Abstract
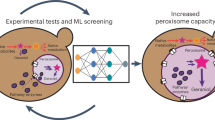
Engineered peroxisomes hold promise as a highly versatile platform for compartmentalizing engineered metabolic pathways, insulating them from native cellular factors to prevent undesired crosstalk. However, native peroxisomes often lack the required substrates and cofactors in their lumen; accordingly, nonnative membrane proteins (MPs) must be recruited to the peroxisomal membrane to support heterologous pathways requiring these molecules. We developed a robust, modular ‘chauffeur’ strategy that enables MP folding in the endoplasmic reticulum (ER) followed by trafficking to the peroxisomal membrane through an engineered interaction in the cytosol with a transmembrane domain natively trafficked from the ER to the peroxisome. We demonstrate the modularity of this strategy by successfully redirecting multiple MP cargoes, including heterologous plant MPs, and observed increased titers for a monoterpene biosynthetic pathway. This strategy overcomes the challenges of misfolding and sorting of MPs to the peroxisome and, accordingly, expands the repertoire of pathways that can be compartmentalized into this organelle.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Plasmids generated in this study were deposited to Addgene (plasmids 239493–239551) and are available upon reasonable request. Data supporting the figures in this study were published to figshare (https://doi.org/10.6084/m9.figshare.28928222.v1)68. Source data are provided with this paper.
References
Guerra-Giraldez, C., Quijada, L. & Clayton, C. E. Compartmentation of enzymes in a microbody, the glycosome, is essential in Trypanosoma brucei. J. Cell Sci. 115, 2651–2658 (2002).
Tenney, K. et al. hex-1, a gene unique to filamentous fungi, encodes the major protein of the Woronin body and functions as a plug for septal pores. Fungal Genet. Biol. 31, 205–217 (2000).
De Bellis, L., Picciarelli, P., Pistelli, L. & Alpi, A. Localization of glyoxylate-cycle marker enzymes in peroxisomes of senescent leaves and green cotyledons. Planta 180, 435–439 (1990).
McCammon, M. T., Veenhuis, M., Trapp, S. B. & Goodman, J. M. Association of glyoxylate and β-oxidation enzymes with peroxisomes of Saccharomyces cerevisiae. J. Bacteriol. 172, 5816–5827 (1990).
Hettema, E. H. et al. The ABC transporter proteins Pat1 and Pat2 are required for import of long‐chain fatty acids into peroxisomes of Saccharomyces cerevisiae. EMBO J. 15, 3813–3822 (1996).
Palmieri, L. et al. Identification and functional reconstitution of the yeast peroxisomal adenine nucleotide transporter. EMBO J. 20, 5049–5059 (2001).
DeLoache, W. C., Russ, Z. N. & Dueber, J. E.Towards repurposing the yeast peroxisome for compartmentalizing heterologous metabolic pathways. Nat. Commun. 7, 11152 (2016).
Dusséaux, S., Wajn, W. T., Liu, Y., Ignea, C. & Kampranis, S. C. Transforming yeast peroxisomes into microfactories for the efficient production of high-value isoprenoids. Proc. Natl Acad. Sci. USA 117, 31789–31799 (2020).
Zhou, Y. J. et al. Harnessing yeast peroxisomes for biosynthesis of fatty-acid-derived biofuels and chemicals with relieved side-pathway competition. J. Am. Chem. Soc. 138, 15368–15377 (2016).
Grewal, P. S., Samson, J. A., Baker, J. J., Choi, B. & Dueber, J. E. Peroxisome compartmentalization of a toxic enzyme improves alkaloid production. Nat. Chem. Biol. 17, 96–103 (2021).
Sparkes, I. A. & Baker, A. Peroxisome biogenesis and protein import in plants, animals and yeasts: enigma and variations? (Review). Mol. Membr. Biol. 19, 171–185 (2002).
Jansen, R. L. M. et al. Novel targeting assay uncovers targeting information within peroxisomal ABC transporter Pxa1. Biochim. Biophys. Acta - Mol. Cell Res. 1870, 119471 (2023).
Subramani, S., Koller, A. & Snyder, W. B. Import of peroxisomal matrix and membrane proteins. Annu. Rev. Biochem. 69, 399–418 (2000).
Chitwood, P. J., Juszkiewicz, S., Guna, A., Shao, S. & Hegde, R. S. EMC is required to initiate accurate membrane protein topogenesis. Cell 175, 1507–1519 (2018).
Hegde, R. S. & Keenan, R. J. The mechanisms of integral membrane protein biogenesis. Nat. Rev. Mol. Cell Biol. 23, 107–124 (2022).
Blobel, G. Intracellular protein topogenesis. Proc. Natl Acad. Sci. USA 77, 1496–1500 (1980).
Dueber, J. E., Yeh, B. J., Chak, K. & Lim, W. A.Reprogramming control of an allosteric signaling switch through modular recombination. Science 301, 1904–1908 (2003).
Sparks, A. B. et al. Distinct ligand preferences of Src homology 3 domains from Src, Yes, Abl, Cortactin, p53bp2, PLCγ, Crk, and Grb2. Proc. Natl Acad. Sci. USA 93, 1540–1544 (1996).
Halbach, A., Rucktächel, R., Rottensteiner, H. & Erdmann, R. The N-domain of Pex22p can functionally replace the Pex3p N-domain in targeting and peroxisome formation. J. Biol. Chem. 284, 3906–3916 (2009).
Kim, H., Melén, K., Österberg, M. & Von Heijne, G. A global topology map of the Saccharomyces cerevisiae membrane proteome. Proc. Natl Acad. Sci. Usa. 103, 11142–11147 (2006).
Burns, G. D. et al. Distinct classes of misfolded proteins differentially affect the growth of yeast compromised for proteasome function. FEBS Lett. 595, 2383–2394 (2021).
Halbach, A. et al. Targeting of the tail-anchored peroxisomal membrane proteins PEX26 and PEX15 occurs through C-terminal PEX19-binding sites. J. Cell Sci. 119, 2508–2517 (2006).
Feng, S. et al. Bright split red fluorescent proteins for the visualization of endogenous proteins and synapses. Commun. Biol. 2, 344 (2019).
Fitzgerald, I. & Glick, B. S. Secretion of a foreign protein from budding yeasts is enhanced by cotranslational translocation and by suppression of vacuolar targeting. Microb. Cell Fact. 13, 1–12 (2014).
Barrero, J. J., Casler, J. C., Valero, F., Ferrer, P. & Glick, B. S. An improved secretion signal enhances the secretion of model proteins from Pichia pastoris. Microb. Cell Fact. 17, 1–13 (2018).
Dueber, J. E. et al. Synthetic protein scaffolds provide modular control over metabolic flux. Nat. Biotechnol. 27, 753–759 (2009).
Schäfer, H., Nau, K., Sickmann, A., Erdmann, R. & Meyer, H. E. Identification of peroxisomal membrane proteins of Saccharomyces cerevisiae by mass spectrometry. Electrophoresis 22, 2955–2968 (2001).
Hefner, T. & Stöckigt, J. Probing suggested catalytic domains of glycosyltransferases by site-directed mutagenesis. Eur. J. Biochem. 270, 533–538 (2003).
Roy, S. K., Chiba, Y., Takeuchi, M. & Jigami, Y. Characterization of yeast Yea4p, a uridine diphosphate-N-acetylglucosamine transporter localized in the endoplasmic reticulum and required for chitin synthesis. J. Biol. Chem. 275, 13580–13587 (2000).
Esther, C. K. et al. Similarities between UDP-glucose and adenine nucleotide release in yeast: Involvement of the secretory pathway. Biochemistry 47, 9269–9278 (2008).
Reguenga, C., Oliveira, M. E. M., Gouveia, A. M. M., Sá-Miranda, C. & Azevedo, J. E. Characterization of the mammalian peroxisomal import machinery. Pex2p, Pex5p, Pex12p, and Pex14p are subunits of the same protein assembly. J. Biol. Chem. 276, 29935–29942 (2001).
Kim, P. K. & Hettema, E. H.Multiple pathways for protein transport to peroxisomes. J. Mol. Biol. 427, 1176–1190 (2015).
Skowyra, M. L. & Rapoport, T. A. PEX5 translocation into and out of peroxisomes drives matrix protein import. Mol. Cell 82, 3209–3225 (2022).
Munakata, R. et al. Parallel evolution of UbiA superfamily proteins into aromatic O-prenyltransferases in plants. Proc. Natl Acad. Sci. USA 118, e2022294118 (2021).
Zhang, Y., Guo, J., Gao, P. & Yan, W. Development of an efficient yeast platform for cannabigerolic acid biosynthesis. Metab. Eng. 80, 232–240 (2023).
Luo, X. et al. Complete biosynthesis of cannabinoids and their unnatural analogues in yeast. Nature 567, 123–126 (2019).
Li, H. et al. A heteromeric membrane-bound prenyltransferase complex from hop catalyzes three sequential aromatic prenylations in the bitter acid pathway. Plant Physiol. 167, 650–659 (2015).
Sasaki, K., Mito, K., Ohara, K., Yamamoto, H. & Yazaki, K. Cloning and characterization of naringenin 8-prenyltransferase, a flavonoid-specific prenyltransferase of Sophora flavescens. Plant Physiol. 146, 1075–1084 (2008).
de Bruijn, W. J. C., Levisson, M., Beekwilder, J., van Berkel, W. J. H. & Vincken, J. P. Plant aromatic prenyltransferases: tools for microbial cell factories. Trends Biotechnol. 38, 917–934 (2020).
Ignea, C., Pontini, M., Maffei, M. E., Makris, A. M. & Kampranis, S. C. Engineering monoterpene production in yeast using a synthetic dominant negative geranyl diphosphate synthase. ACS Synth. Biol. 3, 298–306 (2014).
Baker, J. J. et al. ML-enhanced peroxisome capacity enables compartmentalization of multienzyme pathway. Nat. Chem. Biol. 21, 727–735 (2025).
De Roode, B. M., Oliehoek, L., Van der Padt, A., Franssen, M. C. R. & Boom, R. M. Downstream processing of enzymatically produced geranyl glucoside. Biotechnol. Prog. 17, 881–886 (2001).
Ikemoto, T., Okabe, B. I., Mimura, K. & Kitahara, T. Formation of fragrance materials from odourless glycosidically-bound volatiles by skin microflora (Part 1). Flavour Fragr. J. 17, 452–455 (2002).
Herrmann, A. Controlled release of volatiles under mild reaction conditions: from nature to everyday products. Angew. Chem. Int. Ed. Engl. 46, 5836–5863 (2007).
Zhao, J., Bao, X., Li, C., Shen, Y. & Hou, J. Improving monoterpene geraniol production through geranyl diphosphate synthesis regulation in Saccharomyces cerevisiae. Appl. Microbiol. Biotechnol. 100, 4561–4571 (2016).
Bönisch, F. et al. Activity-based profiling of a physiologic aglycone library reveals sugar acceptor promiscuity of family 1 UDP-glucosyltransferases from grape. Plant Physiol. 166, 23–39 (2014).
Castro, O., Chen, L. Y., Parodi, A. J. & Abeijón, C. Uridine diphosphate-glucose transport into the endoplasmic reticulum of Saccharomyces cerevisiae: in vivo and in vitro evidence. Mol. Biol. Cell 10, 1019–1030 (1999).
Yocum, H. C., Bassett, S. & Da Silva, N. A. Enhanced production of acetyl-CoA-based products via peroxisomal surface display in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 119, e2214941119 (2022).
Perrot, T., Besseau, S., Papon, N. & Courdavault, V. Gaining access to acetyl-CoA by peroxisomal surface display. Synth. Syst. Biotechnol. 8, 224–226 (2023).
Zhang, Y. et al. The construction and optimization of engineered yeast chassis for efficient biosynthesis of 8-hydroxygeraniol. mLife 2, 438–449 (2023).
Ye, C. et al. Metabolic engineering of Pichia pastoris for overproduction of cis-trans nepetalactol. Metab. Eng. 84, 83–94 (2024).
Brown, S., Clastre, M., Courdavault, V. & O’Connor, S. E. De novo production of the plant-derived alkaloid strictosidine in yeast. Proc. Natl Acad. Sci. USA 112, 3205–3210 (2015).
Lee, H., DeLoache, W. C. & Dueber, J. E. Spatial organization of enzymes for metabolic engineering. Metab. Eng. 14, 242–251 (2012).
Trantas, E., Panopoulos, N. & Ververidis, F. Metabolic engineering of the complete pathway leading to heterologous biosynthesis of various flavonoids and stilbenoids in Saccharomyces cerevisiae. Metab. Eng. 11, 355–366 (2009).
Martin, V., Pitera, D., Withers, S., Newman, J. & Keasling, J. Engineering a mevalonate pathway in Escherichia coli for production of terpenoids. Nat. Biotechnol. 21, 796–802 (2003).
Antonenkov, V. D. & Hiltunen, J. K. Transfer of metabolites across the peroxisomal membrane. Biochim. Biophys. Acta Mol. Basis Dis. 1822, 1374–1386 (2012).
Wang, C. et al. Metabolic engineering of Escherichia coli for α-farnesene production. Metab. Eng. 13, 648–655 (2011).
Ma, Y., Shang, Y. & Stephanopoulos, G. Engineering peroxisomal biosynthetic pathways for maximization of triterpene production in Yarrowia lipolytica. Proc. Natl Acad. Sci. USA 121, e2314798121 (2024).
Hamilton, S. R. et al. Humanization of yeast to produce complex terminally sialylated glycoproteins. Science 313, 1441–1443 (2006).
Bayer, T. S. et al. Synthesis of methyl halides from biomass using engineered microbes. J. Am. Chem. Soc. 131, 6508–6515 (2009).
Srinivasan, P. & Smolke, C. D. Engineering cellular metabolite transport for biosynthesis of computationally predicted tropane alkaloid derivatives in yeast. Proc. Natl Acad. Sci. USA 118, e2104460118 (2021).
Higy, M., Junne, T. & Spiess, M. Topogenesis of membrane proteins at the endoplasmic reticulum. Biochemistry 43, 12716–12722 (2004).
Guerriero, C. J. et al. Harmonizing experimental data with modeling to predict membrane protein insertion in yeast. Biophys. J. 117, 668–678 (2019).
Nielsen, H., Tsirigos, K. D., Brunak, S. & von Heijne, G. A brief history of protein sorting prediction. Protein J. 38, 200–216 (2019).
Di Martino, R., Sticco, L. & Luini, A. Regulation of cargo export and sorting at the trans-Golgi network. FEBS Lett. 593, 2306–2318 (2019).
Lee, M. E., DeLoache, W. C., Cervantes, B. & Dueber, J. E.A highly characterized yeast toolkit for modular, multipart assembly. ACS Synth. Biol. 4, 975–986 (2015).
Ng, A. H. et al. Modular and tunable biological feedback control using a de novo protein switch. Nature 572, 265–269 (2019).
Lee, V. & Siu, K.-H. Data supporting Siu et al. in Nat. Chem. Biol. figshare https://doi.org/10.6084/m9.figshare.28928222.v1 (2025).
Acknowledgements
This work was supported by National Science Foundation (NSF) grant MCB 1818307, NSF grant CBET 2104261 and the Center for Cellular Construction, an NSF Science and Technology Center, under grant agreement DBI-1548297. We thank members of the J.E.D. laboratory for valuable discussions and feedback throughout this project and members of the W. Zhang laboratory for assistance with LC–MS, particularly A. Del Rio Flores, Y. Ko and K. Jia.
Author information
Authors and Affiliations
Contributions
K.S. and J.E.D. designed the research. K.S. and V.L. conducted the chauffeuring of various cargoes, bergamottin biosynthesis and the esculin assay. K.S. additionally conducted the peptide ligand optimization assay using split fluorescent protein and biosynthesis of geranyl β-d-glucoside and 8HG. V.L. additionally conducted experiments for the MP orientation in the TEV protease assay. K.S., V.L. and J.E.D. wrote the paper and created the figures.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Bei Gao, Jiazhang Lian and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The chauffeur strategy enables efficient trafficking of various MP cargoes to the peroxisome.
Venus-ePTS1 (YFP) was expressed to confirm the MP and the chauffeur localized to the peroxisome. (a) The yeast vacuolar MP, Vba2p, trafficked to peroxisomal membranes using the chauffeur strategy. In comparison, direct fusion of Vba2p to tPex22 did not result in efficient trafficking to the peroxisome; most of the Vba2p appeared to traffic to the vacuole. (b) CDT1p, a heterologous multispanning cellodextrin transporter from Neurospora crassa, trafficked to the peroxisome using the chauffeur strategy. The tPex22 direct fusion seemed to redirect CDT1p to the peroxisomal membrane; however, the total amount of CFP-fused MP successfully targeted to the peroxisome was much lower than achieved via the chauffeur strategy. Collectively, the results of trafficking Yea4p (Fig. 1b), Vba2p, and CDT1p suggest that the chauffeur strategy is a more robust method to redirect native and heterologous membrane proteins to the peroxisome. All scale bars represent 5 µm.
Extended Data Fig. 2 Activity and localization of trafficked candidate UDP-glucose transporters.
(a) Localization of non-trafficked Yea4p was compared to the ER labeled with a YFP (Venus-HDEL). In the dead SH3 peptide (dSH3) control and cases of inefficient trafficking, Yea4p co-localized with the ER-localized YFP signal as expected. (b) Esculin standard curve. Fluorescence at 460 nm plotted against esculin concentration generated the standard curve used to interpolate esculin titers for esculetin-to-esculin bioconversion assays. (c) Trafficked Yea4p activity was analyzed using the esculetin-to-esculin assay. Yea4p was trafficked via the engineered SH3 domain-SH3 peptide ligand protein-protein interaction. In the direct fusion strategies, the native protein truncations tPex22 and tPex15 were fused to either the N- or C-terminus of Yea4p, respectively. While tPex22-Yea4p and Yea4p-tPex15 resulted in weak and strong peroxisome localization (Fig. 1), respectively, low levels of activity were observed for both fusions. One-way ANOVA was used to compare the sample data between each group of 4 biological replicates. (d) NSTs trafficked to the peroxisome via the chauffeur strategy were tested for their UDP-glucose transport activities on the peroxisome membrane. Previous studies suggest that Yea4p and AtUTR1 exhibit transport activities for UDP-glucose while Hut1p and Ymd8p only exhibit marginal transport activities toward the co-factor. CDT2 is an unrelated membrane protein from Neurospora crassa that transports cellodextrins and is used as a negative control. (e) AtUTR1 trafficked with an active SH3 domain traffics some of the membrane protein to the peroxisome, while the dSH3 control results in protein that does not appear to associate with a yeast membrane. All scale bars represent 5 µm. All data shown are the mean ± s.e.m. of four biological replicates and all subfigures use the following significance: * denotes p-value < 0.05; ** p-value < 0.01; **** p-value < 0.001.
Extended Data Fig. 3 The orientation of chauffeured Yea4p was screened by a TEV protease assay.
Yea4p cargo was tracked by a CFP, and the chauffeur was fused to an RFP. (a) YFP followed by a TEV protease cleavage site was fused to the N-terminus of Yea4p. With and without TEV protease induction (with either 100 nM or 1000 nM β-estradiol), the YFP signal of this N-terminal fusion remained punctate and localized at the peroxisome, consistent with protection from the TEV protease from having a lumen-facing orientation throughout trafficking and within the peroxisome. (b) The C-terminus of Yea4p-2xSH3 was fused with a TEV protease cleavage site followed by a YFP. With this fusion and induction of TEV protease, the release of YFP to the cytoplasm was observed, indicating a cytosolic-oriented C-terminus. Based on traces of Yea4p-CFP signal residing on the ER membrane and the incomplete TEV protease cleavage at 100 nM β-estradiol, the YFP fusion may partially interfere with SH3 chauffeuring and TEV protease cleavage through steric hindrance. A higher concentration of 1000 nM β-estradiol resulted in more complete cleavage of YFP from peroxisomes. All scale bars represent 5 µm.
Extended Data Fig. 4 Trafficking of tCpPT1 using the peroxisome targeting sequence (PTS1) results in inefficient trafficking and non-functional membrane protein.
(a) Trafficking of tCpPT1 (Accession: A0A810JZE7 (https://www.uniprot.org/uniprotkb/A0A810JZE7/entry)) by a peroxisome targeting sequence (PTS1) was visualized using CFP (mTurquoise2) and co-localization with tPex22 fused to RFP (mRuby2). Using the same exposure times and contrast settings as the tCpPT1 trafficked via the chauffeur strategy (Fig. 4c), CFP fluorescence was not detectable (not shown). tCpPT1 signal with greatly enhanced contrast settings remained barely observable as very small CFP punctae that co-localized with RFP, suggesting poor expression and/or membrane protein misfolding and inefficient trafficking to the peroxisome. Scale bar represents 5 µm. (b) Bergamottin titers resulting from trafficking of tCpPT1 via the chauffeur strategy were compared to those from a previously reported method where a peroxisome targeting sequence (PTS1) is fused to the C-terminus of the membrane embedded enzyme. Colors are consistent with those used in Fig. 4c. The titers obtained using the chauffeur strategy are >20-fold higher than those achieved by PTS1 fusion from samples collected 24 hrs after addition of the bergaptol substrate, likely because import via PTS1 tag relies on soluble, cytosolic receptor proteins and has not evolved for functional membrane protein insertion into the peroxisome membrane. “Per” indicates the MP cargo was targeted via Crk SH3 domain-SH3 peptide ligand interaction; “Non-targeted” refers to the use of a non-binding SH3 peptide variant; and “PTS1” refers to the C-terminal fusion of peroxisome targeting sequence (-SKL). Data are shown as the mean ± s.e.m. of three biological replicates.
Extended Data Fig. 5 tCpPT1 localizes to the endoplasmic reticulum.
Fusion of CFP (mTurquoise2) to the N- or C-terminus of tCpPT1 results in localization to the ER. YFP (Venus) was fused to an N-terminal Ost1 signal sequence and a C-terminal HDEL retention sequence for targeting and retention in the ER. All images were taken at the same exposure settings. Controls (“No CFP” and “No YFP”) were performed to ensure there was no overlapping signal that would cause false signals from the fluorescent proteins used. Images were captured 4 h after 50-fold dilution in fresh media. All scale bars represent 5 µm.
Extended Data Fig. 6 Peroxisome-localized bergamottin production results in better host health and bergamottin titers compared to cytosolically-localized production.
(a) mTurquoise2 fluorescence of colonies used for bergamottin production in Fig. 5. As tCpPT1 was expressed as a fusion protein with mTurquoise2, the mTurquoise2 signal can be used as a proxy for protein expression. No significant differences in CpPT1 expression levels were observed with (SH3pep) versus without (dSH3pep) trafficking to the peroxisome. Two-way ANOVA was used. ns denotes not significant. Data are shown as the mean ± s.e.m. of three biological replicates. (b) A growth defect in yeast containing cytosolically-localized bergamottin biosynthesis (Cyt=pex5Δ) is observed when compared to peroxisome-compartmentalized (Per=WT) biosynthesis. OD600 values were taken at 72 h after dilution to an initial OD600 of 0.2. Two-way ANOVA was used. * denotes p-value < 0.05; *** p-value < 0.005. Data are shown as the mean ± s.e.m. of three biological replicates.
Extended Data Fig. 7 Chromatograms of geraniol, geranyl β-D-glucoside, and extracted samples from fermentations of strains expressing the de novo biosynthetic pathway analyzed by LC-MS.
Note the higher product peak with both peroxisomal MVA pathway and peroxisome-targeted Yea4p compared to either non-targeted Yea4p or cytoplasmic MVA pathway.
Extended Data Fig. 8 Trafficked cytochrome P450 and P450 reductase to the peroxisome retain activity in synthesis of (6E)-8-hydroxygeraniol.
(a) Schematic of C. roseus cytochrome P450 geraniol 8-hydroxylase (CrG8H) and C. roseus P450 reductase (CrCPR) trafficking to the peroxisome for 8HG biosynthesis. (b) GCMS data of 8HG titer. One-way ANOVA was used to compare the sample data between each group of 4 biological replicates. * denotes p-value < 0.05; *** p-value < 0.005; ns denotes not significant. Data are shown as the mean ± s.e.m. of four biological replicates. (c) CrG8H and (d) CrCPR were trafficked to the peroxisome using the chauffeur strategy. The p450 and p450 reductases were fused to BFP, and peroxisomes were labeled with a YFP-ePTS1. All scale bars represent 5 µm.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3 and Table 1.
Source data
Source Data Fig. 1
PDF of micrographs.
Source Data Fig. 2
PDF of micrographs and statistical source data.
Source Data Fig. 3
PDF of micrographs and statistical source data.
Source Data Fig. 4
PDF of micrographs and statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
PDF of micrographs.
Source Data Extended Data Fig. 2
PDF of micrographs and statistical source data.
Source Data Extended Data Fig. 3
PDF of micrographs and statistical source data.
Source Data Extended Data Fig. 4
PDF of micrographs and statistical source data.
Source Data Extended Data Fig. 5
PDF of micrographs and statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
PDF of micrographs and statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Siu, KH., Lee, V. & Dueber, J.E. Functional targeting of membrane transporters and enzymes to peroxisomes. Nat Chem Biol 21, 1544–1553 (2025). https://doi.org/10.1038/s41589-025-01948-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41589-025-01948-7
This article is cited by
-
A peroxisomal membrane protein hub
Nature Chemical Biology (2025)