgs1 and ACV-vs-Mock dcOrgs2 had significant positive associations with subtype A (FWER = 1×10-8, 9.6×10-9, 6.4×10-10 and 4×10-7 respectively). Subtype A comprises of 25% and 31% of the LOAD patients from two cohorts115, indicating that the transcriptomic signatures from HSV-1-infected dcOrgs can be used to model transcriptomics from a subset of AD human post-mortem brains.
Replication studies by using additional donor-derived dcOrgs showed high co-abundance of HSV-1 and aducanumab, solanezumab and pTau-181, and transcriptomic enrichment for AD-associated genes
We sought to replicate our results by using dcOrgs that were differentiated from another set of donors. We had reprogrammed 9 donor-derived hiPSC lines from lymphoblastoid cell lines with homozygous APOE ε3/ε3 alleles by using our previously established protocols116,117. We pooled hiPSCs in equal proportions from the 9 donors (and we termed the pool as A3P) and extracted DNA from the A3P hiPSC pool across 4 weeks for sequencing116,118. We used our PoolSeq algorithm to deconvolute donor proportions from the sequencing data118 and observed that at least 5 out of the 9 donors had proportions above the baseline after 4 weeks (Figure S33). We differentiated the A3P hiPSCs into dcOrgs and used 8-month dcOrgs to conduct replication experiments.
We found that there is a strong correlation in co-abundance of HSV-1 and aducanumab in HSV-1-infected A3P dcOrgs, compared to mock dcOrgs (Δ\(r\)=0.73, Figure S34), similar to our prior results from the PGP1 dcOrgs. ACV-treatment reduced the correlation in co-abundance of HSV-1 and aducanumab in treated A3P dcOrgs (Δ\(r\)=0.38). There was a weaker correlation in co-abundance of HSV-1 and solanezumab in HSV-1-infected A3P dcOrgs (Δ\(r\)=0.59). We conducted HSV-1 infection and ACV-treatment on HSV-1-infected A3P dcOrgs for RNA-seq and found that the DEGs in HSV-1-infected A3P dcOrgs versus mock A3P dcOrgs, and HSV-1-infected A3P dcOrgs versus ACV-treated A3P dcOrgs, were enriched for AD-associated genes (Figure S35). These results indicate that the AD-associated Aβ molecular and transcriptomic readouts induced by HSV-1 infection in dcOrgs are likely to replicate across donor lines.
In summary, our HSV-1-induced neuroinflammatory 2D dcOrgs system can recapitulate AD-associated molecular and transcriptomic signatures that can be used as a novel human NAM for neuroinflammation associated with AD. Collectively, the HSV-1-induced neuroinflammatory 2D dcOrgs can complement ongoing research development in the use of 3D cOrgs and animal models in recapitulating certain molecular and transcriptomic signatures associated with subsets of a common, complex disease such as AD.
Discussion
Neuroinflammation is a central mechanism in AD and related dementias, and there is great interest to understand complex cell type interactions in neuroinflammation associated with AD119,120. Herpesvirus infections in complex human in-vitro 3D brain organoids has been reported to recapitulate neuroinflammation-induced AD-associated readouts27,42,53,54,58 and stress factors such as repetitive injury were reported to induce reactivation of HSV-1, leading to AD-associated molecular phenotypes42,43.
In our work, we systematically evaluated the use of HSV-1-induced neuroinflammation in dcOrgs to model molecular and transcriptomic signatures associated with AD, by using high-throughput multi-omics technologies. We observed several key features associated with AD pathology that support the strengths of the 2D HSV-1-infected dcOrgs as a high-throughput, renewable human in-vitro neuroinflammatory NAM for AD, such as, the rapid induction of AD-associated molecular and transcriptomic readouts. Furthermore, dcOrgs provide a renewable source of heterogeneous cell types and can be passaged as 2D cell cultures to obtain large quantities of cells for high-throughput genetic screens or for biobanking of large numbers of donor-derived dcOrgs. Another strength of dcOrgs is that we can model the molecular, cellular and transcriptomic effects of extrinsic factors (pathogens, toxicants, drug treatment) on inherited genetic factors (G×E), to complement results from large-scale genetics studies, human post-mortem brains and animal models. We had conducted extensive quality control and replication methods to identify outlier data that can ensure the robustness of the conclusions drawn using the neuroinflammatory NAM.
We found that HSV-1-infected dcOrgs led to several AD-associated phenotypes, including increased intracellular abundance of Aβ and pTau, decreased ratios of secreted extracellular Aβ42/40, neuronal cell death and increased proportions of astrocytes and microglia, which are hallmarks of AD pathology59,96,121. The intracellular abundance of HSV-1 protein in dcOrgs is strongly correlated with the abundance of aggregated Aβ, pTau-181 and pTau-217, which were reported to be most strongly associated with AD clinical disease markers87. On the other hand, IAV-infected dcOrgs did not exhibit any of these AD-associated phenotypic readouts.
We explored bulk and single-cell transcriptomics of HSV-1-infected dcOrgs and found that the DEGs from HSV-1-infected versus mock dcOrgs were enriched for AD-associated genes identified from GWAS. The observed enrichment was specific for AD-associated genes was driven by PU cells that did not contain much viral transcripts but were exposed to HSV-1-infected cells.
It was previously reported that Aβ and pTau have antimicrobial activities27,122. As such, the cell type changes that we had observed in HSV-1-infected dcOrgs (such as reduced proportions of neurons) are likely to be downstream of Aβ and pTau. Furthermore, we found that immediate ACV treatment on HSV-1-infected dcOrgs led to lower intracellular abundance of Aβ and pTau and rescued the changes in proportions of cell types in treated dcOrgs. However, when there was reduced ACV activity (leading to increased expression of true late HSV-1 viral transcripts), we found that the differential expression of AD-associated GWAS genes in HSV-1-infected and ACV-treated dcOrgs, were not rescued, indicating that some of the transcriptomic changes that we had found were likely to be upstream of Aβ and pTau. A complex system such as dcOrgs can enable us to dissect complex, co-occurring mechanisms associated with AD.
There are exciting avenues to expand the work, such as identifying human-viral transcript interactions that contribute to the AD molecular or transcriptomic signatures observed or by using mutant replication-incompetent HSV-1 strains to dissect cell autonomous effects on AD transcriptomics. It will also be interesting to expand on the system to develop a latency/reactivation model, as well as to dissect cell type interactions such as by incorporating induced microglia, to better model the mechanisms underlying chronic neuroinflammation in AD. It will be exciting to explore additional avenues such as comparisons with other neurotropic viruses associated with AD. There are limitations to our system, such as the absence of a blood-brain barrier in dcOrgs and the short acute infection approach, that poses additional exciting avenues for future research.
There are exciting avenues to expand the work, such as identifying human-viral transcript interactions that contribute to the AD molecular or transcriptomic signatures observed or by using mutant replication-incompetent HSV-1 strains to dissect cell autonomous effects on AD transcriptomics. It will also be interesting to expand on the system to develop a latency/reactivation model, as well as to dissect cell type interactions such as by incorporating induced microglia, to better model the mechanisms underlying chronic neuroinflammation in AD. It will be exciting to explore additional avenues such as comparisons with other neurotropic viruses associated with AD. There are limitations to our system, such as the absence of a blood-brain barrier in dcOrgs and the short acute infection approach, that poses additional exciting avenues for future research.
We demonstrate that the integration of multi-omics data, including multi-transcriptomics data, can be used to dissect complex cell type interactions due to HSV-1-induced neuroinflammation in dcOrgs and that our platform can be used for high-throughput assays such as therapeutic compound screens. Collectively, the 2D dcOrgs can complement 3D human brain organoids and animal models to be used for therapeutic target identification and developing novel biological hypotheses.
Methods
Standard Protocol Approval
Research performed on samples and data of human and viral origin was conducted according to protocols approved by the Institutional Review Board (IRB) and Institutional Biosafety Committee (IBC) of UMass Chan Medical School. HSV-1 and IAV are Biosafety Level (BSL) 2 pathogens. The research was approved by the UMass Chan Medical School IRB (#H00021419) and the IRB determined that the activity is not human subject research as defined by DHHS and FDA regulations.
Source of hiPSCs and maintenance of hiPSCs
The control donor hiPSC line (with APOE ε3/ε3 alleles) used in our study (PGP1 hiPSC) was obtained from Professor George Church’s lab, and we had previously characterized and sequenced the line47,117,118,123. The donor (PGP1) was enrolled in the Harvard Personal Genome Project, PGP (https://my.pgp-hms.org/) which is an open-access, open-consent project established since 2005. All participants have consented to open-access sharing of their data (genotype, phenotype and cell lines). The donor samples for the A3P samples were hiPSCs reprogrammed from donor-derived lymphoblastoid cell lines that were purchased from Coriell Institute, by using our previously described methods117,118 and are available upon request from our groups. We performed flow cytometry (MACSquant VYB) to confirm that >90% of the PGP1 hiPSCs were positive for TRA-1-60 (Novus Biologicals NB100-730F488). hiPSCs were plated using StemFlex media (Thermo Scientific A3349401) with 10 μM Y-27632 (Abcam ab120129) on 6-well plates coated with 0.5 mg of Matrigel Basement Membrane Matrix (Corning 354234) per plate and subsequently dissociated using accutase (BioLegend 423201).
HSV-1 and ACV
The HSV-1 K26GFP KOS strain124,125 used in our study encodes a GFP-fused VP26 gene and was kindly provided by Prashant Desai126. We prepared 200 mM stock solutions of ACV (Sigma PHR1254) by dissolving in UltraPure DNase/RNase-Free Distilled Water (Invitrogen 10977015) with equal molar concentrations of sodium hydroxide. Aliquots were made and kept frozen in −80 °C.
HSV-1 plaque assays
UV-treated and untreated HSV-1 K26GFP stocks were serially diluted in DMEV (high glucose DMEM supplemented with 1% FBS and Pen/Strep). 500 μl of six 10-fold dilutions of virus were added to Vero cell monolayers in a 6-well dish and then incubated at 37 °C with shaking. After 1 h, virus-containing media was removed and replaced with DMEV supplemented with 10 μg/ml pooled human IgG (Sigma catalog #I4506). Seventy-two hours post-infection, Vero cells were fixed with ice-cold methanol and stained with crystal violet. The plates were washed with water, dried overnight, and plaques were counted to determine viral titer.
Influenza A virus
The IAV virus (strain (A/Puerto Rico/8/1934(H1N1))) used in our study127 was purchased from Charles River Laboratories. The virus was propagated in SPF eggs in the allantoic cavity. Clarified allantoic fluid was concentrated and resuspended in Hepes-Saline solution and layered on a sucrose gradient. The interface band was diluted, pelleted and resuspended in a minimal volume of Hepes-Saline solution. Antigen was tested for protein concentrate of 2 mg of protein per ml using a Bio-Rad colorimetric protein assay. The final HA titer per 0.05 ml was 131,072 and the EID50 titer per ml was 109.8. A FITC-conjugated anti-Influenza A NP monoclonal antibody (ThermoFisher MA1-7322) at 1:20 dilution was used to detect viral proteins using flow cytometry.
Cerebral organoid differentiation
We adapted a previously described protocol to generate spontaneously differentiated dcOrgs from PGP1 hiPSCs47,128,129. We re-suspended 900,000 cells in 15 ml of StemFlex with 50μM Y-27632 and seeded the cells across a 96-well ultra-low attachment plate (Corning CLS7007) to form embryoid bodies. After 5 days, each embryoid body was transferred to 24-well ultra-low attachment plates (Corning CLS3473) with 500μl of neural induction media in each well. An additional 500μl of neural induction media was added to each well on Day 8. On Day 10, each organoid was embedded in 40μl of Matrigel (Corning 354234) on parafilm and incubated at 37°C for 15 minutes before they were scraped into the wells containing 2 ml of differentiation media130 using a cell scraper. 1-2 ml of differentiation media with 10% penicillin streptomycin (ThermoFisher 15140122) per well was used to passage the organoids every 3-7 days, and the plates of organoids were placed on an orbital shaker at 90 rpm.
Cerebral organoid dissociation
After 2-4 months of differentiation, we dissociated 168-192 dcOrgs in each batch and dcOrgs were washed twice in ice-cold 1×DPBS (ThermoFisher 14190144) for 10 minutes at 4°C, followed by incubation in 500μL of 0.25% Trypsin-EDTA (ThermoFisher 25200056) for 15 minutes at 37°C and 300 rpm with repeated manual pipetting to dissociate clumps of cells129. To remove cell debris, we further filtered the cells using a 30 μm MACS SmartStrainer (Miltenyi Biotec 130-098-458). Cell counts and viabilities were measured using an automated cell counter (NanoEnTek EVE). Cell viabilities ranged from 80% to 96%, and we obtained 7.8×10-6 to 4.1×10-7 live cells in total. Subsequently, the dissociated cells were passaged in differentiation media for at least a month to allow the cells to recover. We recorded the total time allowed for differentiation and recovery of the cells prior to infection as the age of the dcOrgs. We expect that there would be substantial cellular differences between 3-month and 5-month dcOrgs; as such, experimentation on these dcOrgs would inform us if the cellular or transcriptomic signatures observed could be replicated across dcOrgs with different ages, or if there was a critical age for modeling AD molecular and transcriptomic signatures.
HSV-1 and IAV infection experiments
For HSV-1 infections in 2D dcOrgs, we used an MOI of 2 that resulted in >50% of cells that were positive for GFP (Fig. 1b). We used a higher virus titer (MOI of 4) for hiPSCs. We counted the number of cells needed for each experiment and seeded the cells at a density of 1 × 106 cells/well of a 6-well tissue culture plate coated with 0.5 mg of Matrigel, using 2 mL/well of differentiation media with 10 μM Y-27632. On the following day, cells from one of the wells were counted using an automated cell counter to calculate the amount of virus needed.
Inoculation media containing 1 × DPBS with 0.5% FBS (ThermoFisher 10082147) was prepared, and aliquots of HSV-1 and ACV were kept on ice. Cells in each well were washed with 2 ml of 1 × DPBS. 1 mL of inoculation media was added to each well of mock-infected cells, 1 ml of inoculation media with HSV-1 was added to each well of infected cells, and 1 ml of inoculation media with HSV-1 and 200 μM ACV was added to each well of treated cells. The cells were placed back into an incubator at 37 °C for an hour, and each well of cells were subsequently washed with 2 ml of 1 × DPBS. 2 ml of differentiation media was added to each well of uninfected or infected cells, and 2 ml of differentiation media with 200 μM ACV was added to each well of treated cells. The cells were placed back into an incubator at 37 °C for 23 h. We inspected the cells under a fluorescence microscope (Logos Biosystems CELENA S Digital Imaging System) to visually verify the presence of GFP in the infected and treated cells and that UV-inactivation worked to stop viral replication. A similar protocol was used for IAV infections, except that MOI of 3 was used with a 47-h post-inoculation incubation for the ELISA assays and an MOI of 6 with a 23-h post-inoculation incubation was used for the flow cytometry experiments.
Antibody labeling and flow cytometry experiments
Cells were washed with 1×DPBS through 30 μm filters to ensure that the suspension contained single cells for flow cytometry, followed by pelleting into 1.5 ml microcentrifuge tubes. Each cell pellet was resuspended in 100 μl of 1 × DPBS with 1:500 Zombie Violet (BioLegend 423113) and incubated for 20 min at room temperature in the dark. Cells were then washed with 500 μl of cell staining buffer (BioLegend 420201). For intracellular antibodies, either 100 μl of 4% paraformaldehyde or 250 μl of Cytofix/Cytoperm fixation and permeabilization buffer (BD Biosciences 554714) was then added, and the cells were incubated at 4 °C for 20 min at 300 rpm, then washed once with 500 μl of permeabilization solution. For cell surface antibodies, the cells were then washed with 500 μl of cell staining buffer, then resuspended in 105 μl of cell staining buffer with 1:21 Human TruStain FcX (BioLegend 422301), followed by incubation at 4 °C for 20 min at 300 rpm. Each cell pellet was then resuspended in 50μl of the respective buffer with 1:21 Human TruStain FcX and the antibody of interest, followed by incubation at 4 °C for 60 min at 300 rpm. After antibody labeling, each cell pellet was washed twice with 500 μl of Cell Staining Buffer, resuspended in 300 μl of Cell Staining Buffer and transferred to a 5 mL glass tube. We used a BD Biosciences FACSCelesta Cell Analyzer at the UMass Chan Medical School FACS core facility for our flow cytometry experiments. Using the FACSDiva software, a “usable cell” gate was drawn on the FSC-A/SSC-A plot to minimize the collection of debris, and 20,000–30,000 cells within the usable cell population were analyzed.
List of antibodies used
For quantifying the proportions of cell type markers (e.g., P2RY12) in dcOrgs, we matched the concentrations of the antibodies and control IgG1 antibody for staining the dcOrgs. For the other experiments to compare mock dcOrgs with infected or treated dcOrgs, we stained the dcOrgs using antibody concentrations that were diluted as described below. We acknowledge that the specificity of antibodies remains an issue for the reproducibility of results131 and that extensive validation of antibodies using knockout lines is expensive and dependent on several conditions, such as the antibody concentration and assays conducted (Western blots or flow cytometry). As such, we carefully designed our flow cytometry experiments and compared the correlations in co-abundance of the antibody of interest with viral protein abundance across conditions (e.g., mock, infected and treated), or the fraction of cells that pass an intensity threshold for the antibody of interest across the conditions (e.g., mock, infected or treated). In this way, we evaluated the differences across conditions, rather than the absence or presence of a protein marker, and our results would be less dependent on the specificity of the antibody.
The Aβ and tau antibodies used in our study were at 1:50 dilutions or otherwise stated, and include Alexa Fluor 647-conjugated APP OTI7G9 (Novus Biologicals NBP2-70187AF647), Aβ1-40 11A50-B10 (BioLegend 805401), Alexa Fluor 647-conjugated Aβ1-16 6E10 (BioLegend 803021) at 1:20 dilution, APC-conjugated Aβ MOAB-2 (Novus Biologicals NBP2-13075AF647), Alexa Fluor 647-conjugated aducanumab (Novus Biologicals FAB9710R-100UG), Alexa Fluor 647-conjugated solanezumab (Novus Biologicals FAB9919R), total tau HT7 (Invitrogen MN1000), Alexa Fluor 647-conjugated pTau-181 (Abcam ab309970), pTau-217 E9Y4S (Cell Signaling Technology 51625S), pTau AT8 (Invitrogen MN1020) and pTau-205 (Invitrogen ab307376).
The cell-type antibodies used in our study were used at 1:50 dilutions and conjugated with Alexa 647 or otherwise stated: TRA-1-60 (BioLegend 330605) at 1:20 dilution, Nestin (Novus Biologicals IC1259R-100UG), EOMES (Novus Biologicals IC6166R-100UG), APC-conjugated TuJ1 (BioLegend 801219) at 1:20 dilution, NeuN (Novus Biologicals NBP1-92693AF647), VMAT2 (R&D Systems FAB8327R) at a 1:20 dilution, GFAP (BioLegend 644706) at a 1:20 dilution, APC-conjugated GLAST (Miltenyi 130-123-555), Iba1 (Novus Biologicals 603102), APC-conjugated P2RY12 (BioLegend 392113) at a 1:20 dilution, CD4 (BioLegend 300520) at a 1:20 dilution, CD45 (Invitrogen MA5-38730) at a 1:20 dilution, Olig1/2/3 (Bio-techne FAB2230R-MTO), O4 (R&D Systems FAB1326R) at a 1:20 dilution and O1 (R&D Systems FAB1327R) at a 1:20 dilution. Polyclonal antibodies used included Alexa Fluor 647-conjugated Aβ (Bioss BS-0107R-BF64), Alexa Fluor 647-conjugated pTau-212 (BS-5420R-BF647) and Alexa Fluor 647-conjugated OLIG1 (Bioss bs-8548R-A647).
Isotype controls in our study were used at 1:50 dilutions or otherwise stated and include Alexa Fluor 647-conjugated rabbit IgG (Bioss Antibodies bs-0295P-A647), Alexa Fluor 647-conjugated mouse IgG1 kappa (Novus Biologicals IC002R), Alexa Fluor 647-conjugated mouse IgG2A kappa (Biolegend 400234F2) at a 1:20 dilution and Alexa Fluor 647-conjugated mouse IgG2B kappa (Novus Biologicals IC0041RF3) at a 1:20 dilution.
Flow cytometry data analyses
Flow cytometry data were exported from FlowJo v10 as a.csv file with the per-cell values for the Alexa Fluor 488-A, Alexa Fluor 647-A, Pacific Blue-A, FSC-A, FSC-H, SSC-A, and SSC-H channel intensities. FSC-A and SSC-A gates for each experiment were set at the 5th and 95th percentiles of the control sample and applied to the infected and ACV-treated samples. We next tested for singlets by fitting a linear regression model using FSC-A versus FSC-H intensities for each condition (mock, infected, or treated). We calculated the residuals and cells with residuals greater than the 99th percentile were classified as doublets and were removed from further analyses.
Pacific Blue-A, Alexa Fluor 488-A, and Alexa Fluor 647-A gates were set using an automated pipeline utilizing Gaussian mixture models (GMMs). Pacific Blue-A and Alexa Fluor 647-A gates were set based on the mock samples, while Alexa Fluor 488-A gates were set based on the infected samples. For each distribution, GMMs were initialized with 1 to 4 components, and model fit was evaluated using the Bayesian Information Criterion (BIC), separation index, and component weights. Models were required to meet two criteria: (1) the separation index between the two highest-intensity components had to be ≥1.5, and (2) all components had to have weights ≥0.05. Among models meeting these criteria, the one with the lowest BIC was selected. If the selected model was unimodal, other conditions were inspected for multimodality. If a GMM meeting the criteria above could be identified on a different condition, then the gate was set based on the condition with multimodality instead. If all conditions were unimodal, then the gate was set at the 85th percentile of the designated condition. If a bimodal or multimodal model was selected, the gate was placed at the intersection between the two highest-intensity components, falling between their means. These gates were then applied to all other conditions. In the supplementary figures, the condition that the gate was set on has dotted lines overlaying the GMM chosen and showed the intersection between the top distributions. GMMs and BIC scores were implemented and calculated using the scikit-learn Python package.
UMAP plots
UMAP was performed using the Python package umap-learn. For each flow experiment, the initial UMAP values for Alexa Fluor 488-A, Alexa Fluor 647-A, and Pacific Blue-A were normalized between 0 and 1 across all conditions (uninfected, infected and ACV-treated). The data for these three channels were then fitted to a UMAP model initialized with the number of components = 2 to re-calculate UMAP1 and UMAP2 values. The UMAP was fitted to all viable cells, including cell staining positive for Pacific Blue-A.
Correlations in Alexa Fluor 488 and Alexa Fluor 647 intensities
We calculated Pearson’s correlation \(r\) between the Alexa Fluor 488 or GFP and Alexa Fluor 647 or APC intensities in the mock, infected and treated samples. The final correlations shown in the heatmaps are the differences in the correlation (Δ\(r\)) from the infected dcOrgs minus the correlation from the mock dcOrgs, or the correlation from the treated dcOrgs minus the correlation from the mock dcOrgs. Differences in correlations (Δ\(r\)) were calculated by first applying Fisher’s \(r\)-to-z transformations of the coeffecients, subtracting the difference in z-values, and converting the difference back to \(r\) values using Fisher’s z-to-\(r\) transformation. Mean \(r\) and Δ\(r\) values were calculated by first applying Fisher’s \(r\)-to-z transformations, averaging the values, and transforming them back to \(r\) values using Fisher’s z-to-\(r\) transformation. 95% CI intervals for \(r\) values and differences (Δ\(r\)) were calculated by first applying Fisher’s \(r\)-to-z transformation, performing N = 1000 bootstrap resamples (sampling with replacement), identifying the 2.5th and 97.5th percentiles of the bootstrap distribution and finally transforming these values back to \(r\) values using Fisher’s z-to-\(r\) transformations. All statistical tests were performed in Python using Scipy.stats v 1.14.1132.
Odds ratio calculations from flow cytometry data
Using the machine learning based gating, we grouped the live-gated cells into four quadrants: Q1 where the cells were negative for HSV-1-GFP abundance and positive for abundance of the marker of interest (Alexa Fluor 647 or APC), Q2 where the cells were positive for abundance of both HSV-1 and the marker of interest, Q3 where the cells were positive for abundance of HSV-1 and negative for abundance of the marker of interest, and Q4 where the cells were negative for abundance of both HSV-1 and the marker of interest. Odds ratios were calculated only using viable, live gated (Pacific Blue-A positive) cells. We calculated ORs and Fisher’s Exact Test P values in R133, where:
ELISA assays and analyses
After inoculation, 1 mL of fresh dcOrg differentiation media was dispensed into each well and cells were incubated for 48 h. Conditioned media from each well were collected into two tubes with 500 μl of media in each tube. Each set of conditions (mock dcOrgs, HSV-1-infected dcOrgs, ACV-treated dcOrgs, IAV-infected dcOrgs) had 4–5 experimental replicates. One set of conditioned media was heat-inactivated at 65 °C for an hour and the second set of conditioned media was UV-inactivated. Conditioned media was sent to Harvard on dry ice overnight and extracellular Aβ42/40/38 levels were measured from the conditioned media using the V-PLEX Plus Aβ Peptide Panel 1 (6E10) Kit (MSD K15200G-1) in a blinded experiment, with controls diluted in the given buffer134. The Aβ peptide values were batched normalized to control in the ELISA kit. The concentrations of each Aβ peptide and ratios of Aβ peptides were visualized using ggplot135 in R133. 1-tailed Wilcoxon ranked sum tests were used to evaluate the samples of interest versus matched mock controls.
Bulk RNA extraction, sequencing, and data processing
Cells were fixed using 4% paraformaldehyde and RNA was extracted from the fixed cells using the PureLink FFPE RNA Isolation Kit (ThermoFisher K156002) and treated with Ambion DNase I (ThermoFisher AM2222) to digest genomic DNA according to the manufacturers’ protocols. Each set of experiments was performed in triplicate (technical replicates of the same batch of dcOrgs conducted at the same time across three independent wells). Total RNA was shipped overnight on dry ice to Psomagen for quantification (Data S1), followed by ribosomal RNA depletion and library preparation (Illumina TruSeq Stranded Total RNA Library Prep 20020596), and 151 bp paired-end sequencing with a total of 40 million reads on a NovaSeq6000.
Quality control was performed with the FastQC tool136, and adapter sequences and low-quality base calls were trimmed from FASTQs with Trimmomatic v0.39137. Trimmed FASTQs were aligned to a concatenated GRCh37 human reference genome, human alphaherpesvirus 1 Kos strain reference genome (GenBank: JQ673480.1) and influenza A strain A/Puerto Rico/8/1934(H1N1) reference genome (NCBI BioProject PRJNA485481) using HISAT2 v2.2.1138. SAMtools v1.9 was used to extract reads aligning to the coding sequences of the virus139. Expression quantification and preprocessing were performed using StringTie2 v1.3.6140.
The transcript list was filtered to include only protein-coding transcripts and duplicate genes in the StringTie2 output were removed, resulting in a total of 19,163 unique genes. We compared two approaches for alignment: firstly, we aligned all transcripts to a custom reference comprised of both human and HSV-1 reference sequences and secondly, we aligned all transcripts to human and HSV-1 reference sequences individually. We calculated Pearson’s \(r\) between the raw gene expression using both alignment approaches and found high correlations between both approaches (1−\(r\)2 ≤ 1 × 10−4). As a quality control step, we checked that there was no HSV-1 viral transcript expression in all control uninfected samples. Pearson’s \(r\) was calculated using the fragments per kilobase of exon per million mapped fragments (FPKM) values between sample pairs.
Differential expression analyses
We performed differential expression analyses using the DESeq2 v1.38.2 package in R141. For the PGP1 RNA-seq data, genes were filtered using a cutoff of counts per million (CPM) ≥ 1 across at least two samples. For the A3P RNA-seq data, genes were filtered using a cutoff of CPM≥1 across all samples. CPMs were normalized across samples using DESeq2 median of ratios and models subsequently generated on a categorical design of infection or treatment status. The adjusted P-values from the differential expression analyses used in our study were Benjamini-Hochberg adjusted from the raw P-values. Volcano plots were generated in R with EnhancedVolcano v1.11.3142 and heatmaps were generated in R with ggplot2135.
Gene lists from GWAS catalog and gene set enrichment analyses
Gene lists associated with 21 common diseases were downloaded from the NHGRI-EBI GWAS catalog100 in April 2022. Gene set enrichment analyses were performed using GSEA v4.3.2101,102 to determine the enrichment of DEGs ranked by absolute log2 fold change in decreasing order with the GWAS-associated gene lists. Analyses were performed using 10,000 permutations and a weighted enrichment statistic.
Definitions of rescued, not rescued, exacerbated 1 and 2 transcripts
“Rescued” transcripts were defined as transcripts that were significantly differentially expressed (adjusted P ≤ 0.05) in both the Inf-vs-Mock dcOrgs and Inf-vs-ACV dcOrgs datasets and log2 fold changes in both datasets were in the same direction110. For instance, a positive log2 fold change in Inf-vs-Mock dcOrgs indicated that HSV-1 infection led to increased expression of the transcript. Similarly, a positive log2 fold change in Inf-vs-ACV dcOrgs indicated that ACV treatment of HSV-1-infected dcOrgs lowered the expression of the transcript, with respect to the expression in infected dcOrgs. “Not rescued” transcripts were defined as transcripts that were significantly differentially expressed (adjusted P≤0.05) in Inf-vs-Mock dcOrgs but were not significantly differentially expressed (adjusted P > 0.05) in Inf-vs-ACV dcOrgs.
“Exacerbated 1” transcripts were defined as transcripts that were significantly differentially expressed (P≤0.05) in both the Inf-vs-Mock dcOrgs and Inf-vs-ACV dcOrgs datasets, but log2 fold changes in both datasets were in the opposite direction. For instance, a positive log2 fold change in Inf-vs-Mock dcOrgs indicated that HSV-1 infection led to increased expression of the transcript. However, a negative log2 fold change in Inf-vs-ACV dcOrgs indicated that ACV treatment of HSV-1 infected dcOrgs further increased the expression of the transcript, with respect to the expression in infected dcOrgs.
“Exacerbated 2” transcripts were defined as transcripts that were not significantly perturbed in Inf-vs-Mock dcOrgs (P > 0.05) but were significantly differentially expressed in Inf-vs-ACV dcOrgs (P≤0.05). This indicated that ACV treatment on HSV-1-infected dcOrgs had significantly perturbed the expression of these transcripts when the expression of these transcripts was not perturbed by HSV-1 infection alone in dcOrgs.
Scatter plots to represent rescued, not rescued, exacerbated 1 and 2 transcripts
Scatter plots showed the log2 fold changes in the Inf-vs-Mock dcOrgs versus the log2 fold changes in Inf-vs-ACV dcOrgs for all transcripts that were in common between both datasets. The percentages of transcripts in each category (rescued, not rescued, exacerbated 1, exacerbated 2 or not DEG) were calculated as the number of transcripts in each category divided by the total number of transcripts and multiplied by 100%. Pearson’s \(r\) was calculated between the log2 fold changes in Inf-vs-Mock dcOrgs and Inf-vs-ACV dcOrgs for the transcripts in each category.
Enrichment of disease-associated rescued or exacerbated transcripts
Fisher’s Exact Test was used to calculate the enrichment of the number of disease-associated transcripts in a group (such as the transcripts that were rescued in both dcOrgs1 and dcOrgs2) over the number of disease-associated transcripts that were not in the group, compared to (the total number of transcripts in the group minus the number of disease-associated transcripts in the group) over (the total number of transcripts that were not in the group minus the number of disease-associated transcripts that were not in the group).
scRNA-seq library preparation and sequencing
We prepared 3 replicates for each condition (mock dcOrgs and HSV-1-infected dcOrgs). A single-cell suspension of 900,000 to 1 million cells from each sample was fixed and permeabilized using the Evercode Cell Fixation V2 Kit (Parse Biosciences ECF2001). Following fixation, we performed split-pool combinatorial barcoding using the Evercode WT Mini v2 Kit (Parse Biosciences ECW2010), resulting in two sub-libraries. The cDNA from the sub-libraries was quantified and a total of 129.1 ng of cDNA from each sub-library was sent for 100 bp paired-end sequencing using DNBSEQ, yielding 774,454,526 total reads and an average of 40,044 reads per cell.
scRNA-seq data processing, alignment, and sample deconvolution
Data processing, alignment, sample deconvolution, and count matrix generation were performed using the Parse Biosciences command line pipeline v1.1.0. We generated an indexed reference using a concatenated GRCh38.p12 release 109 human reference genome and human alphaherpesvirus 1 Kos strain reference genome (GenBank: JQ673480.1). Using the pipeline, we integrated two initial sub-libraries from the same library preparation with “--mode combine” option in the Parse pipeline, performed STAR alignment to the indexed reference, and deconvolved the six samples (three mock samples and three HSV-1-infected samples) using SAMtools. Filtered count matrices provided by the Parse Biosciences pipeline output were used for downstream analyses.
Complete human and HSV-1 viral count matrices were preprocessed and clustered using Seurat v4.3143. Samples were filtered by a minimum of 8000 features and a maximum of 40,000 transcript counts per cell based on preliminary quality control. Samples were subsequently normalized using the SCTransform function in Seurat, then integrated into a unified dataset and clustered. We generated pseudobulk groups after establishing a baseline viral count per cell, where cells with <100 summed raw viral transcripts were classified as pseudo-uninfected (PU), cells with ≥100 raw viral transcripts were classified as true infected (TI), and cells from the mock samples were classified as true uninfected (TU). The aggregated cells from TI and PU were classified as infected (I). Genes that had CPM > 1 expression in at least two samples were included in the analyses and differential expression analyses were performed using DESeq2 v1.38.0 on the pseudobulk filtered expression data. Pre-ranked GSEA analyses were then performed with absolute log2 fold change using GSEA v4.3.2. We observed similar correlations across samples from the scRNA-seq data as the bulk RNA-seq data.
scRNA-seq cell type assignment
Cell type assignment was conducted by using the majority consensus vote of three approaches: Seurat143, SingleR144, and scPred145. Initial count matrices were generated for each method utilizing the complete set of cells and features present after quality control pre-processing steps. Reference and query count matrices were normalized using SCTransform in Seurat for use with label transfer. Reference and query count matrices were alternatively log-normalized for use with scPred and SingleR assignment. The reference datasets for cell type assignment, consisting of human brain organoids and fetal brains from Tanaka et al.120 and adult post-mortem brains from Allen (Reference_MTG_RNAseq_final-nuclei.2022-06-07.h5ad)121, were initially down-sampled by using Seurat to 60,000 cells due to memory limitations, prior to integration and normalization. The dataset was processed using Scanpy v1.9.3146. If a raw log-transformed expression matrix was not provided (adata.raw is None), we applied a natural log-transformation to the count matrix using the “sc.pp.log1p” function.
The “TransferData” function in Seurat was used for reference data label transfer across both datasets. SingleR assignments used the Wilcoxon rank sum “de.method” function. scPred assignment models were trained on each reference dataset using a xgbDART-based model, with identity prediction and scores returned using the “scPredict” function. Initial cell type assignment was determined by majority consensus vote between individual assignment methods for each reference dataset, performed with assignment threshold rejections in scPred. Cells that did not have a cell type assignment by majority consensus were assigned by using the highest confidence across the cell type assignments by using the min-max scaled confidence scores from each method.
RT-qPCR experiments
Total RNA was extracted from mock or HSV-1-infected dcOrgs using the PureLink RNA Mini Kit (Thermo Fisher 12183018A) and reverse transcribed using the High-Capacity cDNA Reverse Transcription Kit (Thermo Fisher 4374967). qPCR was performed with a Biorad CFX Opus using the PowerTrack SYBR Green Master Mix (ThermoFisher A46109). Primers were designed using Primer-BLAST. Relative gene expression was calculated using ΔΔCt method and 18s rRNA was used as the housekeeping gene for normalization. Three biological replicates were used per condition and were run in quadruplicates.
Schematic figures
All schematic figures in this manuscript were created using BioRender and can be found at https://BioRender.com/pw4h34p, https://BioRender.com/d29mp4z and https://BioRender.com/22wp3zr.
Data availability
Our scripts have been uploaded to the project GitHub page (https://gitlab.com/elimlab/hsv1-bulk). Our bulk transcriptomics raw data is being uploaded to https://www.synapse.org/Synapse:syn69762030.
References
Green, G. S. et al. Cellular communities reveal trajectories of brain ageing and Alzheimer’s disease. Nature 633, 634–645, https://doi.org/10.1038/s41586-024-07871-6 (2024).
Mathys, H. et al. Single-cell multiregion dissection of Alzheimer’s disease. Nature 632, 858–868, https://doi.org/10.1038/s41586-024-07606-7 (2024).
Gabitto, M. I. et al. Integrated multimodal cell atlas of Alzheimer’s disease. Nat. Neurosci. https://doi.org/10.1038/s41593-024-01774-5 (2024).
Wang, Q. et al. Single cell transcriptomes and multiscale networks from persons with and without Alzheimer’s disease. Nat. Commun. 15, 5815. https://doi.org/10.1038/s41467-024-49790-0 (2024).
Kosoy, R. et al. Genetics of the human microglia regulome refines Alzheimer’s disease risk loci. Nat. Genet. 54, 1145–1154, https://doi.org/10.1038/s41588-022-01149-1 (2022).
Sun, N. et al. Human microglial state dynamics in Alzheimer’s disease progression. Cell 186, 4386–4403.e4329. https://doi.org/10.1016/j.cell.2023.08.037 (2023).
Efthymiou, A. G. & Goate, A. M. Late onset Alzheimer’s disease genetics implicates microglial pathways in disease risk. Mol. Neurodegener. 12, 43, https://doi.org/10.1186/s13024-017-0184-x (2017).
Wozniak, M. A., Mee, A. P. & Itzhaki, R. F. Herpes simplex virus type 1 DNA is located within Alzheimer’s disease amyloid plaques. J. Pathol. 217, 131–138, https://doi.org/10.1002/path.2449 (2009).
Jamieson, G. A., Maitland, N. J., Craske, J., Wilcock, G. K. & Itzhaki, R. F. Detection of herpes simplex virus type 1 DNA sequences in normal and Alzheimer’s disease brain using polymerase chain reaction. Biochem Soc. Trans. 19, 122S, https://doi.org/10.1042/bst019122s (1991).
Itzhaki, R. F. et al. Herpes simplex virus type 1 in brain and risk of Alzheimer’s disease. Lancet 349, 241–244, https://doi.org/10.1016/S0140-6736(96)10149-5 (1997).
Wozniak, M. A., Shipley, S. J., Combrinck, M., Wilcock, G. K. & Itzhaki, R. F. Productive herpes simplex virus in brain of elderly normal subjects and Alzheimer’s disease patients. J. Med. Virol. 75, 300–306, https://doi.org/10.1002/jmv.20271 (2005).
Lin, W. R., Wozniak, M. A., Cooper, R. J., Wilcock, G. K. & Itzhaki, R. F. Herpesviruses in brain and Alzheimer’s disease. J. Pathol. 197, 395–402, https://doi.org/10.1002/path.1127 (2002).
Carbone, I. et al. Herpes virus in Alzheimer’s disease: relation to progression of the disease. Neurobiol. Aging 35, 122–129, https://doi.org/10.1016/j.neurobiolaging.2013.06.024 (2014).
De Chiara, G. et al. Recurrent herpes simplex virus-1 infection induces hallmarks of neurodegeneration and cognitive deficits in mice. PLoS Pathog. 15, e1007617, https://doi.org/10.1371/journal.ppat.1007617 (2019).
Itzhaki, R. F., Golde, T. E., Heneka, M. T. & Readhead, B. Do infections have a role in the pathogenesis of Alzheimer's disease? Nat. Rev. Neurol. 16, 193–197, https://doi.org/10.1038/s41582-020-0323-9 (2020).
Wainberg, M. et al. The viral hypothesis: how herpesviruses may contribute to Alzheimer’s disease. Mol. Psychiatry 26, 5476–5480, https://doi.org/10.1038/s41380-021-01138-6 (2021).
Bu, X. L. et al. A study on the association between infectious burden and Alzheimer’s disease. Eur. J. Neurol. 22, 1519–1525, https://doi.org/10.1111/ene.12477 (2015).
Itzhaki, R. F., Wozniak, M. A., Appelt, D. M. & Balin, B. J. Infiltration of the brain by pathogens causes Alzheimer’s disease. Neurobiol. Aging 25, 619–627, https://doi.org/10.1016/j.neurobiolaging.2003.12.021 (2004).
Lin, W. R., Shang, D., Wilcock, G. K. & Itzhaki, R. F. Alzheimer’s disease, herpes simplex virus type 1, cold sores and apolipoprotein E4. Biochem. Soc. Trans. 23, 594S, https://doi.org/10.1042/bst023594s (1995).
Tzeng, N. S. et al. Anti-herpetic medications and reduced risk of dementia in patients with herpes simplex virus infections—a nationwide, population-based cohort study in Taiwan. Neurotherapeutics 15, 417–429, https://doi.org/10.1007/s13311-018-0611-x (2018).
Bae, S. et al. Association of herpes zoster with dementia and effect of antiviral therapy on dementia: a population-based cohort study. Eur. Arch. Psychiatry Clin. Neurosci. 271, 987–997, https://doi.org/10.1007/s00406-020-01157-4 (2021).
Tsai, M. C. et al. Increased risk of dementia following herpes zoster ophthalmicus. PLoS One 12, e0188490, https://doi.org/10.1371/journal.pone.0188490 (2017).
Chen, V. C. et al. Herpes zoster and dementia: a nationwide population-based cohort study. J. Clin. Psychiatry 79. https://doi.org/10.4088/JCP.16m11312 (2018).
Itzhaki, R. F. & Lathe, R. Herpes viruses and senile dementia: first population evidence for a causal link. J. Alzheimers Dis. 64, 363–366, https://doi.org/10.3233/JAD-180266 (2018).
Levine, K. S. et al. Virus exposure and neurodegenerative disease risk across national biobanks. Neuron. https://doi.org/10.1016/j.neuron.2022.12.029 (2023).
Readhead, B. et al. Multiscale analysis of independent Alzheimer’s cohorts finds disruption of molecular, genetic, and clinical networks by human herpesvirus. Neuron 99, 64–82.e67, https://doi.org/10.1016/j.neuron.2018.05.023 (2018).
Eimer, W. A. et al. Alzheimer’s disease-associated beta-amyloid is rapidly seeded by herpesviridae to protect against brain infection. Neuron 99, 56–63.e53, https://doi.org/10.1016/j.neuron.2018.06.030 (2018).
Rybak-Wolf, A. et al. Modelling viral encephalitis caused by herpes simplex virus 1 infection in cerebral organoids. Nat. Microbiol. 8, 1252–1266, https://doi.org/10.1038/s41564-023-01405-y (2023).
Bruno, F. et al. Alzheimer’s disease as a viral disease: revisiting the infectious hypothesis. Ageing Res. Rev. 91, 102068. https://doi.org/10.1016/j.arr.2023.102068 (2023).
Jang, H. et al. Highly pathogenic H5N1 influenza virus can enter the central nervous system and induce neuroinflammation and neurodegeneration. Proc. Natl. Acad. Sci. USA 106, 14063–14068, https://doi.org/10.1073/pnas.0900096106 (2009).
De Chiara, G. et al. Infectious agents and neurodegeneration. Mol. Neurobiol. 46, 614–638, https://doi.org/10.1007/s12035-012-8320-7 (2012).
Jamieson, G. A., Maitland, N. J., Wilcock, G. K., Craske, J. & Itzhaki, R. F. Latent herpes simplex virus type 1 in normal and Alzheimer’s disease brains. J. Med. Virol. 33, 224–227, https://doi.org/10.1002/jmv.1890330403 (1991).
Readhead, B. P. et al. Alzheimer’s disease-associated CD83(+) microglia are linked with increased immunoglobulin G4 and human cytomegalovirus in the gut, vagal nerve, and brain. Alzheimers' Dement. https://doi.org/10.1002/alz.14401 (2024).
Field, H. J., Bell, S. E., Elion, G. B., Nash, A. A. & Wildy, P. Effect of acycloguanosine treatment of acute and latent herpes simplex infections in mice. Antimicrob. Agents Chemother. 15, 554–561, https://doi.org/10.1128/AAC.15.4.554 (1979).
Weidung, B. et al. VALZ-Pilot: High-dose valacyclovir treatment in patients with early-stage Alzheimer’s disease. Alzheimer's Dement. 8, e12264. https://doi.org/10.1002/trc2.12264 (2022).
Devanand, D. P. et al. Antiviral therapy: Valacyclovir Treatment of Alzheimer’s Disease (VALAD) trial: protocol for a randomised, double-blind,placebo-controlled, treatment trial. BMJ Open 10, e032112, https://doi.org/10.1136/bmjopen-2019-032112 (2020).
Klysik, K., Pietraszek, A., Karewicz, A. & Nowakowska, M. Acyclovir in the treatment of herpes viruses—a review. Curr. Med. Chem. 27, 4118–4137, https://doi.org/10.2174/0929867325666180309105519 (2020).
Lathe, R., Tzeng, N. S. & Itzhaki, R. Herpes infections and dementia: rebutting alternative fact. Neurotherapeutics 16, 176–179, https://doi.org/10.1007/s13311-018-00700-5 (2019).
Lehrer, S. & Rheinstein, P. H. Herpes zoster vaccination reduces risk of dementia. In Vivo 35, 3271–3275, https://doi.org/10.21873/invivo.12622 (2021).
Lophatananon, A. et al. The association of herpes zoster and influenza vaccinations with the risk of developing dementia: a population-based cohort study within the UK Clinical Practice Research Datalink. BMC Public Health 23, 1903. https://doi.org/10.1186/s12889-023-16768-4 (2023).
Tang, E., Ray, I., Arnold, B. F. & Acharya, N. R. Recombinant zoster vaccine and the risk of dementia. Vaccine 46, 126673. https://doi.org/10.1016/j.vaccine.2024.126673 (2024).
Cairns, D. M., Itzhaki, R. F. & Kaplan, D. L. Potential involvement of Varicella Zoster virus in Alzheimer’s disease via reactivation of quiescent herpes simplex virus type 1. J. Alzheimers Dis. 88, 1189–1200, https://doi.org/10.3233/JAD-220287 (2022).
Cairns, D. M. et al. Repetitive injury induces phenotypes associated with Alzheimer’s disease by reactivating HSV-1 in a human brain tissue model. Sci. Signal 18, eado6430. https://doi.org/10.1126/scisignal.ado6430 (2025).
Velasco, S. et al. Individual brain organoids reproducibly form cell diversity of the human cerebral cortex. Nature. https://doi.org/10.1038/s41586-019-1289-x (2019).
Mariani, J. et al. FOXG1-dependent dysregulation of GABA/glutamate neuron differentiation in autism spectrum disorders. Cell 162, 375–390, https://doi.org/10.1016/j.cell.2015.06.034 (2015).
Quadrato, G. et al. Cell diversity and network dynamics in photosensitive human brain organoids. Nature 545, 48–53, https://doi.org/10.1038/nature22047 (2017).
Lim, E. T. et al. Orgo-Seq integrates single-cell and bulk transcriptomic data to identify cell-type-specific-driver genes associated with autism spectrum disorder. Nat. Commun. 13, 3243. https://doi.org/10.1038/s41467-022-30968-3 (2022).
Pasca, S. P. et al. A framework for neural organoids, assembloids and transplantation studies. Nature 639, 315–320, https://doi.org/10.1038/s41586-024-08487-6 (2025).
He, Z. et al. An integrated transcriptomic cell atlas of human neural organoids. Nature 635, 690–698, https://doi.org/10.1038/s41586-024-08172-8 (2024).
Renner, M. et al. Self-organized developmental patterning and differentiation in cerebral organoids. EMBO J. 36, 1316–1329, https://doi.org/10.15252/embj.201694700 (2017).
Zhao, J. et al. APOE4 exacerbates synapse loss and neurodegeneration in Alzheimer’s disease patient iPSC-derived cerebral organoids. Nat. Commun. 11, 5540. https://doi.org/10.1038/s41467-020-19264-0 (2020).
Choi, S. H. et al. A three-dimensional human neural cell culture model of Alzheimer’s disease. Nature 515, 274–278, https://doi.org/10.1038/nature13800 (2014).
Cairns, D. M. et al. A 3D human brain-like tissue model of herpes-induced Alzheimer’s disease. Sci. Adv. 6, eaay8828. https://doi.org/10.1126/sciadv.aay8828 (2020).
D’Aiuto, L. et al. Modeling Herpes Simplex Virus 1 infections in human central nervous system neuronal cells using two- and three-dimensional cultures derived from induced pluripotent stem cells. J. Virol. 93. https://doi.org/10.1128/JVI.00111-19 (2019).
Ezzat, K. et al. The viral protein corona directs viral pathogenesis and amyloid aggregation. Nat. Commun. 10, 2331. https://doi.org/10.1038/s41467-019-10192-2 (2019).
Wang, Y. et al. Dysregulation of cofilin-1 activity-the missing link between herpes simplex virus type-1 infection and Alzheimer’s disease. Crit. Rev. Microbiol 46, 381–396, https://doi.org/10.1080/1040841X.2020.1794789 (2020).
Krenn, V. et al. Organoid modeling of Zika and herpes simplex virus 1 infections reveals virus-specific responses leading to microcephaly. Cell Stem Cell 28, 1362–1379.e1367, https://doi.org/10.1016/j.stem.2021.03.004 (2021).
Hyde, V. R. et al. Anti-herpetic tau preserves neurons via the cGAS-STING-TBK1 pathway in Alzheimer’s disease. Cell Rep. 115109. https://doi.org/10.1016/j.celrep.2024.115109 (2024).
Ijezie, E. C. et al. Herpes simplex virus-1 infection alters microtubule-associated protein Tau splicing and promotes Tau pathology in neural models of Alzheimer’s disease. Brain Pathol. e70006. https://doi.org/10.1111/bpa.70006 (2025).
Sundstrom, J. et al. Herpes Simplex Virus 1 Infection of Human Brain Organoids and Pancreatic Stem Cell-Islets Drives Organoid-Specific Transcripts Associated with Alzheimer’s Disease and Autoimmune Diseases. Cells 13, 1978. https://doi.org/10.3390/cells13231978 (2024).
Otero-Garcia, M. et al. Molecular signatures underlying neurofibrillary tangle susceptibility in Alzheimer’s disease. Neuron 110, 2929–2948.e2928, https://doi.org/10.1016/j.neuron.2022.06.021 (2022).
Funes, S. et al. Expression of ALS-PFN1 impairs vesicular degradation in iPSC-derived microglia. Nat Commun. 15, 2497, https://doi.org/10.1038/s41467-024-46695-w (2024).
Liu, S. et al. Iterative transcription factor screening enables rapid generation of microglia-like cells from human iPSC. Nat Commun. 16, 5136, https://doi.org/10.1038/s41467-025-59596-3 (2025).
Ormel, P. R. et al. Microglia innately develop within cerebral organoids. Nat. Commun. 9, 4167. https://doi.org/10.1038/s41467-018-06684-2 (2018).
Bodnar, B. et al. Novel scalable and simplified system to generate microglia-containing cerebral organoids from human induced pluripotent stem cells. Front. Cell Neurosci. 15, 682272. https://doi.org/10.3389/fncel.2021.682272 (2021).
Samudyata Oliveira, A.O. et al. SARS-CoV-2 promotes microglial synapse elimination in human brain organoids. Mol. Psychiatry 27,3939–3950, https://doi.org/10.1038/s41380-022-01786-2 (2022).
Gumbs, S. B. H. et al. Characterization of HIV-1 infection in microglia-containing human cerebral organoids. Viruses 14. https://doi.org/10.3390/v14040829 (2022).
Fagerlund, I. et al. Microglia-like cells promote neuronal functions in cerebral organoids. Cells 11, https://doi.org/10.3390/cells11010124 (2021).
Dos Santos, S. E. et al. Similar microglial cell densities across brain structures and mammalian species: implications for brain tissue function. J. Neurosci. 40, 4622–4643, https://doi.org/10.1523/JNEUROSCI.2339-19.2020 (2020).
Imfeld, P., Toovey, S., Jick, S. S. & Meier, C. R. Influenza infections and risk of Alzheimer’s disease. Brain Behav. Immun. 57, 187–192, https://doi.org/10.1016/j.bbi.2016.03.014 (2016).
Lim, E. T. et al. Orgo-Seq integrates single-cell and bulk transcriptomic data to identify cell-type specific-driver genes associated with autism spectrum disorder. Nat. Commun. 13, 3243. https://doi.org/10.1038/s41467-022-30968-3 (2022).
Wu, X. et al. Intrinsic immunity shapes viral resistance of stem cells. Cell 172, 423–438.e425, https://doi.org/10.1016/j.cell.2017.11.018 (2018).
Swartzendruber, D. E. & Lehman, J. M. Neoplastic differentiation: interaction of simian virus 40 and polyoma virus with murine teratocarcinoma cells in vitro. J. Cell Physiol. 85, 179–187, https://doi.org/10.1002/jcp.1040850204 (1975).
Friedrich, R. P. et al. Mechanism of amyloid plaque formation suggests an intracellular basis of Abeta pathogenicity. Proc. Natl. Acad. Sci. USA 107, 1942–1947, https://doi.org/10.1073/pnas.0904532106 (2010).
Huang, L. et al. Intracellular amyloid toxicity induces oxytosis/ferroptosis regulated cell death. Cell Death Dis. 11, 828. https://doi.org/10.1038/s41419-020-03020-9 (2020).
Caramello, A. et al. Intracellular accumulation of amyloid-ss is a marker of selective neuronal vulnerability in Alzheimer’s disease. Nat. Commun. 16, 5189. https://doi.org/10.1038/s41467-025-60328-w (2025).
Sasmita, A. O. et al. Oligodendrocytes produce amyloid-beta and contribute to plaque formation alongside neurons in Alzheimer’s disease model mice. Nat. Neurosci. 27, 1668–1674, https://doi.org/10.1038/s41593-024-01730-3 (2024).
Spires-Jones, T. L. & Hyman, B. T. The intersection of amyloid beta and tau at synapses in Alzheimer’s disease. Neuron 82, 756–771, https://doi.org/10.1016/j.neuron.2014.05.004 (2014).
Sollvander, S. et al. Accumulation of amyloid-beta by astrocytes result in enlarged endosomes and microvesicle-induced apoptosis of neurons. Mol. Neurodegener. 11, 38, https://doi.org/10.1186/s13024-016-0098-z (2016).
Tcw, J. et al. Cholesterol and matrisome pathways dysregulated in astrocytes and microglia. Cell 185, 2213–2233 e2225, https://doi.org/10.1016/j.cell.2022.05.017 (2022).
Tcw, J. & Arranz, A. M. hiPSC-based models to decipher the contribution of human astrocytes to Alzheimer’s disease and potential therapeutics. Mol. Neurodegener. 18, 19, https://doi.org/10.1186/s13024-023-00612-9 (2023).
Knopman, D. S., Jones, D. T. & Greicius, M. D. Failure to demonstrate efficacy of aducanumab: An analysis of the EMERGE and ENGAGE trials as reported by Biogen, December 2019. Alzheimers Dement. 17, 696–701, https://doi.org/10.1002/alz.12213 (2021).
Sevigny, J. et al. The antibody aducanumab reduces Abeta plaques in Alzheimer’s disease. Nature 537, 50–56, https://doi.org/10.1038/nature19323 (2016).
Budd Haeberlein, S. et al. Two randomized phase 3 studies of aducanumab in early Alzheimer’s disease. J. Prev. Alzheimer's Dis. 9, 197–210, https://doi.org/10.14283/jpad.2022.30 (2022).
Sperling, R. A. et al. Trial of solanezumab in preclinical Alzheimer’s disease. N. Engl. J. Med. 389, 1096–1107, https://doi.org/10.1056/NEJMoa2305032 (2023).
Honig, L. S. et al. Trial of solanezumab for mild dementia due to Alzheimer’s disease. N. Engl. J. Med. 378, 321–330, https://doi.org/10.1056/NEJMoa1705971 (2018).
Barthelemy, N. R. et al. A soluble phosphorylated tau signature links tau, amyloid and the evolution of stages of dominantly inherited Alzheimer’s disease. Nat. Med. 26, 398–407, https://doi.org/10.1038/s41591-020-0781-z (2020).
Mila-Aloma, M. et al. Plasma p-tau231 and p-tau217 as state markers of amyloid-beta pathology in preclinical Alzheimer’s disease. Nat. Med. 28, 1797–1801, https://doi.org/10.1038/s41591-022-01925-w (2022).
Jonaitis, E. M. et al. Plasma phosphorylated tau 217 in preclinical Alzheimer’s disease. Brain Commun. 5, fcad057. https://doi.org/10.1093/braincomms/fcad057 (2023).
Xia, Y., Prokop, S. & Giasson, B. I. Don’t Phos Over Tau”: recent developments in clinical biomarkers and therapies targeting tau phosphorylation in Alzheimer’s disease and other tauopathies. Mol. Neurodegener. 16, 37, https://doi.org/10.1186/s13024-021-00460-5 (2021).
Ashton, N. J. et al. Diagnostic accuracy of a plasma phosphorylated Tau 217 immunoassay for Alzheimer's disease pathology. JAMA Neurol. 81, 255–263, https://doi.org/10.1001/jamaneurol.2023.5319 (2024).
Arranz, J. et al. Diagnostic performance of plasma pTau(217), pTau(181), Abeta(1-42) and Abeta(1-40) in the LUMIPULSE automated platform for the detection of Alzheimer disease. Alzheimers Res Ther. 16, 139, https://doi.org/10.1186/s13195-024-01513-9 (2024).
Lewczuk, P. et al. Cerebrospinal fluid Abeta42/40 corresponds better than Abeta42 to Amyloid PET in Alzheimer’s disease. J. Alzheimers Dis. 55, 813–822, https://doi.org/10.3233/JAD-160722 (2017).
Constantinides, V. C. et al. CSF Abeta42 and Abeta42/Abeta40 ratio in Alzheimer’s disease and frontotemporal dementias. Diagnostics 13. https://doi.org/10.3390/diagnostics13040783 (2023).
Osborn, L. M., Kamphuis, W., Wadman, W. J. & Hol, E. M. Astrogliosis: an integral player in the pathogenesis of Alzheimer’s disease. Prog. Neurobiol. 144, 121–141, https://doi.org/10.1016/j.pneurobio.2016.01.001 (2016).
DeTure, M. A. & Dickson, D. W. The neuropathological diagnosis of Alzheimer’s disease. Mol. Neurodegener. 14, 32, https://doi.org/10.1186/s13024-019-0333-5 (2019).
Beach, T. G., Walker, R. & McGeer, E. G. Patterns of gliosis in Alzheimer’s disease and aging cerebrum. Glia 2, 420–436, https://doi.org/10.1002/glia.440020605 (1989).
Serrano-Pozo, A., Frosch, M. P., Masliah, E. & Hyman, B. T. Neuropathological alterations in Alzheimer's disease. Cold Spring Harb. Perspect. Med. 1, a006189, https://doi.org/10.1101/cshperspect.a006189 (2011).
Walker, D. G. et al. Patterns of expression of purinergic receptor P2RY12, a putative marker for non-activated microglia, in aged and alzheimer’s disease brains. Int. J. Mol. Sci. 21. https://doi.org/10.3390/ijms21020678 (2020).
Buniello, A. et al. The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 47, D1005–D1012, https://doi.org/10.1093/nar/gky1120 (2019).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550, https://doi.org/10.1073/pnas.0506580102 (2005).
Mootha, V. K. et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273, https://doi.org/10.1038/ng1180 (2003).
Mangold, C. A., Rathbun, M. M., Renner, D. W., Kuny, C. V. & Szpara, M. L. Viral infection of human neurons triggers strain-specific differences in host neuronal and viral transcriptomes. PLoS Pathog. 17, e1009441, https://doi.org/10.1371/journal.ppat.1009441 (2021).
Gulen, M. F. et al. cGAS-STING drives ageing-related inflammation and neurodegeneration. Nature 620, 374–380, https://doi.org/10.1038/s41586-023-06373-1 (2023).
Delhaye, S. et al. Neurons produce type I interferon during viral encephalitis. Proc. Natl. Acad. Sci. USA 103, 7835–7840, https://doi.org/10.1073/pnas.0602460103 (2006).
Ruhl, C. R. et al. Mycobacterium tuberculosis sulfolipid-1 activates nociceptive neurons and induces cough. Cell 181, 293–305 e211, https://doi.org/10.1016/j.cell.2020.02.026 (2020).
Chiu, I. M. et al. Bacteria activate sensory neurons that modulate pain and inflammation. Nature 501, 52–57, https://doi.org/10.1038/nature12479 (2013).
Heath, J. R. & Dembowski, J. A. Fashionably late: temporal regulation of HSV-1 late gene transcription. PLoS Pathog. 18, e1010536, https://doi.org/10.1371/journal.ppat.1010536 (2022).
Pesola, J. M., Zhu, J., Knipe, D. M. & Coen, D. M. Herpes simplex virus 1 immediate-early and early gene expression during reactivation from latency under conditions that prevent infectious virus production. J. Virol. 79, 14516–14525, https://doi.org/10.1128/JVI.79.23.14516-14525.2005 (2005).
Sundstrom, J. et al. Herpes simplex virus 1 infection of human brain organoids and pancreatic stem cell-islets drives organoid-specific transcripts associated with alzheimer’s disease and autoimmune diseases. Cells 13, 1978 (2024).
Tanaka, Y., Cakir, B., Xiang, Y., Sullivan, G. J. & Park, I. H. Synthetic analyses of single-cell transcriptomes from multiple brain organoids and fetal brain. Cell Rep. 30, 1682–1689.e1683, https://doi.org/10.1016/j.celrep.2020.01.038 (2020).
Pfefferkorn, C. et al. Abortively infected astrocytes appear to represent the main source of interferon beta in the virus-infected brain. J. Virol. 90, 2031–2038, https://doi.org/10.1128/JVI.02979-15 (2016).
Embry, A. & Gammon, D. B. Abortive infection of animal cells: what goes wrong. Annu. Rev. Virol. 11, 193–213, https://doi.org/10.1146/annurev-virology-100422-023037 (2024).
Feige, L., Zaeck, L. M., Sehl-Ewert, J., Finke, S. & Bourhy, H. Innate immune signaling and role of glial cells in herpes simplex virus- and rabies virus-induced encephalitis. Viruses 13. https://doi.org/10.3390/v13122364 (2021).
Neff, R. A. et al. Molecular subtyping of Alzheimer’s disease using RNA sequencing data reveals novel mechanisms and targets. Sci. Adv. 7. https://doi.org/10.1126/sciadv.abb5398 (2021).
Smullen, M. et al. Modeling of mitochondrial genetic polymorphisms reveals induction of heteroplasmy by pleiotropic disease locus 10398A>G. Sci. Rep. 13, 10405. https://doi.org/10.1038/s41598-023-37541-y (2023).
Smullen, M. et al. Reliable multiplex generation of pooled induced pluripotent stem cells. Cell Rep. Methods 3, 100570. https://doi.org/10.1016/j.crmeth.2023.100570 (2023).
Chan, Y. et al. Enabling multiplexed testing of pooled donor cells through whole-genome sequencing. Genome Med 10, 31. https://doi.org/10.1186/s13073-018-0541-6 (2018).
Kinney, J. W. et al. Inflammation as a central mechanism in Alzheimer’s disease. Alzheimers Dement. 4, 575–590, https://doi.org/10.1016/j.trci.2018.06.014 (2018).
Heneka, M. T. et al. Neuroinflammation in Alzheimer’s disease. Lancet Neurol. 14, 388–405, https://doi.org/10.1016/S1474-4422(15)70016-5 (2015).
Rodriguez, J. J., Olabarria, M., Chvatal, A. & Verkhratsky, A. Astroglia in dementia and Alzheimer’s disease. Cell Death Differ. 16, 378–385, https://doi.org/10.1038/cdd.2008.172 (2009).
Eimer, W. A. et al. Phosphorylated tau exhibits antimicrobial activity capable of neutralizing herpes simplex virus 1 infectivity in human neurons. Nat. Neurosci. https://doi.org/10.1038/s41593-025-02157-0 (2025).
Chan, Y. et al. An unbiased index to quantify participant’s phenotypic contribution to an open-access cohort. Sci. Rep. 7, 46148. https://doi.org/10.1038/srep46148 (2017).
Macdonald, S. J., Mostafa, H. H., Morrison, L. A. & Davido, D. J. Genome sequence of herpes simplex virus 1 strain KOS. J. Virol. 86, 6371–6372, https://doi.org/10.1128/jvi.00646-12 (2012).
Desai, P. & Person, S. Incorporation of the green fluorescent protein into the herpes simplex virus type 1 capsid. J. Virol. 72, 7563–7568, https://doi.org/10.1128/JVI.72.9.7563-7568.1998 (1998).
Colgrove, R. C. et al. History and genomic sequence analysis of the herpes simplex virus 1 KOS and KOS1.1 sub-strains. Virology 487, 215–221, https://doi.org/10.1016/j.virol.2015.09.026 (2016).
Brownlee, G. G. & Fodor, E. The predicted antigenicity of the haemagglutinin of the 1918 Spanish influenza pandemic suggests an avian origin. Philos. Trans. R. Soc. Lond. B Biol. Sci. 356, 1871–1876, https://doi.org/10.1098/rstb.2001.1001 (2001).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379, https://doi.org/10.1038/nature12517 (2013).
Dawes, P. et al. oFlowSeq: a quantitative approach to identify protein coding mutations affecting cell type enrichment using mosaic CRISPR-Cas9 edited cerebral organoids. Hum Genet. https://doi.org/10.1007/s00439-023-02534-4 (2023).
Lancaster, M. A. & Knoblich, J. A. Generation of cerebral organoids from human pluripotent stem cells. Nat. Protoc. 9, 2329–2340, https://doi.org/10.1038/nprot.2014.158 (2014).
Lund-Johansen, F. A strong case for third-party testing. Elife 12. https://doi.org/10.7554/eLife.93329 (2023).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272, https://doi.org/10.1038/s41592-019-0686-2 (2020).
R Development Core Team, V.A. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2010).
Lagomarsino, V. N. et al. Stem cell-derived neurons reflect features of protein networks, neuropathology, and cognitive outcome of their aged human donors. Neuron 109, 3402–3420 e3409, https://doi.org/10.1016/j.neuron.2021.08.003 (2021).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis. ISBN 978-3-319-24277-4, http://ggplot2.org (Springer-Verlag New York, 2016).
Andrews, S. FastQC: a quality control tool for high-throughput sequence data. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (2020).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120, https://doi.org/10.1093/bioinformatics/btu170 (2014).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915, https://doi.org/10.1038/s41587-019-0201-4 (2019).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079, https://doi.org/10.1093/bioinformatics/btp352 (2009).
Kovaka, S. et al. Transcriptome assembly from long-read RNA-seq alignments with StringTie2. Genome Biol. 20, 278. https://doi.org/10.1186/s13059-019-1910-1 (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550. https://doi.org/10.1186/s13059-014-0550-8 (2014).
Blighe, K., Rana, S. & Lewis, M. EnhancedVolcano: Publication-ready volcano plots with enhanced colouring and labeling. https://github.com/kevinblighe/EnhancedVolcano (2018).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502, https://doi.org/10.1038/nbt.3192 (2015).
Aran, D., Looney, A.P. & Liu, L. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat Immunol 20, 163–172, https://doi.org/10.1038/s41590-018-0276-y (2019).
Alquicira-Hernandez, J., Sathe, A., Ji, HP., Nguyen, Q. & Powell, JE. scPred: accurate supervised method for cell-type classification from single-cell RNA-seq data. Genome Biol 20, 264, https://doi.org/10.1186/s13059-019-1862-5 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15. https://doi.org/10.1186/s13059-017-1382-0 (2018).
Acknowledgements
This study was supported by the National Institutes of Health (NIH) grants U01AG061835 (PI: B.R.; Sub-contract PIs: George M. Church, E.T.L., Y.C.), R01AG083881 (PI: E.T.L.), U01AG088673 (PI: Y.C.), P01AI09681 (PI: D.M.K), T32AI007349 (Student: A.R.O.) and a grant from the PolyBio Foundation (PIs: Y.C., E.T.L.). We thank Aimee Alyward and Tracy Young-Pearse for running the ELISA assays. We are grateful to Manuel Garber and Khashayar Afshari for discussions on optimizing our flow cytometry protocols, as well as Fiachra Humphries, Douglas Golenbock, Michael Heneka and Katherine Fitzgerald for suggestions, feedback and generating pilot data.
Author information
Authors and Affiliations
Contributions
E.T.L. and Y.C. conceived the project. N.J.B., P.D., C.S., V.R.U., Q.W., B.R., Y.C. and E.T.L. designed and performed the computational analyses and methods development. M.N.O., L.F., S.M.C., K.T., A.R.O., J.S., L.F.M., Y.C. and E.T.L. designed and performed the cerebral organoid differentiation and dissociation, RNA extraction and quality control. H.S.O., D.M.K., Y.C. and E.T.L. designed and performed the initial HSV-1 experiments at Harvard Medical School. Y.C. designed and performed the initial IAV experiments at Harvard Medical School. H.S.O. and D.M.K. provided stocks of HSV-1 and acyclovir to our group at UMass Chan Medical School. M.N.O., L.F., S.M.C., K.T., A.R.O., N.J.B., J.S., L.F.M., Y.C. and E.T.L. designed and performed the flow cytometry experiments and analyses. L.F.M. and E.T.L. designed the ELISA experiments and analyses. K.T., N.J.B., P.D., Y.H., L.F.M., Y.C. and E.T.L. designed and conducted the scRNA-seq experiments, sample preparation, library preparation and analyses. J.M.J. and M.H.O. conducted the plaque assays with UV-HSV-1. L.F., K.T., C.S., V.R.U. conducted experiments and analyses with the A3P dcOrgs. M.N.O., S.M.C., K.T., A.R.O., N.J.B., P.D., Y.C. and E.T.L. wrote and revised the manuscript with edits from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial or nonfinancial interests. E.T.L. and Y.C. are Guest Editors of Cells/Genetic Mechanisms in Cellular Genomics and Therapeutics. E.T.L. is an Associate Editor of Frontiers in Genetics/Genetics of Common and Rare Diseases. E.T.L. and Y.C. were not involved in the journal’s review of, or decisions related to, this manuscript.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Olson, M.N., Barton, N.J., Feng, L. et al. A high-throughput, quantitative platform using 2D dissociated human cerebral organoids to model neuroinflammation in Alzheimer’s disease. npj Dement. 2, 20 (2026). https://doi.org/10.1038/s44400-026-00066-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44400-026-00066-y








