Abstract
The aim of this study was to investigate the mechanisms of acquired resistance to anti-epidermal growth factor receptor (EGFR) therapy in metastatic colorectal cancer (mCRC), with a focus on the role of KRAS secondary mutations. We sought to evaluate how these mutations contribute to resistance in patients treated with chemotherapy alone or in combination with cetuximab, and to explore the impact of treatment duration and disease progression on mutation rates. We retrospectively collected data from 50 mCRC patients with wild-type KRAS who received either chemotherapy alone or chemotherapy combined with anti-EGFR therapy (cetuximab). Following treatment, tumor samples were recollected through surgery, biopsy, or colonoscopy. The secondary mutation status of KRAS, NRAS, and BRAF was determined via an amplification refractory mutation system. Based on the nature of the chemotherapy regimen and the presence or absence of tumor progression at the time of the second KRAS mutation analysis, patients were stratified into three groups: Group A (adjuvant chemotherapy and recurrence), Group B (first-line chemotherapy without progression), and Group C (first-line chemotherapy with progression). Secondary KRAS mutations were assessed in relation to therapeutic regimens and progression profiles to investigate the mechanisms of acquired resistance to anti-EGFR therapy. The overall rate of secondary KRAS mutations was 6.0% (3/50). These mutations were not observed in patients treated with chemotherapy alone (0/24), but occurred in 11.5% (3/26) of patients receiving chemotherapy combined with cetuximab. Stratification by treatment regimens and tumor progression showed mutation rates of 0% (0/13) in Group A, 0% (0/16) in Group B, and 14.3% (3/21) in Group C. In Group C, the mutation rate rose from 0% (0/5) in patients receiving chemotherapy alone to 18.8% (3/16) in those receiving combination therapy. Furthermore, in patients receiving chemotherapy combined with cetuximab of Group B and Group C, longer anti-EGFR treatment duration was associated with a higher mutation rate: 5.9% (1/17) in those treated for ≤ 10 months versus 25.0% (2/8) in those treated for > 10 months. Including one case of secondary BRAF mutation, the cumulative mutation rate reached 37.5% (3/8) in patients with prolonged cetuximab exposure. This study innovatively revealed from multiple perspectives that the crucial role of secondary KRAS mutations in the resistance of mCRC patients to EGFR therapy, emphasizes the importance of continuous genomic monitoring, and provides new ideas for future combination targeted therapies.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is one of the most common cancers worldwide, with an annual incidence of approximately 1.9 million cases, and is a leading cause of cancer-related mortality1. The incidence of CRC increases with age, particularly among individuals aged 50 years and older. Additionally, lifestyle factors such as smoking, alcohol consumption, and inadequate dietary fiber intake, are contributing to a rising incidence of CRC in younger populations. As a result, CRC has become a major cause of cancer-related deaths among young adults in both developed and developing countries2,3. A significant proportion of patients are diagnosed at an advanced stage, and approximately 15–30% of individuals with early-stage CRC experience tumor recurrence after curative surgery, which adversely affects their quality of life and overall survival4. The median survival time for patients with metastatic colorectal cancer (mCRC) is only 3 years, indicating a poor prognosis5. In many societies, CRC is emerging as an increasingly urgent public health challenge, particularly as populations age, underscoring the critical need for effective screening and treatment strategies.
Cytotoxic chemotherapy regimens incorporating oxaliplatin or irinotecan have traditionally been the cornerstone of treatment for patients with mCRC6. However, recent advancements in tumor molecular biology have led to significant and transformative shifts in the mCRC treatment landscape. These changes are increasingly driven by biomarker-based strategies, including the evaluation of microsatellite instability (MSI) or mismatch repair (MMR) status, as well as the identification of genomic alterations in genes such as RAS and BRAF7. For patients with MSI-high mCRC, the combination of cytotoxic chemotherapy and immunotherapy has emerged as the most effective treatment regimen8,9. Additionally, the epidermal growth factor receptor (EGFR) has become a critical therapeutic target in mCRC patients with RAS/BRAF wild-type status. Specifically, for anatomically left-sided tumors, the use of cytotoxic chemotherapy combined with anti-EGFR monoclonal antibodies has been established as the preferred treatment modality, significantly improving patient survival outcomes10,11. In contrast, clinical guidelines recommend against the use of anti-EGFR therapy in mCRC individuals with RAS or BRAF mutations due to the limited efficacy observed in this subset of patients.
EGFR, a member of the ERBB protein family, plays a critical role in regulating cellular growth and migration12. Anti-EGFR monoclonal antibodies—such as cetuximab, an IgG1 chimeric antibody containing both human and murine components, and panitumumab, a fully human IgG2 antibody—competitively bind to the extracellular ligand-binding domain of EGFR. This binding inhibits ligand interaction and subsequent receptor phosphorylation, thereby blocking downstream signaling cascades, particularly the RAS/RAF/MAPK and PI3K/AKT/mTOR pathways. As a result, these therapies affect key cellular processes including proliferation, apoptosis, angiogenesis, and migration13. Although combining cytotoxic chemotherapy with EGFR-targeted therapy has led to improved survival outcomes in patients with mCRC harboring wild-type RAS/RAF, nearly all tumors eventually develop secondary resistance to anti-EGFR treatment14. This resistance typically arises after initial exposure to agents like cetuximab, where tumors that initially respond subsequently undergo clonal evolution and acquire molecular heterogeneity over time and across different tumor regions15. Such adaptive changes enable tumors to evade therapeutic pressure, ultimately rendering EGFR-targeted therapies ineffective.
Understanding the mechanisms of resistance is essential, and both genomic and non-genomic mechanisms underlying resistance to EGFR inhibitors remain the focus of ongoing investigation16. Under the selective pressure of EGFR-targeted therapies, approximately 40–50% of patients acquire genetic alterations. These include mutations in the RAS, BRAF, and PIK3CA genes, mutations in the extracellular domain of EGFR, and HER2 amplification—each contributing significantly to the development of resistance to EGFR inhibitors17,18. The RAS oncogene family—including KRAS, NRAS, and HRAS—belongs to the small GTPase superfamily and functions as a crucial molecular switch in the regulation of cell proliferation and survival through downstream signaling pathways19. These oncogenes are recognized as central contributors to acquired resistance against anti-EGFR monoclonal antibodies. Notably, primary KRAS mutations are detected in nearly 40% of CRC patients, while NRAS and HRAS mutations are relatively rare20. Importantly, anti-EGFR antibodies exert selective pressure that can induce secondary mutations in initially wild-type KRAS, leading to its aberrant activation independent of upstream EGFR signaling. This activation involves interactions with downstream effectors, such as RAF kinases, sustaining continuous signaling through proliferative pathways21. As a result, tumors become resistant to EGFR-targeted therapy, promoting unchecked cancer cell growth and metastasis.
Circulating tumor DNA (ctDNA) is released into the bloodstream through the apoptotic and necrotic processes of tumor cells originating from primary or metastatic lesions.
This ctDNA can be detected using methodologies such as polymerase chain reaction (PCR) and next-generation sequencing (NGS)22,23. Liquid biopsy, which analyzes ctDNA, offers a noninvasive method for dynamically monitoring genetic alterations in plasma, presenting a promising alternative to traditional tissue biopsy for assessing tumor genomic profiles24. Studies have shown that among patients with RAS wild-type mCRC treated with EGFR inhibitors, 46.7% of plasma samples exhibited secondary RAS mutations detected via PCR25. Another study reported that ctDNA testing conducted before and after anti-EGFR antibody therapy identified genetic alterations in RAS, BRAF, and other genes in 62.0% of patients18. Despite its potential, liquid biopsy has notable limitations. ctDNA constitutes only a small fraction of the total cell-free DNA in circulation, and approximately 15% of cases may lack detectable levels of ctDNA. Furthermore, studies have reported up to a 20% discrepancy in RAS mutation detection between tumor tissue and plasma-derived ctDNA26. As such, tissue biopsy remains the gold standard for determining tumor clonality and conducting comprehensive genomic analyses. However, due to the practical difficulties in obtaining tissue samples, most studies rely on liquid biopsies, with fewer incorporating tumor tissue to explore resistance mechanisms to anti-EGFR therapy. Moreover, many of these studies lack chemotherapy control groups, which limits the strength of their conclusions.
In this study, we performed genetic analyses of tumor tissues collected both before and after anti-EGFR therapy, including a chemotherapy control group for comparison. Subgroup stratification was based on progression status following combined chemotherapy and EGFR inhibitor treatment. The primary objective was to elucidate the molecular mechanisms underlying acquired resistance to EGFR blockade, with the goal of identifying potential combination targeted therapy strategies to overcome resistance and ultimately improve patient survival outcomes.
Materials and methods
Patient and protocol
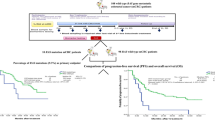
In this study, we retrospectively collected data from fifty patients with mCRC harboring wild-type KRAS, who received either chemotherapy alone or in combination with anti-EGFR therapy at our institution over the past decade. Following treatment, these patients underwent tumor rebiopsy for a second KRAS mutation analysis. Patients were stratified into three groups based on the nature of chemotherapy regimen received and the presence or absence of tumor progression at the time of the second KRAS mutation analysis: Group A (adjuvant chemotherapy with recurrence), Group B (first-line chemotherapy without progression), and Group C (first-line chemotherapy with progression). Recurrence was defined as the reappearance of tumor lesions or metastases during the follow-up period after adjuvant chemotherapy, confirmed through clinical evaluation or pathological biopsy. Progressive disease (PD) was characterized by a ≥ 20% increase in the sum of the largest diameters of target lesions during follow-up after first-line chemotherapy, or by the emergence of new lesions. Treatment regimens in Group A included FOLFOX, XELOX, and other protocols. Groups B and C received regimens consisting of FOLFOX, XELOX, FOLFIRI, or other chemotherapy combinations, with or without the addition of the anti-EGFR monoclonal antibody cetuximab. Cetuximab was administered either biweekly at a dose of 500 mg/m2 or weekly, starting with an initial dose of 400 mg/m2 followed by 250 mg/m2 for subsequent treatments. The study was conducted in accordance with the principles outlined in the Declaration of Helsinki, and all experimental protocols were approved by the Institutional Review Board of West China Hospital.
Test method for gene mutation
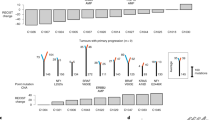
Tumor samples were collected prior to the commencement of drug therapy, with all patients verified to possess wild-type KRAS at the initiation of therapy. Following treatment with chemotherapy or a regimen combining chemotherapy and anti-EGFR therapy, additional tumor specimens, encompassing primary tumors or metastases, were obtained via surgical resection, biopsy, or colonoscopy. DNA was subsequently obtained from either paraffin-embedded or fresh tissue samples. The amplification refractory mutation system (ARMS) was employed to identify mutations in the KRAS, BRAF, and NRAS genes.
The ARMS technique is a highly sensitive and specific polymerase chain reaction (PCR)-based method designed to selectively amplify mutant alleles in the presence of an excess of wild-type DNA. The ARMS assay can detect mutations with a sensitivity as high as 0.1–1%, making it suitable for identifying low-frequency mutations in heterogeneous tumor samples. The specificity of the assay is ensured by the use of allele-specific primers that discriminate single nucleotide polymorphisms (SNPs) through selective amplification, minimizing non-specific binding and amplification of wild-type sequences. The ARMS assay was performed according to manufacturer guidelines, and data were analyzed using standard threshold settings for mutation detection. Mutations were confirmed by independent sequencing in cases where ambiguity was observed.
Clinical assessments
Tumor response was evaluated using computed tomography (CT) every 1–2 months, following the Response Evaluation Criteria in Solid Tumors (RECIST) version 1.1, until disease progression was observed. Response categories included complete response (CR), partial response (PR), stable disease (SD), and PD.
Statistical methods
Qualitative variables were presented as counts and percentages, while continuous variables were summarized using medians and ranges. Differences in the rates of secondary KRAS mutations among the cohorts were evaluated using the chi-square test. A p value of less than 0.05 was considered statistically significant. All statistical analyses were performed using SPSS software, version 22.
Results
Patient characteristics
The specific clinical features of the 50 individuals are presented in Table 1. Group A included 13 patients (4 women and 9 men) with a median age of 52 years (range 30–72 years). Treatment regimens included FOLFOX in 10 patients, XELOX in 2, and XELOX plus cetuximab in 1 patient. The median number of treatment cycles was 8.7 (range 4–12), with a median treatment duration of 5.5 months (range 2.3–10.6 months). The median interval between disease recurrence and KRAS mutation detection was 58.4 days (range 5–336 days). Group B included 16 patients (5 women and 11 men), with a median age of 51.5 years (range 30–73 years). Of the patients, 7 received chemotherapy alone, while 9 were treated with chemotherapy combined with cetuximab. The median number of treatment cycles was 5.4 (range 3–12), and the median treatment duration was 3.8 months (range 1.6–6.8 months). Among these patients, 10 achieved a PR, while 6 demonstrated SD. Group C consisted of 21 patients (6 women, 15 men), with a median age of 51 years (range 27–75 years). Of the 21 patients, 5 were treated with chemotherapy alone and 16 with chemotherapy plus cetuximab. The median number of treatment cycles administered was 17.5 (range 2–55). For patients receiving chemotherapy plus cetuximab, the median number of cetuximab cycles was 16 (range 4–37). The median treatment duration was 14.9 months (range 1.4–50.1 months). Among these patients, one achieved a CR, 11 achieved a PR, 5 maintained SD, and 4 experienced PD. The median interval from disease progression to KRAS mutation detection was 21 days (range 1–230 days).
Secondary KRAS mutation rate
Among the cohort of 50 patients, 3 developed secondary KRAS mutations, resulting in an overall secondary KRAS mutation rate of 6.0% (3/50). The secondary KRAS mutation rates for Group A, Group B, and Group C were 0% (0/13), 0% (0/16), and 14.3% (3/21), respectively (p = 0.11) (Table 1). Detailed information on secondary KRAS mutations in patients from Groups B and C treated with combined chemotherapy and cetuximab is presented in Table 2. In contrast, Table 3 outlines the specifics of secondary KRAS mutations in patients from the same groups who received chemotherapy alone.
In the subgroup analysis stratified by therapeutic regimens with or without cetuximab, the incidence of secondary KRAS mutations was 0% (0/24) in the chemotherapy-alone subgroup and 11.5% (3/26) in the chemotherapy plus cetuximab subgroup. Within Group A, the mutation rate was 0% in both the chemotherapy-alone subgroup (0/12) and the chemotherapy plus cetuximab subgroup (0/1). Similarly, in Group B, no secondary KRAS mutations were detected in either the chemotherapy-alone subgroup (0/7) or the chemotherapy plus cetuximab subgroup (0/9). In contrast, within Group C, the mutation rate remained 0% in the chemotherapy-alone subgroup (0/5), while the chemotherapy plus cetuximab subgroup exhibited a notably higher incidence of secondary KRAS mutations at 18.8% (3/16) (Table 4).
Among patients in Groups B and C, 25 individuals received cetuximab-based therapy, with a median duration of targeted treatment of 5.6 months (range 1.4–43.9 months). Subgroup analysis stratified by targeted treatment duration revealed a secondary KRAS mutation rate of 5.9% (1/17) in patients with a treatment duration of less than 10 months, compared to 25.0% (2/8) in those treated for more than 10 months (p = 0.23). When one case of secondary BRAF mutation was included, the overall secondary mutation rate in the > 10-month subgroup increased to 37.5% (3/8), with a trend toward statistical significance (p = 0.08) (Table 5).
BRAF and NRAS mutation rates
Among the cohort of 50 patients, 36 underwent post-therapy testing for BRAF mutations. Of these, 34 were identified as BRAF wild-type, while 2 exhibited BRAF mutations. Notably, one of the two mutated cases had been classified as wild-type prior to therapy. The overall post-therapy BRAF mutation rate was 5.6% (2/36). Within this subgroup of 36 patients, 6 individuals with BRAF wild-type status were assigned to Group A; Group B consisted of 13 patients with BRAF wild-type and 1 with a BRAF mutation; Group C included 15 BRAF wild-type patients and 1 who acquired a secondary BRAF mutation post-therapy.
Of the original 50 patients, 34 underwent post-therapy testing for NRAS mutations. All tested individuals were confirmed to be NRAS wild-type. The distribution of these patients across the three groups was as follows: 6 in Group A, 14 in Group B, and 14 in Group C.
Discussion
The study finds that secondary KRAS mutations occur in 6% of cases overall. The frequency of these mutations varies with the treatment regimen—no mutations were observed in the chemotherapy-alone group, whereas a frequency of 11.5% (3/26) was detected in patients receiving chemotherapy combined with cetuximab. Based on stratification by the nature of the chemotherapy regimen and the presence or absence of tumor progression at the time of the second KRAS mutation analysis, the secondary KRAS mutation rates increased from 0% (0/13) in Group A and 0% (0/16) in Group B to 14.3% (3/21) in Group C. In Group C, the secondary KRAS mutation rate rose from 0% (0/5) in patients receiving chemotherapy alone to 18.8% (3/16) in those treated with chemotherapy combined with cetuximab. Furthermore, in the chemotherapy plus cetuximab subgroups of Group B and Group C, stratified by anti-EGFR therapy duration, the secondary KRAS mutation rate increased from 5.9% (1/17) in the subgroup with a duration of 10 months or less to 25.0% (2/8) in the subgroup with a duration of more than 10 months. Including one additional case of secondary BRAF mutation, the cumulative secondary mutation rate in the subgroup with anti-EGFR therapy duration exceeding 10 months rose to 37.5% (3/8). This study innovatively revealed from multiple perspectives that secondary KRAS mutations, which play a significant role in the development of resistance to anti-EGFR treatment in mCRC patients, are more frequently observed in those treated with cetuximab, particularly with longer treatment durations.
Previous studies have demonstrated that secondary KRAS mutations are key contributors to acquired resistance to EGFR inhibitor therapy in mCRC patients, accounting for more than half of resistance cases27. However, most of this knowledge is derived from research focused on later-line single-agent anti-EGFR therapies, with limited data available regarding resistance mechanisms during first-line combination treatment with cytotoxic chemotherapy27. One study reported that secondary mutations were significantly more frequent with anti-EGFR monotherapy (46%) compared to combination therapy (9%)14. Similarly, other research has shown that while 62.0% of patients receiving later-line EGFR inhibitor monotherapy developed genomic alterations, only 6.6% of those treated with first-line cetuximab-based chemotherapy exhibited such changes18. Our study provides novel insights into the development of acquired resistance to anti-EGFR therapy in mCRC by systematically analyzing secondary KRAS mutation rates across stratified patient subgroups. While prior research has identified secondary KRAS mutations as a resistance mechanism, our study is to comprehensively stratify these mutations by treatment modality (chemotherapy alone vs. chemotherapy plus cetuximab), disease progression status, and duration of anti-EGFR exposure. Notably, our findings reveal that secondary KRAS mutations were absent in all patients receiving chemotherapy alone but occurred in 11.5% of those receiving combination therapy, and reached as high as 18.8% in progressive patients treated with cetuximab. Furthermore, we observed a significant increase in mutation rates with longer cetuximab exposure, rising from 5.9% in patients treated ≤ 10 months to 25.0% in those treated > 10 months. These stratified analyses offer a more nuanced understanding of resistance development, surpassing the scope of previous studies that often lacked such temporal and treatment-based resolution. Our findings advance the field by underscoring the dynamic nature of tumor evolution under prolonged anti-EGFR therapy. Unlike earlier studies that primarily focused on baseline mutations, our results highlight the importance of longitudinal genomic monitoring, particularly in patients receiving extended cetuximab treatment. Previous studies have commonly reported KRAS exon 2 mutations (specifically at codons 12 and 13), which is consistent with our findings, although we did not quantify mutations at specific exons or codons28. Sackstein et al. further demonstrated that KRAS, EGFR, and BRAF mutations frequently arise in patients undergoing anti-EGFR therapy, highlighting the adaptive capacity of tumor cells under selective therapeutic pressure15. In our study, we also observed a case of secondary BRAF mutation, although EGFR gene testing was not performed. Additionally, our findings are in line with those of Topham et al., who reported that secondary NRAS mutations were not detected in mCRC patients treated with a combination of anti-EGFR therapy and chemotherapy14.
Investigating the relationship between resistance mechanisms to anti-EGFR therapy and treatment regimens or duration is of significant value. Parseghian et al. reported that resistance mechanisms differ depending on the type and combination of therapies14.
Specifically, transcriptomic alterations are more commonly associated with combined cytotoxic chemotherapy and anti-EGFR therapy, while MAPK pathway mutations are more frequently observed with anti-EGFR monotherapy14. Their study also demonstrated that first-line combination therapies are associated with fewer acquired genomic alterations compared to later-line treatments. In our study, secondary mutations of KRAS, NRAS, and BRAF were absent in both the adjuvant and first-line chemotherapy groups without anti-EGFR treatment. These findings suggest that cytotoxic chemotherapy alone does not affect common MAPK pathway mutations. Unfortunately, we did not perform an analysis of transcriptomic mechanisms. In Group C of our study, one patient underwent anti-EGFR therapy for 10.3 months, whereas two other patients exceeded 41 months, suggesting that longer treatment may increase secondary genomic alterations. This finding indicates a link between secondary mutations and resistance to anti-EGFR therapy, which typically emerges around 10 months, coinciding with the median progression-free survival (mPFS) of first-line chemotherapy combined with cetuximab. Another patient developed secondary mutations after only 1.9 months of treatment, suggesting that tumor heterogeneity may have been present prior to therapy initiation. Clonal evolution driven by anti-EGFR treatment remains a major challenge in the management of mCRC29. The emergence of secondary genomic alterations serves as evidence of this evolutionary process. We further show that not only do secondary KRAS mutations increase with time, but additional resistance alterations (e.g., BRAF mutations) may also emerge, with a cumulative secondary mutation rate of 37.5% in patients treated with cetuximab for over 10 months. This suggests a multifactorial and time-dependent evolution of resistance. Thus, our study supports a shift toward adaptive treatment strategies informed by ongoing molecular profiling, with the potential to improve patient outcomes by anticipating and counteracting resistance.
ctDNA is increasingly utilized to map resistance in mCRC due to its advantages in dynamic monitoring, minimal invasiveness, safety, and cost-effectiveness28,30,31,32. Recent studies have shown that ctDNA analysis can detect acquired genomic alterations—such as mutations in RAS, BRAF, and the EGFR ectodomain—in approximately 6.6% of mCRC patients undergoing first-line chemotherapy combined with cetuximab18. Topham et al. further demonstrated the evolution of resistance mechanisms through longitudinal ctDNA profiling24. Our study, based on pre- and post-EGFR therapy tissue biopsies, complements this finding by providing a comprehensive molecular characterization of resistance. Although ctDNA offers a noninvasive alternative for monitoring, our results underscore the continued necessity of tissue biopsies for accurately assessing tumor heterogeneity and resistance evolution. Notably, a 20% discrepancy in RAS mutation detection between ctDNA and tissue biopsy highlights the limitations of relying on a single method26. Therefore, combining tissue biopsy with plasma ctDNA testing could enhance the accuracy of resistance profiling and better capture tumor heterogeneity. Continuous genomic monitoring using both methods is essential for understanding resistance dynamics during anti-EGFR rechallenge33,34,35. Integrating ctDNA analysis into routine clinical practice holds the potential to improve the early detection of resistance mechanisms and guide more personalized treatment strategies. The integration of ctDNA and tissue-based real-time genomic analyses to elucidate resistance mechanisms will be a key focus of our future research.
Our study identified key molecular mechanisms underlying secondary resistance to anti-EGFR treatment in mCRC patients, highlighting the crucial role of secondary KRAS mutations in therapy resistance. The absence of secondary gene mutations in the nonprogressive group, which received the same combination therapy, suggests that intact wild-type KRAS and BRAF genes may contribute to maintaining treatment efficacy. In contrast, cytotoxic chemotherapy alone does not induce MAPK pathway mutations, indicating that prolonged anti-EGFR therapy exerts selective pressure, driving clonal evolution and the emergence of resistant subclones. This mechanism supports the notion of clonal evolution, where treatment pressure selects for resistant clones, leading to tumor heterogeneity and ultimately treatment failure29. Furthermore, our study demonstrated the dynamic nature of resistance mechanisms, as evidenced by the variation in genetic test results before and after disease progression.
The presence of secondary BRAF mutations underscores the multifactorial nature of anti-EGFR tolerance in mCRC, with multiple genetic alterations enabling cancer cells to escape treatment.
Our study has several limitations. The small sample size limits the generalizability of our findings, and the exclusion of patients receiving anti-EGFR monotherapy further narrows the applicability of the results. Additionally, the absence of genetic testing for key genes such as EGFR, HER2, and MET restricts the comprehensiveness of our resistance profiling. As our analysis is based solely on tissue biopsies, it also has inherent limitations in detecting low-frequency mutations and fully capturing tumor heterogeneity. Future studies should include larger and more diverse patient cohorts across different therapy stages to validate and extend our findings. Comprehensive genomic profiling, encompassing a broader range of resistance-related genes, is essential to better elucidate the mechanisms underlying therapy resistance. Furthermore, real-time dynamic genomic monitoring that combines tissue biopsies with ctDNA analysis should be employed to track the evolution of tumor resistance and enable timely treatment adaptations. Exploring the potential of anti-EGFR rechallenge strategies in combination with other targeted therapies or immunotherapies may also provide new avenues for overcoming resistance. Overall, our study underscores the clinical importance of continuous genomic monitoring in the management of mCRC, particularly given the prominent role of secondary KRAS mutations as a resistance mechanism. A deeper understanding of resistance pathways will support the development of targeted combination therapies, highlighting the need for an integrated, adaptive approach to improve patient outcomes.
Conclusions
Our findings underscore the pivotal role of secondary KRAS mutations in conferring resistance to EGFR inhibition in mCRC. Continuous genomic monitoring is imperative for elucidating and managing resistance mechanisms. Future research should encompass larger and more diverse cohorts. These insights are vital for the development of targeted combination therapies aimed at enhancing survival outcomes in mCRC patients.
Data availability
When there is a reasonable request for raw data, the first author can share it.
References
Siegel, R. L., Wagle, N. S., Cercek, A., Smith, R. A. & Jemal, A. Colorectal cancer statistics, 2023. CA Cancer J. Clin. 73, 233–254. https://doi.org/10.3322/caac.21772 (2023).
Morgan, E. et al. Global burden of colorectal cancer in 2020 and 2040: Incidence and mortality estimates from GLOBOCAN. Gut 72, 338–344. https://doi.org/10.1136/gutjnl-2022-327736 (2023).
Klimeck, L., Heisser, T., Hoffmeister, M. & Brenner, H. Colorectal cancer: A health and economic problem. Best Pract. Res. Clin. Gastroenterol. 66, 101839. https://doi.org/10.1016/j.bpg.2023.101839 (2023).
Väyrynen, V. et al. Incidence and management of patients with colorectal cancer and synchronous and metachronous colorectal metastases: A population-based study. BJS Open 4, 685–692. https://doi.org/10.1002/bjs5.50299 (2020).
Yoshino, T. et al. Pan-Asian adapted ESMO clinical practice guidelines for the diagnosis, treatment and follow-up of patients with metastatic colorectal cancer. Esmo Open 8, 10–32. https://doi.org/10.1016/j.esmoop.2023.101558 (2023).
Morris, V. K. et al. Treatment of metastatic colorectal cancer: ASCO guideline. J. Clin. Oncol. 41, 678–700. https://doi.org/10.1200/jco.22.01690 (2023).
Ciardiello, F. et al. Clinical management of metastatic colorectal cancer in the era of precision medicine. CA Cancer J. Clin. 72, 372–401. https://doi.org/10.3322/caac.21728 (2022).
Lenz, H.-J. et al. First-line nivolumab plus low-dose ipilimumab for microsatellite instability-high/mismatch repair-deficient metastatic colorectal cancer: The phase II checkmate 142 study. J. Clin. Oncol. 40, 161–170. https://doi.org/10.1200/jco.21.01015 (2022).
Taieb, J. et al. Deficient mismatch repair/microsatellite unstable colorectal cancer: Diagnosis, prognosis and treatment. Eur. J. Cancer 175, 136–157. https://doi.org/10.1016/j.ejca.2022.07.020 (2022).
Leowattana, W., Leowattana, P. & Leowattana, T. Systemic treatment for metastatic colorectal cancer. World J. Gastroenterol. 29, 1569–1588. https://doi.org/10.3748/wjg.v29.i10.1569 (2023).
Watanabe, J. et al. Panitumumab vs bevacizumab added to standard first-line chemotherapy and overall survival among patients with RAS wild-type, left-sided metastatic colorectal cancer. JAMA 329, 1271–1282. https://doi.org/10.1001/jama.2023.4428 (2023).
Koveitypour, Z. et al. Signaling pathways involved in colorectal cancer progression. Cell Biosci. 9, 97. https://doi.org/10.1186/s13578-019-0361-4 (2019).
Kasi, P. M., Afable, M. G., Herting, C., Lukanowski, M. & Jin, Z. H. Anti-EGFR antibodies in the management of advanced colorectal cancer. Oncologist 28, 1034–1048. https://doi.org/10.1093/oncolo/oyad262 (2023).
Parseghian, C. M. et al. Resistance mechanisms to anti-epidermal growth factor receptor therapy in RAS/RAF wild-type colorectal cancer vary by regimen and line of therapy. J. Clin. Oncol. 41, 460–471. https://doi.org/10.1200/jco.22.01423 (2023).
Sackstein, P. E. et al. Overcoming the hurdles: Surmounting acquired resistance to anti-EGFR therapy in metastatic colorectal cancer. Clin. Adv. Hematol. Oncol. 21, 572–583 (2023).
Ríos-Hoyo, A., Monzonís, X., Vidal, J., Linares, J. & Montagut, C. Unveiling acquired resistance to anti-EGFR therapies in colorectal cancer: A long and winding road. Front. Pharmacol. 15, 1398419. https://doi.org/10.3389/fphar.2024.1398419 (2024).
Tan, E. E. S. et al. Prognostic and predictive value of mutations in metastatic colorectal cancer. Target. Oncol. 17, 483–492. https://doi.org/10.1007/s11523-022-00898-7 (2022).
Raghav, K. et al. Acquired genomic alterations on first-line chemotherapy with cetuximab in advanced colorectal cancer: Circulating tumor DNA analysis of the CALGB/SWOG-80405 trial (alliance). J. Clin. Oncol. 41, 472–478. https://doi.org/10.1200/jco.22.00365 (2023).
Janssens, K. et al. New developments in treating mutated metastatic colorectal cancer. Curr. Treat. Opt. Oncol. 24, 965–987. https://doi.org/10.1007/s11864-023-01095-y (2023).
Muzny, D. M. et al. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337. https://doi.org/10.1038/nature11252 (2012).
Parseghian, C., Eluri, M., Kopetz, S. & Raghav, K. Mechanisms of resistance to EGFR-targeted therapies in colorectal cancer: more than just genetics. Front. Cell Dev. Biol. 11, 11766570. https://doi.org/10.3389/fcell.2023.1176657 (2023).
Malla, M., Loree, J. M., Kasi, P. M. & Parikh, A. R. Using circulating tumor DNA in colorectal cancer: Current and evolving practices. J. Clin. Oncol. 40, 2846–2857. https://doi.org/10.1200/jco.21.02615 (2022).
Mauri, D. et al. Next-generation sequencing of circulating tumor DNA can optimize second-line treatment in RAS wild-type metastatic colorectal cancer after progression on anti-EGFR therapy: Time to rethink our approach. Oncol. Res. Treat. 45, 216–220. https://doi.org/10.1159/000521845 (2022).
Topham, J. T. et al. Circulating tumor DNA identifies diverse landscape of acquired resistance to anti-epidermal growth factor receptor therapy in metastatic colorectal cancer. J. Clin. Oncol. 41, 485–496. https://doi.org/10.1200/jco.22.00364 (2023).
Pinheiro, M. et al. KRAS and NRAS mutational analysis in plasma ctDNA from patients with metastatic colorectal cancer by real-time PCR and digital PCR. Int. J. Colorect. Dis. 37, 895–905. https://doi.org/10.1007/s00384-022-04126-6 (2022).
Normanno, N. et al. testing of liquid biopsy correlates with the outcome of metastatic colorectal cancer patients treated with first-line FOLFIRI plus cetuximab in the CAPRI-GOIM trial. Ann. Oncol. 29, 112–118. https://doi.org/10.1093/annonc/mdx417 (2018).
Misale, S. et al. Emergence of KRAS mutations and acquired resistance to anti-EGFR therapy in colorectal cancer. Nature 486, 532–536. https://doi.org/10.1038/nature11156 (2012).
Zarkavelis, G. et al. Exploring RAS mutation incidence and temporal heterogeneity in metastatic colorectal cancer patients—A single-institution experience utilising circulating tumour DNA. Współcz. Onkol. 28, 45–50. https://doi.org/10.5114/wo.2024.138899 (2024).
Anupriya, S., Chakraborty, A. & Patnaik, S. Clonal evolution and expansion associated with therapy resistance and relapse of colorectal cancer. Mut. Res. Rev. Mut. Res. 790, 108445. https://doi.org/10.1016/j.mrrev.2022.108445 (2022).
Sagawa, T. et al. Case report: Longitudinal monitoring of clonal evolution by circulating tumor DNA for resistance to anti-EGFR antibody in a case of metastatic colorectal cancer. Front. Oncol. 13, 1203296. https://doi.org/10.3389/fonc.2023.1203296 (2023).
Cheng, B. et al. Correlation between NGS panel-based mutation results and clinical information in colorectal cancer patients. Heliyon 10, e29299. https://doi.org/10.1016/j.heliyon.2024.e29299 (2024).
Lin, C. Y., Shen, M. Y., Chen, W. T. L. & Yang, C. A. Evaluation of the prognostic value of low-frequency KRAS mutation detection in circulating tumor DNA of patients with metastatic colorectal cancer. J. Pers. Med. 13, 1051. https://doi.org/10.3390/jpm13071051 (2023).
Ciardiello, D. et al. Anti-EGFR rechallenge in patients with refractory ctDNA RAS/BRAF wt metastatic colorectal cancer. JAMA Netw. Open 7, e245635. https://doi.org/10.1001/jamanetworkopen.2024.5635 (2024).
Martini, G. et al. Cetuximab as third-line rechallenge plus either irinotecan or avelumab is an effective treatment in metastatic colorectal cancer patients with baseline plasma RAS/BRAF wild-type circulating tumor DNA: Individual patient data pooled analysis of CRICKET and CAVE trials. Cancer Med. 12, 9392–9400. https://doi.org/10.1002/cam4.5699 (2023).
Ciardiello, D. et al. The role of anti-EGFR rechallenge in metastatic colorectal cancer, from available data to future developments: A systematic review. Cancer Treat. Rev. 124, 102683. https://doi.org/10.1016/j.ctrv.2024.102683 (2024).
Acknowledgements
We thank Dr. Li Bai, MD, First Department of Medical Oncology, Chinese People’s Liberation Army General Hospital, Beijing 100853, People’s Republic of China, for providing assistance and guidance.
Funding
This work received support from Sichuan Provincial Science and Technology Program Project (2024YFHZ0051) and Sichuan University’s Innovation Project from 0 to 1 (2022SCUH0032).
Author information
Authors and Affiliations
Contributions
Writing—original draft, conceptualization, data curation, Xueming Xia; formal analysis Wei Du; writing—review and editing Qiheng Gou.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Institutional review board statement
The research was carried out following the guidelines of the Declaration of Helsinki. All experimental protocols were approved by west China hospital’s review board.
Informed consent
Patient informed consent was obtained during this research.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xia, X., Du, W. & Gou, Q. Retrospective stratified analysis of resistance mechanisms to anti-EGFR therapy in mCRC using tumor tissue samples. Sci Rep 15, 19607 (2025). https://doi.org/10.1038/s41598-025-03684-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-03684-3
Keywords
This article is cited by
-
Exploratory biomarker analysis of RAS/BRAF somatic mutations and gene expression signatures for predicting treatment effects of aflibercept in the velour trial
npj Precision Oncology (2026)
-
Multiple drugs
Reactions Weekly (2025)