Abstract
Cancer metastasis is the leading cause of cancer-related death. While organs such as the lung are hotspots for metastases, others -like skeletal muscle- remain rarely colonized, a phenomenon that remains poorly understood. In this study, we show that EO771 breast cancer cells proliferated robustly when co-cultured with MLg lung stromal cells, whereas their proliferation was restrained when maintained in direct contact with differentiated C2C12 skeletal muscle myotubes. Notably, these effects were not cell-type-specific, as similar results were obtained with 4T1 breast cancer cells and Sol8 myotubes. After two days of co-culture, both cancer and host cells (MLg and C2C12) exhibited distinct niche-specific transcriptional remodeling. Strikingly, the poorly proliferative EO771 cells co-cultured with C2C12 myotubes acquired a hypoxia-associated gene-expression signature despite normoxic conditions (~20% O₂), showing that muscle cells reprogram cancer cells into a hypoxic, anti-proliferative state. Under hypoxic conditions, we confirmed that the depletion of oxygen allows C2C12 cells to nearly abolish EO771 proliferation. Neither exogenous lactate, culture acidosis, their combination, altered glucose levels, nor conditioned medium could reproduce the suppressive environment created by C2C12 myotubes. In contrast, MLg cells induced minimal transcriptional changes in EO771 cells and were themselves broadly reprogrammed by the cancer cells. Moreover, hypoxia enhanced EO771 proliferation in MLg co-cultures, emphasizing the permissive nature of the MLg environment. Collectively, these findings uncover a unique, paradoxical, muscle-induced pseudo-hypoxic program that restricts cancer cell proliferation. They also highlight the need for caution in targeting hypoxia signaling in anti-metastatic therapies, as such interventions could weaken skeletal muscle’s natural defense against tumor colonization.
Similar content being viewed by others
Introduction
Cancer metastasis, the spread of cancerous tumor cells from the primary tumor site to distant tissues, is responsible for over 90% of cancer-related deaths [1]. For successful metastases, cancer cells must navigate a complex series of biological processes - such as metabolic adaptation [2,3,4,5,6], interaction with the extracellular matrix [7,8,9,10,11], and immune evasion [12,13,14] - to survive, migrate, and establish colonies in distant organs. While most studies have focused on how metastatic tumor cells can colonize permissive sites like the lung, far less is known about the incapacity of cancer cells to adapt to certain tissues. Exploring these resistant environments could uncover novel metastasis-suppressive mechanisms and provide new therapeutic horizons.
While most organs, such as the lung, liver, or brain, are common sites for cancer cell colonization, skeletal muscle stands out as one of the least colonized, despite representing ~30–40% of total body weight [15]. Among 3827 individuals with 41 different primary cancers, only 16 patients (0.4%) revealed skeletal muscle metastases [16]. Additionally, carcinoma metastasis to skeletal muscle was observed in 4 and 15 patients over a 15- and 19-year period, respectively [17, 18]. Despite this striking rarity, the mechanisms underlying the resistance of skeletal muscle to metastatic colonization remain poorly understood, and only a very small number of studies have been performed in this research field.
Blood flow and hemodynamics have been proposed as factors limiting the ability of circulating tumor cells to colonize skeletal muscle [19]. However, this explanation is unlikely, as cancer cells like melanoma cells can reach skeletal muscle in vivo [20]. Contraction-derived mechanical forces have been proposed to impede metastatic colonization [21], but their episodic, activity-dependent nature makes durable protection uncertain. Additionally, myokines with anti-cancer activity have been suggested as contributors to the tumor-suppressive milieu of muscle [22]. More recently, elevated reactive oxygen species (ROS) in skeletal muscle have been proposed to contribute to cancerous tumor suppression [23], findings that remain to be confirmed. Interestingly, there also exists evidence that metastatic tumor cells undergo transcriptional reprogramming to adapt to their new environments [24, 25]. However, it is still unclear how the transcriptomes of cancer cells are influenced by permissive (e.g., lung) versus hostile (e.g., skeletal muscle) environments - and conversely, how tumor cells influence the transcriptomic landscapes of host cells. Deciphering these reciprocal interactions may uncover novel tissue-specific mechanisms of metastatic resistance and, more broadly, advance our understanding of cancer metastasis.
In this study, we employed a co-culture model in which cancer cells are maintained directly on either MLg cells (lung-derived) or C2C12 myotubes (skeletal muscle-derived). Our approach was to use cell lines to facilitate comparison between different tissue-derived cells, distinguishing permissive versus hostile environments for cancer cells. We selected the myoblast line C2C12 as the skeletal muscle model because these cells retain a robust capacity to proliferate as myoblasts and differentiate into multinucleated myotubes, recapitulating key aspects of skeletal muscle biology, and are widely used in skeletal muscle research. We show that EO771 breast cancer cells undergo extensive niche-dependent transcriptomic reprogramming, with the most profound changes occurring in the proliferation-restrictive environment of C2C12 myotubes. Notably, the C2C12 myotubes cause the cancer cells to acquire a pseudo-hypoxic gene-expression reprogramming, and under hypoxia, EO771 proliferation was almost completely abolished on C2C12 myotubes. In contrast to the EO771 cancer cells, host cells display more modest transcriptional modifications, though with varying degrees of resilience: C2C12 myotubes maintain a relatively stable transcriptomic identity, whereas MLg cells undergo broader reprogramming in response to tumor cell interaction, indicative of a more permissive and reactive environment. Taken together, our findings reveal that skeletal muscle cells can reprogram cancer cells toward a hypoxic, anti-proliferative state -an unexpected outcome given hypoxia’s well-established pro-metastatic role in most tissues.
Material and methods
Cells
C2C12 mouse myoblasts (CRL-1772), Sol8 mouse myoblasts (CRL-2174), and MLg mouse lung stroma cells (CCL-206) were obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA). EO771 (C57BL6 genetic background) and 4T1 (BALB/c genetic background) mouse breast cancer cells tagged with GFP were a gift from Dr Cyrus M Ghajar (Cancer Research Center, Seattle, WA, USA). All cell lines were tested for mycoplasma contamination using a MycoAlert Mycoplasma Detection kit (LT07-318, Lonza, Basel, Switzerland).
Co-cultures
C2C12, Sol8 and MLg cells were maintained in Dulbecco’s Modified Eagle Medium (DMEM, #11995-065, Gibco, Grand Island, NY, USA) supplemented with 10% Fetal Bovine Serum (#10091148, Invitrogen, Waltham, MA, USA) and 1% penicillin-streptomycin (Pen/Strep; #15140-122, Gibco) at 37 °C for 2 days. At 80–90% confluency, cells were switched to differentiation medium [DMEM with 2% horse serum (#16050-122, Gibco) and 1% Pen/Strep] for 4 days to induce C2C12 and Sol8 myotube formation. Both C2C12, Sol8, and MLg cells were cultured in 2% horse serum to avoid experimental variability between the different cell types. Then, GFP-tagged EO771 or 4T1 breast cancer cells were seeded onto the differentiated C2C12/Sol8 myotubes or confluent MLg cells in DMEM. After 15 min at room temperature, 20% (final concentration) Cultrex basement membrane extract PathClear (#3433-005-01, Bio-Techne, Minneapolis, MN, USA), diluted in DMEM, was added to each well. The matrix was allowed to settle at room temperature for 15 min before polymerizing at 37 °C. Co-cultures were maintained for 2 days in DMEM, with the medium replaced after 24 h. An overview of the co-cultures is shown in Fig. 1A.
Monocultures
Parental C2C12 and MLg cells were cultured individually under the same conditions used in the co-culture experiments. Briefly, cells were maintained in growth medium for 2 days, differentiation medium for 4 days, and in DMEM with 20% Cultrex for 2 additional days. Parental EO771 cells were cultured in DMEM with 20% Cultrex for 2 days. Phase-contrast microscopy images of monocultures were acquired using an Eclipse Ts2 microscope (Nikon, Tokyo, Japan), operated with NIS-Elements software (version 5.42.06, Nikon).
Cell staining
After 4 days of differentiation, the cell medium was discarded, and C2C12 and MLg cells were rinsed twice with Phosphate-Buffered Saline (PBS, #P4417, Sigma-Aldrich, Saint-Louis, MO, USA). Then, cells were fixed with methanol (#1.06009.2511, Sigma-Aldrich) for 6 min with gentle agitation, rinsed with PBS before being stained with May-Grünwald (#63590, Sigma-Aldrich) and Giemsa (#109204, Sigma-Aldrich). Phase-contrast microscopy images were acquired using a CKX53 microscope with a UC90 color camera (Olympus, Tokyo, Japan) and a 10x objective, operated with Olympus cellSens software (version 2.2, Olympus).
Quantification of cancer cell attachment and outgrowth
Imaging of 96-well plates was performed using the IncuCyte Live-Cell Imaging System (IncuCyte S3, Sartorius, Göttingen, Germany) equipped with a 10× objective in the green fluorescence channel. Four images were acquired per well, starting 3 h after seeding GFP-labeled EO771 cells onto MLg cells or C2C12 myotubes, and subsequently at 24 h intervals. The initial attachment of EO771 cells at day 0 (D0) was quantified as the number of fluorescent cells per mm², whereas the outgrowth of GFP-labeled EO771 or 4T1 cells was quantified as the percentage of fluorescent area per well. Unless otherwise stated in the figure legends, all area measurements were normalized to D0, corresponding to the time of tumor cell attachment. Image acquisition and analysis were conducted using IncuCyte software (version 2021A).
FACS
Monoculture and co-culture experiments were performed in 6-well plates with C2C12 and MLg (200,000 cells/well) and EO771 (120,000 cells/well). Two days after the seeding of EO771, cells were washed with PBS and Cultrex removed with Cultrex Organoid Harvesting Solution (#3700-100-01, Bio-Techne), before being harvested with Trypsin-EDTA (#T3924, Sigma-Aldrich). Cells were then diluted in PBS to 2 million cells per ml before sorting with a FACSMelody cell sorter (BD Biosciences, Franklin Lakes, NJ, USA) using a 488 nm laser to detect GFP and a 100 \(\mathrm{\mu m}\) nozzle. The distinct cell populations were determined by GFP area (GFP-A) and side scatter area (SSC-A). Sorting efficiency (number of sorted cells/number of identified cells x100) consistently exceeded 80%. To confirm the absence of cross-contamination, a fraction of sorted cells was cultured for 3 days to verify that green fluorescence was only observed in the isolated GFP-labeled EO771 cells. Phase-contrast and fluorescence (LED unit: 470 nm) microscopy images of the cells were acquired by an Eclipse Ts2 microscope with a 10× objective. Sorted cells were used for RNA isolation and RNA-seq, as described below.
RNA isolation
FACS-isolated cells were immediately pelleted by centrifugation, and total RNA was extracted using the RNeasy Mini Kit (#74106, Qiagen, Hilden, Germany) according to the manufacturer’s instructions. RNA purity was assessed using a NanoDrop (Thermo Fisher Scientific), and RNA integrity was evaluated with the TapeStation 4150 system (Agilent Technologies, Santa Clara, CA, USA). All samples had RNA Integrity Number (RIN) values greater than 8.5, indicating high-quality RNA suitable for downstream transcriptomic analysis. For each sample, total RNA was diluted to obtain 500 ng RNA in a final volume of 100 µl.
RNA-seq and analyses
mRNA was prepared for sequencing from total RNA using the TruSeq Stranded mRNA Library Kit (Illumina, San Diego, CA, USA). Sequencing was performed using the NovaSeqXPlus platform with 150 bp paired-end reads at the Norwegian Sequencing Center. RNA-seq reads were filtered to remove low-quality reads by fastp (v0.23.2) [26]. Filtered reads were aligned to the mouse genome GRCm38vM21 with STAR (v2.7.10a) [27] using the Gencode annotation (gencode.vM21). Transcript abundance was estimated using featureCounts in Subread v2.0.1 [28]. DESeq2 (v 1.36.0) [29], as implemented in SARTools [30], was used to normalize the raw counts, apply exploratory analysis (e.g., principal component analysis, PCA), and to perform differential expression gene analysis. Gene set enrichment analysis (GSEA) was performed on normalized read counts [31]. Gene ranking was generated across each comparison with the Pearson correlation metric and analyzed against the mSigDB Hallmark v2024.1.Mm gene sets [32].
Gene expression
Analyses were performed based on [33]. Briefly, cDNA was prepared from RNA using the High-Capacity cDNA Reverse Transcription kit (4368814, Thermo Fisher Scientific). Gene expression was determined by qRT-PCR with Bio-Rad iQTMSYBR® Green supermix (1708886, Bio-Rad, Hercules, CA, USA) using CFX connectTM Real time system and CFX Maestro 2.2 (Version 5.2.008.0222, Bio-Rad). Samples were run in duplicate. Reactions yielding Ct values above 40 in both duplicates were excluded from further analyses. The efficiency corrected method with Rplpo as a reference gene was used for analyses. Primer sequences are shown in Table S7.
Hypoxia
Monocultured and co-cultured cells were grown in a cell incubator under hypoxic conditions (10%, 3%, or 1% O2) for 2 days.
Western blot
Cells were lysed in SDS buffer (25 mM Tris, 1% SDS, 10% glycerol), sonicated with BioRuptor (Diagenode, Belgium), and centrifuged (12,000 × g) for 5 min. Protein concentration was determined with the BCA Protein Assay Kit (#23225, Thermo Fisher Scientific). Samples were diluted in Laemmli buffer (64 mM Tris-HCl pH6.8, 2% SDS, 10% glycerol, 5% β-mercaptoethanol, 0.02% bromophenol blue) and 10 μg protein was loaded on a 4-20% SDS-gel (#5671093, Bio-Rad). After gel electrophoresis, proteins were transferred onto Immobilon-P PVDF membrane (IPFL0010, Merck Millipore, Burlington, MA, USA). Membranes were blocked for 1 h with 5% non-fat milk and incubated overnight at 4 °C with hypoxia inducible factor-1α (HIF-1α) rabbit primary antibody (1:1000, #36169, Cell Signaling, Danvers, MA, USA) or GAPDH primary antibody (1:2000, #2118S, Cell Signaling) dissolved in Tris-Buffered Saline with 0.1% Tween20 (TBS-T) and 3% BSA. After washing in TBS-T, membranes were incubated with HRP-conjugated goat anti-rabbit secondary antibody (1:10,000, #111-035-144, Nordic Biosite, Täby, Sweden) in TBS-T with 1% non-fat milk for 1 h. Chemiluminescence was detected with SuperSignal West Pico PLUS (#34578, Thermo Fisher Scientific) on a ChemiDoc imaging system (version 2.4.0.03, Bio-Rad).
Lactate and glucose measurement
Extracellular lactate levels were measured in culture supernatants using the L-Lactate Assay Kit (ab65331, Abcam, Cambridge, UK), following the manufacturer’s instructions. Extracellular glucose levels were measured using a glucometer (Freestyle Precision Neo, Abbott, Chicago, IL, USA).
pH measurement
pH levels were measured in the same culture supernatants used for lactate analysis using a pH meter (OrionStarA111, Thermo Fisher Scientific). To obtain sufficient volume, supernatants from multiple wells were pooled to a total volume of at least 2 ml. To avoid fluctuations in pH measurements, supernatants were maintained at room temperature and at atmospheric CO₂ level for approximately one hour before pH assessment, allowing the bicarbonate buffering system to equilibrate and minimizing artificial pH shifts caused by temperature variation and CO2 degassing from the medium [34].
Lactate, glucose, and pH treatment
DMEM supplemented with Na-L-lactate (#L14500.06, Thermo Fisher Scientific), NaCl (#1.06404, Supelco, Saint-Louis, MO, USA), D-(+)-Glucose (#G7528-250, Sigma-Aldrich), or adjusted to a reduced pH was prepared at room temperature under atmospheric CO₂ conditions. Both Na-L-lactate, NaCl, and D-(+)-Glucose were dissolved in PBS (vehicle control). Monocultured and co-cultured cells were exposed to DMEM containing Na-L-lactate or NaCl at either normal or reduced pH, and DMEM containing D-(+)-Glucose at normal pH. Treatment media were refreshed every 24 h.
Conditioned medium
Parental C2C12 and MLg cells were maintained as described previously in Monocultures. Briefly, cells were cultured in growth medium for 2 days, followed by differentiation medium for 4 days. Subsequently, cells were incubated for an additional 2 days in serum-free DMEM supplemented with 20% Cultrex. The culture medium collected after this 2-day serum-free incubation period was defined as conditioned medium. Conditioned medium was centrifuged (2000 × g) for 10 min to remove cell debris, passed through a 0.2 µm filter (514-0073, VWR, Radnor, PA, USA) to ensure sterility while retaining soluble factors, and immediately used to culture parental EO771 cells.
Statistical analysis
Data were analyzed with GraphPad Prism software (version 10.4.1). All data are represented as mean ± standard error of the mean (SEM). Normality was tested with the Shapiro–Wilk test. Statistical significance in monocultures and co-cultures with two or more variables was determined using a two-way ANOVA with Šidák (only two groups) or Tukey’s (more than two groups) multiple comparisons. Comparisons between two groups with one variable were done using a two-tailed unpaired Student’s t-test (for normalized data) and Mann–Whitney test (for non-normalized data). For all t-tests, equality of variances was tested by F-tests. P < 0.05 was considered statistically significant. No statistical methods were used to predetermine sample sizes.
Results
C2C12 myotubes restrain breast cancer cell outgrowth compared to MLg cells
We started our investigations by establishing a co-culture model based on [23], and examining whether C2C12 myotubes suppress tumor cell outgrowth relative to MLg cells (Fig. 1A). To this end, we seeded EO771 cells - a breast cancer cell line with metastatic potential commonly utilized in cancer research [35] - onto fully confluent C2C12 myotubes or MLg cells (Fig. 1B). After 3 h, the number of EO771 cells adhering to C2C12 myotubes and MLg cells was comparable (Fig. 1C), indicating uniform seeding and attachment across the two niche environments.
A Schematic representation of co-cultures. C2C12 and MLg cells were submitted to similar culture conditions for 6 days. At day 0 (D0), when C2C12 myotubes and MLg cells were fully confluent, EO771 cancer cells were seeded onto the cells. B Representative phase-contrast microscopy images of C2C12 and MLg stained with May-Grünwald and Giemsa at D0. Scale bar = 200 µm. C Number of EO771 cells per mm2 attached onto C2C12 myotubes (C2C12 + EO771) and MLg cells (MLg + EO771) 3 h post-seeding at D0. D Outgrowth of EO771 cells (100 cells per well in 96-well plate) co-cultured with C2C12 myotubes (C2C12 + EO771) or MLg cells (MLg + EO771) for 2 days. EO771 area represents the percentage of fluorescent cells normalized to D0 (n = 21–24 from three independent experiments). Representative fluorescence microscopy images of EO771 cell outgrowth on C2C12 or MLg at day 2 (D2). Scale bar = 200 µm. E Outgrowth of EO771 cells in co-cultures with C2C12 myotubes or MLg cells when seeded at 500, 1000, 4000, and 8000 cells per well in a 96-well plate at day 0. Green fluorescence of EO771 was measured over 2 days without normalization (n = 24 from two independent experiments). F Representative fluorescence microscopy images of EO771 cell outgrowth (4000 cells per well in 96-well plate) on C2C12 or MLg. Scale bar = 200 µm. Data are mean ± SEM. Normality was tested with the Shapiro–Wilk test. Comparison between groups was done using the Mann–Whitney test (comparing two groups with one variable) or two-way ANOVA adjusted for multiple testing with the Šidák method (comparing two groups with two variables). ****P < 0.0001.
Both C2C12 + EO771 and MLg + EO771 co-cultures were maintained similarly in supplement-free DMEM to avoid experimental variability and confounders that could potentially mask the proliferation-suppressive effects of the host cells. When EO771 cells were seeded (100 cells per well in a 96-well plate) onto C2C12 myotubes or MLg cells, we observed that EO771 cells proliferated significantly less in co-culture with C2C12 myotubes than with MLg cells (Fig. 1D). After one and two days, EO771 cell outgrowth on MLg cells was over twice as high as on C2C12 myotubes (Fig. 1D).
Then, we asked whether this suppressive effect was maintained when co-culturing myotubes with higher EO771 seeding densities. Even when EO771 input was increased to 500, 1000, 4000, or 8000 cells per well (96-well plate), C2C12 myotubes consistently refrained their proliferation relative to MLg cells (Fig. 1E). Across all seeding densities, EO771 outgrowth in co-culture with C2C12 myotubes was reduced by approximately 40–60% at both day 1 and day 2 compared with MLg co-cultures, demonstrating that the anti-proliferative effect of C2C12 myotubes remained proportionally consistent regardless of EO771 input. Additionally, EO771 cells adopted a typical elongated, myotube-like shape (Fig. 1F), whereas EO771 cells grown on MLg cells retained a more typical round morphology and were evenly intermixed (Fig. 1F). Further investigations showed that C2C12 myotubes and MLg cells before and after 2 days in co-cultures maintained their morphological structure (Fig. S1).
To determine whether these findings were reproducible with another breast cancer cell line, we repeated the co-culture experiments with 4T1 cells (Fig. S2A). Similar to EO771 cells, MLg cells supported robust 4T1 outgrowth (Fig. S2B). In contrast, 4T1 cell outgrowth was significantly suppressed in co-culture with C2C12 myotubes and accompanied by notable morphological changes (Fig. S2B, C). In additional experiments, we further confirmed that the suppression of EO771 cell outgrowth was not C2C12-cell specific, as the independent mouse skeletal muscle line, Sol8 (soleus-derived), suppressed cancer cell outgrowth relative to the MLg + EO771 co-culture (Fig. S3).
Thus, using a model that allows for controlled growth factor conditions and eliminates confounding influences such as physical barriers and immune cells, we demonstrated that myotubes create a hostile niche for breast cancer cell colonization, in contrast to MLg cells, which serve as a permissive environment.
Co-cultured cancer and host cells undergo distinct transcriptional reprogramming
To explore the molecular basis underlying the divergent proliferation of cancer cells in muscle versus lung environments, we performed RNA-seq on both tumor and host cells isolated from co-cultures (Fig. 2A). In addition, we included monocultured parental cells to identify baseline transcriptional profiles and distinguish co-culture-induced changes (Fig. 2A). To ensure comparability across all cell types, both co-cultured and monocultured cells were subjected to identical experimental conditions, including FACS-based isolation, prior to RNA-seq analysis (Fig. 2A). An overview of cell isolation and absence of cross-contamination is shown in Fig. S4.
A Schematic representation of FACS sorting of monocultures and co-cultures. B PCA plot of the different cell groups identified after mRNA sequencing (n = 4). C Scatter dot plot showing the distribution of up- and downregulated genes in each comparison (number of DEGs is shown in the figure). D–I Dot plots of enriched hallmarks from gene set enrichment analysis (GSEA). The color of the dot indicates statistical significance.
We addressed two main questions focused on cancer and host cell adaptations. First, do the EO771 cells co-cultured with C2C12 (EO771-C) or MLg (EO771-M) retain a cancer-like profile similar to the parental EO771 (EO771-P) cells or shift toward the identity of the host cells? Second, do the C2C12 and MLg host cells, co-cultured with EO771 cells (C2C12-E and MLg-E, respectively), maintain the transcriptomes of their parental cells (C2C12-P and MLg-P) or shift toward a cancer profile?
Using our RNA-seq data, PCA revealed seven distinct clusters corresponding to the different cell types and culture conditions (Fig. 2B). Regarding cancer cell adaptations, the PCA showed a shift in the co-cultured EO771 cells towards their host cells - C2C12 myotubes and MLg cells. The highest number of differentially expressed genes (DEGs) was found in EO771-C vs. EO771-P, with 5649 genes affected (Fig. 2C). In contrast, only 2035 DEGs were observed in EO771-M vs. EO771-P, less than half the number observed in EO771-C vs. EO771-P (Fig. 2C). Thus, compared to their parental cells, EO771 cells appear to undergo more extensive differential gene regulation when co-cultured with C2C12 myotubes than with MLg cells. A direct comparison between EO771-M vs. EO771-C confirmed that a large number of genes were differentially regulated in the EO771 cells (Fig. 2C), showing that the host cells, C2C12 and MLg, caused specific transcriptional reprogramming of cancer cells. GSEA further showed that EO771-C vs. EO771-P had 23 significantly enriched hallmark pathways (P < 0.01), including myogenesis and hypoxia as the top hits (Fig. 2D). In contrast, EO771-M had only four enriched hallmark pathways (interferon alpha response, angiogenesis, epithelial mesenchymal transition and interferon gamma response), indicating limited transcriptional reprogramming (Fig. 2E). We also noticed that all four EO771-M hallmark pathways were present in EO771-C (Fig. 2D), implying that the C2C12 niche induces both shared and unique adaptations. A direct comparison between EO771-C and EO771-M further revealed 15 differentially regulated pathways in EO771-C (Top 3: myogenesis, hypoxia, glycolysis) while EO771-M was characterized by four cell-cycle-related hallmark sets (Myc v1 and v2, E2F targets, and G2M checkpoint) (Fig. 2F), consistent with their robust proliferation on MLg cells (Fig. 1B).
Heatmaps of selected hallmark pathways further illustrated the distinct reprogramming of EO771 cells in muscle versus lung environments. EO771-C showed strong activation of myogenesis, hypoxia, and glycolysis gene clusters-signatures absent in EO771-M and EO771-P (Fig. S5A–C and Tables S1–S3), consistent with a shift in metabolic state. In contrast, hallmark pathways associated with proliferation -Myc targets, E2F targets, and G2M checkpoint- were overall downregulated in EO771-C but maintained in EO771-M compared to EO771-P (Fig. S5D–F and Tables S4–S6).
Then, we examined the transcriptional adaptations of the host cells when co-cultured with EO771 cancer cells. The PCA showed that both C2C12 and MLg cells co-cultured with EO771 cells (C2C12-E and MLg-E, respectively) diverged slightly from their parental monocultured cells, suggesting modest but distinct transcriptional reprogramming (Fig. 2B). Additional analysis revealed that C2C12-E displayed 1449 DEGs compared to C2C12-P, whereas MLg-E exhibited a broader transcriptional shift, with 2290 DEGs (+58%) compared to MLg-P (Fig. 2C). Furthermore, the distribution of DEGs showed a higher proportion of genes with large log2 fold change in MLg-E than in C2C12-E, suggesting that C2C12 myotubes maintain greater transcriptional stability upon interaction with tumor cells. Consistently, GSEA identified fewer significantly enriched hallmark pathways in C2C12-E (Fig. 2G) compared to MLg-E (Fig. 2H), when each co-cultured cell was analyzed relative to their parental cells (13 vs. 21 hallmark pathways, respectively). While most pathways were shared between both C2C12-E and MLg-E, the latter showed specific hallmarks, including complement, IL2-STAT5 signaling, p53 pathway, oxidative phosphorylation, apoptosis, UV response up, coagulation, and reactive oxygen species pathway (Fig. 2H). In line with the induction of inflammatory response and apoptosis hallmarks observed in MLg-E compared to MLg-P (Fig. 2H), we confirmed by qPCR that mRNA levels of several related markers were increased in MLg-E cells (Fig. S6A). Similarly, for C2C12 cells, the mRNA expression of inflammatory markers such as Tnfα was enhanced in C2C12-E relative to C2C12-P (Fig. S6B), whereas apoptosis marker expression was not increased, confirming the inflammatory response activation in C2C12-E (Fig. 2G). Finally, in a direct comparison between C2C12-E and MLg-E, C2C12-E retained hallmark pathways related to myogenic identity, whereas MLg-E showed significantly regulated pathways associated with cell-cycle regulation and proliferation, such as E2F targets, G2M checkpoint, MYC targets V1 and V2 (Fig. 2I). These findings suggest that C2C12-E myotubes retain a relatively stable transcriptomic identity upon co-culture with cancer cells, in contrast to MLg-E cells, which undergo more comprehensive transcriptional reprogramming. Heatmaps of genes further supported this interpretation, showing that expression patterns in C2C12-E and MLg-E clustered closely with their respective parental controls, with transcriptional changes being more extensive in MLg cells (Fig. S5A–F).
Thus, after two days of co-cultures, EO771 cancer cells undergo niche-specific transcriptomic reprogramming, with the most pronounced shift occurring in the proliferative-restrictive environment of C2C12 myotubes. In contrast, host cells exhibit more modest transcriptional adaptations, albeit with differing levels of resilience: C2C12 myotubes largely preserve their parental transcriptomic identity, whereas MLg cells display broader reprogramming in response to cancer cell interaction, likely reflecting a more permissive and reactive niche.
Hypoxia boosts the non-permissive capacity of C2C12 myotubes
Given that the proliferative-restricted EO771 cells co-cultured with C2C12 myotubes were characterized by a pseudo-hypoxic gene reprogramming, we hypothesized that oxygen deprivation could further potentiate the anti-metastatic properties of muscle cells – an effect that would stand in stark contrast to the recognized role of hypoxia in driving cancerous tumor proliferation [36, 37].
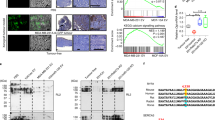
To test this hypothesis, we subjected our cell cultures to a reduced level of O₂. Under 3% O₂, HIF-1α protein levels increased robustly in monocultures of parental EO771, MLg, and C2C12 cells, as well as in co-cultures (C2C12 + EO771 and MLg + EO771), compared with normoxia (Fig. 3A), confirming an appropriate hypoxic response across all conditions. We then examined the effect of hypoxia in conditions that replicated our co-culture experiments (Fig. 3B); EO771 cells were seeded at 4000 cells per well (96-well plate) on day 0 (D0), whereas MLg and C2C12 cells were seeded at 8000 cells per well six days prior to D0 to allow full differentiation of C2C12 myoblasts into myotubes. Due to inherent differences in cell number and growth kinetics among the three cell types, we focused our analysis on the relative changes induced by hypoxia versus normoxia within each individual cell type. In C2C12 myotubes, hypoxia significantly increased lactate production (Fig. 3C), glucose consumption (Fig. 3D) and was also associated with a trend toward reduced medium pH (P = 0.0984, Fig. 3E). In contrast, EO771 and MLg cells showed no significant changes in extracellular lactate levels, glucose consumption, or medium pH under hypoxic condition (Fig. 3C–E).
A HIF-1α protein level in normoxic and hypoxic (6 h at 3% O2) conditions in monocultures and co-cultures. The full Western blot membrane images are shown in Fig. S7. B Schematic representation of monocultures. C Extracellular lactate concentration (n = 7), D Extracellular glucose concentration (n = 3) and E Medium pH measurements performed at room temperature (n = 2, each replicate represents pooled supernatants from 3 to 4 wells of a 12-well plate) in supernatants from monocultures of EO771 (40,000 cells per well seeded in 12-well plate at D0), MLg and C2C12 (80,000 cells per well seeded in 12-well plate) after 2 days of incubation under normoxic or hypoxic conditions. F Schematic representation of co-cultures. G Outgrowth of EO771 alone and EO771 in co-cultures with either C2C12 myotubes (C2C12 + EO771) or MLg cells (MLg + EO771) under normoxic or hypoxic conditions over 2 days. Experiments were performed in 96-well plates. EO771 area represents the percentage of fluorescent cells normalized to day 0 (D0) (n = 34–40). Representative fluorescence microscopy images of EO771 (4000 cells per well in 96-well plate) cell outgrowth on C2C12 or MLg at day 2 (D2). Scale bar = 200 µm. H Outgrowth of EO771 (4000 cells per well in 96-well plate) in co-cultures with C2C12 myotubes under normoxic or hypoxic conditions over 2 days. EO771 area represents the percentage of fluorescent cells normalized to D0 (n = 30–36). I Extracellular lactate concentration (n = 8), J Extracellular glucose concentration (n = 3–4), and K Medium pH measurements performed at room temperature (n = 2, each replicate represents pooled supernatants from 20 wells of a 96-well plate) in supernatants from co-cultures under normoxic or hypoxic conditions after 2 days. Data are mean ± SEM. In each figure, samples were collected from two independent experiments (except glucose). Comparison between groups was done using two-way ANOVA with Šidák (only two groups) or Tukey’s (more than two groups) comparisons. **P < 0.01; ****P < 0.0001.
Next, we performed co-culture experiments under both normoxic and hypoxic conditions and included monocultured parental EO771 cells as an additional group (Fig. 3F). Under 3% O2, the outgrowth of EO771 onto C2C12 myotubes was strongly -and almost completely- abolished, whereas MLg cells exhibited the opposite response with EO771 outgrowth increasing significantly (Fig. 3G). No noticeable changes in cell morphology were observed when comparing normoxic and hypoxic conditions (Fig. 3G). By testing additional oxygen levels (1% and 10% O2), we further found that C2C12 myotubes restrict EO771 outgrowth in an oxygen dose-dependent manner (Fig. 3H). Furthermore, co-cultures of EO771 with either MLg or C2C12 showed relatively similar levels of hypoxia-induced lactate production (Fig. 3I) and glucose consumption (Fig. 3J), with no significant changes in medium pH (Fig. 3K).
Altogether, these findings indicate that hypoxic conditions have an unexpectedly cell host-specific effect, exacerbating the non-permissive environment of C2C12 myotubes, while increasing the permissive environment of MLg cells.
Exogenous lactate and glucose, culture acidosis, and conditioned medium do not reproduce the suppressive environment created by C2C12 myotubes
As lactate and medium pH were both affected by hypoxia, we explored whether these two factors, individually or in combination, could be responsible for the anti-metastatic response of C2C12 myotubes. In our investigations, we used Na-L-Lactate because L-lactate is the predominant and biologically active enantiomer of lactate. Moreover, the Na-L-lactate was selected as it is the most common form of extracellular lactate. As controls, we used cells treated with PBS (the solvent of Na-L-Lactate) and NaCl (to compensate for Na). Using exogenous Na-L-lactate treatment in monocultures (Fig. S8A), we first confirmed that the extracellular lactate levels increased proportionally with the administered lactate concentrations (10 and 20 mM) (Fig. S8B), supporting the robustness of our experimental design. Then, we tested the impact of Na-L-Lactate in co-cultures (Fig. 4A) and found that neither 10 mM nor 20 mM Na-L-Lactate treatment altered EO771 outgrowth in C2C12 myotubes or MLg cells (Fig. 4B). In additional experiments, we examined the impact of medium pH changes on EO771 proliferation in co-cultured cells maintained in DMEM at pH 8.2 (standard DMEM) or adjusted to 8.0 (reflecting normoxic C2C12 medium; Figs. 3E), 7.5 (close to the pH of hypoxic C2C12; Fig. 3E), or 7.0 (more acidic). For these pH experiments, media were made at room temperature to ensure reproducibility between experiments. After one and two days of co-cultures, lowering the medium pH had no effect on EO771 outgrowth in C2C12 or MLg co-cultures (Fig. 4C). Furthermore, we tested whether combining Na-L-lactate treatment with reduced medium pH would influence EO771 proliferation. Compared to the control condition (DMEM pH 8.2 + PBS), treatment with Na-L-lactate and reduced medium pH had no major effect on EO771 outgrowth onto C2C12 myotubes or MLg cells (Fig. 4D and Fig. S9).
A Schematic representation of co-cultures. B EO771 (4000 cells per well in 96-well plate) outgrowth in co-culture with C2C12 myotubes (C2C12 + EO771) or MLg cells (MLg + EO771) with added Na-L-lactate (n = 12), C Altered medium pH (n = 10–12), or D Added Na-L-lactate and altered medium pH (n = 9–10). All pH adjustments and measurements were done at room temperature. For (B–D), representative pictures can be found in Fig. S9. E Outgrowth of EO771 cells cultured for 2 days in medium with different glucose concentrations (n = 24–27). F Outgrowth of EO771 cells co-cultured for 2 days with C2C12 myotubes (C2C12 + EO771) or MLg cells (MLg + EO771) in medium with 5.5 mM or 25 mM glucose (n = 12). G Outgrowth of EO771 cells cultured in conditioned medium from C2C12 myotubes or MLg cells (n = 21). The EO771 area represents the percentage of fluorescent cells normalized to day 0 (D0). Data are mean ± SEM from two independent experiments. Comparison between groups was done using two-way ANOVA with Šidák (only two groups) or Tukey’s (more than two groups) comparisons. *P < 0.05; **P < 0.01; ****P < 0.0001.
Since extracellular glucose levels tended to be lower in C2C12 than in MLg cells (Fig. 3D), we next examined whether glucose availability affected EO771 proliferation by using culture medium with different glucose concentrations. EO771 cells showed no difference in outgrowth when exposed to 5.5, 10, 15, 20 or 25 mM glucose (Fig. 4E). Moreover, repeating the co-cultures under both 5.5 mM and 25 mM glucose conditions showed that glucose concentration did not modify the permissive environment provided by MLg cells or the non-permissive environment created by C2C12 myotubes (Fig. 4F). Finally, to assess whether C2C12 myotubes exert their suppressive effect on EO771 proliferation through a soluble factor other than lactate, pH, or glucose, we cultured EO771 cells with conditioned medium from either C2C12 myotubes or MLg cells. Under these conditions, EO771 outgrowth remained relatively comparable between the two groups (Fig. 4G), further excluding the involvement of a secreted factor in the suppressive effect of C2C12 myotubes on EO771 cells.
Discussion
Cancer metastases have the unique capacity to invade distant tissues, disrupting normal organ functions and accounting for the majority of cancer-related deaths. Understanding how these tumor cells adapt, or not, to foreign niches has remained a formidable challenge in cancer research.
Here, to bypass the complexity of in vivo metastasis processes -including steps such as pre-metastatic niche formation, intravasation, and circulation [38], we used co-culture systems that allow a focused exploration of cancer-cell host interactions. In our experimental settings, MLg cells supported robust proliferation of EO771 breast cancer cells, whereas C2C12 myotubes created a non-permissive environment, characterized by sparse and poorly proliferative EO771 cells. This inhibitory effect was consistent across multiple time points, a broad range of tumor cell densities, and observed in two distinct murine breast cancer cell lines, EO771 and 4T1. Additionally, we found that the suppression of cancer cell outgrowth is also present in Sol8 myotubes, supporting a general anti-cancer cell proliferation property of skeletal muscle cell and confirming the remarkable capacity of skeletal muscle to resist metastatic invasion previously reported by others [20,21,22,23]. Notably, this tumor-suppressive effect of muscle cells appears to be broad, as muscle cells not only inhibit the proliferation of murine breast cancer cells, but also suppress melanoma and Lewis lung carcinoma cells [20] as well as various human breast cancer cell lines, including MDA-MB-231, Hs578T, BT549, HCC70 and MDA-MB-468 [23]. This is consistent with clinical observations in humans, where skeletal muscle is among the least frequent sites of metastatic colonization, in stark contrast to organs such as lung, liver, and bone [16,17,18]. Overall, our data reinforce the idea that skeletal muscle harbors intrinsic properties that provide a unique, potent, and generalized defense against metastatic tumor colonization.
Since the transcriptional landscape of cancer metastases undergoes dynamic adaptations when colonizing new environments [24, 25], we reasoned that early profiling transcriptional changes of cancer and host cells when interacting with each other could offer novel insights into the anti-metastatic effects exerted by skeletal muscle cells. Therefore, we developed a FACS-based strategy to perform RNA-seq on both cancer and host cells maintained in monoculture (parental cells) and co-culture conditions for two days, enabling cell-type–specific analysis of context-dependent transcriptional responses. Our approach revealed two major findings. Firstly, at the cancer cell level, EO771 cells undergo extensive and niche-specific transcriptional reprogramming, with the most pronounced changes occurring in the presence of C2C12 myotubes. Indeed, EO771 cells co-cultured with C2C12 myotubes (EO771-C) exhibited over twice as many DEGs compared to those co-cultured with MLg cells (EO771-M). This reprogramming of EO771 cells was characterized by the upregulation of myogenesis, hypoxia, and glycolysis pathways, reflecting the barriers encountered by EO771 cells when attempting to colonize C2C12 myotubes. In contrast, EO771-M cells retained proliferation-associated signatures, including MYC targets, E2F targets, and G2M checkpoint pathways, suggesting that EO771 cells proliferate on MLg cells with minimal transcriptional remodeling. These findings indicate that the muscle environment not only halts EO771 cell proliferation but also induces a fundamental shift in cancer cell identity - potentially toward a more dormant state. Secondly, at the host cell level, C2C12 myotubes remained relatively transcriptionally stable upon interaction with EO771 cells, implying that limited cancer cell proliferation fails to disrupt muscle cell homeostasis. In contrast, MLg cells underwent broader transcriptomic changes, indicating that proliferating EO771 cells substantially perturb the MLg environment. This divergence in host cell plasticity may be a key determinant of metastatic compatibility, reinforcing the concept of skeletal muscle as a “metastatic desert”. Altogether, these findings underscore how hostile and permissive niches impose fundamentally different transcriptional demands on both cancer and host cells.
One of the most intriguing findings of our study is the robust pseudo-hypoxic-related transcriptional program that distinguished EO771 cells co-cultured with C2C12 myotubes (EO771-C) from those co-cultured with MLg cells (EO771-M). This observation was unexpected given the conventional view of hypoxia as a driver of metastatic progression [36, 37]. To determine the functional significance of this finding, we conducted experiments under hypoxic conditions, testing the provocative assumption that oxygen deprivation could enhance the anti-proliferative capacity of C2C12 myotubes. Strikingly, exposure to hypoxia nearly abolished EO771 outgrowth in co-cultures with C2C12 myotubes, while conversely enhancing cancer cell outgrowth in MLg co-cultures. These findings underscore the context-specific nature of hypoxia signaling in the metastatic setting, and importantly, show that C2C12 myotubes can couple hypoxia to an anti-proliferative response - effectively transforming a typically pro-tumorigenic stimulus into a barrier to cancer cell expansion. From a therapeutic standpoint, these findings carry important implications. Broad inhibition of hypoxia signaling - for example, through hypoxia-inducible factor (HIF) inhibitors [39] - could inadvertently compromise skeletal muscle integrity or weaken its exceptional resistance to metastatic cell proliferation.
The hypoxia-related transcriptional program found in EO771-C was particularly intriguing as this hypoxic signature emerged despite the co-cultures being maintained under normoxic conditions. Such a pseudo-hypoxic response has been previously linked to metabolic stress or redox imbalance in tumor environments and can profoundly influence tumor cell fate [40, 41]. To explore the possibility that C2C12 myotubes impose a metabolic competition on EO771 cells, we investigated whether lactate accumulation and medium acidification caused by C2C12 myotube metabolism could account for the anti-proliferative effect of muscle cells. Lactate and acidosis are generally recognized for promoting, rather than suppressing, tumor progression, as they support cancer cell proliferation, immune evasion, and metastasis -hallmarks often associated with the Warburg effect and altered tumor metabolism [42, 43]. Nevertheless, whether these same factors could contribute to the anti-metastatic environment characteristic of skeletal muscle remains unexplored.
We therefore treated EO771 cells with Na-L-lactate, the biologically active L-enantiomer and predominant extracellular form of lactate, while including both PBS (vehicle control) and NaCl (osmolarity/sodium control) in our experimental design. This choice was motivated by previous evidence showing that the anti-obesity effects of Na-L-lactate in mice were abolished when compared to NaCl-treated mice, and that Ucp1 induction in brown adipocytes could be equally triggered by NaCl and Na-L-lactate [44], raising concerns about the specificity of lactate’s biological effects and underscoring the need for careful control conditions. Under these rigorously controlled conditions, neither Na-L-lactate supplementation nor medium acidification altered EO771 outgrowth in co-cultures with either C2C12 or MLg cells. Likewise, varying glucose concentrations in the culture medium had no measurable effect. Collectively, these findings indicate that the hypoxic and glycolytic transcriptional signatures observed in EO771-C cells are unlikely to arise from a metabolic competition between EO771 and C2C12 cells.
Instead, these transcriptional changes may reflect an adaptive response associated with the induction of a quiescent or dormant-like phenotype in EO771 cells [45]. Notably, recent studies have shown that the entry of tumor cells into dormancy can itself elicit a pseudo-hypoxic state characterized by mitochondrial downregulation even under normoxic conditions, thereby driving glycolytic and hypoxia-associated gene expression [46,47,48]. This suggests that the hypoxic and glycolytic transcriptional programs observed in EO771-C cells may represent an intrinsic metabolic rewiring linked to dormancy induction. In this context, C2C12 myotubes could promote tumor cell quiescence and thereby indirectly trigger a hypoxia-like metabolic adaptation, consistent with the low-proliferative, anti-metastatic phenotype typically associated with muscle cells.
Next, we evaluated whether paracrine cues could contribute to the anti-metastatic effects of C2C12 myotubes. To this end, we cultured EO771 cells with conditioned media from either C2C12 or MLg cells. Conditioned media from C2C12 failed to reduce EO771 proliferation, demonstrating that the suppressive effect of C2C12 myotubes is not mediated by soluble factors. These results instead suggest that direct cell–cell interactions are required for C2C12 myotubes to exert their inhibitory influence on EO771 proliferation. One plausible mechanism involves trogocytosis, a process by which membrane fragments or surface proteins are exchanged between neighboring cells, which has been implicated in modulating signaling pathways and cellular fate in other contact-dependent systems [49, 50]. We therefore propose that such close physical interactions between myotubes and cancer cells could underlie the tumor-suppressive microenvironment imposed by skeletal muscle.
Recent evidence has implicated sustained oxidative stress in skeletal muscle as a critical barrier to tumor outgrowth. Indeed, MDA-MB-231 breast cancer cells exhibit markedly increased oxidative stress when co-cultured with primary human myotubes, demonstrating that muscle-derived ROS can impose a proliferative blockade [23]. In our hands, however, the global oxidative-stress gene-expression signature of EO771 cells co-cultured with C2C12 myotubes (EO771-C) did not significantly differ from that of EO771 cells onto MLg fibroblasts (EO771-M) after two days, suggesting an absence of acute transcriptional adaptation to ROS within this timeframe. Moreover, the parental transcriptomes of C2C12 and MLg cells showed no notable differences in oxidative-stress-related pathways, further indicating that baseline ROS-associated gene expression does not significantly distinguish the two environments. Thus, under our experimental conditions, oxidative stress did not appear to be the primary driver of muscle-induced growth suppression of EO771 cells.
Although in vitro models offer a controlled framework to uncover fundamental mechanisms, they inevitably simplify the complexity of in vivo processes. For instance, oxygen tensions in skeletal muscle in vivo are typically below 3% [51], whereas in vitro hypoxic conditions of 3% O₂ likely represent the higher end of physiological muscle oxygenation. Conversely, standard culture conditions at 20% O₂ are likely hyperoxic for muscle cells, although appropriate for cell types normally exposed to high oxygen levels, such as lung cells. Similarly, the lactate concentrations (10–20 mM) used in our experiments may exceed typical physiological levels in resting muscle but can reflect conditions observed during physical exercise [52] or pathological stress [53]. Moreover, our study was conducted in a two-dimensional (2D) co-culture system, which, while advantageous for dissecting direct cell–cell and paracrine interactions, does not fully reproduce the three-dimensional (3D) architecture and mechanical constraints of the in vivo tumor microenvironment. Cancer cells within a 3D matrix experience spatial gradients of oxygen, nutrients, and metabolites that profoundly influence their proliferative and metabolic behavior. Nonetheless, 2D co-cultures provide a valuable reductionist approach to identify key cellular interactions and signaling events that can subsequently be validated in 3D or in vivo models. These considerations underscore that our findings, although mechanistically informative, should be interpreted in the context of the physiological and spatial constraints that characterize the in vivo skeletal muscle–tumor interface.
In conclusion, our study highlights how hostile and permissive niches impose fundamentally different transcriptional demands on both cancer and host cells. We identify a hypoxia-related transcriptional program induced by C2C12 myotubes as an anti-metastatic mechanism, offering new insights into how cancer cells adapt -or fail to adapt- to new environments. Moreover, our findings reveal that hypoxia - a cue typically linked to tumor progression - was associated with reduced EO771 proliferation in C2C12 myotubes, indicating that hypoxia can be coupled to an anti-proliferative output by the muscle milieu. Importantly, these findings also caution against broadly targeting hypoxia signaling in anti-metastatic therapies, as such interventions could inadvertently undermine the tumor-suppressive functions of skeletal muscle.
Data availability
The RNA-seq data generated during and/or analyzed during the current study are available at the Gene Expression Omnibus under the accession number GSE299422 and following link: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE299422.
References
Steeg PS. Tumor metastasis: mechanistic insights and clinical challenges. Nat Med. 2006;12:895–904.
Dupuy F, Tabaries S, Andrzejewski S, Dong Z, Blagih J, Annis MG, et al. PDK1-dependent metabolic reprogramming dictates metastatic potential in breast cancer. Cell Metab. 2015;22:577–89.
Tasdogan A, Faubert B, Ramesh V, Ubellacker JM, Shen B, Solmonson A, et al. Metabolic heterogeneity confers differences in melanoma metastatic potential. Nature. 2020;577:115–20.
Sullivan MR, Mattaini KR, Dennstedt EA, Nguyen AA, Sivanand S, Reilly MF, et al. Increased serine synthesis provides an advantage for tumors arising in tissues where serine levels are limiting. Cell Metab. 2019;29:1410–1421.e1414.
Knott SRV, Wagenblast E, Khan S, Kim SY, Soto M, Wagner M, et al. Asparagine bioavailability governs metastasis in a model of breast cancer. Nature. 2018;554:378–81.
Gupta GP, Nguyen DX, Chiang AC, Bos PD, Kim JY, Nadal C, et al. Mediators of vascular remodelling co-opted for sequential steps in lung metastasis. Nature. 2007;446:765–70.
Oskarsson T, Acharyya S, Zhang XH, Vanharanta S, Tavazoie SF, Morris PG, et al. Breast cancer cells produce tenascin C as a metastatic niche component to colonize the lungs. Nat Med. 2011;17:867–74.
Zhang H, Freitas D, Kim HS, Fabijanic K, Li Z, Chen H, et al. Identification of distinct nanoparticles and subsets of extracellular vesicles by asymmetric flow field-flow fractionation. Nat Cell Biol. 2018;20:332–43.
Barkan D, El Touny LH, Michalowski AM, Smith JA, Chu I, Davis AS, et al. Metastatic growth from dormant cells induced by a col-I-enriched fibrotic environment. Cancer Res. 2010;70:5706–16.
Boudreau N, Sympson CJ, Werb Z, Bissell MJ. Suppression of ICE and apoptosis in mammary epithelial cells by extracellular matrix. Science. 1995;267:891–3.
Petersen OW, Ronnov-Jessen L, Howlett AR, Bissell MJ. Interaction with basement membrane serves to rapidly distinguish growth and differentiation pattern of normal and malignant human breast epithelial cells. Proc Natl Acad Sci USA. 1992;89:9064–8.
Pommier A, Anaparthy N, Memos N, Kelley ZL, Gouronnec A, Yan R, et al. Unresolved endoplasmic reticulum stress engenders immune-resistant, latent pancreatic cancer metastases. Science. 2018;360:eaao4908.
Pantel K, Schlimok G, Kutter D, Schaller G, Genz T, Wiebecke B, et al. Frequent down-regulation of major histocompatibility class I antigen expression on individual micrometastatic carcinoma cells. Cancer Res. 1991;51:4712–5.
Malladi S, Macalinao DG, Jin X, He L, Basnet H, Zou Y, et al. Metastatic latency and immune evasion through autocrine inhibition of WNT. Cell. 2016;165:45–60.
Janssen I, Heymsfield SB, Wang ZM, Ross R. Skeletal muscle mass and distribution in 468 men and women aged 18-88 yr. J Appl Physiol. 2000;89:81–88.
Disibio G, French SW. Metastatic patterns of cancers: results from a large autopsy study. Arch Pathol Lab Med. 2008;132:931–9.
Sudo A, Ogihara Y, Shiokawa Y, Fujinami S, Sekiguchi S. Intramuscular metastasis of carcinoma. Clin Orthop Relat Res. 1993;213–7.
Herring CL, Jr., Harrelson JM, Scully SP. Metastatic carcinoma to skeletal muscle. A report of 15 patients. Clin Orthop Relat Res. 1998;272–81.
Erwing J. Neoplastic disease: a treatise on tumors. Philadelphia and London, WB Saunders Company, Second Edition, 1922.
Parlakian A, Gomaa I, Solly S, Arandel L, Mahale A, Born G, et al. Skeletal muscle phenotypically converts and selectively inhibits metastatic cells in mice. PLoS ONE. 2010;5:e9299.
Weiss L. Biomechanical destruction of cancer cells in skeletal muscle: a rate-regulator for hematogenous metastasis. Clin Exp Metastasis. 1989;7:483–91.
Djaldetti M, Sredni B, Zigelman R, Verber M, Fishman P. Muscle cells produce a low molecular weight factor with anti-cancer activity. Clin Exp Metastasis. 1996;14:189–96.
Crist SB, Nemkov T, Dumpit RF, Dai J, Tapscott SJ, True LD, et al. Unchecked oxidative stress in skeletal muscle prevents outgrowth of disseminated tumour cells. Nat Cell Biol. 2022;24:538–53.
Sheraj I, Guray NT, Banerjee S. A pan-cancer transcriptomic study showing tumor specific alterations in central metabolism. Sci Rep. 2021;11:13637.
Wang R, Li J, Zhou X, Mao Y, Wang W, Gao S, et al. Single-cell genomic and transcriptomic landscapes of primary and metastatic colorectal cancer tumors. Genome Med. 2022;14:93.
Chen S, Zhou Y, Chen Y, Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884–i890.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21.
Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014;30:923–30.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Varet H, Brillet-Gueguen L, Coppee JY, Dillies MA. SARTools: a DESeq2- and EdgeR-based R pipeline for comprehensive differential analysis of RNA-seq data. PLoS ONE. 2016;11:e0157022.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Liberzon A, Birger C, Thorvaldsdottir H, Ghandi M, Mesirov JP, Tamayo P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015;1:417–25.
Bresson SE, Ruzzin J. Persistent organic pollutants disrupt the oxidant/antioxidant balance of INS-1E pancreatic beta-cells causing their physiological dysfunctions. Environ Int. 2024;190:108821.
Freshney RI. Culture of animal cells: a manual of basic technique and specialized applications. 7th ed. Wiley-Blackwell, 2015.
Le Naour A, Rossary A, Vasson MP. EO771, is it a well-characterized cell line for mouse mammary cancer model? Limit and uncertainty. Cancer Med. 2020;9:8074–85.
Eales KL, Hollinshead KE, Tennant DA. Hypoxia and metabolic adaptation of cancer cells. Oncogenesis. 2016;5:e190.
Rankin EB, Giaccia AJ. Hypoxic control of metastasis. Science. 2016;352:175–80.
Fares J, Fares MY, Khachfe HH, Salhab HA, Fares Y. Molecular principles of metastasis: a hallmark of cancer revisited. Signal Transduct Target Ther. 2020;5:28.
Mustafa M, Rashed M, Winum JY. Novel anticancer drug discovery strategies targeting hypoxia-inducible factors. Expert Opin Drug Discov. 2025;20:103–21.
Paredes F, Williams HC, San Martin A. Metabolic adaptation in hypoxia and cancer. Cancer Lett. 2021;502:133–42.
Augustin RC, Delgoffe GM, Najjar YG. Characteristics of the tumor microenvironment that influence immune cell functions: hypoxia, oxidative stress, metabolic alterations. Cancers. 2020;12:3802.
Qiao Y, Liu Y, Ran R, Zhou Y, Gong J, Liu L, et al. Lactate metabolism and lactylation in breast cancer: mechanisms and implications. Cancer Metastasis Rev. 2025;44:48.
Corbet C, Feron O. Tumour acidosis: from the passenger to the driver’s seat. Nat Rev Cancer. 2017;17:577–93.
Lund J, Breum AW, Gil C, Falk S, Sass F, Isidor MS, et al. The anorectic and thermogenic effects of pharmacological lactate in male mice are confounded by treatment osmolarity and co-administered counterions. Nat Metab. 2023;5:677–98.
Endo H, Inoue M. Dormancy in cancer. Cancer Sci. 2019;110:474–80.
Aguirre-Ghiso JA. Models, mechanisms and clinical evidence for cancer dormancy. Nat Rev Cancer. 2007;7:834–46.
Fluegen G, Avivar-Valderas A, Wang YR, Padgen MR, Williams JK, Nobre AR, et al. Phenotypic heterogeneity of disseminated tumour cells is preset by primary tumour hypoxic microenvironments. Nat Cell Biol. 2017;19:120–32.
Pranzini E, Raugei G, Taddei ML. Metabolic features of tumor dormancy: possible therapeutic strategies. Cancers. 2022;14:547.
Yang X, Huang X, Chen Z, Zhang W, Liu O, Wu J, et al. Trogocytosis at crossroads: navigating the duality in tumor-immune cell interactions. Biochim Biophys Acta Rev Cancer. 2025;1880:189431.
Marcarian HQ, Sivakoses A, Bothwell ALM. Molecular dynamics of trogocytosis and other contact-dependent cell trafficking mechanisms in tumor pathogenesis. Cancers. 2025;17:2268.
Richardson RS, Noyszewski EA, Kendrick KF, Leigh JS, Wagner PD. Myoglobin O2 desaturation during exercise. Evidence of limited O2 transport. J Clin Invest. 1995;96:1916–26.
Hermansen L, Vaage O. Lactate disappearance and glycogen synthesis in human muscle after maximal exercise. Am J Physiol. 1977;233:E422–429.
Liang Z, Zhao M, Liu K, Liang W, Luo S, Guan J, et al. Lactate trajectories in early survivors of sepsis: a new lens on mortality risk. Shock. 2025;64:386–96.
Acknowledgements
The sequencing service was provided by the Norwegian Sequencing Centre (www.sequencing.uio.no), a national technology platform hosted by the University of Oslo and Oslo University Hospital, and supported by the Research Council of Norway and the Southeastern Regional Health Authorities. Mycoplasma testing was performed by the Norwegian Stem Cell Core Facility. We thank Dr Cyrus M Ghajar (Cancer Research Center, Seattle) for providing GFP-labeled mouse breast cancer cells and advice on co-cultures. We thank all ChroMet members as well as Prof. Heidi Kiil Blomhoff and the Flow Cytometry Core Facility (Oslo University Hospital) for discussion and advice. This work was supported by internal funding from the University of Oslo and UNIFOR. AA and CC were supported by an internal grant from the University of Oslo.
Author information
Authors and Affiliations
Contributions
Methodology and experiments: AA, CC, MA; Data analyses: AA, CC, MA; Visualization: AA, CC, MA; Writing—original draft: AA, CC, JR; Writing—review and editing: AA, CC, MA, JR; Conceptualization, Funding acquisition, and Project supervision: JR.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
No ethics approval or consent to participate were required in the herein paper, as all experiments were performed using established mouse cell lines exclusively. All methods were performed in accordance with the relevant guidelines and regulations.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Aunan, A., Claeyssen, C., Abdelhalim, M. et al. Transcriptomic profiling of co-cultured cancer-host cells identifies hypoxia as a driver of the skeletal muscle cell’s anti-proliferative effect on cancer cells. Oncogenesis 15, 7 (2026). https://doi.org/10.1038/s41389-026-00601-9
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41389-026-00601-9