Abstract
Gamma delta (γδ) T cells, which reside in mucosal and epithelial tissues, are integral to immune responses and are involved in various cancers, autoimmune, and infectious diseases. To study human γδ T cells to a translational level, we developed γδ humanized TCR-T1 (HuTCR-T1) mice using our TruHumanization platform. We compared the metabolomic profiles from plasma samples of wild-type (WT), γδ HuTCR-T1 mice, and humans using UHPLC-MS/MS. Untargeted metabolomics and lipidomics were used to screen all detectable metabolites. Principal component analysis revealed that the metabolomic profiles of γδ HuTCR-T1 mice closely resemble those of humans, with a clear segregation of metabolites between γδ HuTCR-T1 and WT mice. Most humanized γδ metabolites were classified as lipids, followed by organic compounds and amino acids. Pathway analysis identified significant alterations in the metabolism of tryptophan, tyrosine, sphingolipids, and glycerophospholipids, shifting these pathways towards a more human-like profile. Immunophenotyping showed that γδ HuTCR-T1 mice maintained normal proportions of both lymphoid and myeloid immune cell populations, closely resembling WT mice, with only a few exceptions. These findings demonstrate that the γδ HuTCR-T1 mouse model exhibits a metabolomic profile that is remarkably similar to that of humans, highlighting its potential as a relevant model for investigating the role of metabolites in disease development and progression. This model also offers an opportunity to discover therapeutic human TCRs.
Similar content being viewed by others
Introduction
Gamma delta (γδ) T cells are ‘unconventional’ T cells that comprise 0.5–5% of the T lymphocytes in peripheral blood mononuclear cells1,2,3. They orchestrate an immune surveillance critical in innate and adaptive immunity4,5,6. The principal function of γδ T cells is to present immediate homeostatic responses to maintain immune and tissue integrity7,8. In addition, γδ T cells play a critical role in recognizing, coordinating, and protecting against malignant cancer, atherosclerosis, and autoimmune and infectious diseases. Also, γδ T cells are critical in sustaining essential host defense and barrier functions against foreign/indigenous antigens9,10,11,12,13,14.
γδ T cells perform these functions through their unique tissue distribution. Conventional αβ T cells are primarily found in lymphoid organs, while γδ T cells reside in the mucosal and epithelial tissues of peripheral organs. Notably, in the absence of a specific activation signal, γδ T cells are generally not under the surveillance of conventional αβ T cells. The rapid γδ T cell activation occurs through the direct recognition of ligands by its T cell receptor (TCR) in a major histocompatibility (MHC) complex independent manner, which distinguishes γδ T cell from conventional αβ T cells15. In addition, γδ T cells are a unique lineage that expresses TCRs derived from separate variable (V), joining (J), and constant (C) for T cell receptor gamma (TRG) and additional diversity (D) for T cell receptor delta (TRD) gene loci.
Despite significant strides in understanding the diverse roles of γδ T cells in health and disease, a critical gap exists in research – the absence of a suitable humanized γδ T cell animal model. To address this limitation and comprehensively unravel the metabolic aspects of γδ T cell function with translational implications, we have generated γδ humanized (Hu) TCR-T1 (HuTCR-T1) mice and undertaken metabolomic analysis in these mice. Our approach aims to provide crucial insights into the metabolic underpinnings of γδ T cell activity, offering potential avenues for translational research and therapeutic development.
Metabolomics and lipidomics are analytical profiling techniques for determining metabolites from mammalian specimens in healthy and pathological conditions. These techniques can identify potential biomarkers, metabolic profiles, and/or pathways.
Various metabolites can be evaluated depending on the biological samples’ nature16,17. Recent findings demonstrated that alterations in the number of metabolites and pathway profiles related to nucleic acids, organic compounds, lipids, and amino acids significantly impact cancer, autoimmune, and infectious diseases18,19,20,21,22.
We hypothesized that metabolomic profiles revealed from our γδ HuTCR-T1 mice could provide new insights into investigating and mitigating human disease conditions. We further hypothesized that identifying associated metabolic biomarkers will delineate pathways and molecular targets for developing therapeutics/vaccines targeting metabolites/pathways perturbed by pathological conditions. Here, we employed untargeted metabolomics and lipidomics to identify metabolites and metabolic pathways present in the plasma of γδ HuTCR-T1 mice compared to those of wild type (WT) mice and human samples. Through these comparisons, we aim to offer a strategic approach to bridge basic research and clinical applications. These comparative analyses may be essential for conserving immunological functions, translational relevance, identifying differences, insights into γδ T cell biology, and discovering potential biomarkers and therapeutic targets. We also discuss the potential strategy of utilizing a γδ HuTCR-T1 mouse model to study human health and pathological conditions.
Materials and methods
Generation of gamma delta (γδ) humanized T cell receptor (HuTCR)-T1 mice
All animal experiments were performed per IACUC protocol approved by the inGenious protocol committee, all the experiments were performed in accordance with relevant guidelines and regulations, and all methods are reported in accordance with ARRIVE guidelines. For the generation of humanized γδ mice, our laboratory employed inGenious’ TruHumanization technology large bacterial artificial chromosome (BAC) genomic DNA platform (> 100 kb). This model is a gene replacement in which the mouse sequence is eliminated and replaced with a human genomic sequence.
Generation of humanized TCR γ mouse
Mouse Vγ1 (TRGV1) and J4 segments were replaced with human Vγ9–11 and Js segments in a targeted fashion. Human V and J segments were then fused with multiple mice C4 regions for appropriate signaling.
Generation of humanized TCR δ mouse
Mouse TRDV4, Ds, and Js segments were replaced with human Vδ1 – Vδ8, Ds, and Js segments in a targeted fashion. Human TCR, V, D, and J segments were then fused with a mouse C region for appropriate signaling. γδ TCR diversity was confirmed in the spleen and blood of humanized TCR mice (unpublished observation). Finally, humanized γ and δ mice were bred to double homozygosity to establish inGenious γδ humanized T cell receptor-T1 (HuTCR-T1) mice, also referring to inGenious γδ humanized TCR version 1 mice. The patent application related to HuTCR-T1 mice has been accepted. The patent, entitled ‘Genetically modified non-human having humanized gamma and delta TCR variable genes’, the patent has been published under patent number US20240114883A1 and filed by inGenious Targeting Laboratory, Ronkonkoma, NY.
Isolation of mononuclear cells (MNCs), flow cytometry analysis, and immunoglobulin (Ig) G ELISA assay
At the end of the experimentation, the mice were humanely euthanized using carbon dioxide inhalation and various tissues were collected. The composition of immune cells in different tissues was analyzed to compare γδ HuTCR-T1 with WT mice. Spleen, lung, lymph node, blood, thymus, liver, and ileum were harvested from mice and washed twice with wash media [DMEM (Gibco, cat. # 12430-047) with 1% penicillin/streptomycin (Gibco, cat. # 15140-122)]. To prepare single-cell suspensions, spleen, lymph node, and thymus tissues were minced into 1 cm x 1 cm pieces, pressed through tissue collectors with a plunger, then passed through a 70 μm cell strainer, and the supernatants were collected. Cells were then centrifuged at 800× g for 5 min at 4oC, supernatants were discarded, and MNCs were resuspended in cell culture media [(DMEM containing 10% FBS (Cytiva Hyclone, cat. # SH30070.03) and 1% penicillin/streptomycin)].
The liver was pressed through tissue collectors with a plunger, passed through a 70 μm cell strainer, and the resulting single-cell suspension was diluted with an equal amount of PBS containing 2% FBS. The cell suspension was then layered over Lymphoprep (Stem cell technologies, cat. # 07801) and centrifuged at 800× g for 20 min at RT without brakes23. MNCs were collected at the plasma: lymphoprep interface, washed with wash media, and centrifuged at 800× g for 5 min at 4oC. The supernatant was discarded, and the MNCs were resuspended in cell culture media.
Peripheral blood was collected and mixed with a 10% citrate dextrose solution/ACD. MNCs were isolated using the Lymphoprep method as described above. The isolated MNCs were then resuspended in cell culture media.
Lung tissues were minced into a homogeneous paste (< 1 mm in size) in PBS containing 2% FBS and transferred to a sterile 50 ml conical tube containing dissociation medium [RPMI 1640 (Stem cell, cat. # 36750), collagenase/hyaluronidase (Stem cell, cat. # 07912, 1×), DNase I solution (Stem cell, cat. # 07900, 1 mg/ml)]24. The mixture was then incubated at 37 °C for 20 min on a shaking platform. The digested lung tissues were filtered through a 70 μm cell strainer and centrifuged at 300× g for 10 min. The supernatant was discarded and the MNCs were resuspended in cell culture media.
Ileal tissues were harvested, and intestinal contents were flushed with wash media. The tissues were cut longitudinally and incubated with DTT (Sigma, cat. # 10197777001, 2mM) in HBSS media (Sigma, cat. # H2387) on a shaking platform at 37 °C for 20 min25. Subsequently, the tissues were centrifuged at 300× g for 10 min at RT. The supernatant was discarded, and the tissues were minced into 1 cm pieces, followed by digestion with collagenase (Sigma, cat. # COLLA-RO, 2.4 mg/ml) at 37 °C for 25 min on a shaking platform. The mixture was then filtered through a 70 μm cell strainer and centrifuged at 300× g for 10 min at RT. Subsequently, the supernatant was discarded, and cells were diluted with 43% Percoll (Bioworld cat. # 65455-52-9) and under-layered with 70% Percoll, then centrifuged at 300× g for 25 min at RT without brakes. MNCs were collected at the interface, washed once with wash medium, centrifuged at 300× g for 5 min, and resuspended in cell culture medium.
Erythrocytes (if any) were lysed with RBC lysing buffer (Sigma, cat. # R7757), centrifuged at 300× g for 5 min, and MNCs were then resuspended in a cell culture medium. The Trypan blue (0.002%) exclusion method was used to determine the quantity and viability of the MNCs.
Freshly isolated MNCs were stained to determine the frequencies of immune cells. Following anti-mouse antibodies were purchased from Biolegend: anti-CD3e (PE, cat. # 100308, clone 145-2C11), -CD19 (PB, cat. # 115523, clone 6D5), -CD4 (FITC, cat. # 10050, clone RM4-5), -CD8a (APC, cat. # 100712, clone 53 − 6.7), -CD25 (APC, cat. # 102012, clone PC61), -FoxP3 (PE, cat. # 126403, clone MF-14), -γδ (PE, cat. # 107508, clone UC7-13D5), -NKp46 (APC/Cy7, cat. # 137645, clone 29A1.4), -Ly6C (APC, cat. # 128016, clone HK1.4), -CD11b (APC, cat. # 101212, clone M1/70), -F4/80 (FITC, cat. # 123108, clone BM8). Anti-MHC-II (FITC, cat. # 130-112-229, clone REA813) and anti-CD11c (APC, cat. # 130-110-702, clone REA754) anti-mouse antibodies were obtained from Miltenyi Biotech. Anti-CD68 (PE, cat. # 566386, clone FA/11) was purchased from BD Pharmigen. Anti-Ly6G (FITC, cat. # 11–5931-82, clone RB-BC5) and CD161/NK1.1 (APC, MABF1488, clone PK136) anti-mouse antibodies were purchased from Invitrogen and Sigma-Aldrich, respectively. Appropriate isotype-matched controls were included in all analyses. Subsequently, 50,000 events were acquired per sample using Novocyte® Flow Cytometer (ACEA Biosciences Inc, San Diego, CA, USA). Data were analyzed using Novo Express® software version 1.5.0 (www.agilent.com). Mouse IgG isotype titers in serum were detected by an enzyme-linked immunosorbent (ELISA) assay per manufacturer’s instructions (Thermo Fisher Scientific, cat #88–50400-22).
Sample collection and metabolomics analyses
Blood samples were collected via cheek bleed from (n = 5) age matching and genetic background matching (129/C57/B6 strain) γδ HuTCR-T1 and wild type (WT). National Institute of Standards and Technology (NIST) standardized human plasma (n= 3) samples were procured from Sigma Aldrich (Saint Louis, MO). Blood samples were mixed with 10% acid citrate dextrose to prevent blood clotting and centrifuged at 3,000×g for 10 min at 4 °C to separate plasma from blood cells. Plasma samples (supernatant) were collected and stored at −80°C until further processing. To extract metabolites from plasma samples, 3–4 volumes of pre-chilled methanol were added to each plasma sample to precipitate proteins. Samples were vortexed vigorously for 30 s and incubated on ice for 10 min to allow complete protein precipitation. Samples were then centrifuged at 12,000× g for 10 min at 4 °C to pellet the precipitated proteins and supernatants were transferred to new tubes, reconstituted in 5% ACN with 0.1% formic acid and placed in glass liquid chromatography vial for injection26.
The samples, blanks, and a pooled QC sample were injected at 10 µL onto a Thermo Scientific Vanquish Horizon using an Agilent Poroshell 120 Aq-C18 column (2.1 × 100 mm, 2.7 μm particle size) with a solvent system of 100% water with 0.1% formic acid for solvent A and 100% ACN with 0.1% formic acid for solvent B. A flow rate of 300 µL/min was run throughout with an initial gradient of 5% solvent B until 0.5 min followed by a linear increase to 95% B ending at minute 14, holding 95% until minute 17, going back down to 5% B at minute 17.1 min, and ending the run at 5% solvent B at minute 20. Detection was performed on a Thermo Scientific Exploris 480 Orbitrap mass spectrometer with ionization of samples following LC separation with a Heated Electrospray ionization (HESI) source with runs performed in positive mode with ESI voltages of 3.5 kV with a sheath gas of 20, an auxiliary gas of 10, sweep gas of 1, and capillary and vaporizer temperatures of 325 and 350 oC, respectively. MS data collection was performed using Data-Dependent Analysis with a mass range of 70–800 m/z at 50% RF lens, an MS resolution of 60,000, an MSMS resolution of 15,000, 30/50/70V HCD collision energy selection, and 2.5 second dynamic exclusion with a 0.5 second cycle time26.
Metabolites were identified by automated comparison of the positive ion features in the experimental samples to a reference library of chemical standard entries that included retention time, the mass-to-charge ratio (m/z), preferred adducts, as well as associated MS spectra, and were quality-controlled using pooled sample alignment using the Progenesis QI metabolomics package. The sample set was randomized with periodic QC samples to both correct for batch shifts and peak alignment and all samples were run in triplicate within 24 of run time so no significant batch correct was needed. All samples were normalized to QC samples using a log10 ratio method and normalized to all compounds. For peak annotation, the Human Metabolite Database was used to compare to features with cut-offs of 0.05 p-value, CV < 30 for pooled samples, and features were only annotated with MSMS fragmentation scores above 20% for Progenesis QI matching algorithm were selected. Additionally, features had to be present in the NIST plasma group to be selected. This resulted in 143 features annotated and additionally analyzed using MetaboAnalyst to perform statistical analysis (one-factor) and pathway analysis ( by viewing metabolomes on all matched pathways according to the pvalues from the pathway enrichment analysis and pathway impact values from the pathway topology analysis)26. Samples were analyzed at the Mass Spectrometry and Proteomics Facility of the Ohio State University, Columbus, OH (www.ccic.osu.edu/MSP).
Statistical analysis
Statistical analysis was done using GraphPad Prism 8.0.2 (GraphPad Software, Inc., www.graphpad.com, San Diego, CA, USA), and Two-way ANOVA was performed, and p values < 0.05 were considered as a significant metabolite. Data was normalized by sum and auto-scaled using MetaboAnalyst 5.0 (www.metaboanlyst.ca) and used the Human Kyoto Encyclopedia of Genes and Genomes (KEGG) and Human Metabolome Databases (HMBD) 5.0 in metaboanalyst that matched putative metabolites and/or pathways27,28,29,30.
Results
Non-targeted metabolomics revealed several resembling metabolites between human and gamma-delta (γδ) T cell receptor (TCR)-T1 mice while differently altered in wild-type (WT) mice plasma samples
The human, HuTCR-T1, and WT mice plasma samples were analyzed using the UHPLC/MS-MS method to investigate the metabolic profiles. The resulting relative peak intensities from positive ion modes were considered to generate tables, and multivariate analysis was performed. A total of 6,558 putative metabolites were detected in plasma metabolomics; only 143 metabolites were found statistically significant (p-value < 0.05) among the three groups. Among these significant metabolites, 71 metabolites were observed among the three groups (γδ HuTCR-T1 mice, WT mice, and human). We identified 35 metabolites that exhibited resemblance and/or no statistically significant difference between γδ HuTCR-T1 mice and human plasma samples (Table 1, Supplementary Table 1), while 36 metabolites exhibited statistical (p value < 0.05) significant difference between γδ HuTCR-T1 mice and humans. On the contrary, 72 metabolites exhibited statistical (p value < 0.05) significance differences between γδ HuTCR-T1 and WT mice groups. This data set indicates the yielding of human-like metabolites in γδ HuTCR-T1 mouse model and strayed from WT mice profiles.
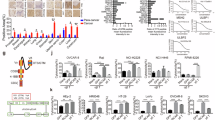
Multivariate analysis of plasma metabolic profiles also demonstrated a close resemblance between humans and γδ HuTCR-T1 mice. In contrast, WT mice showed remarkable segregation from HuTCR-T1 and humans, as shown in the heatmap (Fig. 1) and at component 1 in a two-dimensional (Fig. 2) and three-dimensional (supplementary Fig. 1) principal component analysis (PCA) plot. The analysis also identified metabolites specific to WT mice and human plasma, as illustrated in the heatmap (Supplementary Fig. 2) and detailed in the list of metabolites (Supplementary Table 2), which includes metabolites found in human plasma or WT mouse plasma samples. The metabolites contributing to the classification of humans, γδ HuTCR-T1 mice, and WT mice were also identified using the variable importance in projection (VIP) scores. Metabolites with greater VIP scores (> 1.0, p value < 0.05) contribute significantly to the separation among the three groups, while metabolites with smaller VIP scores (< 1.0) exhibit less influence on the model (Fig. 3).
Determination of non-targeted metabolomics and lipidomics in plasma samples. A heatmap of top 100 significant metabolites in plasma samples visualizes the intensity of metabolites. The magnitude of a metabolite is arranged to show their relatedness in the cluster heatmap. In addition, the more intense yellow color represents the higher magnitude, while the red color represents the lower magnitude of a metabolite. Humanized gamma-delta (γδ) transgenic mice were generated using BAC genomic DNA platform (> 100 kb). Plasma samples were prepared using the cold methanol extraction method, analyzed with UHPLC-MS/MS, and analyzed using MetaboAnalyst 5.0. Human Kyoto Encyclopedia of Genes and Genomes (KEGG) and Human Metabolome Database (HMBD) were used to match putative metabolites and/or pathways.
Determination of non-targeted metabolomics and lipidomics in plasma samples. Principal-component analysis (PCA) indicates segregation of gamma-delta (γδ), human, and wild type mice metabolites. γδ humanized TCR-T1 mice were generated using BAC genomic DNA platform (> 100 kb). The PCA plot is generated from the peak intensities obtained from liquid chromatography-mass spectrometry (LC-MS), which are used as the input data. Plasma samples were prepared using the cold methanol extraction method, analyzed with UHPLC-MS/MS, and analyzed using MetaboAnalyst 5.0. Human Kyoto Encyclopedia of Genes and Genomes (KEGG) and Human Metabolome Database (HMBD) were used to match putative metabolites and/or pathways.
Determination of non-targeted metabolomics and lipidomics in plasma samples. Variable importance in projection (VIP) scores indicates the most critical metabolites contributing to the discrimination of metabolic profiles among different groups. The relative abundance of metabolites is indicated by a colored scale from red to blue, representing the high to low, respectively. Comparisons were performed among gamma delta (GD), human plasma, and wild type (WT) (A), between GD and human plasma (B), and between GD and WT (C). The colored boxes on the right indicate each group’s relative concentrations of the corresponding metabolite. GD humanized TCR-T1 mice were generated using BAC genomic DNA platform (> 100 kb). Plasma samples were prepared using the cold methanol extraction method, analyzed with UHPLC-MS/MS, and analyzed using MetaboAnalyst 5.0. Human Kyoto Encyclopedia of Genes and Genomes (KEGG) and Human Metabolome Database (HMBD) were used to match putative metabolites and/or pathways.
Metabolites such as 5-(4-Hydroxybenzyl) thiazolidine-2,4-dione; DL-2,5-Dihydrophenylalanine; oxonal; Ethyl 2-furanacrylate; dehydro-beta-Ionone; and butyl salicylate played an essential role in the discrimination of HuTCR-T1 mice metabolic profiles from WT mice. On the contrary, (6E)−8-Methylnon-6-enoylcarnitine; DG(18:1n7/0:0/20:5n3); 20:0 Cholesterol ester; ximaosarcophytol B; 21 H-Biline-8,12-dipropanoicacid, 3,18-diethenyl-1,19,22,24-tetrahydro-2,7,13,17-tetramethyl-1,19-dioxo; (10Z,12E)-Pentadeca-10,12-dienoylcarnitine; Pregnanetriol; 3-Allyl-1-cyclohex; didesethyl chloroquine; 1-hydroxyisoquinoline played an essential role in the discrimination of WT metabolic profiles from HuTCR-T1 mice (Fig. 3a, c). Cer(d18:1(4E)/15:0(2OH)); MG(20:3(6,8,11)-OH(5)/0:0/0:0); N-Stearoyl Glutamic acid; 1alpha-hydroxy-24-methylvitamin D2; 8,11,14-nonadecatriynoic acid; Ananstrep A; 4beta-Hydroxydesogestrel; (6E)−8-Methylnon-6-enoylcarnitine played an essential role in segregating metabolomic profiles from γδ HuTCR-T1 mice and human plasma. On the other hand, 5-(4-Hydroxybenzyl) thiazolidine-2,4-dione; Cytosine; Trigoforin; 10-Hydroxy-9-ketooctadecanoic acid; (7b,10a)−3-Hydroxy-1,3,5-cadinatrien-9-one; Vitamin A3 played an essential role in segregation of human metabolomic profiles from γδ HuTCR-T1 mice (Fig. 3b).
Metabolite set enrichment and metabolic pathways associated with HuTCR-T1 mice
The quantitative metabolite set enrichment and metabolic pathway analyses were performed using MetaboAnalyst 5.0 to investigate metabolic pathways among three groups (Fig. 4). The quantitative metabolite enrichment analysis employed 99 metabolite sets based on normal human metabolic pathways. Nine biochemical pathways were identified by the analysis that significantly affected metabolite set among the groups (p < 0.05), including alpha-linolenic acid and linoleic acid metabolism, retinol metabolism; sphingolipid metabolism; tryptophan metabolism, phosphatidylcholine biosynthesis; catecholamine biosynthesis; phospholipid biosynthesis; tyrosine metabolism; and purine metabolism.
The distributing diagram of metabolites sets enrichment. Metabolic set enrichment using metabolite concentrations shows the most enriched pathways among different groups. Gamma delta (γδ) humanized TCR-T1 mice were generated using BAC genomic DNA platform (> 100 kb). Plasma samples were prepared using the cold methanol extraction method, analyzed with UHPLC-MS/MS, and analyzed using MetaboAnalyst 5.0. Human Kyoto Encyclopedia of Genes and Genomes (KEGG) and Human Metabolome Database (HMBD) were used to match putative metabolites and/or pathways.
The significant metabolic pathway impact values were calculated from topology pathway analysis. The analysis identified four matched metabolic pathways: glycerophospholipid metabolism, tryptophan metabolism, sphingolipid, and tyrosine (Fig. 5).
Determination of metabolic pathways in plasma samples. The bubble chart of the Human Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis reveals matched pathways are shown as circles. The color and size of each circle are based on significant p-value and pathway impact values, respectively. Human KEGG and Human Metabolome Database (HMBD) were used to match putative metabolites and/or pathways.
Normal immune cell development in HuTCR-T1 mice
HuTCR-T1 mice possess humanized genes encoding components of the immune synapse. To investigate the impact this gene targeting (engineering) had on the development of the immune system, we performed comprehensive immunophenotyping of naïve HuTCR-T1 mice. We compared the results to WT controls of a similar strain background (Figs. 6 and 7).
Normal lymphoid immune cell proportions in HuTCR-T1 mice. Representative flow cytometry analysis of HuTCR-T1 and wild-type (WT) mice. Mean frequencies (%) (A) B cells (CD19+CD3e−), (B) T cells (CD3e+), (C) helper T cells (CD3+CD4+), (D) cytotoxic T cells (CD3+CD8+), (E) regulatory T cells (CD4+CD25+FoxP3+), and (F) γδ T cells (CD3+γδ+). Statistical analysis was performed using Two-way ANOVA followed by Bonferroni posttest and data are shown as means ± SEM (n = 3), *p < 0.05, and **p < 0.01.
Normal lymphoid and myeloid immune cell proportions in HuTCR-T1 mice. Representative flow cytometry analysis of HuTCR-T1 and wild-type (WT) mice. Mean frequencies (%) (A) natural killer T cells (CD3e+CD161/NK1.1+), (B) natural killer cells (NKp46+CD3e−), (C) dendritic cells (CD11c+MHC-II+), (D) monocytes (Ly6C+CD11b+), (E) macrophages (F4/80+CD68+), and (F) neutrophils (Ly6G+CD11b+). Statistical analysis was performed using Two-way ANOVA followed by Bonferroni posttest and data are shown as means ± SEM (n = 3), ****p < 0.0001.
The proportion of B cells was comparable between HuTCR-T1 and WT mice across various tissues and IgG antibody titers in serum (Fig. 8); HuTCR-T1 mice exhibited a significantly higher percentage of B cells in the spleen and liver, suggesting alterations in B cell development and distribution in HuTCR-T1 mice.
Proportions of T cells were comparable between HuTCR-T1 and WT mice in systemic tissues; marginally higher in lung, ileum, and lymph nodes of HuTCR-T1 mice; numerically elevated in the liver of WT mice; and significantly higher in the thymus of HuTCR-T1 mice compared to WT mice, indicates possible alterations in T cells development or localization in thymus of HuTCR-T1 mice.
Helper, cytotoxic, regulatory, γδ, and NK T cell proportions were comparable between HuTCR-T1 and WT mice across all the tissues. In blood, liver, and thymus, NK cells were comparable between HuTCR-T1 and WT mice. However, proportions were marginally higher in the lung, ileum, and spleen of WT mice while modestly elevated in lymph nodes in HuTCR-T1 mice. These data indicate generally normal development or distribution of lymphoid lineage in HuTCR-T1 mice.
Proportions of dendritic cells were comparable between HuTCR-T1 and WT mice across the various tissues while significantly elevated in the spleen of WT mice. Monocytes and macrophages were comparable between HuTCR-T1 and WT mice. Neutrophils were comparable in blood, liver, lymph nodes, and thymus of HuTCR-T1 and WT mice; numerically elevated in the ileum of HuTCR-T1 mice; and significantly elevated in the lung and spleen of WT mice. These data suggest overall normal development or distribution of myeloid lineage in HuTCR-T1 mice. In summary, HuTCR-T1 mice exhibited normal proportions of lymphoid and myeloid immune cell populations among various tissues and are comparable with WT mice with fewer exceptions,
Discussion
γδ T cells play critical roles in early immune responses, epithelial barrier protection, direct cytotoxicity, immune regulation, and tumor surveillance, through mechanisms that remain less well understood. To investigate these mechanisms and to develop a clinically translatable animal model, we generated HuTCR-T1 mice by replacing the mouse γδ TCRs with human γδ TCRs. The resulting mice exhibited normal immune cell development and profiles, indicating that human γδ TCRs are well tolerated by the mouse immune system. However, to our surprise, the introduction of human γδ TCRs had a significant impact on the metabolite profiles of the mice. Notably, many of the mouse metabolites were altered to resemble human metabolic profiles. This is the first report demonstrating a strong relationship between human γδ TCRs and host metabolic regulation. Since metabolites play key roles in various human diseases, and no current animal models are well-suited to study human metabolic responses. Due to the close alignment of their metabolomic and lipidomic profiles with human plasma, our analysis revealed that γδ HuTCR-T1 shares a significant number of comparable metabolites with human plasma approximately 50%, as shown in Table 1. The following metabolites are particularly noteworthy:
Dihydroferulic acid 4-O-glucuronide is classified as an organic compound and contains antioxidant, anti-inflammatory properties and also has the potential to inhibit cancer cell proliferation and treat neurological disorders31,32. In our investigations, the relative intensity of dihydroferulic acid 4-O-glucuronide was observed at similar levels in γδ HuTCR-T1 mice as in human plasma.
Vitamin A is indispensable for appropriate innate and adaptive immune function, and influences gut microbiome composition33,34,35,36,37. Similarly, the relative intensity of vitamin A3 in γδ HuTCR-T1 mice mirrored that in human plasma, supporting their use in research on vitamin A status, microbiome dysbiosis, and immune responses.
Bradykinin 1–5 is a bioactive metabolite of bradykinin and is formed by the proteolytic activity of angiotensin-converting enzyme (ACE) II. It also exhibits a role in inflammation, pain perception, vasodilates the arterial system, inhibits thrombin-induced platelet aggregation38,39, and promote proliferation of cervical and gastric cancers40,41. Both humans and γδ HuTCR-T1 mice have elevated levels of bradykinin 1–5.
Cholesterol ester, aminoethyl oleate, and diglyceride are implicated in cardiovascular diseases42,43. In the course of our study, the levels of these metabolites were comparable between γδ HuTCR-T1 mice and human samples while significantly different from WT mice.
Dihydrophenylalanine is an antimicrobial compound44 and we observed elevated levels of dihydroxyphenylalanine in γδ HuTCR-T1 mice, making them suitable for investigating antimicrobial activities.
Aminopicolinic acid is a catabolite of the tryptophan through the kynurenine pathway45and implicated in moderating acne vulgaris and exhibits neuroprotective and immunological activities46,47. In our studies, the intensity of aminopicolinic acid was observed in both human and γδ HuTCR-T1 mice highlighting their potential for research into immunological, neurological, and dermatological conditions.
Terbutaline is implicated for the management of pulmonary diseases48and elevated levels in γδ HuTCR-T1 mice indicates its relevance for studying pulmonary diseases. Similarly, higher intensity of N-octanoyl-L-homoserine lactone49,50,51, in γδ HuTCR-T1 mice suggests their applicability in infection prevention and cystic fibrosis research52,53.
Pregnanetriol intensity was elevated in γδ HuTCR-T1 mice, which signifies the importance of these mice for investigation of congenital adrenal hyperplasia54 or other endocrinological conditions.
Hippuric acid levels are significantly lower in patients with inflammatory bowel disease compared to healthy individuals55. We observed a comparable intensity of hippuric acid in both human plasma and γδ HuTCR-T1 mice.
LysoPC is a phospholipid known for its role as an immune activator and implicated in the pathogenesis of atherosclerosis56, colorectal cancer57, renal cell carcinoma58, and digestive tract tumors59 in humans. Higher intensity of LysoPC in γδ HuTCR-T1 mice compared to WT mice suggests that the humanized γδ TCR might influence LysoPC metabolism or its regulation.
Sphingosine-1-phosphate (sphingolipid) is a bioactive lipid mediator and promotes cellular proliferation60, vascular maturation61, immune-cell trafficking62, and angiogenesis63,64, and maintains vascular integrity65. In our data set, the intensity of sphingosine-1-phosphate was observed comparable in γδ HuTCR-T1 mice as in human plasma, supporting the usefulness of these mice in oncogenesis.
Other metabolites, such as phenylpyridine, and 2E,4E,8Z,10E-dodecatetraenoic acid, suggest the utility of γδ HuTCR-T1 mice in researching antioxidant and immunological studies66. Elevated valerylcarnitine levels in γδ HuTCR-T1 mice support their use in studying cancer and cardiovascular diseases67.
Several metabolic pathways are observed in γδ HuTCR-T1 mice comparable to humans
Tryptophan metabolism regulates inflammation, brain function, and energy homeostasis. Moreover, metabolites of tryptophan metabolism drive aging and elicit the pathophysiology of age-related diseases68. The gastrointestinal microbiome also affects tryptophan metabolism, directly or indirectly, affecting behavior and cognitive function69. Interestingly, tryptophan metabolites, indoleamine-2,3-dioxygenase, and kynurenine influenced γδ T cell cytotoxicity against ductal pancreatic adenocarcinoma cells70. Thus, the tryptophan metabolic pathway in γδ HuTCR-T1 mice may exhibit potential future therapeutic targets for various cancers.
Spingolipid metabolism is correlated with the aging, growth, differentiation, and apoptosis of cancer cells71. Ceramide (Cer) is the most critical sphingolipid, and it regulates cell growth, survival, vascular and epithelial integrity, and immune cell trafficking. It also plays a crucial role in inflammation and cancer72. Morad et al. (2012) demonstrated that in cultured triple-negative breast cancer cells, nanoliposomal tamoxifen enhances nanoliposomal ceramide cytotoxicity73. Similarly, ceramide was implicated in the apoptosis of leukemic natural killer cells74, and hepatic tumors75,76. The specificity of ceramide in γδ HuTCR-T1 mice calls for additional investigations into the role of sphingolipid metabolism in the cytotoxicity ability of γδ mouse T cells and the anti-tumor capacity of γδ mouse T cells. These inquiries aim to enhance our understanding and potentially improve the efficacy of immunotherapies against solid tumors that have proven ineffective with current state-of-the-art clinical practices.
Glycerophospholipid metabolism is implicated in biological membrane lipids and plays a vital role in cellular signaling and energy storage77. These membrane lipids bind antigen-presenting molecules CD1d in mice and CD1a-d in humans, presenting them to T cells that subsequently affect immune cells78,79. Moreover, glycerophospholipids profoundly impact phagocytosis, innate and adaptive immunity, and inflammation80. Therefore, γδ HuTCR-T1 mice may assist in identifying biological targets that will be imperative for creating new therapies against pathologies initiated by endogenous and/or exogenous agents.
Tyrosine metabolism synthesizes various biologically important compounds such as dopamine (catecholamines), norepinephrine, epinephrine, melanin, and thyroid hormones. In addition, several inherited disorders are associated with tyrosine metabolism, such as tyrosinemia type I-III, alkaptonuria, and hawkinsinuria81. Moreover, Tyrosine catabolic genes were down-regulated in patients with hepatocellular carcinoma82. This demonstrates that γδ HuTCR-T1 mice may support investigations of biochemical disorders and liver cancer.
Branched-chain amino acids (valine, isoleucine, and leucine) are implicated in energy expenditure and their supplementation benefits health83. However, inconsistently increased circulating branched-chain amino acid levels are associated with diabetes and obesity84,85,86. Interestingly, in our data set, the relative intensities of isoleucine and leucine were markedly reduced (although not statistically significant) in γδ HuTCR-T1 mice compared with WT mice, indicating that humanized γδ mice can be employed to investigate metabolic signaling, diabetes mellitus, hypercholesterolemia, obesity, and insulin resistance.
Potential mechanisms underlying the metabolic effects of human γδ TCR replacement in mice
Here can be few potential mechanisms: (1) Replacement of γδ can lead to modifications in enzyme activity or regulatory proteins, potentially impacting metabolic pathways. Such alterations may affect plasma levels of metabolites, such as lysoPC, by influencing the activity of key enzymes including Phospholipase A1, A2, B, and Lysophosphatidylcholine acyltransferase. These enzymes play critical roles in lysoPC metabolism, and their activity modulation may result in significant metabolic shifts, providing insight into the molecular consequences of γδ TCR replacement on cellular processes. (2) Replacing human γδ TCRs may alter the metabolic rate of fats, proteins, and carbohydrates by influencing thyroid hormones (T3 and T4) levels. (3) Changes in inflammatory cytokines like TNF-α and IL-6 can impair insulin sensitivity, leading to metabolic dysregulation which has impact on lipid metabolism, and glucose homeostasis87. (4) The composition of the microbiome may have undergone changes. γδ T cells are known to play a crucial role in shaping gut composition through bidirectional interactions with the microbiota, influencing tissue homeostasis10. Although the full extent of these interactions is not yet fully understood, they are essential for maintaining immune tolerance and regulating intestinal balance. These interactions can also drive metabolic shifts that affect other organs, underscoring the importance of γδ T cells in cross-organ metabolic regulation.
We acknowledge several limitations in our study: (1) The sample size of γδ double-homozygous mice (n= 5) was limited, restricting our ability to perform in-depth analyses such as examining variables like sex, age, stress, microbiome composition, and infection status. (2) Our metabolomic analysis was not extended to additional sample types, so the data should be interpreted with caution. To gain a deeper understanding of the roles of γδ TCR and their effects on metabolic profiles, further studies are essential to clarify the function of γδ T cells and their potential clinical applications. (3) The NIST human plasma used as a reference material consists of pooled plasma from various healthy donors, making it suitable for method comparison but insufficient to capture the normal variability of human plasma metabolomes. Our ongoing research aims to validate these findings using alternative assays and identify human TCRs associated with the observed metabolomic changes.
In conclusion, it is uncommon to see changes in TCR lead to such a wide array of metabolic changes. The PCA plot and VIP score analyses revealed significant segregation of unique metabolites between γδ HuTCR-T1 and WT mice. Notably, γδ HuTCR-T1 mice displayed a range of metabolites resembling human profiles. Pathway analysis highlighted the involvement of tryptophan, sphingolipid, tyrosine, and glycerophospholipid metabolism in γδ HuTCR-T1 mice. These findings provide metabolic insights that can guide future research using γδ HuTCR-T1 mice. Our analysis identified lipid profiles associated with various diseases, underscoring the potential of humanized γδ mice as a valuable model for exploring specific aspects of cancer and infectious disease.
Data availability
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.
References
Bank, I. et al. A functional T3 molecule associated with a novel heterodimer on the surface of immature human thymocytes. Nat. 1986 Jul 10–16 ;322(6075):179–81. PubMed PMID: 3487737.
Brenner, M. B. et al. Identification of a putative second T-cell receptor. Nat. 1986 Jul 10–16;322(6075):145–149. PubMed PMID: 3755221.
Chien, Y. H., Iwashima, M., Kaplan, K. B., Elliott, J. F. & Davis, M. M. A new T-cell receptor gene located within the alpha locus and expressed early in T-cell differentiation. Nature. 1987 Jun 25-Jul 1;327(6124):677 – 82. PubMed PMID: 2439914.
Hayday, A. C. Gammadelta T cell update: Adaptate Orchestrators of Immune Surveillance. J. Immunol. 203 (2), 311–320 (2019). PubMed PMID: 31285310.
Holtmeier, W. & Kabelitz, D. Gammadelta T cells link innate and adaptive immune responses. Chem. Immunol. Allergy. 86, 151–183 (2005). PubMed PMID: 15976493.
Papadopoulou, M., Sanchez Sanchez, G. & Vermijlen, D. Innate and adaptive gammadelta T cells: how, when, and why. Immunol. Rev. 298 (1), 99–116 (2020). PubMed PMID: 33146423.
Ribot, J. C., Lopes, N. & Silva-Santos, B. Gammadelta T cells in tissue physiology and surveillance. Nat. Rev. Immunol. 21 (4), 221–232 (2021). PubMed PMID: 33057185.
Kohlgruber, A. C. et al. Gammadelta T cells producing interleukin-17A regulate adipose regulatory T cell homeostasis and thermogenesis. Nat. Immunol. 19 (5), 464–474 (2018). PubMed PMID: 29670241. Pubmed Central PMCID: 8299914.
Nielsen, M. M., Witherden, D. A. & Havran, W. L. Gammadelta T cells in homeostasis and host defence of epithelial barrier tissues. Nat. Rev. Immunol. 17 (12), 733–745 (2017). PubMed PMID: 28920588. Pubmed Central PMCID: 5771804.
Papotto, P. H., Yilmaz, B. & Silva-Santos, B. Crosstalk between gammadelta T cells and the microbiota. Nat. Microbiol. 6 (9), 1110–1117 (2021). PubMed PMID: 34341528.
Silva-Santos, B., Serre, K. & Norell, H. Gammadelta T cells in cancer. Nat. Rev. Immunol. 15 (11), 683–691 (2015). PubMed PMID: 26449179.
Zhao, Y. et al. Protective role of gammadelta T cells in different Pathogen infections and its potential clinical application. J. Immunol. Res. 2018, 5081634 (2018). PubMed PMID: 30116753. Pubmed Central PMCID: 6079409.
Bank, I. & Marcu-Malina, V. Quantitative peripheral blood perturbations of gammadelta T cells in human disease and their clinical implications. Clin. Rev. Allergy Immunol. 47 (3), 311–333 (2014). PubMed PMID: 24126758.
Bank, I. The role of Gamma Delta T Cells in Autoimmune Rheumatic diseases. Cells ;9(2). (2020). PubMed PMID: 32085540. Pubmed Central PMCID: 7072729.
Hayday, A. C. & Vantourout, P. The innate biologies of adaptive Antigen receptors. Annu. Rev. Immunol. 38, 487–510 (2020). PubMed PMID: 32017636.
Shahisavandi, M., Wang, K., Ghanbari, M. & Ahmadizar, F. Exploring metabolomic patterns in type 2 diabetes Mellitus and response to glucose-lowering medications-review. Genes ;14(7). (2023). PubMed PMID: 37510368. Pubmed Central PMCID: 10379356.
Potenza, A. et al. Lipidomic approaches in Common and Rare Cerebrovascular diseases: the Discovery of unconventional lipids as novel biomarkers. Int. J. Mol. Sci. ;24(16). (2023). PubMed PMID: 37628924. Pubmed Central PMCID: 10454673.
Alreshidi, M. M. Selected metabolites profiling of Staphylococcus aureus following exposure to low temperature and elevated Sodium Chloride. Front. Microbiol. 11, 834 (2020). PubMed PMID: 32457719. Pubmed Central PMCID: 7225588.
Pang, Z., Zhou, G., Chong, J. & Xia, J. Comprehensive Meta-Analysis of COVID-19 Global Metabolomics Datasets. Metabolites ;11(1). (2021). PubMed PMID: 33435351. Pubmed Central PMCID: 7827862.
Zhou, C. X. et al. Investigation of urine metabolome of BALB/c mouse infected with an avirulent strain of Toxoplasma Gondii. Parasites Vectors. 15 (1), 271 (2022). PubMed PMID: 35906695. Pubmed Central PMCID: 9338554.
Corsale, A. M. et al. Metabolic changes in Tumor Microenvironment: how could they affect gammadelta T cells functions? Cells ;10(11). (2021). PubMed PMID: 34831116. Pubmed Central PMCID: 8616133.
De Preter, V. Metabolomics in the clinical diagnosis of inflammatory bowel disease. Dig. Dis. 33 (Suppl 1), 2–10 (2015). PubMed PMID: 26368862.
Chen, Y. et al. Acute myeloid leukemia-induced remodeling of the human bone marrow niche predicts clinical outcome. Blood Adv. 4 (20), 5257–5268 (2020). PubMed PMID: 33108453. Pubmed Central PMCID: 7594397 interests.
Jungblut, M., Oeltze, K., Zehnter, I., Hasselmann, D. & Bosio, A. Standardized preparation of single-cell suspensions from mouse lung tissue using the gentleMACS Dissociator. J. Vis. Exp. Jul 2(29). (2009). PubMed PMID: 19574953. Pubmed Central PMCID: 2798855.
Lee, J. S. et al. Improved isolation methods for mucosal leukocytes from small and large intestines in rats. Biosci. Biotechnol. Biochem. 73 (8), 1732–1740 (2009). PubMed PMID: 19661699.
Michael, H. et al. The combined Escherichia coli Nissle 1917 and Tryptophan Treatment modulates Immune and Metabolome responses to human Rotavirus infection in a human infant fecal microbiota-transplanted malnourished Gnotobiotic Pig Model. mSphere 7 (5), e0027022 (2022). PubMed PMID: 36073800. Pubmed Central PMCID: 9599269.
Wishart, D. S. et al. HMDB 5.0: the human metabolome database for 2022. Nucleic Acids Res. 50 (D1), D622–D31 (2022). PubMed PMID: 34986597. Pubmed Central PMCID: 8728138.
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28 (1), 27–30 (2000). PubMed PMID: 10592173. Pubmed Central PMCID: 102409.
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51 (D1), D587–D92 (2023). PubMed PMID: 36300620. Pubmed Central PMCID: 9825424.
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28 (11), 1947–1951 (2019). PubMed PMID: 31441146. Pubmed Central PMCID: 6798127.
Hassani, S. & Esmaeili, A. The neuroprotective effects of ferulic acid in toxin-induced models of Parkinson’s disease: a review. Ageing Res. Rev. 97, 102299 (2024). PubMed PMID: 38604452.
Singh Tuli, H. et al. Ferulic acid: a natural phenol that inhibits neoplastic events through modulation of Oncogenic Signaling. Molecules ;27(21). (2022). PubMed PMID: 36364478. Pubmed Central PMCID: 9654319.
Amimo, J. O. et al. Immune Impairment Associated with vitamin A Deficiency: insights from Clinical studies and Animal Model Research. Nutrients ;14(23). (2022). PubMed PMID: 36501067. Pubmed Central PMCID: 9738822.
Gombart, A. F., Pierre, A. & Maggini, S. A review of micronutrients and the Immune System-Working in Harmony to reduce the risk of infection. Nutrients ;12(1). (2020). PubMed PMID: 31963293. Pubmed Central PMCID: 7019735.
Wolowczuk, I. et al. Feeding our immune system: impact on metabolism. Clin. Dev. Immunol. 2008;2008:639803. PubMed PMID: 18350123. Pubmed Central PMCID: 2266987.
Chen, B. W., Zhang, K. W., Chen, S. J., Yang, C. & Li, P. G. Vitamin A Deficiency exacerbates gut microbiota dysbiosis and cognitive deficits in Amyloid Precursor Protein/Presenilin 1 transgenic mice. Front. Aging Neurosci. 13, 753351 (2021). PubMed PMID: 34790112. Pubmed Central PMCID: 8591312.
Huda, M. N. et al. Neonatal vitamin a supplementation and vitamin a Status are Associated with gut Microbiome Composition in Bangladeshi infants in early infancy and at 2 years of age. J. Nutr. 149 (6), 1075–1088 (2019). PubMed PMID: 31006815. Pubmed Central PMCID: 6543205.
Murphey, L. J. et al. Bradykinin and its metabolite bradykinin 1–5 inhibit thrombin-induced platelet aggregation in humans. J. Pharmacol. Exp. Ther. 318 (3), 1287–1292 (2006). PubMed PMID: 16772538.
Moreau, M. E. et al. The kallikrein-kinin system: current and future pharmacological targets. J. Pharmacol. Sci. 99 (1), 6–38 (2005). PubMed PMID: 16177542.
Wang, W. et al. Bradykinin promotes proliferation, migration, and invasion of cervical cancer cells through STAT3 signaling pathways. Oncol. Rep. 42 (6), 2521–2527 (2019). PubMed PMID: 31638249. Pubmed Central PMCID: 6859440.
Wang, G., Sun, J., Liu, G., Fu, Y. & Zhang, X. Bradykinin promotes Cell Proliferation, Migration, Invasion, and Tumor Growth of Gastric Cancer through ERK Signaling Pathway. J. Cell. Biochem. 118 (12), 4444–4453 (2017). PubMed PMID: 28464378.
Tosi, M. R. & Tugnoli, V. Cholesteryl esters in malignancy. Clinica Chimica acta; international journal of clinical chemistry. ;359(1–2):27–45. (2005). PubMed PMID: 15939411.
Jeong, S. M. et al. Effect of change in total cholesterol levels on Cardiovascular Disease among Young adults. J. Am. Heart Association ;7(12). (2018). PubMed PMID: 29899019. Pubmed Central PMCID: 6220545.
Genghof, D. S. 2,5-dihydrophenylalanine as an inhibitor of microbial growth. Can. J. Microbiol. 16 (6), 545–547 (1970). PubMed PMID: 4912492.
Grant, R. S., Coggan, S. E. & Smythe, G. A. The physiological action of picolinic acid in the human brain. Int. J. Tryptophan Res. : IJTR ;2: (2009). 71 – 9. PubMed PMID: 22084583. Pubmed Central PMCID: 3195224.
Tan, L., Yu, J. T. & Tan, L. The kynurenine pathway in neurodegenerative diseases: mechanistic and therapeutic considerations. J. Neurol. Sci. 323 (1–2), 1–8 (2012). PubMed PMID: 22939820.
Heffernan, M. P., Nelson, M. M. & Anadkat, M. J. A pilot study of the safety and efficacy of picolinic acid gel in the treatment of acne vulgaris. Br. J. Dermatol. 156 (3), 548–552 (2007). PubMed PMID: 17300246.
Cayley, W. E. Jr. Beta2 agonists for Acute Cough or a clinical diagnosis of Acute Bronchitis. Am. Family Phys. 95 (9), 551–552 (2017). PubMed PMID: 28671384.
Kuo, A., Blough, N. V. & Dunlap, P. V. Multiple N-acyl-L-homoserine lactone autoinducers of luminescence in the marine symbiotic bacterium Vibrio fischeri. J. Bacteriol. 176 (24), 7558–7565 (1994). PubMed PMID: 8002580. Pubmed Central PMCID: 197213.
Lithgow, J. K. et al. The regulatory locus cinRI in Rhizobium leguminosarum controls a network of quorum-sensing loci. Mol. Microbiol. 37 (1), 81–97 (2000). PubMed PMID: 10931307.
McClean, K. H. et al. Quorum sensing and Chromobacterium violaceum: exploitation of violacein production and inhibition for the detection of N-acylhomoserine lactones. Microbiology 143 (Pt 12), 3703–3711 (1997). PubMed PMID: 9421896.
Riedel, K. et al. N-acylhomoserine-lactone-mediated communication between Pseudomonas aeruginosa and Burkholderia cepacia in mixed biofilms. Microbiology 147 (Pt 12), 3249–3262 (2001). PubMed PMID: 11739757.
Winson, M. K. et al. Multiple N-acyl-L-homoserine lactone signal molecules regulate production of virulence determinants and secondary metabolites in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. U.S.A. 92 (20), 9427–9431 (1995). PubMed PMID: 7568146. Pubmed Central PMCID: 40998.
Itonaga, T., Izawa, M., Hamajima, T. & Hasegawa, Y. First Morning Pregnanetriol and 17-Hydroxyprogesterone correlated significantly in 21-Hydroxylase Deficiency. Front. Endocrinol. 12, 808254 (2021). PubMed PMID: 35140686. Pubmed Central PMCID: 8820395.
Williams, H. R. et al. Characterization of inflammatory bowel disease with urinary metabolic profiling. Am. J. Gastroenterol. 104 (6), 1435–1444 (2009). PubMed PMID: 19491857.
Matsumoto, T., Kobayashi, T. & Kamata, K. Role of lysophosphatidylcholine (LPC) in atherosclerosis. Curr. Med. Chem. 14 (30), 3209–3220 (2007). PubMed PMID: 18220755.
Zhao, Z. et al. Plasma lysophosphatidylcholine levels: potential biomarkers for colorectal cancer. J. Clin. Oncol. 25 (19), 2696–2701 (2007). PubMed PMID: 17602074.
Sullentrop, F. et al. 31P NMR spectroscopy of blood plasma: determination and quantification of phospholipid classes in patients with renal cell carcinoma. NMR Biomed. 15 (1), 60–68 (2002). PubMed PMID: 11840554.
Kuliszkiewicz-Janus, M., Janus, W. & Baczynski, S. Application of 31P NMR spectroscopy in clinical analysis of changes of serum phospholipids in leukemia, lymphoma and some other non-haematological cancers. Anticancer Res. 1996 May-Jun ;16(3B):1587–1594. PubMed PMID: 8694531.
Goetzl, E. J., Kong, Y. & Mei, B. Lysophosphatidic acid and sphingosine 1-phosphate protection of T cells from apoptosis in association with suppression of Bax. J. Immunol. 162 (4), 2049–2056 (1999). PubMed PMID: 9973477.
Liu, Y. et al. Edg-1, the G protein-coupled receptor for sphingosine-1-phosphate, is essential for vascular maturation. J. Clin. Investig. 106 (8), 951–961 (2000). PubMed PMID: 11032855. Pubmed Central PMCID: 314347.
Dorsam, G. et al. Transduction of multiple effects of sphingosine 1-phosphate (S1P) on T cell functions by the S1P1 G protein-coupled receptor. J. Immunol. 171 (7), 3500–3507 (2003). PubMed PMID: 14500646.
Watson, C. et al. High expression of sphingosine 1-phosphate receptors, S1P1 and S1P3, sphingosine kinase 1, and extracellular signal-regulated kinase-1/2 is associated with development of tamoxifen resistance in estrogen receptor-positive breast cancer patients. Am. J. Pathol. 177 (5), 2205–2215 (2010). PubMed PMID: 20889557. Pubmed Central PMCID: 2966780.
Neubauer, H. A. et al. An oncogenic role for sphingosine kinase 2. Oncotarget 7 (40), 64886–64899 (2016). PubMed PMID: 27588496. Pubmed Central PMCID: 5323123.
Baek, D. J. et al. Structure-activity relationships and molecular modeling of sphingosine kinase inhibitors. J. Med. Chem. 56 (22), 9310–9327 (2013). PubMed PMID: 24164513. Pubmed Central PMCID: 3848335.
Dietz, B., Heilmann, J. & Bauer, R. Absorption of dodeca-2E,4E,8Z,10E/Z-tetraenoic acid isobutylamides after oral application of Echinacea purpurea tincture. Planta Med. 67 (9), 863–864 (2001). PubMed PMID: 11745026.
Dambrova, M. et al. Acylcarnitines: nomenclature, biomarkers, therapeutic potential, drug targets, and clinical trials. Pharmacol. Rev. 74 (3), 506–551 (2022). PubMed PMID: 35710135.
Sorgdrager, F. J. H., Naude, P. J. W., Kema, I. P., Nollen, E. A. & Deyn, P. P. Tryptophan Metabolism in Inflammaging: from biomarker to therapeutic target. Front. Immunol. 10, 2565 (2019). PubMed PMID: 31736978. Pubmed Central PMCID: 6833926.
Roth, W., Zadeh, K., Vekariya, R., Ge, Y. & Mohamadzadeh, M. Tryptophan metabolism and Gut-Brain Homeostasis. Int. J. Mol. Sci. ;22(6). (2021). PubMed PMID: 33804088. Pubmed Central PMCID: 8000752.
Jonescheit, H. et al. Influence of Indoleamine-2,3-Dioxygenase and its Metabolite Kynurenine on gammadelta T cell cytotoxicity against Ductal pancreatic adenocarcinoma cells. Cells ;9(5). (2020). PubMed PMID: 32384638. Pubmed Central PMCID: 7290398.
Li, R. Z. et al. The key role of sphingolipid metabolism in cancer: New therapeutic targets, diagnostic and prognostic values, and anti-tumor immunotherapy resistance. Front. Oncol. 12, 941643 (2022). PubMed PMID: 35965565. Pubmed Central PMCID: 9364366.
Hannun, Y. A. & Obeid, L. M. Principles of bioactive lipid signalling: lessons from sphingolipids. Nat. Rev. Mol. Cell Biol. 9 (2), 139–150 (2008). PubMed PMID: 18216770.
Morad, S. A. et al. Ceramide–antiestrogen nanoliposomal combinations–novel impact of hormonal therapy in hormone-insensitive breast cancer. Mol. Cancer Ther. 11 (11), 2352–2361 (2012). PubMed PMID: 22962326. Pubmed Central PMCID: 3495995.
Watters, R. J. et al. Targeting glucosylceramide synthase synergizes with C6-ceramide nanoliposomes to induce apoptosis in natural killer cell leukemia. Leuk. Lymphoma. 54 (6), 1288–1296 (2013). PubMed PMID: 23181473. Pubmed Central PMCID: 4296321.
Li, G. et al. Nanoliposome C6-Ceramide increases the Anti-tumor Immune Response and slows growth of liver tumors in mice. Gastroenterology 154 (4), 1024–1036 (2018). e9. PubMed PMID: 29408569. Pubmed Central PMCID: 5908238.
Wang, S. B., Ma, Y. Y., Chen, X. Y., Zhao, Y. Y. & Mou, X. Z. Ceramide-Graphene Oxide nanoparticles enhance cytotoxicity and decrease HCC Xenograft Development: a Novel Approach for targeted Cancer Therapy. Front. Pharmacol. 10, 69 (2019). PubMed PMID: 30800068. Pubmed Central PMCID: 6376252.
van Meer, G., Voelker, D. R. & Feigenson, G. W. Membrane lipids: where they are and how they behave. Nat. Rev. Mol. Cell Biol. 9 (2), 112–124 (2008). PubMed PMID: 18216768. Pubmed Central PMCID: 2642958.
De Libero, G. & Mori, L. Structure and biology of self lipid antigens. Curr. Top. Microbiol. Immunol. 314, 51–72 (2007). PubMed PMID: 17593657.
Brennan, P. J., Brigl, M. & Brenner, M. B. Invariant natural killer T cells: an innate activation scheme linked to diverse effector functions. Nat. Rev. Immunol. 13 (2), 101–117 (2013). PubMed PMID: 23334244.
Di Gioia, M. & Zanoni, I. Dooming phagocyte responses: Inflammatory effects of Endogenous oxidized phospholipids. Front. Endocrinol. 12, 626842 (2021). PubMed PMID: 33790857. Pubmed Central PMCID: 8005915.
Matthews, D. E. An overview of phenylalanine and tyrosine kinetics in humans. J. Nutr. 137 (6 Suppl 1), 1549 (2007). discussion 73S-75S. PubMed PMID: 17513423. Pubmed Central PMCID: 2268015.S-55S.
Nguyen, T. N., Nguyen, H. Q. & Le, D. H. Unveiling prognostics biomarkers of tyrosine metabolism reprogramming in liver cancer by cross-platform gene expression analyses. PloS One. 15 (6), e0229276 (2020). PubMed PMID: 32542016. Pubmed Central PMCID: 7295234.
Martinho, D. V. et al. Oral branched-chain amino acids supplementation in athletes: a systematic review. Nutrients ;14(19). (2022). PubMed PMID: 36235655. Pubmed Central PMCID: 9571679.
Wang, T. J. et al. Metabolite profiles and the risk of developing diabetes. Nat. Med. 17 (4), 448–453 (2011). PubMed PMID: 21423183. Pubmed Central PMCID: 3126616.
Huffman, K. M. et al. Relationships between circulating metabolic intermediates and insulin action in overweight to obese, inactive men and women. Diabetes care. 32 (9), 1678–1683 (2009). PubMed PMID: 19502541. Pubmed Central PMCID: 2732163.
Newgard, C. B. et al. A branched-chain amino acid-related metabolic signature that differentiates obese and lean humans and contributes to insulin resistance. Cell Metabol. 9 (4), 311–326 (2009). PubMed PMID: 19356713. Pubmed Central PMCID: 3640280.
Meng, Z. C., Guangchao; Yang, Q., Yang, H. & Hao, J. Yin, Zhinan. Metabolic control of γδ T cell function. Infect. Microbes Dis. ;3(3). (2021).
Acknowledgements
We thank Sulma P. Claros and Andrew Magnuson for their technical and animal care assistance.
Author information
Authors and Affiliations
Contributions
Conceptualized and designed the experiments, H. M., and W.W. Data collection, H.M., G.W.W., E.C., M.M.V., D.L. Analyzed the data and wrote the paper, H.M.; critically revised the paper, W.W., and P.S. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
H.M., G.W.W., M.M.V, D.L., E.C., P.S., W.W. are employed by InGenious Targeting Laboratory, Ronkonkoma, NY. The authors declared that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Michael, H., Weng, G.W., Vallas, M.M. et al. Metabolomics analysis reveals resembling metabolites between humanized γδ TCR mice and human plasma. Sci Rep 14, 29321 (2024). https://doi.org/10.1038/s41598-024-81003-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-81003-y