Abstract
Induced pluripotent stem cell (iPSC)-derived mesenchymal stromal cells (iMSCs) offer a promising alternative to primary mesenchymal stromal cells (MSCs) and their derivatives, particularly extracellular vesicles (EVs), for use in advanced therapy medicinal products. In this study we evaluated the immunomodulatory and regenerative potential of iMSCs as well as iMSC-EVs, alongside primary human umbilical cord-derived mesenchymal stromal cells (hUCMSCs). Our findings demonstrate that iMSCs exhibit comparable abilities to hUCMSCs in regulating lymphocyte proliferation and inducing an anti-inflammatory phenotype in monocytes. We also observed decreased TNFα levels and increased IL-10 induction, indicating a potential mechanism for their immunomodulatory effects. Furthermore, iMSC-EVs also showed effective immunomodulation by inhibiting T cell proliferation and inducing macrophage polarization similar to their parental cells. Additionally, iMSC-EVs exhibited pro-regenerative potential akin to hUCMSC-EVs in in vitro scratch assays. Notably, priming iMSCs with pro-inflammatory cytokines significantly enhanced the immunomodulatory potential of iMSC-EVs. These results underscore the considerable promise of iMSCs and iMSCs-EVs as an alternate source for MSC-derived therapeutics, given their potent immunomodulatory and regenerative properties.
Similar content being viewed by others
Introduction
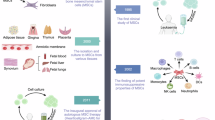
Mesenchymal stromal cells (MSCs) represent a stem cell population with promising therapeutic effects in a broad range of diseases involving tissue regeneration and the immune system1. However, complexities related to donor variability, different tissue sources and the need for extensive in vitro expansion continue to limit the use of MSCs as an advanced therapy medicinal product2. To overcome these limitations, it is essential to identify different, accessible sources of MSCs. The generation of induced pluripotent stem cell (iPSC)-derived MSCs (iMSCs) appears to provide an opportunity to effectively address most of these limitations3,4,5. iPSCs, as an alternative source of pluripotent stem cells, can be generated from patient-specific adult somatic cells, such as skin fibroblasts, peripheral blood cells, or other tissues through transcriptional factor-induced reprogramming6. Moreover, iPSCs are considered an inexhaustible source of iMSCs that can meet the high clinical demand7. Bloor et al. found that one iPSC bank can produce 29 million clinical doses of iMSC therapy8. In addition, iMSCs induced from a single iPSC cell or clone are theoretically much more homogeneous8,9. As a potential unlimited source of MSCs, research groups have been engaged in differentiation studies of iPSCs into iMSCs to explore their therapeutic effects in multiple diseases10,11,12. Importantly, iMSCs closely resemble their primary tissue-derived MSC counterparts in morphology, immunophenotype, and tree-lineage differentiation capacity11,13,14. Furthermore, it has been shown that iMSCs possess superior proliferation capacities, as well as advantages in tissue repair, immunomodulation and differentiation applications compared to tissue-derived MSCs3,11,15,16.
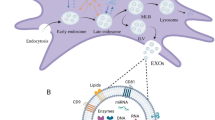
In the past two decades, several studies have demonstrated that MSCs possess potent potential for clinical applications due to their strong immunomodulatory effects via cell-to-cell interactions and release of soluble factors and extracellular vesicles (EVs)17,18,19. EVs are a family of small membranous nanoparticles that carry a wide range of biomolecules, including proteins, lipids, nucleic acids, and small molecules20. Interestingly, EVs play important roles in various physiological and pathological processes, such as cell-to-cell communication and immune regulation, and are thought to be involved in the progression of several diseases, including cancer, neurodegenerative and cardiovascular diseases21,22,23. Recently, there is increasing appeal in using MSC-derived EVs as a therapeutic alternative to MSCs24 since EV-based therapies may mitigate the safety concerns associated with the use of MSCs25,26. iMSC therapy offers further benefits over primary MSC therapy as iMSCs have been reported to be superior in producing microenvironment-modulating EVs15,27,28. The therapeutic effect of iMSC-derived EVs is explored in many diseases and is best studied in wound healing, cardiovascular disease, and musculoskeletal pathology28,29,30. Although MSC-derived EVs seem to play a key role in inflammation-related complications, studies to evaluate the immunomodulatory potential of iMSC-EVs in this context are still in early stages and further research is needed to describe their functionality31,32,33.
To comprehensively compare the therapeutic efficacy of iMSCs and primary MSCs, it is imperative to elucidate their functionality and the underlying mechanisms. Only one study to date has compared primary MSCs against iMSCs and iMSC-EVs in an in vivo inflammatory model. In this study, the authors observed that iPSC-MSCs responded similarly to BM-MSCs in the NOD mouse model of Sjögren’s syndrome (SS). Additionally, they found that iPSC-MSC EVs were as functional as their parental cells34. To further explore this area and address the existing gap, we aimed to develop a comprehensive cell-based assay platform. This platform designed to assess the immunomodulatory potential of iMSCs and iMSC-EVs includes several specific components to comprehensively evaluate their effects: T cell proliferation and macrophage polarization assays, alongside the regenerative potential of iMSC-EVs in a scratch-wound assay. Macrophage polarization refers to the process by which macrophages differentiate into distinct functional phenotypes, M1 (pro-inflammatory) and M2 (anti-inflammatory). The scratch assay is an in vitro technique used to study cell migration and wound healing. As a benchmark, we conducted a comparative analysis with primary umbilical cord-derived MSCs (hUCMSCs). By integrating these analysis, the platform provides a robust assessment of the immunomodulatory potential of iMSCs and iMSC-EVs, offering valuable insights into their mechanisms of action and therapeutic potential. Furthermore, we explored the impact of priming MSCs with pro-inflammatory cytokines on the anti-inflammatory effect of iMSC-EVs. We propose that further functional studies will unveil the mechanisms of action of iMSCs and iMSC-EVs in immunomodulation and regenerative processes, enhancing our understanding of their therapeutic potential.
Results
iMSC characterization
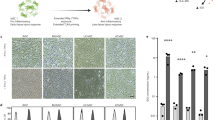
In this study, we first optimized a protocol to differentiate iPSCs into iMSCs. Throughout the study, we also included human umbilical cord MSCs (hUCMSCs) to compare their characteristics and functionality. iMSCs showed typical fibroblast-like and spindle-shaped appearance after 4 weeks of differentiation (Fig. 1A, right panel), and no morphological differences between hUCMSCs and iMSCs were found (Fig. 1A, left panel). Afterwards, the immunophenotype was evaluated by flow cytometry assays. iMSCs displayed a typical MSC phenotype as shown by high expression of CD90, CD105, CD73 and HLA-ABC, and absence of hematopoietic markers CD34, CD45 and HLA-DR (Fig. 1B)35. However, even though a high expression of the cell surface marker CD90 was measured in iMSCs, a significant differential expression with hUCMSCs was observed (Fig. 1B).
Characterization of hUCMSCs and iMSCs primary cells. (A) Representative morphology of two different confluences for hUCMSCs (left panel) and iMSCs (right panel) in culture, scale bar 500 μm. (B) Immunophenotypic characterization of hUCMSCs (blue bars) and iMSCs (orange bars) represented as percentage expression of a subset of classical cell surface markers, including CD90, CD105, CD73, HLA-ABC, CD34, CD45 and HLA-DR (n = 3). All data are presented as means ± SD. ns: not significant, ***: p < 0.001 by two-way ANOVA followed by Sisak’s multiple comparisons test.
iMSCs show an immunomodulatory effect on T lymphocyte proliferation
In order to assess iMSC-driven immunomodulatory responses, we first established a co-culture immune assay using peripheral blood mononuclear cells (PBMCs) as previously described36. We tested the iMSC suppressive potential by exposing anti-CD2, -CD3, -CD28 coated bead-stimulated PBMCs to iMSCs for 5 days in two culture formats: direct cell-to-cell contact and indirect contact using pore transwell inserts (Supplemental Fig. 1A, left panel). From the CD3 + counts, we calculated the inhibition (expressed as percentage) of T cell proliferation by iMSCs, taking as reference the CD3 + counts in activated PBMCs. Importantly, in the presence of iMSCs we observed a suppression of T cell proliferation with different percentages of inhibition between the two co-culture setups (Fig. 2A, right panel). iMSCs inhibited T cell proliferation by 94.6% ± 3.9% and 49.3% ± 2.4% in cell-to-cell contact and contact-independent cultures, respectively. The efficiency of the suppression of PBMCs proliferation by iMSCs was also compared with the cell primary hUCMSCs (Fig. 2A, left panel). Remarkably, iMSCs exhibited the ability to regulate the proliferation of activated lymphocytes in a similar response pattern to that of primary hUCMSCs cells both in direct (94.6% ± 3.9% vs. 96.5 ± 2.1%) and in indirect (49.3% ± 2.4% vs. 50.0 ± 3.3%) conditions.
To further elucidate the iMSC-mediated immune suppression on T cells, we employed ELISA to assess the expression level of the pro-inflammatory cytokine TNFα. This molecule is a well-known soluble factor that contributes to the immunosuppressive effects of MSCs derived from several sources36. Firstly, the high concentration of TNFα measured in the supernatants from activated PBMCs (Fig. 2B, grey bar) confirmed that, under the experimental conditions assayed, a strong inflammatory environment was induced. Secondly, we observed that the concentration of TNFα significantly decreased in the supernatants from the indirect co-cultures with iMSCs, compared to the control sample (bead-activated PBMCs). In presence of iMSCs the concentration of TNFα cytokine decreased by 91.9% (480.1 pg/ml ± 28.7 pg/ml vs. 38.7 pg/ml ± 6.9 pg/ml) (Fig. 2B, grey and orange bar). Similar results were obtained with the primary cells hUCMSCs (24.28 pg/ml ± 5.9 pg/ml) (Fig. 2B, blue bar). Altogether, these results confirm that under bead-induced immune activation, iMSCs trigger potent and robust immune suppression, either in a cell-to-cell interaction or in cell-contact independent setting. Moreover, the results suggest that the pro-inflammatory cytokine TNFα is playing a role in the immunomodulatory effect on T-cells by iMSCs, possibly leading to the suppression of T cell proliferation.
iMSCs possess an immunomodulatory effect on macrophage polarization
Macrophages are reported to play critical roles in the therapeutic effects of MSCs in several diseases37. Moreover, secretome and transcriptome data have suggested a pivotal role of inflammation-related cytokines (CKs) and chemokines that directly influence the differentiation and activation of myeloid cells in hUCMSCs-mediated immunomodulation36. Therefore, we assessed the differentiation and activation status of monocyte populations in the context of MSC-driven immunomodulation as previously described36. For that purpose, we investigated whether exposure to an immunomodulatory environment driven by iMSCs would alter monocyte (CD14+) differentiation and activation. We first depleted CD14+ monocytes from PBMCs (CD14−) and performed iMSC co-cultures under bead-activated T-cells. Isolated CD14+ monocytes were seeded in transwells and exposed to bead-activated CD14− (CD14−*) or CD14−*/iMSC direct co-cultures (CD14−*/iMSC). The expression of CD80, CD206 and CD163 markers were evaluated after 5 days (Supplemental Fig. 1A, right panel). Monocyte co-cultures with CD14−*/iMSCs and CD14−*/hUCMSCs showed significant levels of CD206 (p < 0.0001) and CD163 (p = 0.0012) markers, as compared to CD14+ exposed to activated CD14− (CD14++CD14−*) (Fig. 2C, left and middle panel). We also found that CD80 showed increased levels in the CD14+ cells conditioned with CD14−*/iMSC, however, the fold change was much lower than that of CD206 and CD163 (1.4 vs. 3.3 and 2.6, respectively) (Fig. 2C, right panel). Taken together, these results validate that monocytes are polarized toward the regulatory M2-phenotype under the iMSC-driven immunomodulatory environment. Simultaneously, we confirmed that a strong inflammatory environment was induced as we observed an increase in CD3+ counts in activated CD14-depleted PBMCs (CD14*, fold change of 8.3) compared to non-activated CD14-depleted PBMCs (CD14−, Supplemental Fig. 1B, uncoloured and dotted bar). We also observed a significant reduction of T cell proliferation in presence of iMSCs (80.1% inhibition) (Supplemental Fig. 1B, orange bar).
Finally, we measured the levels of cytokines present in the supernatants of CD14+ monocytes following conditioning with activated CD14− or CD14−*/iMSC cocultures. Secretion of the M1-type cytokine TNFα (p = 0.0072) was significantly reduced in the co-cultures CD14++CD14−*/iMSC (Fig. 2D, left panel) as compared to CD14+ exposed to activated CD14−. Conversely, CD14+ monocytes exposed to CD14−*/iMSC co-cultures, displayed enhanced amounts of the M2-type cytokine IL-10 (p < 0.0001) (Fig. 2D, right panel). Thus, under inflammatory stimuli, iMSCs exert an immunomodulatory effect on monocytes to induce an anti-inflammatory response, in which the secretome (cytokines) could be playing a crucial role.
iMSCs have an immunomodulatory effect on T lymphocytes and macrophage polarization. (A) Percentage of CD3+ T cell suppressed in PBMCs*/MSCs co-cultures (n = 8) under cell-to-cell contact (direct) or in a transwell system (indirect) with hUCMSCs (left panel) or iMSCs (right panel). PBMCs were stimulated with beads (PBMNC*) in presence of hUCMSCs or iMSCs (PBMNC*+hUCMSC or iMSC). After 5 days cells were collected and stained with an anti-CD3 antibody. (B) Concentration of the pro-inflammatory cytokine TNFα in supernatants (n = 6) of stimulated PBMCs (PBMNC*) alone or in indirect co-culture with hUCMSCs (blue bar) or iMSCs (orange bar). After 5 days in co-culture, supernatants were collected and analysed by ELISA. (C) Expression of the M1 marker CD80 (n = 3) and M2 markers CD206 (n = 6) and CD163 (n = 6) in monocytes (CD14+) co-cultured with hUCMSCs (blue bar) or iMSCs (orange bar) under an inflammation condition (CD14−*). CD14+ cells were purified, seeded in the upper chamber of transwell inserts and exposed to activated CD14− cells (CD14−*) in co-culture with MSCs (CD14++CD14−*/hUCMSC or iMSC) for 5 days. CD14++CD14−* co-cultures were used as a reference sample. (D) Concentration of the cytokines TNFα and IL-10 in supernatants (n = 6) of CD14+/CD14−* and CD14+/CD14−*+MSC (hUCMSCs or iMSCs). After 5 days in co-culture, supernatants were collected and analysed by ELISA. All data are presented as mean. **: p < 0.01, ****: p < 0.0001 by one-way ANOVA followed by Dunnett´s test.
iMSC-EV characterization
We hypothesized that pre-conditioning of iMSCs with pro-inflammatory cytokines (CK) might enhance the production and functionality of iMSC-derived EVs. To evaluate our hypothesis, conditioned medium (CM) was harvested from iMSCs after 48 h in culture with serum deprivation (starvation) or after priming with two pro-inflammatory cytokines (CKs), IL-1β and TNFα. Subsequently, EVs were isolated from the CM via differential centrifugation followed by tangential flow filtration (Fig. 3A). The basic characteristics of these EVs were assessed in comparison with those of EVs collected from hUCMSCs. Quantitative analysis of the particle concentration and size distribution of iMSC-EVs from the two culture conditions showed no significant differences (Fig. 3B and C, respectively). Regarding the starvation condition, we observed a mean size of 156.6 nm ± 17.1 nm and a concentration of 5.00 × 1010 ± 0.96 (x1010) particles/ml. Similarly, the particle size and yield of the pro-inflammatory condition showed an average of 143.6 nm ± 9.7 nm and 6.47 ± 1.02 (x1010) particles/ml. Interestingly, hUCMSCs-derived EVs displayed similar results in both of the culture conditions. However, the particle concentration was lower for both culture conditions compared to iMSC-EVs, pointing to iMSCs as more efficient EV-producing cells (Fig. 3C).
The expression of EV markers (Alix and CD81) in iMSC-EVs and hUCMSC-EVs was confirmed by western blot (Fig. 3D, original blot and image showed in Supplemental Fig. 2). A low level of ApoAI, a common contaminant in EV preparations, was also detected. We further characterized the expression of the three tetraspanins CD9, CD81 and CD63, which are all recognized molecular markers for EVs38, in iMSC-EVs (Fig. 3E). Exoview analysis showed that iMSC-EVs expressed all three of the exosomal markers (Fig. 3E, right panel), however tetraspanin CD81 is more highly expressed in the iMSC-EVs compared to other tetraspanins and, is co-localized with CD63. Notably, iMSC-EVs cultured under starvation or proinflammation conditions displayed no significant differences in expression levels of the three markers. In contrast, a different pattern was measured for all three exosomal markers in hUCMSC-EVs, where it is clear that the three tetraspanins have similar expression levels with minor differences between the EVs prepared in different culture conditions (Fig. 3E, left panel).
Finally, TEM analysis showed that the isolated iMSC vesicles had a spherical morphology surrounded by a bilipid membrane, as expected (Fig. 3F). These results suggest that iMSC-EVs have the general characteristics of EVs.
Biophysical characterization of hUCMSCs and iMSCs-derived EVs. (A) Schematic illustration of EVs isolation from conditioned media (CM) using tangential flow filtration (TFF) method. After harvesting CM, sequential stepwise centrifugation was performed. The cleared CM was filtered through a 0.45 μm PES device before EV concentration using a TFF-easy device. (B) Analysis of particle concentration for hUCMSCs (n = 3) and iMSC-EVs (n = 4) measured by Nanoparticle Tracking Analysis (NTA). Concentration is displayed as a media. (C) Analysis of size distribution for hUCMSCs (n = 3) and iMSC-EVs (n = 4) measured by NTA. Size distributions are displayed as a media. (D) Immunoblotting for CD81, Alix, and APO A I in hUCMSCs and iMSC-EVs. The original image was cropped from different parts of the blot and the original is shown in Supplemental Fig. 2. (E) Expression of membrane markers CD9, CD63 and CD81 measured by Exoview. Tetraspanin colocalization of hUCMSCs and iMSC-EVs using three fluorescent channels and overlay of fluorescent images. Data shown represents the respective tetraspanin and colocalization fraction (%) out of all detected EV. (F) Morphological characterization of EVs isolated from iMSCs by Cryo electron microscopy. Representative images of iMSCs-derived EVs.
The proliferative ability of stimulated T lymphocytes is reduced by iMSC-derived EVs
To assess the biological activity of iMSC-EVs, we aimed to determine their effect on the proliferation rate of lymphocyte subsets. For that, a total of 5.0 × 105 PBMCs were stimulated with anti-CD2-CD3-CD28 coated beads at a bead-to-cell ratio of 1:20, and co-cultured with iMSC-EVs for 5 days. This suboptimal bead-to-cell ratio was selected after preliminary tests demonstrated that the standard recommended 1:2 ratio generated a maximum level of total T-cell stimulation, making it more difficult to reveal the immunomodulatory activity of iMSC-EVs. The proliferation activity was initially determined by calculating the inhibition (expressed as percentage) of T cell proliferation by iMSC-EVs, as described previously in the cell-based assay (Fig. 2). As shown in Fig. 4A, when in vitro stimulated lymphocytes were cultured in the presence of iMSC-EVs, proliferation was decreased in the CD3+ cell population (7.8%), and in both CD4+ (6.3%) and CD8+ (12.3%) T-cell subtypes. Interestingly, iMSC-EVs derived from preconditioned iMSCs displayed significantly enhanced inhibitory effects on activated CD3+ T cells as compared to starvation EVs (24.7% vs. 7.8%, respectively) (Fig. 4A, left panel). Likewise, the proinflammatory condition demonstrated significantly greater inhibition on both CD4+ (27.7% vs. 6.35%) and CD8+ (28.5 vs. 12.3%) T-cell subtypes compared to starvation EVs (Fig. 4A, middle and right panel, respectively). Thus, inflammatory preconditioning of iMSCs greatly improved the immunomodulatory function of EVs. Similar results were obtained with the EVs derived from hUCMSCs on the CD3+, CD4+ and CD8+ T cell populations (Fig. 4A, grey bars).
To further elucidate the biological activity of iMSC-EVs, we assessed the proliferation behaviour of T lymphocytes by CFSE dilution. The CFSE staining was performed before seeding the cells with iMSC-EVs. Stimulated PBMCs without EVs constituted the positive control (activated), and non-stimulated PBMCs served as the negative control. Cells were initially gated to select a viable lymphocyte population. From this population, cells were gated to select the CD3 + T cell population, followed by gating for CD4 + and CD8 + T cell subpopulations. CFSE fluorescence for each subpopulation was analysed using a flow cytometer. A total of seven cell divisions were detected for the CD4 + T cell subtype by CFSE fluorescence (Fig. 4B, upper panel). When stimulated lymphocytes were cultured in the presence of starvation and proinflammation condition iMSC-EVs, the proliferation profile changed in both cases (Fig. 4B, upper panel, left box). Interestingly, a large number of cells presented a low number of cell divisions (G1 to G5), conversely a lower percentage of cells reached the highest number of cell divisions (G6 and G7). A detailed representation showing cell counts in each division cycle is provided in Fig. 4B (upper panel, middle box), and a representative histogram is also provided (upper panel, right box). Here, firstly, it can be seen that under starvation and pro-inflammation condition, iMSC-EVs significantly decreased the cell number of the undivided cell population by 36.3% and 31.4%, respectively (Fig. 4B, middle box, black bars). We suggest that this could be explained by early mechanisms of cell death, cell cycle arrest or both. Secondly, the distribution of cell counts across generations appears to show that iMSC-EVs are arresting CD4 + T cells and preventing their progress through cell division compared to the control (activated). Cell counts in the late generations (G5, G6 and G7) were significantly decreased by 10.6% (G5), 29.5% (G6) and 51.8% (G7) when starvation condition iMSC-EVs were added, compared to the control. A higher effect was observed by addition of pro-inflammation condition iMSC-EVs; a 13.2% (G5), 37.5% (G6) and 58.8% (G7) decrease in cells was seen compared to control. Regarding the CD8 + T cell population, a total of six divisions were detected by CFSE fluorescence (Fig. 4B, down panel). Likewise, the proliferation profile of T lymphocytes changed in the presence of iMSC-EVs from both stimulation conditions (Fig. 4B, down panel, left box). In particular, cell counts in the last generations (G4, G5 and G6) decreased significantly, but the undivided cell population was not affected. Starvation condition iMSC-EVs decreased CD8 + cell counts by 49.5% (G4) and 32.6% (G5) compared to control. Pro-inflammation condition EVs did not show a major effect for these generations, but for the G6 generation the number of cells decreased by 19.8% (Fig. 4B, down panel, middle box). Remarkedly, inflammatory preconditioning of iMSCs enhanced the immunomodulatory function of EVs on CD4 + and CD8 + T cell subpopulations. Similar results were obtained with the hUCMSCs-derived EVs after tracking the proliferation of CD4 + and CD8 + T cell subpopulations (Supplemental Fig. 3).
The proliferative ability of stimulated T lymphocytes is reduced by iMSCs-derived EVs. (A) Percentage of CD3+, CD4+ and CD8+ T cells suppressed by hUCMSC-EVs (n = 4) or iMSCs-EVs (n = 9). PBMCs underwent staining with carboxyfluorescein succinimidyl ester (CFSE), then were subsequently stimulated with beads and cultivated alongside EVs obtained from conditioned medium harvested following starvation or pro-inflammation culture conditions. After 5 days cells were stained with anti-CD3, anti-CD4 and anti-CD8 antibodies and quantified for CFSE dilution by flow cytometry. All data are presented as mean. *: p < 0.05, **: p < 0.01, ****: p < 0.0001 by one-way ANOVA followed by Tukey ‘s multiple comparison test. (B) Change in the proliferation profile of CD4+ T cells and CD8+ T cells by iMSC-EVs (n = 3). A detailed representation is showing the percentage of the total population in each cell division cycle (indicated as G#, left panel), as well as total cell counts (middle panel) and representative histograms (right panel). Red histograms represent PBMCs activated with beads and blue histograms, PBMCs activated and in coculture with iMSC-EVs. All data are presented as mean ± SD. *: p < 0.05, **: p < 0.01, ****: p < 0.0001 by two-way ANOVA followed by Tukey´s multiple comparisons test.
iMSCs-derived EVs regulate macrophage polarization to M2-like phenotype
To further investigate the regulatory effects of iMSC-EVs on macrophage polarization, we isolated monocytes (CD14+) from PBMCs and differentiated them over 4 days into resting macrophages (M0) using macrophage colony-stimulating factor (M-CSF). To determine which type of polarization could be induced by iMSC-EVs, M0 macrophages were treated with the vesicles for an additional 48 h before M1- and M2-like phenotypes were assessed by flow cytometry. M0 macrophages without iMSC-EVs treatment were used as reference. Additionally, M1- and M2-like macrophages (controls) were induced by treatment of M0 with LPS and IFNγ for M1, and IL-4 for M2, following the manufacturer’s protocol. Consistently, flow cytometry analysis revealed that the percentage of CD80 positive cells (M1 macrophages) increased and CD206 positive (M2 macrophage) did not change in the M1-like control and, conversely, CD80 did not change and CD206 increased for the M2-like control compared to M0 (Fig. 5B, dark grey bars). These control results indicate that a reliable in vitro model of macrophage polarization was established. Following iMSC-EV treatment, macrophages showed a slight increase of CD80 positive cells with significant differences for the iMSC-EVs isolated from the pro-inflammatory condition. Interestingly, a greater increase of CD206 positive cells were observed compared to M0, with strong differences between starvation and pro-inflammation condition EVs (Fig. 5B, uncoloured bars). The experimental setup showed a PBMCs donor-dependence for both controls and iMSC-EVs treatments (Fig. 5B). Therefore, to circumvent PBMCs-donor dependence, a M2/M1 ratio was calculated based on flow cytometry data. Surprisingly, data were highly reproducible and consistently showed a significant increase in the M2/M1 ratio after treatment with iMSC-EVs produced in a pro-inflammatory environment. This result was supported by the same, expected, response in the M2-like control (Fig. 5A, orange bars). Taken together, these results suggest that iMSC-EVs successfully induce macrophage polarization toward the anti-inflammatory M2 phenotype, potentially inhibiting inflammation. Unfortunately, we did not observe changes in the concentration of the pro-inflammatory cytokine TNFα and the anti-inflammatory cytokine IL-10 following iMSC-EV treatment. However, it was notable that the pro-inflammatory cytokine TNFα did not increase when M0 macrophages were incubated with iMSC-EVs unlike the increase seen in the M1-like control. This result indicates that iMSC-EVs do not induce an inflammatory environment (Supplemental Fig. 4A). Similar results were obtained when macrophages were incubated with hUCMSCs-derived EVs (Fig. 5A, blue bars).
Based on our first experiments, iMSC-EVs were unable to repolarize M1 macrophages into M2 macrophages. However, further experiments are essential since we found a different response when M0 macrophages were first incubated with EVs, and then induced to the M1-like phenotype with corresponding cytokines after a few hours. Therefore, this incubation time needs to be systematically tested (Supplemental Fig. 4B, C). Furthermore, analysing flow cytometry data, we observed a significant increase in the M2/M1 ratio when M0 macrophages polarized to the M1-like phenotype were pre-incubated with hUCMSC-EVs from the pro-inflammatory condition (Supplemental Fig. 4B, blue bars). This result revealed that EVs derived from primary MSCs, such as hUCMSCs, contain the immunomodulatory potential to repolarize M1 to M2 macrophages and therefore we hypothesize a similar functional potential for iMSC-EVs.
iMSCs-derived EVs modulate macrophage polarization to M2-like phenotype. (A) Analysis of the shift in the proportion of M1 and M2 populations, represented as M2/M1 ratio, after macrophages were cultured in the presence of hUCMSC-EVs (n = 3–4, blue bars) or iMSC-EVs (n = 5, orange bars). Monocyte cells (CD14+) were purified from PBMCs and differentiated into macrophages. After 4 days, macrophages were exposed to EVs obtained from conditioned medium harvested following starvation and pro-inflammation culture conditions. After 48 h, cells were stained with anti-CD80 and anti-CD206 antibodies. All data are presented as means. *: p < 0.05, **: p < 0.01 by one-way Brown-Forsythe and Welch ANOVA test followed by Dunnett´s T3 multiple comparisons test. (B) Analysis of the expression for the M1 marker CD80 and M2 marker CD206 for two PBMCs donors (donor 1 and 2). Changes in fluorescence were compared to the marker’s expression in macrophages M0. For activation controls (M1- and M2-like), macrophages were activated with specific cytokines according to a predefined following protocol. All data are presented as mean. *: p < 0.05, **: p < 0.01, ****: p < 0.0001 by one-way ANOVA followed by Dunnett´s test.
iMSC-EVs enhance fibroblast migration in a wound healing assay
Recent research efforts have focused on harnessing EVs as a powerful therapeutic tool in tissue repair and regeneration. We studied the ability of iMSC-EVs to induce dermal fibroblast migration during wound healing by using the scratch assay method. Dermal fibroblasts (HDFa) were plated to confluence. Prior to creation of the scratch, cells were treated with vehicle (PBS, negative control, NC), EGF (50 ng/ml, positive control, PC) or iMSC-EVs. After creation of the scratch, treatments remained on the cells for the entire imaging time course of90 h. The scratch assay revealed that iMSC-EVs treatment yielded faster wound healing over time compared to the negative control (PBS) (Fig. 6A, orange and black line, respectively). Furthermore, the pro-regenerative potential of iMSC-EVs was similar to hUCMSC-EVs (Fig. 6, orange and blue line, respectively). Interestingly, the pro-regenerative potential of iMSC-EVs was not increased by priming iMSCs with pro-inflammatory cytokines prior to EVs collection and purification (Fig. 6, purple line). Representative light microscopy images of scratch wound at beginning, middle, and end of experiment are shown (Fig. 6B).
Migration capacity of HDFa cells is increased after iMSC-derived EV treatment. Wound healing assay of skin cells treated with EVs. (A) Analysis of relative wound density after hUCMSC-EV (n = 9, blue line) or iMSC-EV (n = 9, orange and purple lines) treatment. HDFa cells were incubated with hUCMSC-EVs or iMSC-EVs and imaged every 6 h for 90 h. The cell confluence within the wound was normalized against that obtained at 0 h. All data are presented as mean ± SD. ns: no significant difference by two-way ANOVA followed by Tukey’s multiple comparisons test. (B) Representative light microscopy images of scratch wound at beginning, middle, and end of experiment. The relative wound density was calculated using the Incucyte Scratch Wound Analysis Software Module. NC: negative control (starvation media with PBS); PC: Positive control (starvation media supplemented with EGF). Scale bars are 200 nm.
Discussion
The generation of induced pluripotent stem cell (iPSC)-derived MSCs (iMSCs) has opened an exceptional opportunity to solve most of the hindrances limiting the broad application of MSCs as advanced cell therapy39. To shed light on the potential of iMSCs and iMSC-derived EVs for biomedical applications, extensive research of their functionality is still under way. In the present study, we compared the therapeutic efficacy of iMSCs and their EVs with human primary umbilical cord-derived MSCs (hUCMSCs) in in vitro cellular assays. We initially confirmed that GMP-compliant iPSCs could be used to generate iMSCs which fulfil the characterisation criteria of primary MSCs, and that these release bona fide-EVs. Subsequently, we found that our iMSCs effectively exert an immunomodulatory potential to inhibit T cell proliferation and induce macrophage polarization toward M2-like phenotype as hUCMSCs. Moreover, we found that iMSC-EVs exhibited substantial immunomodulatory properties similar to iMSCs cells, further showing similar pro-regenerative potential as hUCMSC-derived EVs.
Primary hUCMSCs are the second major source used for clinical purposes, mainly due to their easy access and stronger immunomodulatory effects in vitro and in vivo than MSCs from classical sources such as bone marrow or adipose tissue40,41. Our study demonstrates that generated iMSCs exhibit a comparable ability to regulate the proliferation of activated lymphocytes through direct cell-to-cell interactions and cell-contact independent mechanisms. Therefore, we highlight the potent immunomodulatory effect of iMSCs generated from iPSCs produced from umbilical cord blood cell progenitors, following the developed protocol. These data are consistent with studies reporting similar findings for MSC-like populations generated from embryonic stem cells (hESC-MSCs)42, as well as different human iPSC-MSC lines generated from periodontal ligament and gingival tissue5. Of note, studies have shown differences in the immunomodulatory properties between various iPSC-MSCs lines2,5, suggesting that they may be attributed to tissue of origin43.
iMSC populations have been found to modulate immune cells through various mechanisms including secretion of different cytokines and soluble factors to suppress B and T cell proliferation, inhibit monocyte maturation, and induce the generation of regulatory T cells and M2 macrophages44,45,46,47,48. In this context, reports have suggested that iMSC populations derived from pluripotent cells use cytokines and soluble factors similar to those of bone marrow-derived MSCs to suppress T lymphocyte proliferation in allogenic mixed lymphocyte reaction assays49,50. In our experimental setting we detected a reduced level of the pro-inflammatory cytokine TNFα in the cell-contact independent iMSCs/PBMCs co-cultures, suggesting that TNFα plays a role in the immunomodulatory effect of iMSCs on T lymphocytes. A previous study also reported other soluble factors that are implied to be involved in the immunomodulatory potential of iMSCs. Ng et al., reported a general trend of elevated IDO1 and IL-6 transcript levels in iPSC-MSC lines and their respective primary cells co-cultured with activated PBMCs, albeit at varying levels5. By contrast, we observed a comparable secretion dynamic of the cytokine TNFα between iMSCs and primary hUCMSCs in co-cultures with activated PBMCs. These data imply that TNFα may be a common factor utilized by different mesenchymal cell populations to induce their immunomodulatory effects. Moreover, our data provide additional findings from the secretome of iMSCs under an inflammatory environment, and support the notion that iPSC-derived MSCs modulate a cocktail of different pro- and anti-inflammatory cytokines, along with soluble factors to regulate different immune cell populations5,51,52.
Monocyte/macrophages are reported to play critical roles in the therapeutic effects of MSCs in several diseases37. Accordingly, we evaluated the differentiation and activation status of monocyte populations under iMSCs-driven immunomodulation. We confirmed that iMSCs cultured under inflammation conditions induced the reprogramming of monocytes (CD14 + cells) toward the regulatory M2-phenotype. Moreover, we observed comparable results with primary hUCMSCs, strongly supporting the potent immunomodulatory effect of iMSCs generated from iPSCs. Similar to our study, Kim et al. directly compared the therapeutic efficacy of iPSC-MSCs (iMSCs) with BM-MSCs in NOD mice with secondary Sjögren’s syndrome (sSS) and found that iMSCs inhibited the onset of lymphocyte infiltration into salivary glands in the NOD mouse model of SS in the same way as BM-MSCs34.
So far, secretome and transcriptome data have suggested a critical role of inflammation-related cytokines and chemokines in the differentiation and activation of myeloid cells in hUCMSC-mediated immunomodulation36. Similar to that reported for hUCMSCs, we observed a decrease in the secretion of the M1-type cytokine, TNFα, and an increase in the M2-type cytokine, IL-10 under iMSC-driven immunomodulation, with variable levels compared to hUCMSCs. Therefore, we suggest that different mechanisms are likely to dictate the efficiency of immunosuppression exhibited by iMSCs and primary hUCMSCs toward the monocytes/macrophage population. Even if the few factors examined in this study limit the repertoire of probable mechanisms used by immunomodulatory cells, these analyses support the hypothesis that iMSCs can also induce a specific immunomodulatory milieu that reprograms monocytes toward an M2-like phenotype. Consequently, monocytes become more tolerogenic in a cytokine microenvironment highly influenced by iMSCs. Our in vitro findings are consistent with an in vivo study which investigated the neuroprotective effects of iPSC-MSCs in a rat model of cardiac arrest (CA). The study demonstrated that iPSC-MSCs protect rats against ischemia/reperfusion injury by promoting macrophage transition towards the M2 phenotype after CA53. Until now, we have not found in vitro studies performed to investigate the interaction between iMSCs and myeloid cells, thus our study demonstrates for first time the immunomodulatory potential of iMSCs in an in vitro model of macrophage polarization. We suggest that cytokines are one part of the mechanisms used by iMSCs to reprogram monocytes towards an immune tolerance environment. These findings will help improve the comprehension of the molecular mechanisms underlying iMSCs-mediated immunomodulation and provide strategies leading to the clinical translation of iMSCs for the treatment of immune-mediated diseases.
It has been suggested that the main mechanism of stem cell transplant therapy depends on paracrine activity. Among these paracrine bioactive substances, EVs, secreted by cells, have attracted increasing attention. MSC-EVs have become promising immunomodulators as they possess similar properties to their parental MSCs54,55,56. In this study, we aimed to investigate the immunomodulatory role of an enriched fraction of iMSC-EVs on T cell subsets and macrophage polarization with in vitro settings. Our study demonstrated EV-enriched fractions isolated from iMSCs triggered similar immunomodulatory potency as hUCMSC-derived EVs, in an in vitro model of inflammation. Although this is a preliminary study, our results showed that iMSC-EVs significantly abrogated the proliferation of CD4+ and CD8+ T cells. Moreover, we suggest that iMSC-EVs differentially affect T cell subsets and that the mechanisms underlying the inhibition might be related to cell death or cell cycle arrest, or both. We also noticed that the iMSC-EV-driven inhibition of CD3+ T cells was less strong than the inhibition in the contact-independent cultures with iMSCs cells (24.7% vs. 49.3%), suggesting that almost half of lymphocyte proliferation inhibition is driven by iMSC-EVs, and that other molecules and nanoparticles present in the secretome contribute to the entire immunomodulatory effect. Afterward, we wanted to investigate whether iMSC-EVs can affect the balance between M1 and M2 macrophages. We demonstrated that iMSC-EVs could induce the M2 phenotype in macrophages as indicated by the reduction of the level of CD80, a M1 cell surface marker, and the upregulation of the CD206, a M2 marker. These results suggest that iMSC-EVs successfully induced macrophage polarization toward the anti-inflammatory phenotype M2 and consequently could inhibit inflammation.
Similar to our purpose, a previous study evaluated the immunomodulatory effect of iMSC-EVs on macrophages using a diverse in vitro setting. The authors employed the murine macrophage cell line RAW 264.7 and their results suggest that large iMSC-EVs (lEVs) could repolarize the macrophages from M1 phenotype towards M2 phenotype32. Published studies using iMSC-EVs in several models of disease have also addressed the immunomodulation potential for these nanoparticles. Ye et al. demonstrated that large iMSC-EVs possessed anti-inflammatory functions and analgesic effects in the rat tendinopathy model by regulating macrophage heterogeneity32. A further study, demonstrated that treatment with small EVs (iMSC-sEV) attenuated sepsis pulmonary inflammation in an experimental rat model and exerted anti-inflammatory effects on alveolar macrophages (AMs) in vitro, through AM internalization of iMSCs-sEV which alleviated the release of inflammatory factors31.
We further aimed to boost the potency of EVs by conditioning the EV-producer iMSCs. We observed that inflammatory preconditioning of iMSCs enhanced the immunomodulatory function of EVs to inhibit CD4+ and CD8+ T cell proliferation and polarized macrophages to the M2 phenotype, similar to the enhanced immunomodulation widely described for EVs derived from primary MSCs. Regardless of the PBMNC donor, iMSC-EV treatment increased the M2/M1 ratio in the macrophages. Accumulating evidence has suggested that an inflammatory microenvironment induces MSC activation to exert their powerful immunomodulatory responses48,57,58,59,60. Taken together, our results strongly suggest for the first time that an inflammatory stimulus may be fundamental in inducing the release of immunotherapeutic EVs from iMSCs.
The therapeutic effects of iMSC-derived EVs have been explored in many diseases but have been best studied in wound healing30,61,62, cardiovascular disease63,64,65 and musculoskeletal pathology66,67. Therefore, we also explored whether EVs derived from iMSCs can exert comparable functions to those from hUCMSCs in promoting the migration of skin cells in a wound healing assay. We demonstrated that iMSC-EVs accelerated the migration of HDFa cells in a similar way to hUCMSC-EVs. Our results are in line with previous studies pointing to promising therapeutic effects of iMSC-EVs in a wide range of preclinical models of tissue repair, including kidney, liver, lung, myocardial infarction, cerebral artery occlusion, and skin wound healing68,69. Excitingly, a recent study from Zhang et al. showed that iMSC-derived EVs accelerate the process of wound repair via enhancing collagen synthesis and angiogenesis resulting in stimulation of proliferation and migration of human dermal fibroblast (HDFa cells) and human umbilical vein endothelial cells (HUVECs)30. Furthermore, iMSC-derived EV therapy in diabetic ulcer mice demonstrated faster wound healing and closure rate61. Studies have also suggested common mechanisms in the activation of skin cells between adult MSCs and iMSC-derived EVs. However, some differential effects exist and are thought to be due to distinct biological cargos, for example, surface mitogens and mRNA62,70,71. Therefore, more research is needed to further our mechanistic understanding. Until now, only a few reports have begun to explore the underlying mechanisms iMSC-EV application in various disease models. Interestingly, these studies have established that iMSC-EVs encapsulate diverse bioactive components including RNAs and proteins from the parental cell iMSC, which contribute to cell-to-cell communication and regulate cell behaviours31,32.
Overall, we have shown that iMSCs can have immunomodulatory and regenerative effects however our study does have several limitations: (1) Different concentrations of iMSC-EVs should be investigated in dose/response experiments, which are used to evaluate the dosage of clinical medication. (2) Testing batch-to-batch variations in their ability to modulate functional assays is necessary. In this regard, some studies have demonstrated that EVs from early-passage iMSCs have better immunomodulatory potency than EVs from late-passage iMSCs in TLR4-stimulated splenocytes72. Therefore, it is imperative to perform batch-to batch functional testing for downstream applications.
In conclusion, our findings provide additional scientific support regarding the immunomodulatory potential of iMSCs and validate that iMSC-EVs demonstrated significant properties akin those of iMSCs in mediating anti-inflammatory and regenerative processes. In agreement with other authors, we strongly promote the development of iMSC-EVs as a novel cell-free medicinal product for the treatment of immune-related diseases that required simultaneous modulation of tissue repair. Furthermore, this study provides for the first-time evidence on pro-inflammatory preconditioning of iMSCs as a feasible and compelling strategy to enhance the immunomodulatory effects of iMSC-EVs on human immune cells.
Materials and methods
Expansion of hUCMSCs
Human umbilical cord-derived MSCs (hUCMSCs) obtained from RoosterBio (RoosterVial-hUC-1 M-XF, Part N. C43001UC, RoosterBio, Inc., Frederick, MD USA) were expanded and cultured according the manufacturer’s protocol. Cells were cultured in T75 CellBIND Flasks (Corning, NY USA) with the medium RoosterNourish-MSC-XF (Part Number SU-005/022 + SU-016, RoosterBio, Inc., Frederick, MD USA) supplemented with 1% penicillin/streptomycin 10,000 U/mL (ThermoFisher Scientific, Waltham, MA USA) at 37 °C in 5% CO2. Upon reaching > 80% confluency, cells were detached with TrypLE Express (ThermoFisher Scientific, Waltham, MA, USA) and split into culture flasks using a seeding density of 3,000 cells/cm2 (min. > 2,000 cells/cm2).
Generation of iMSCs
The hiPSCs line TC-1133 was purchased from Lonza. hiPSCs were seeded at 1.0 × 103 cells/cm2 in T25-flasks coated with Vitronectin XF (StemCell Technologies, Germany, Cat. 07180) in Essential 8 (E8, ThermoFisher Scientific, Waltham, MA USA, Cat. A1517001) media containing 10 µM Y27632 Rock-Inhibitor (StemCell Technologies, Germany, Cat. 72302). hiPSCs were maintained in E8 for two additional days and then induced to differentiate at “day 0” with differentiation media consisting of 10 ng/mL bFGF (Cat. 78003), 4 µM SB431542 TGFβ inhibitor (Cat. 72232) and 4 µM WNT agonist CHIR99021 (Cat. 72052) (all from StemCell Technologies, Germany) in Essential 6 media (E6, ThermoFisher Scientific, Waltham, MA USA, Cat. A1516401). Differentiation media was changed daily for the next 5 days. On day 6, the differentiated neural crest stem cells (NCSCs) were plated as “multipotent passage 0” (MP0) on Vitronectin coated T75-flasks in serum-free MSC culture media NutriStem (MSC NutriStem XF Basal Medium, NutriStem XF Supplement Mix, Sartorius, Germany) at a density of 4.0 × 104 cells/cm2. When 90% confluency was reached, the cells were replated at a density of 2.6 × 104 cells/cm2 for MP1–MP2. From MP3 to MPX cells were plated w/o Vitronectin on CellBIND flasks (Corning, NY USA) in NutriStem supplemented with 2.5% human Platelet Lysate (hPL, StemCell Technologies, Germany) and were passaged when 90% confluency was reached. All the experiments were performed with iMSCs in early passage (12 to 15).
Immunophenotyping of hUCMSCs and iMSCs
MSCs were characterized by immunophenotyping using the membrane markers CD90 APC (SE10), CD73 PE/Cy7 (AD2), CD105 PE (SN6h), HLA-ABC FITC (W6/32), CD34 PerCP/Cy5.5 (561), CD45 APC/Cy7 (2D1) and HLA-DR Pacific Blue (L243) antibodies or corresponding isotype controls (all from BioLegend, Inc., CA USA) and analysed by flow cytometry (Cytoflex LX, Beckman Coulter Life Sciences, IN USA). Cells were trypsinised and washed twice prior to resuspension in FACS buffer containing PBS 1X, 0.5% BSA and 2 mM EDTA. Cells were adjusted to 1.0 × 105 in 100 µL of FACS buffer. For cell surface labelling, cell suspensions were incubated at 4 °C for 30 min with the corresponding titrated antibody.
Production and isolation of EV
Upon reaching 70% confluency, MSCs were further cultured for 48 h under two conditions: starvation and pro-inflammation. Starvation consisted of DMEM low glucose (Gibco, ThermoFisher Scientific, Waltham, MA USA, Cat.11885084,) or MSC NutriStem XF Basal Medium (Sartorius, Germany) without supplements for hUCMSCs and iMSCs, respectively. Pro-inflammation consisted of DMEM or NutriStem medium supplemented with human recombinant TNF-α and IL-1β 10 ng/mL each (premium grade, Miltenyi Biotec, Germany). After incubation, conditioned media (CM) were collected and sequentially centrifuged for 5 min at 106 xg, 10 min at 425 xg and 30 min at 1,699 xg at 4 °C. Resulting supernatants (SN) were filtered through a 0.45 μm PES filter device and subjected to Tangential Flow Filtration (TFF-Easy, HansaBiomed/Lonza, Tallinn, Estonia) for EV concentration. The EV concentration process follows the manufacturer’s protocol. Each 150 mL of CM was concentrated to 1.5 mL in PBS 1X filtered by 0.45 μm CA.
Characterization of EV
Particle size distribution and concentration were determined by Nanoparticle tracking analysis (NTA) using NanoSight NS300 (Malvern Instruments, Malvern, UK). A syringe pump with a speed of 20 µL/min was used and cell temperature was set at 25 °C. The embedded laser wavelength was 488 nm, and the particles were imaged three times with a 60 s acquisition time. Samples were diluted (1:100-1:200) in 1X PBS (filtered through 0.45 μm CA) to ensure an accurate measurement with camera levels and detection thresholds kept the same between EV samples. Data was analysed using NTA software 2.3 (https://www.malvernpanalytical.com/en/support/ product-support/software/nanosight-nta-software-update-v3-2).
The morphology of EV was analysed by cryo-EM. Briefly, freshly glow-discharged carbon grids (EMResolutions) were placed inside the chamber of the EM GP2 Automatic Plunge Freezing (Leica Company). Four microliters of the sample were dropped onto the grid for 30 s, blotted and the grid was abruptly plunged into a liquid ethane bath, automatically set to -184 °C. The vitrified grid was removed from the plunger and imaged with JEM-2200FS/CR (JEOL Europe) transmission electron microscope. The images were recorded using serialEM software 4.0 (https://bio3d.colorado.edu/SerialEM/) at nominal magnifications of 25,000 X with a pixel size of 0.473 nm. Different particles types were catalogued manually and used for training and auto-picking using TOPAZ software v3.2.0 (https://www.topazlabs.com/down loads).
The expression of EV typical markers was determined by using western-blot analysis. EV samples were analysed with the following antibodies, all at a 1:1000 dilution: anti-CD81 (clone M38; Invitrogen, Carlsbad, CA; Cat. #10630D), anti-Alix (EP23653-32; Abcam, Cambridge, UK; Cat. #ab275377) and anti-Apolipoprotein A I (EP1368Y; Abcam, Cambridge, UK; Cat. #ab52945). Briefly, EV samples in PBS 1X were prepared with 6X sample buffer with (reducing) or without (non-reducing) 50 mM DTT to a final 1X concentration and heated at 70 °C for 10 min. Samples were then subjected to SDS-PAGE and followed by transfer to a PVDF membrane (ThermoFisher Scientific, Waltham, MA, Cat. #88518). PVDF membranes were blocked and incubated with primary antibodies in 1% milk in TBS-T overnight at 4 °C. Washed membrane was probed with a mouse and rabbit HRP-conjugated secondary antibodies at a 1:10,000 dilution (Abcam, Cambridge, UK; Cat. ab97051 and ab6789). Signals were developed using the Pierce ECL Western Blotting Substrate (ThermoFisher Scientific, Waltham, MA, Cat. #32106) and imaged using a Vilber Fusion Fx Spectra imaging system.
ExoView (ExoView R100, NanoView Biosciences, Boston, MA, USA) was used to detect the tetraspanin markers CD9, CD81 and CD63 (EV-TETRA-C). Briefly, microarray chips coated with antibodies against anti-CD9 (HI9a), -CD63 (H5C6) and -CD81(JS-81) were pre-scanned according to the manufacturer’s instructions. All samples were diluted in incubation solution and 150 µL of the diluted samples were placed on the chips overnight. The next day, a detection antibody cocktail containing anti-CD9 (CF-488-labeled), -CD63 (CF-647-labeled) and -CD81 (CF-555-labeled) in blocking buffer, was added to the chips. After incubation at room temperature for 1 h, chips were subjected to three cycles of washing. The chips were then dried and scanned using the ExoView Scanner software version 3.1. The total particle counts and immunofluorescence of the EVs were analysed using the ExoView Analyzer software version 3.1 (https://www.unchainedlabs.com/leprechaun/).
In Vitro T cell assay
Human peripheral blood mononuclear cells (PBMCs) from healthy donors were obtained from StemCell (StemCell Technologies, Germany, Catalog #70025). For the assay with cells (hUCMSCs and iMSCs), two MSCs/PBMCs coculture settings were evaluated: direct (cell-to-cell contact) and indirect (transwell) co-cultures. PBMCs were thawed and cultured for 24 h in RPMI-1640 medium supplemented with 10% FBS (Gibco, ThermoFisher Scientific, Waltham, MA USA) and 1% penicillin/ streptomycin 10,000 U/mL (ThermoFisher Scientific, Waltham, MA USA). In parallel, MSCs were seeded at 5.0 × 104 cells/well in a 24-well plate and cultured overnight in the corresponding media. Next day, PBMCs were stimulated using anti-CD2, -CD3, -CD28 T cell activation beads (Miltenyi Biotec, Germany, Ref. 130-091-441) in a ratio 1:2 (bead: PBMCs) following manufacturer’s instructions. Immediately, MSC medium was removed and 5.0 × 105 PBMCs were added. For indirect co-cultures, MSCs were seeded at the bottom of the well and pre-stimulated PBMCs seeded on top of transwell inserts 0.4 μm PET membrane (Falcon, Corning, NY, USA). Following 120 h of incubation, supernatants were collected and stored at -80 °C for subsequent ELISA assays, and the cells were harvested for CD3+ staining and analysis by flow cytometry. Cells were initially gated to select a viable lymphocyte population. From this population, cells were gated to select the CD3 + population.
For the assay with MSC-EVs, T cell proliferative response was evaluated as follow: PBMCs were thawed and pre-labelled with 0.5 µM carboxyfluorescein diacetate succinimidyl ester (CFSE, ThermoFisher, Waltham, MA USA, Ref. C34554,). The CFSE dye concentration was standardized at 0.5 µM after testing a range of 0.1 µM to 10 µM as this was determined to be the best experimental condition to avoid losing proliferation ability and to be able to track most generations. After labelling, PBMCs were stimulated using anti-CD2, -CD3, -CD28 T cell activation beads in a ratio 1:20 (bead: PBMCs). In 24-well flat-bottomed plates, 5.0 × 105 bead-activated PBMCs were seeded per well and incubated with freshly prepared EVs using a ratio of 1: 8,000 (PBMCs: EVs), in a total volume of 500 µL per well. Cells were incubated for 5 days at 37 °C with 5% CO2. Following 120 h of incubation, supernatants were collected and stored at -80 °C for the subsequent ELISA assays, and cells were harvested for T cell subtypes analysis and tracking proliferation by flow cytometry. Cells were initially gated to select a viable lymphocyte population. From this population, cells were gated to select the CD3 + population, followed by gating for the CD4 + and CD8 + subpopulations.
In Vitro Macrophage polarization assay
For the assay with cells (hUCMSCs and iMSCs), MSCs were seeded at 5.0 × 104 cells/well in a 24-well plate and cultured overnight in the corresponding media. Next day, PBMCs were thawed and the CD14+ population (CD14 positive fraction, CD14+) was isolated by positive selection using magnetic anti-CD14 microbeads (Miltenyi Biotec, Germany). Subsequently, the monocyte-depleted-PBMCs (CD14 negative fraction, CD14-) were resuspended in RPMI 1640 Very Low Endotoxin medium (PAN Biotech, Germany) supplemented with 10% FBS and stimulated with anti-CD2, -CD3, -CD28 T cell activation beads in a 1:2 ratio (bead: CD14-) (bead-activated CD14 negative fraction, CD14-*). Immediately, MSC medium was removed and 5.0 × 105 CD14-* cells added. Furthermore, 3.5 × 105 isolated CD14+ were added on top of transwell inserts with a 0.4 μm PET membrane (Falcon, Corning, NY, USA). CD14++CD14-* co-cultures were used as a reference sample. Assays were incubated for 120 h at 37 °C with 5% CO2. After incubation, pre-conditioned CD14+ (reference and MSCs-conditioned) cells were harvested for phenotype characterization and supernatants collected and stored at -80 °C for the subsequent ELISA assays.
For the assay with MSC-EVs, macrophage polarization was evaluated as follow: CD14+ monocytes were isolated as described above and 2.5 × 105 cells/well were differentiated to macrophages. Macrophage differentiation was performed using the ImmunoCult-SF Macrophage Medium (StemCell Technologies, Germany) supplemented with 50ng/mL human recombinant M-CSF (StemCell Technologies, Germany) following manufacture´s protocol. After incubation for 96 h, freshly prepared EVs were added in a ratio of 1:6,500 (CD14+: EVs) and assays were incubated for additional 48 h. For M1 and M2 activation controls, macrophages were activated with 10 ng/mL LPS (InvivoGen, Toulouse France, Ref. tlrl-3pelps) and 50 ng/mL IFNγ (StemCell Technologies, Germany) for M1-like phenotype, and with 10 ng/mL IL-4 (StemCell Technologies, Germany), for M2-like phenotype, following manufacture´s protocol (StemCell Technologies, Germany). Following incubation, cells were harvested for phenotype characterization and supernatants collected and stored at -80 °C for the subsequent ELISA assays. Initially, the cells were gated to select a viable population, ensuring that only live cells were included in the subsequent analysis. From this viable population, further gating was performed to identify and select cells expressing CD80+, CD163 + and CD206+.
In vitro wound scratch assay
Adult human Dermal Fibroblasts (HDFa; ThermoFisher Scientific, Waltham, MA, USA, Cat. C0135C) were seeded in complete media (DMEM (ThermoFisher Scientific, Waltham, MA USA, Cat. 41966029) supplemented with 10% FBS and 1% penicillin/streptomycin (P/S), at a density of 7.0 × 103 cells/well in Incucyte Imagelock 96-well culture plates (Sartorius, Germany, Cat. BA-04856). The next day, media was changed to starvation conditions (DMEM + 1% P/S) and cells were cultivated for an additional 48 h. Freshly prepared EVs in PBS were diluted in starvation media to a final concentration of 3.0 × 109 particles/ml and added to wells 1–2 h prior to creating a scratch. Positive (EGF 50 ng/ml, StemCell Technologies, Germany) and negative (PBS) controls were also prepared in starvation media. A uniform scratch was made in each well with the Incucyte Woundmaker Tool (Sartorius, Germany, cat. 4563). Images were obtained every 6 h for a total of 90 h using an Incucyte SX5 (Sartorius, Germany) and automated image analysis was completed using the Incucyte Scratch Wound Analysis Software Module (Sartorius, Germany, https://shop.sartorius.com/de/p/incucyte-scratch-wound-analysis-software-module/9600-0012).
Flow cytometry analysis
For T cell subset immune phenotyping, cells were stained with CD3-PE/Cyanine7 (HIT3a), CD4-Brilliant Violet 421 (OKT4) and CD8-APC (SK1) (all from BioLegend, Inc., CA USA). For macrophage immune phenotyping, cells were stained with CD80-APC (2D10) and CD206-PE/Cyanine7 (15 − 2) antibodies. For all flow cytometry analyses, isotype controls were included. A Cytoflex LX instrument (Beckman Coulter Life Sciences, IN USA) was used for data collection and data sets were analysed using FlowJo v10.8.1 data analysis software (TreeStar, Inc., San Carlos, CA, USA, https://www.flowjo.com/solutions/flowjo/downloads).
Enzyme-linked immunosorbent assay (ELISA)
The protein levels of TNFα (sensitivity 3.5 pg/mL) and IL-10 (sensitivity 2pg/mL) in cell culture undiluted supernatants were detected using the corresponding human ELISA kit according to the manufacturer’s instructions (BioLegend, Inc., CA USA).
Statistical analysis
GraphPad Prism 10 (La Jolla, CA, USA) was used for statistical analysis. All data are expressed as mean ± SD unless otherwise indicated. Two-way ANOVA followed by Sisak’s multiple comparisons test was used for MSC immunophenotyping. One-way ANOVA followed by Dunnett´s test was used for T cell and macrophage polarization assays with MSC. One-way ANOVA followed by Tukey’s multiple comparison test was used for T cell subtypes assays with MSC-EV. Two-way ANOVA followed by Tukey’s multiple comparisons test was used for tracking T cell proliferation with MSC-EVs. One-way Brown-Forsythe and Welch ANOVA test followed by Dunnett´s T3 multiple comparisons test was used for macrophage assays with MSC-EVs. And two-way ANOVA followed by Tukey’s multiple comparisons test was used for scratch assays with MSC-EVs. P-value ≤ 0.05 were considered significant.
Data availability
Data supporting the findings of this work are available within the paper and Supplemental Information files.
References
Squillaro, T., Peluso, G. & Galderisi, U. Clinical trials with mesenchymal stem cells: an update. Cell. Transpl. 25 (5), 829–848 (2016).
Hynes, K., Menicanin, D., Mrozik, K., Gronthos, S. & Bartold, P. M. Generation of functional mesenchymal stem cells from different induced pluripotent stem cell lines. Stem Cells Dev. 23 (10), 1084–1096 (2014).
Lian, Q. et al. Functional mesenchymal stem cells derived from human induced pluripotent stem cells attenuate limb ischemia in mice. Circulation. 121 (9), 1113–1123 (2010).
Robinton, D. A. & Daley, G. Q. The promise of induced pluripotent stem cells in research and therapy. Nature. 481 (7381), 295–305 (2012).
Ng, J. et al. Immunomodulatory properties of Induced Pluripotent Stem cell-derived mesenchymal cells. J. Cell. Biochem. 117 (12), 2844–2853 (2016).
Dupuis, V. & Oltra, E. Methods to produce induced pluripotent stem cell-derived mesenchymal stem cells: mesenchymal stem cells from induced pluripotent stem cells. World J. Stem Cells. 13 (8), 1094–1111 (2021).
Zhou, A. K. et al. Using Pre-clinical studies to explore the potential clinical uses of Exosomes secreted from Induced Pluripotent Stem cell-derived mesenchymal stem cells. Tissue Eng. Regen Med. 20 (6), 793–809 (2023).
Bloor, A. J. C. et al. Production, safety and efficacy of iPSC-derived mesenchymal stromal cells in acute steroid-resistant graft versus host disease: a phase I, multicenter, open-label, dose-escalation study. Nat. Med. 26 (11), 1720–1725 (2020).
Saetersmoen, M. L., Hammer, Q., Valamehr, B., Kaufman, D. S. & Malmberg, K. J. Off-the-shelf cell therapy with induced pluripotent stem cell-derived natural killer cells. Semin Immunopathol. 41 (1), 59–68 (2019).
McGrath, M. et al. GMP-compatible and xeno-free cultivation of mesenchymal progenitors derived from human-induced pluripotent stem cells. Stem Cell. Res. Ther. 10 (1), 11 (2019).
Fernandez-Rebollo, E. et al. Senescence-Associated Metabolomic phenotype in primary and iPSC-Derived mesenchymal stromal cells. Stem Cell. Rep. 14 (2), 201–209 (2020).
Luo, L. et al. Feeder-free generation and transcriptome characterization of functional mesenchymal stromal cells from human pluripotent stem cells. Stem Cell. Res. 48, 101990 (2020).
Ozay, E. I. et al. Cymerus™ iPSC-MSCs significantly prolong survival in a pre-clinical, humanized mouse model of graft-vs-host disease. Stem Cell. Res. 35, 101401 (2019).
Chang, Y. H., Wu, K. C. & Ding, D. C. Induced Pluripotent Stem Cell-differentiated chondrocytes repair cartilage defect in a rabbit osteoarthritis model. Stem Cells Int. 2020, 8867349 (2020).
Sabapathy, V. & Kumar, S. hiPSC-derived iMSCs: NextGen MSCs as an advanced therapeutically active cell resource for regenerative medicine. J. Cell. Mol. Med. 20 (8), 1571–1588 (2016).
Gao, W-X. et al. Effects of mesenchymal stem cells from human induced pluripotent stem cells on differentiation, maturation, and function of dendritic cells.
Colter, D. C., Class, R., DiGirolamo, C. M. & Prockop, D. J. Rapid expansion of recycling stem cells in cultures of plastic-adherent cells from human bone marrow. Proc. Natl. Acad. Sci. U S A. 97 (7), 3213–3218 (2000).
Zuk, P. A. et al. Human adipose tissue is a source of multipotent stem cells. Mol. Biol. Cell. 13 (12), 4279–4295 (2002).
Miranda, R. A., Galván Cabrera, J. A. & de León Delgado, J. Propiedades inmunomoduladoras de las células madre mesenquimales. Revista Cubana de Hematología, Inmunología y Hemoterapia. ;31:20–31. (2015).
Raposo, G. & Stoorvogel, W. Extracellular vesicles: exosomes, microvesicles, and friends. J. Cell. Biol. 200 (4), 373–383 (2013).
Nawaz, M. et al. Extracellular vesicles: evolving factors in Stem Cell Biology. Stem Cells Int. 2016, 1073140 (2016).
Wen, S. et al. Mesenchymal stromal cell-derived extracellular vesicles rescue radiation damage to murine marrow hematopoietic cells. Leukemia. 30 (11), 2221–2231 (2016).
de Godoy, M. A. et al. Mesenchymal stem cells and cell-derived extracellular vesicles protect hippocampal neurons from oxidative stress and synapse damage induced by amyloid-beta oligomers. J. Biol. Chem. 293 (6), 1957–1975 (2018).
Phinney, D. G. & Pittenger, M. F. Concise Review: MSC-Derived exosomes for cell-free therapy. Stem Cells. 35 (4), 851–858 (2017).
Chang, M. G. et al. Proarrhythmic potential of mesenchymal stem cell transplantation revealed in an in vitro coculture model. Circulation. 113 (15), 1832–1841 (2006).
Breitbach, M. et al. Potential risks of bone marrow cell transplantation into infarcted hearts. Blood. 110 (4), 1362–1369 (2007).
Giuliani, M. et al. Human mesenchymal stem cells derived from induced pluripotent stem cells down-regulate NK-cell cytolytic machinery. Blood. 118 (12), 3254–3262 (2011).
Zhu, Y. et al. Comparison of exosomes secreted by induced pluripotent stem cell-derived mesenchymal stem cells and synovial membrane-derived mesenchymal stem cells for the treatment of osteoarthritis. Stem Cell. Res. Ther. 8 (1), 64 (2017).
Lai, R. C. et al. Exosome secreted by MSC reduces myocardial ischemia/reperfusion injury. Stem Cell. Res. 4 (3), 214–222 (2010).
Zhang, J. et al. Exosomes released from human induced pluripotent stem cells-derived MSCs facilitate cutaneous wound healing by promoting collagen synthesis and angiogenesis. J. Transl Med. 13, 49 (2015).
Peng, W. et al. Small extracellular vesicles secreted by iPSC-Derived MSCs ameliorate pulmonary inflammation and lung Injury Induced by Sepsis through Delivery of miR-125b-5p. J. Immunol. Res. 2023, 8987049 (2023).
Ye, T. et al. Large extracellular vesicles secreted by human iPSC-derived MSCs ameliorate tendinopathy via regulating macrophage heterogeneity. Bioact Mater. 21, 194–208 (2023).
Kim, H. et al. Identification of molecules responsible for Therapeutic effects of Extracellular vesicles produced from iPSC-Derived MSCs on Sjo¨gren’s syndrome. Aging Dis. 12 (6), 1409–1422 (2021).
Hai, B., Shigemoto-Kuroda, T., Zhao, Q., Lee, R. H. & Liu, F. Inhibitory effects of iPSC-MSCs and their extracellular vesicles on the onset of Sialadenitis in a mouse model of Sjögren’s syndrome. Stem Cells Int. 2018, 2092315 (2018).
Dominici, M. et al. Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy. 8 (4), 315–317 (2006).
Cruz-Barrera, M. et al. Integrated Analysis of Transcriptome and Secretome from umbilical cord mesenchymal stromal cells reveal New mechanisms for the modulation of inflammation and Immune Activation. Front. Immunol. 11, 575488 (2020).
Németh, K. et al. Bone marrow stromal cells attenuate sepsis via prostaglandin E(2)-dependent reprogramming of host macrophages to increase their interleukin-10 production. Nat. Med. 15 (1), 42–49 (2009).
Théry, C., Amigorena, S., Raposo, G. & Clayton, A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr. Protoc. Cell. Biol. ; (2006). Chap. 3:Unit 3.22.
Zhang, J. et al. Induced Pluripotent Stem cell-derived mesenchymal stem cells hold Lower Heterogeneity and Great Promise in Biological Research and clinical applications. Front. Cell. Dev. Biol. 9, 716907 (2021).
Nagamura-Inoue, T. & He, H. Umbilical cord-derived mesenchymal stem cells: their advantages and potential clinical utility. World J. Stem Cells. 6 (2), 195–202 (2014).
Ma, D. et al. Immunomodulatory effect of human umbilical cord mesenchymal stem cells on T lymphocytes in rheumatoid arthritis. Int. Immunopharmacol. 74, 105687 (2019).
Kimbrel, E. A. et al. Mesenchymal stem cell population derived from human pluripotent stem cells displays potent immunomodulatory and therapeutic properties. Stem Cells Dev. 23 (14), 1611–1624 (2014).
Polo, J. M. et al. Cell type of origin influences the molecular and functional properties of mouse induced pluripotent stem cells. Nat. Biotechnol. 28 (8), 848–855 (2010).
Di Nicola, M. et al. Human bone marrow stromal cells suppress T-lymphocyte proliferation induced by cellular or nonspecific mitogenic stimuli. Blood. 99 (10), 3838–3843 (2002).
Meisel, R. et al. Human bone marrow stromal cells inhibit allogeneic T-cell responses by indoleamine 2,3-dioxygenase-mediated tryptophan degradation. Blood. 103 (12), 4619–4621 (2004).
Beyth, S. et al. Human mesenchymal stem cells alter antigen-presenting cell maturation and induce T-cell unresponsiveness. Blood. 105 (5), 2214–2219 (2005).
Wada, N., Gronthos, S. & Bartold, P. M. Immunomodulatory effects of stem cells. Periodontol 2000. 63 (1), 198–216 (2013).
Sivanathan, K. N., Gronthos, S., Rojas-Canales, D., Thierry, B. & Coates, P. T. Interferon-gamma modification of mesenchymal stem cells: implications of autologous and allogeneic mesenchymal stem cell therapy in allotransplantation. Stem Cell. Rev. Rep. 10 (3), 351–375 (2014).
Fu, X. et al. Comparison of immunological characteristics of mesenchymal stem cells derived from human embryonic stem cells and bone marrow. Tissue Eng. Part. A. 21 (3–4), 616–626 (2015).
Schnabel, L. V. et al. Induced pluripotent stem cells have similar immunogenic and more potent immunomodulatory properties compared with bone marrow-derived stromal cells in vitro. Regen Med. 9 (5), 621–635 (2014).
Ranganath, S. H., Levy, O., Inamdar, M. S. & Karp, J. M. Harnessing the mesenchymal stem cell secretome for the treatment of cardiovascular disease. Cell. Stem Cell. 10 (3), 244–258 (2012).
Ankrum, J. A., Ong, J. F. & Karp, J. M. Mesenchymal stem cells: immune evasive, not immune privileged. Nat. Biotechnol. 32 (3), 252–260 (2014).
Yu, Y. et al.
Kourembanas, S. Exosomes: vehicles of intercellular signaling, biomarkers, and vectors of cell therapy. Annu. Rev. Physiol. 77, 13–27 (2015).
Zhang, B. et al. Mesenchymal stem cells secrete immunologically active exosomes. Stem Cells Dev. 23 (11), 1233–1244 (2014).
Mitsialis, S. A. & Kourembanas, S. Stem cell-based therapies for the newborn lung and brain: possibilities and challenges. Semin Perinatol. 40 (3), 138–151 (2016).
Wang, Y., Chen, X., Cao, W. & Shi, Y. Plasticity of mesenchymal stem cells in immunomodulation: pathological and therapeutic implications. Nat. Immunol. 15 (11), 1009–1016 (2014).
Ren, G. et al. Mesenchymal stem cell-mediated immunosuppression occurs via concerted action of chemokines and nitric oxide. Cell. Stem Cell. 2 (2), 141–150 (2008).
Song, Y. et al. Exosomal miR-146a contributes to the enhanced therapeutic efficacy of Interleukin-1β-Primed mesenchymal stem cells against Sepsis. Stem Cells. 35 (5), 1208–1221 (2017).
Ti, D. et al. LPS-preconditioned mesenchymal stromal cells modify macrophage polarization for resolution of chronic inflammation via exosome-shuttled let-7b. J. Translational Med. ;13(1). (2015).
Kobayashi, H. et al. <Editors’ Choice> effects of exosomes derived from the induced pluripotent stem cells on skin wound healing. Nagoya J. Med. Sci. 80 (2), 141–153 (2018).
Kim, S., Lee, S. K., Kim, H. & Kim, T. M. Exosomes secreted from Induced Pluripotent Stem cell-derived mesenchymal stem cells accelerate skin cell proliferation. Int. J. Mol. Sci. ;19(10). (2018).
Santoso, M. R. et al. Exosomes from Induced Pluripotent Stem Cell-Derived cardiomyocytes promote autophagy for myocardial repair. J. Am. Heart Assoc. 9 (6), e014345 (2020).
Gao, L. et al. Exosomes secreted by hiPSC-derived cardiac cells improve recovery from myocardial infarction in swine. Sci. Transl Med. ;12(561). (2020).
Wang, Y. et al. Exosomes/microvesicles from induced pluripotent stem cells deliver cardioprotective miRNAs and prevent cardiomyocyte apoptosis in the ischemic myocardium. Int. J. Cardiol. 192, 61–69 (2015).
Qi, X. et al. Exosomes secreted by Human-Induced pluripotent stem cell-derived mesenchymal stem cells repair critical-sized bone defects through enhanced angiogenesis and Osteogenesis in osteoporotic rats. Int. J. Biol. Sci. 12 (7), 836–849 (2016).
Liu, X. et al. Exosomes secreted from Human-Induced pluripotent stem cell-derived mesenchymal stem cells prevent osteonecrosis of the femoral head by promoting angiogenesis. Int. J. Biol. Sci. 13 (2), 232–244 (2017).
Rani, S., Ryan, A. E., Griffin, M. D. & Ritter, T. Mesenchymal stem cell-derived Extracellular vesicles: toward cell-free therapeutic applications. Mol. Ther. 23 (5), 812–823 (2015).
El-Tookhy, O. S., Shamaa, A. A., Shehab, G. G., Abdallah, A. N. & Azzam, O. M. Histological evaluation of experimentally Induced critical size defect skin wounds using Exosomal Solution of mesenchymal stem cells derived microvesicles. Int. J. Stem Cells. 10 (2), 144–153 (2017).
Sharma, G. D., He, J. & Bazan, H. E. p38 and ERK1/2 coordinate cellular migration and proliferation in epithelial wound healing: evidence of cross-talk activation between MAP kinase cascades. J. Biol. Chem. 278 (24), 21989–21997 (2003).
Shabbir, A., Cox, A., Rodriguez-Menocal, L., Salgado, M. & Van Badiavas, E. Mesenchymal stem cell exosomes induce Proliferation and Migration of normal and chronic wound fibroblasts, and enhance Angiogenesis in Vitro. Stem Cells Dev. 24 (14), 1635–1647 (2015).
Tertel, T., Dittrich, R., Arsène, P., Jensen, A. & Giebel, B. EV products obtained from iPSC-derived MSCs show batch-to-batch variations in their ability to modulate allogeneic immune responses in vitro. Front. Cell. Dev. Biol. 11, 1282860 (2023).
Acknowledgements
J.C.B. received a PhD scholarship from Universidad del Rosario. This research was funded by Curexsys GmbH. We thank the CIC bioGUNE platform staff for their assistance in sample vitrification, data collection and analysis.
Author information
Authors and Affiliations
Contributions
J.G., S.S. and J.C.B. conceived the project; J.C.B., A.B, S.M. and G.K., performed experiments and analysed primary data; J.C.B wrote the manuscript; J.G. and S.S provided permanent direction during experiments, data analysis and manuscript preparation. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
All authors are employed in Curexsys GmbH.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Buitrago, J.C., Morris, S.L., Backhaus, A. et al. Unveiling the Immunomodulatory and regenerative potential of iPSC-derived mesenchymal stromal cells and their extracellular vesicles. Sci Rep 14, 24098 (2024). https://doi.org/10.1038/s41598-024-75956-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-75956-3