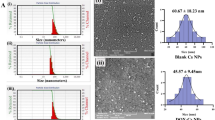
Abstract
Cisplatin is a commonly used chemotherapy drug that improves survival in many cancer patients but often causes severe neurotoxicity. Quercetin, a flavonoid antioxidant, has neuroprotective properties that can reduce cisplatin-induced neurotoxicity, but its clinical application is limited by poor bioavailability. This study aimed to combine nanomedicine and network pharmacology to develop Tween 80-functionalized chitosan-selenium nanoparticles containing quercetin (Q-Tw-Cs@Se NPs) and evaluate their protective effects against cisplatin-induced neurotoxicity in PC12 cells. Q-Tw-Cs@Se NPs were synthesized in a core–shell structure and characterized for size, morphology, optical properties, and chemical composition. Hemocompatibility, scavenging activity, and reducing power were assessed. The protective effects of the nanoparticles against cisplatin-induced toxicity in PC12 cells, used as a neuronal cell model, were evaluated using MTT assays and flow cytometry. Furthermore, network pharmacology was employed to predict hub targets and probable cellular mechanisms responsible for the protective effects of the nanoparticles. Q-Tw-Cs@Se NPs had an average size of 110 nm, a zeta potential of + 11.5 mV, and a quercetin encapsulation efficiency of ~ 50%. Pre-exposure to Q-Tw-Cs@Se NPs (1.25 µg/mL) for 24 h increased cell viability from 50% to nearly 80% in PC12 cells treated with cisplatin at its IC50 concentration. The nanoparticles also inhibited cisplatin-induced apoptosis and necrosis. Network pharmacology analysis identified BCL2, CASP3, AKT1, EGFR, STAT3, TNF, HSP90AA1, CTNNB1, IL-1β, and BCL2L1 as main targets enriched in apoptotic regulation. Altogether, Q-Tw-Cs@Se NPs show potential as neuroprotective adjuvants to reduce cisplatin-induced neurotoxicity. However, further in vitro and in vivo studies are required to confirm their therapeutic value.
Similar content being viewed by others
Introduction
Cisplatin is one of the most widely used chemotherapy drugs, which can dramatically improve the survival rate of cancer patients. Although cisplatin is very effective in cancer therapy, it can cause severe toxicities in high doses, such as nephrotoxicity, neurotoxicity, and ototoxicity1. The neurotoxicity caused by cisplatin is cumulative and dose-dependent, limiting the administered dose in chemotherapy. This highlights the importance of finding neuroprotective agents against this cisplatin side effect2.
One of the main mechanisms through which cisplatin exerts its cytotoxic effects involves the generation of reactive oxygen species (ROS) and the suppression of the cellular antioxidant defense system. This can lead to oxidative damage and consequent apoptotic cell death in brain and peripheral nervous system tissues3. Cisplatin enhances ROS generation, leading to mitochondrial dysfunction, lipid peroxidation, and DNA damage within neurons4. This redox imbalance triggers inflammatory responses and activates apoptotic pathways, leading to neuronal cell death and cognitive impairment. Additionally, cisplatin alters glutamate receptor activity, causing excitotoxicity and further mitochondrial damage5. These mechanisms collectively underlie the neurotoxic effects associated with cisplatin chemotherapy.
Selenium is considered an essential micronutrient, playing a crucial role in the pathophysiology of certain neurodegenerative diseases, including epilepsy, Parkinson’s disease, and Alzheimer’s disease6,7,8. The neuroprotective effects of selenium primarily result from its incorporation as selenocysteine in many enzymes involved in the redox system, protecting cell components against oxidative stress9. Besides regulating oxidative stress, selenium can modulate Ca2+ influx through ion channels and suppress apoptosis10. Notably, in addition to its neuroprotective properties, selenium can also exert antitumor activity against tumor cells11. Therefore, it can be a promising supplement for cancer patients who have received cisplatin for chemotherapy. Among different forms of selenium, Se0 has the lowest toxicity; however, its narrow therapeutic index limits its application in the clinic12,13. This highlights the significance of identifying a safe and effective form of selenium with minimal toxicity, which is essential for selenium-based adjuvant therapy. Recently, nanotechnology has drawn considerable interest due to the benefits of nanoparticles (NPs), which include their small size, high surface-to-volume ratio, straightforward chemical modification, enhanced bioavailability, and reduced toxicity compared to selenium-based compounds14. According to the available evidence, Se NPs can reduce the high-dose-associated toxicity of selenium and exhibit better biocompatibility than inorganic and organic selenium compounds15.
Despite the numerous benefits of using Se NPs, one of the most significant challenges is their high tendency to aggregate and form large particles due to their small size. Therefore, fabricating stable NPs is crucial, as the size of NPs can significantly influence their biological activities. Indeed, the enlargement of NPs can negatively influence their effectiveness in biological environments9,16,17. A wide variety of compounds have been suggested as stabilizers for Se NPs. In this regard, polysaccharides have received considerable attention due to their energy-saving and eco-friendly properties18,19.
Chitosan (Cs), a semisynthetic polysaccharide, is derived from chitin through alkaline deacetylation. The positive charge of chitosan increases the interactions with the negatively charged cell membrane, thereby enhancing cellular uptake20. Chitosan offers considerable potential as a natural polymer in developing drug delivery systems. Of note, several studies have demonstrated the neuroprotective properties of chitosan and its derivatives21,22,23.
Quercetin is a bioflavonoid that has been granted Generally Recognized As Safe (GRAS) status by the Food and Drug Administration (FDA). Quercetin is known for its numerous pharmacological attributes, including antioxidant, anti-inflammatory, potential anticancer, cardioprotective, and neuroprotective effects24,25. Its neuroprotective effects have been proven in several in vitro studies, in vivo animal studies, and clinical trials. Quercetin has the potential to protect organisms from neurotoxic compounds and prevent the progression of neuronal injury and neurodegeneration26. However, the clinical application of quercetin has been limited due to its low bioavailability, caused by poor solubility in water, extensive first-pass metabolism, and restricted permeability of the BBB27,28.
In the development of NPs targeting the brain, coating NPs with Tween 80 is a simple yet effective approach to improve their penetration into cerebral tissue. It has been demonstrated that Tween 80-coated NPs can adsorb Apoprotein E onto their surface. This modification allows them to interact with lipoprotein receptors present on the luminal side of the BBB. As a result, these functionalized NPs can undergo receptor-mediated transcytosis, facilitating their passage across the BBB and into the brain parenchyma29.
In this study, the PC12 cell line was selected as a neuronal model. PC12 cells are widely used in neuroscience research, including studies on cisplatin-induced neurotoxicity3,30,31,32. Derived from rat adrenal pheochromocytoma, PC12 cells have been extensively characterized for their neurogenic properties, such as neurosecretion (norepinephrine, catecholamines, and dopamine) and the expression of ion channels and neurotransmitter receptors33. In addition to their neurogenic properties, PC12 cells offer several practical advantages. These include their high versatility for pharmacological manipulation, ease of culture, a more uniform and reproducible population compared to primary sensory neurons, and extensive background knowledge on their proliferation and differentiation34. Together, these features enable detailed mechanistic investigations, including studies on cell survival and neuroprotective interventions. Therefore, although cisplatin primarily affects sensory neurons derived from the dorsal root ganglia, PC12 cells are among the most suitable models for studying cisplatin-induced neurotoxicity.
According to the unique properties of Se NPs, chitosan, and Tween 80, this study aims to develop a novel nanoformulation of quercetin based on Tween 80-functionalized chitosan Se NPs and to evaluate its protective effects against neurotoxicity induced by cisplatin in PC12 cells. As mentioned above, these compounds have protective properties, and their combination may have considerable neuroprotective effects. To the best of our knowledge, no study has yet evaluated the protective effects of this combination against cisplatin-induced toxicity in PC12 cells.
Materials and methods
Material
Sodium selenite pentahydrate (Na2SeO3⋅5H2O), ascorbic acid, quercetin, potassium ferricyanide K3[Fe(CN)6], 1,1-diphenyl-2-picrylhydrazyl (DPPH), trichloroacetic acid (TCA), and ferric chloride (FeCl3) were obtained from Merck Chemicals (Darmstadt, Germany). Medium molecular weight-chitosan (190–310 kDa, degree of deacetylation:75 ~ 85%) and MTT were provided by Sigma–Aldrich (St. Louis, MO, USA). Pen/Strep (100X), FBS, DMEM medium were purchased from Borna Pouyesh Gene Company (BPGene Co., Kerman, Iran).
Synthesis of chitosan-selenium nanoparticles (Cs@Se NPs)
Chitosan-selenium nanoparticles (Cs@Se NPs) were prepared due to our previous study with slight modifications35. Briefly, to synthesize Se NPs stabilized with chitosan, Na2SeO3 solution (0.2 mM) and ascorbic acid solution (1 mM) were added dropwise into 95 mL of chitosan solution prepared by dissolving 0.1 g of chitosan into 95 mL of acetic acid (0.01% w/w). Afterward, the resultant solution was stirred for 1 h and then incubated overnight at 4 °C. The synthesized NPs were ultimately collected by centrifugation (13841 g; 30 min) using a Thermo Fisher centrifuge equipped with an 8.6 cm radius fixed-angle rotor. Then, the NPs were washed twice with an acetic acid solution (0.01% w/w).
Preparation of tween 80-functionalized chitosan-selenium nanoparticles (Tw-Cs@Se NPs)
To cover the surface of synthesized NPs from the previous step with Tween 80, Cs@Se NPs were added dropwise into the acetic acid solution containing Tween 80 (2% w/w) under sonication for 10 min. The resultant solution was then stirred (200 rpm; 2 h). Finally, Tw-Cs@Se NPs were collected through centrifugation (13845 g; 30 min).
Preparation of quercetin-loaded tween 80 functionalized chitosan-selenium nanoparticles (Q-Tw-Cs@Se NPs)
Quercetin solution was prepared by dissolving quercetin (1 mg) in absolute ethanol (10 mL) to reach a concentration of 0.1 mg/mL. Afterward, the quercetin solution was added dropwise to Tw-Cs@Se NPs dispersed in an acetic acid solution (0.01% w/w), and the resultant mixture was stirred for 2 h at 200 rpm and 40 °C. After 2 h of stirring, the prepared NPs were collected by centrifugation (13845 g; 30 min) and washed twice with an acetic acid solution (0.01% w/w). The purified NPs were then used for further studies. Figure 1 provides a schematic summary of Q-Tw-Cs@Se NPs synthesis process.
Characterization of the NPs
Firstly, the optical properties of the nanostructures were investigated using UV-Vis spectrophotometry (UV-1800, Shimadzu CO, Japan) over a wavelength range of 220 to 800 nm. Subsequently, the hydrodynamic size and zeta potential of the nanostructures were measured using a dynamic light scattering analyzer (VASCOTM, Cordouan Technologies, France) and a zeta sizer (Nano-ZS ZEN 3600, Malvern Instruments, UK), respectively. After drying the nanostructures (40 °C; 24 h), their Fourier-transform infrared (FTIR) spectra were recorded using an FTIR spectrophotometer (Bruker Optics, MA, USA) in the 4000–500 cm-1 range. Elemental analysis was conducted using energy-dispersive X-ray spectroscopy (EDX). Finally, the morphology of the NPs was evaluated using High-resolution Transmission Electron Microscopy (HR-TEM) (Tecnai G2 20 TEM, FEI, USA) and Field Emission Scanning Electron Microscopy (FE-SEM) (ZEISS Sigma 300, Germany).
Determination of encapsulation efficiency and drug loading capacity
To calculate encapsulation efficiency (EE) and drug loading capacity (LC), the concentration of unloaded quercetin in the supernatant after centrifugation of the NPs was measured by UV-visible spectrophotometry using a standard curve. Briefly, a quercetin solution in absolute ethanol with a concentration of 10 µg/mL was prepared and scanned at wavelengths from 200 to 800 nm using a UV spectrophotometer. After determining the maximum absorption wavelength (λmax = 374 nm), standard solutions of quercetin in the concentration range of 1.25 to 20 µg/mL were prepared on different days, and their absorption was measured at 374 nm to plot a suitable calibration curve (R² = 0.9981). The concentration of quercetin in the test samples was then determined by interpolating the calibration curve. Finally, encapsulation efficiency and loading capacity were calculated as follows:
Evaluation of DPPH scavenging activity
The antioxidant activity of the prepared NPs was evaluated by the DPPH scavenging activity assay, according to Shakibaie et al36.. For this purpose, in a 96-well microplate, 50 µL of Tw-Cs@Se NPs, Q-Tw-Cs@Se NPs, and quercetin solution, with concentrations ranging from 5 to 320 µg/mL, were added separately. Each of these was mixed with 150 µL of freshly prepared 1 mM DPPH solution in methanol. After a 30-minute incubation of the microplate at 25 °C in a dark environment, the absorbance was measured at 517 nm. As the standard and control, quercetin solution and water were used, respectively. The scavenging effect was determined using the following equation:
Abs Sample is the absorbance of the sample, Abs Blank is the absorbance of the blank solution, and Abs Control is the absorbance measured for the control. All experiments were conducted in triplicate, and the data are presented as mean ± SD.
Evaluation of reducing power
For measuring the reducing power of the NPs, various concentrations of Tw-Cs@Se NPs, Q-Tw-Cs@Se NPs, and quercetin (10–1280 µg/mL) were prepared, and 500 µL of each concentration was separately added to 250 µL of potassium phosphate buffer (0.2 M, pH 6.6). Thereafter, 250 µL of potassium ferricyanide (1% w/v) was added to the reaction solution. Following 20 min incubation at 50 °C, the reaction solution was treated with 1 mL of TCA solution (10% w/v). After centrifugation (865 g; 10 min), 0.5 mL of deionized water and 0.1 mL of FeCl3 solution (6 mM) were added to 0.5 mL of the supernatant. Finally, the absorbance was determined at a wavelength of 700 nm36. All experiments were performed in triplicate, and the results are expressed as mean ± SD.
Hemolysis assay
Hemolysis assay was performed based on previous studies to assess the impact of NPs on red blood cell (RBC) integrity35,37. For the hemolysis assay, fresh blood from a healthy 28-year-old female donor was collected in K3EDTA tubes. RBCs were isolated through centrifugation (1541 g, 3 min, 4 C°) and washed three times using Alsever’s solution (NaCl 0.071 M, sodium citrate 0.027 M, citric acid 0.002 M, and dextrose 0.116 M, pH 6.4). Afterward, RBCs were diluted with Alsever’s solution and mixed with different concentrations of NP. The samples were centrifuged after incubation under controlled conditions (37 °C; 30 min) and mild shaking.
Following centrifugation of the samples (1541 g; 3 min), the supernatant was analyzed spectrophotometrically to quantify hemoglobin release at 415 nm. Alsever’s solution and deionized water were used as negative and positive controls, respectively. All experiments were conducted in duplicate. Results are reported as mean ± SD, and the degree of hemolysis was calculated as follows:
Abs Sample refers to the absorbance measured for the sample, Abs Negative control is the absorbance of the negative control, and Abs Positive control is the absorbance of the positive control.
Evaluation of cytocompatibility
PC12 cells were purchased from the Pasteur Institute of Iran (Tehran, Iran) and cultured in DMEM medium with 10% FBS under 95% humidity and 5% CO2 at 37 °C. For the MTT assay, cells were seeded at a density of 2 × 104 cells per well in a 96-well microplate and allowed to attach. Different concentrations of the NPs, quercetin, and cisplatin were also prepared in DMEM before being applied to cell cultures. After 24 h of incubation, cells were treated with Cs@Se NPs, Tw-Cs@Se NPs, Q-Tw-Cs@Se NPs, and quercetin at concentrations ranging from 0.16 to 40 µg/mL. The next day, each sample was treated with 5 µl of MTT dye (5 mg/mL) and incubated at 37 °C for 4 h. After removing the media, the purple formazan crystal sediment was dissolved in 100 µL of DMSO. Finally, the absorbance of each sample was measured at a wavelength of 570 nm. Cell viability percentage was calculated as follows:
Abs Treated refers to the absorbance measured for the treated cells, Abs Blank is the absorbance of the blank, and Abs Untreated is the absorbance of untreated cells (control group).
In the next step, an MTT assay was performed to evaluate the cytotoxicity of cisplatin on PC12 cells within a concentration range of 5 to 640 µg/mL, and to determine the IC50 concentration. After identifying the IC50 concentration (80 µg/mL), we assessed the protective effects of Q-Tw-Cs@Se NPs against cisplatin-induced toxicity in the PC12 cell line. Cells were first pretreated with the NPs (0.16–20 µg/mL) for 24 h, followed by exposure to cisplatin at the IC50 concentration. After 24 h of incubation, the percentage of viable cells was then calculated. All experiments were conducted in triplicate, and results are expressed as mean ± SD.
Flow cytometry
PC12 cells were pretreated with Q-Tw-Cs@Se NPs (1.25 µg/mL) 24 h prior to the addition of cisplatin at its IC50 concentration. After 24 h, cells were rinsed with cold PBS, detached using trypsin, and resuspended in Annexin binding buffer to a concentration of 1 × 106 cells/mL. The suspension was then mixed with 5 µL of Annexin V-FITC with Propidium Iodide (PI) according to the manufacturer’s protocol (Miltenyi Biotec, Germany). Following a 15-minute dark incubation at room temperature, the samples were analyzed using a flow cytometer (BD-Biosciences, USA) and FlowJo v7.6.1 software. Cell death was assessed by measuring the rates of apoptosis and necrosis across a population of 10,000 events.
Network pharmacology analysis
Prediction of potential targets
To identify genes related to quercetin, the SMILES structure of quercetin from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/) was input into the SwissTargetPrediction database (http://www.swisstargetprediction.ch/). Moreover, the keyword “quercetin” was searched in the Comparative Toxicogenomics Database (CTD) (http://ctdbase.org/) and the GeneCards database (https://www.genecards.org/). Finally, duplicates were removed to provide a clean dataset. The same process was applied for the keywords “cisplatin” and “selenium nanoparticle” in the CTD and GeneCards databases. To identify genes associated with peripheral neuropathy, this keyword was searched in the GeneCards database. Finally, intersection targets were identified using a Venn diagram (http://bioinfogp.cnb.csic.es/tools/venny/index.html)38.
Protein-protein interaction (PPI) network and topological analysis
A protein-protein interaction (PPI) network illustrates the direct connections among input proteins, forming a network in which proteins are depicted as nodes and their interactions are shown as edges linking these nodes. The PPI network was constructed by inputting overlapped genes in the STRING database (https://string-db.org/). The parameters were set to a high level of confidence (0.7) and “Homo sapiens” to construct the PPI network. Finally, the constructed PPI network was imported into Cytoscape 3.10.3 for visualization. Next, the cytoHubba plugin in Cytoscape was used to score each node based on topological metrics. Nodes were then ranked according to their degree (k). The parameter k indicates how many links a node has with other nodes within the network. Nodes with a high k value play a crucial role in the transmission of information, as they exert a greater influence over the flow of information throughout the network. Ultimately, to identify important gene clusters within the molecular interaction network, the PPI network was clustered using Molecular Complex Detection (MCODE) analysis, which is a plugin of Cytoscape39.
Functional enrichment analysis
Gene Ontology (GO) and Reactome pathway enrichment analyses of overlapping target genes were performed using the R package “enrichR” to explore the molecular functions (MF), cellular components (CC), biological processes (BP), and potential signaling pathways associated with these target genes40.
Statistical analysis
Statistical analysis of the DPPH scavenging effect, reducing power, hemolysis assay, and MTT assay results obtained from cells pretreated with NPs was conducted using two-way ANOVA followed by Tukey’s correction for multiple comparisons. For cytocompatibility analysis, one-way ANOVA followed by Dunnett’s correction was used to compare groups. All statistical analyses were performed using GraphPad Prism 10 software (GraphPad Inc., San Diego, CA, USA).
Results and discussion
Preparation & characterization of the NPs
To ensure the synthesis of Se NPs, the UV-Vis absorption spectrum of the Cs@Se NPs was evaluated. As shown in Fig. 2A, the UV-Vis spectrum of the Cs@Se NPs exhibits a clear peak in the 200–400 nm range, indicating the successful synthesis of Se NPs41. Moreover, the appearance of a single peak in the UV-Vis absorption spectrum of the NPs indicates that they are likely spherical42. When comparing the UV-Vis spectra of Cs@Se NPs, Tw-Cs@Se NPs, and Q-Tw-Cs@Se NPs, a slight red shift is observed in the peak of Q-Tw-Cs@Se NPs, indicating an increase in particle size. This can be explained by the fact that larger particles have a smaller band gap energy, which leads to absorption at longer wavelengths43. According to Fig. 2B, the orange hue of the NPs resulted from their surface plasmon resonance (SPR) excitation, suggesting the presence of Se NPs44,45.
Firstly, Cs@Se NPs were synthesized with a hydrodynamic size of 91.3 nm and a zeta potential of 26.1 mV. To enhance the encapsulation efficiency of the NPs for loading quercetin, Cs@Se NPs were functionalized with Tween 80, resulting in NPs with a size of 105 nm and a zeta potential of 14 mV. Ultimately, Tw-Cs@Se NPs were loaded with quercetin, achieving an encapsulation efficiency of almost 50% and a loading capacity of 4%. As presented in Table 1, the size of the NPs increased from 91.3 nm to 110.6 nm, while the zeta potential decreased from +26 mV to +11.5 mV, confirming the successful functionalization of Cs@Se NPs and the effective encapsulation of quercetin.
According to HR-TEM and FE-SEM images, Q-Tw-Cs@Se NPs were spherical, with a size of around 70 nm (Fig. 3A–D). To examine the elemental composition of Q-Tw-Cs@Se NPs, an EDX assay was carried out. According to the obtained results, in Q-Tw-Cs@Se NPs, the atomic percentages were found to be 45% for selenium, 27.7% for carbon, 24.2% for oxygen, and 3% for nitrogen (Fig. 3E).
FTIR analysis was conducted to investigate surface modifications and conjugation (Fig. 4). According to the FTIR spectrum of chitosan, the broad peak around 3400 cm−1 is attributed to the O–H and N–H stretching vibrations of chitosan. The peak at 1653 cm−1 is ascribed to the stretching vibration mode of amide I. The peak at 1376 cm−1 corresponds to the COO⁻ group, whereas the peak at 1077 cm−1 is associated with the stretching vibration of the C–O–C bond in the glucose ring structure46,47. Cs@Se NPs’ spectrum shows characteristic peaks of chitosan functional groups, including a broad band at 3425 cm−1 from the O–H and N–H groups, asymmetric C–H stretch at 2920, a symmetric C–H stretch at 2853 cm−1, the − NH₂ scissoring vibration at 1551 cm−1, and a C=O stretch at 1064 cm−1. Furthermore, the 1623 cm−1 and 566 cm−1 peaks are ascribed to the stretching and bending vibrations of Se–O bonds, respectively48. With respect to the FTIR spectrum of Tween 80, the characteristic peaks at 1109 cm−1 and 1734 cm−1 are assigned to the asymmetric C–O and C=O stretching vibrations, respectively49. The absorption bands at 1105 cm−1 and 1735 cm−1 in the Tw-Cs@Se NPs’ spectrum confirm the coverage of the NPs with Tween 80. In the spectrum of quercetin, the peak at 1382 cm−1 corresponds to the in-plane bending vibration of the phenolic O–H group, while the peak at 1663 cm−1 is associated with the C=O stretching vibration of the aryl ketonic group50,51,52. The spectrum of Q-Tw-Cs@Se NPs, compared to Tw-Cs@Se NPs, shows a slight red shift in the band corresponding to the O–H group from 3417 to 3415, indicating an interaction between the NPs and the O–H group of quercetin53. Moreover, in the Q-Tw-Cs@Se NPs spectrum, the red shift in the C=O peak from 1663 cm−1 to 1636 cm−1 confirms the presence of quercetin. The peak at 1559 cm−1 is attributed to the aromatic group of quercetin in the spectrum of the NPs54.
Antioxidant activity of the NPs
To investigate the antioxidant activity of the prepared NPs, DPPH and reducing power assays were performed. Firstly, different concentrations of Tw-Cs@Se NPs, Q-Tw-Cs@Se NPs, and quercetin, ranging from 5 to 320 µg/mL, were compared for their ability to inhibit the DPPH radical. According to Fig. 5A, free quercetin at a concentration of 5 µg/mL exhibited around 70% scavenging activity. At concentrations ranging from 5 µg/mL to 320 µg/mL, it showed more than 80% DPPH scavenging activity. Of note, there is no significant difference in the inhibition of DPPH over the concentration range of 10 to 320 µg/mL (P > 0.05). Therefore, the DPPH scavenging effect of quercetin was not dose-dependent in this concentration range. In the current study, Tw-Cs@Se NPs demonstrated a scavenging effect of approximately 30% at the lowest concentration. In contrast, a previous study reported that Se NPs did not improve antioxidant parameters measured by DPPH assays55. This might have resulted from the antioxidant effect of chitosan itself56. Moreover, the presence of stabilizers can enhance the antioxidant activity of NPs by improving their solubility and stability in the aqueous environment, which in turn increases the surface area of the NPs exposed to free radicals57.
(A) Free radical scavenging capacities of Tw-Cs@Se NPs, Q-Tw-Cs@Se NPs, and quercetin. (B) Reducing power ability of Tw-Cs@Se NPs, Q-Tw-Cs@Se NPs, and quercetin. Each value is represented as mean ± SD, n = 3. A two-way ANOVA was used for statistical comparison among groups, followed by Tukey’s post-test and statistical significance is denoted as follows: *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001.
The free radical scavenging activity of Tw-Cs@Se NPs was considerably lower than that of free quercetin and reached 50% at the highest concentration. At concentrations less than 320 µg/mL, there is no significant difference between Tw-Cs@Se NPs and Q-Tw-Cs@Se NP. However, at the concentration of 320 µg/mL, there is a significant difference between quercetin-free NPs and quercetin-loaded NPs (P < 0.0001). At 320 µg/mL concentration, Q-Tw-Cs@Se NPs displayed around 80% scavenging effect, the same as free quercetin (P > 0.05). According to the results, quercetin loaded into the NPs did not demonstrate antioxidant activity at lower concentrations, leading to similar results to quercetin-free NPs. This result is consistent with a previous study in which iron oxide NPs containing quercetin exhibited no significant antioxidant activity at concentrations below 100 µg/mL; however, a sudden increase in scavenging activity was observed at 1000 µg/mL58. This might have resulted from insufficient time for the release of quercetin from the NPs, and we predict that the antioxidant activity of the NPs will increase over time due to quercetin release58. It is noteworthy that drug release is not solely a function of time but also of NP concentration. At high concentration (320 µg/mL), the collective surface area of the NPs in the assay medium is vastly greater. This increases the probability of interactions and the total number of quercetin molecules beginning to diffuse out simultaneously, leading to a more pronounced initial burst release. This initial rapid release partly results from unattached or loosely bound drug molecules on NP surfaces, which increase at higher NP concentrations and are quickly released, especially in DPPH medium containing ethanol, the quercetin solvent. At lower concentrations (80 and 160 µg/mL), this collective effect is reduced, and the released quercetin may be below the critical concentration threshold of the assay, explaining why their results are similar to each other and to the quercetin-free NPs59. Moreover, NPs are complex systems, and at certain concentrations and under specific medium conditions, NPs can undergo conformational and surface chemistry changes that alter their total surface area, release profiles, accessibility of redox-active sites, and ultimately the reaction kinetics. For instance, a recent study reported that DPPH can adsorb onto the surface of NPs, hindering drug-free diffusion and reducing the antioxidant activity of NPs within the assay timeframe60. All these factors collectively can explain the nonlinear correlation observed between the concentration of NPs and their scavenging effect.
As shown in Fig. 5A, at the highest concentration, not only did released quercetin show considerable scavenging activity, but its effect might also be intensified in the presence of the NPs since quercetin-free NPs indicated around 50% scavenging effect at a similar concentration. In fact, Tw-Cs@Se NPs and quercetin showed a synergistic effect at this concentration. Consistent with the previous study regarding curcumin-loaded Se NPs, it has been demonstrated that the effect of DPPH free radical scavenging between Se NPs and curcumin in the Se NPs@Cur was strongly synergistic61.
Alongside the DPPH assay, to evaluate their reducing power, the NPs and free quercetin were compared at concentrations ranging from 10 to 1280 µg/mL based on their ability to reduce Fe3+ ions to Fe2+. According to Fig. 5B, at all concentrations, the reducing power ability of free quercetin is considerably higher than that of the NPs (P < 0.001). Within the concentration range of 10 to 640 µg/mL, there is no considerable difference among Cs@Se NPs, Tw-Cs@Se NPs, and Q-Tw-Cs@Se NPs. However, at the highest concentration (1280 µg/mL), Q-Tw-Cs@Se NPs exhibited greater reducing power compared to quercetin-free NPs. In fact, at this concentration, the quercetin released from the NPs was sufficient to cause a significant difference between quercetin-free NPs and NPs containing quercetin.
Hemocompatibility of the NPs
When NPs are exposed to RBCs, they may cause rupture and release hemoglobin into the bloodstream. There is no standardized animal study to evaluate nanoparticle-induced hemolysis. Therefore, hemocompatibility tests should be considered a part of toxicity testing62. In this study, we examined the hemolytic activity of Cs@Se NPs, Tw-Cs@Se NPs, and Q-Tw-Cs@Se NPs based on the Standard Practice for the Assessment of Hemolytic Properties of Materials (ASTM F 756-08), which defines compounds as non-hemolytic if they exhibit a hemolysis rate below 2%63,64,65. As shown in Fig. 6A and B, the hemolysis percentage of Q-Tw-Cs@Se NPs remained below 2% at all tested concentrations. In contrast, Tw-Cs@Se NPs at the highest concentration (640 µg/mL) exhibited nearly 2% hemolysis, while Cs@Se NPs at the same concentration caused RBC lysis exceeding 2%. It is noteworthy that Q-Tw-Cs@Se NPs demonstrated significantly lower hemolysis than quercetin-free NPs at concentrations above 160 µg/mL, with the most significant difference observed at 640 µg/mL (P < 0.0001).
(A) Comparison of hemolytic activity among Cs@Se NPs, Tw-Cs@Se NPs, Q-Tw-Cs@Se NPs. (B) Visual inspection of tubes containing the NPs post-centrifugation. Each value is represented as mean ± SD, n = 2. A two-way ANOVA was used for statistical comparison among groups, followed by Tukey’s post-test and statistical significance is denoted as follows: *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001.
Based on these results, Q-Tw-Cs@Se NPs are non-hemolytic within the concentration range of 5 to 640 µg/mL. The lower surface positive charge of Q-Tw-Cs@Se NPs compared to the other formulations may explain its reduced hemolytic activity. In contrast, Cs@Se NPs, which caused the highest level of hemolysis, had the highest surface positive charge, suggesting that these NPs have a stronger tendency to interact with negatively charged cell membranes of RBCs66.
Cytocompatibility of the NPs
Prior to evaluating the protective effects of the NPs on PC12 cells, it was necessary to assess their cytocompatibility. The cellular toxicity of Cs@Se NPs, Tw-Cs@Se NPs, Q-Tw-Cs@Se NPs, and free quercetin was first evaluated within the concentration range of 0.16 to 40 µg/mL after 24 h in the PC12 cell line. According to previous studies, Se NPs exhibit protective effects at low concentrations, whereas at higher concentrations they can induce cytotoxicity67,68,69,70. Since the goal of the current study was to evaluate the potential protective effects of NPs, it was essential to select a concentration range in which NPs would not induce toxicity, maintaining cell viability above 90%. Based on previous studies, we chose concentrations below 40 µg/mL67,68,69. Moreover, because a similar study reported the highest protective effect of quercetin-loaded NPs at 1 µg/mL, we decided to set the lowest concentration in our experiments at 0.16 µg/mL58.
According to Fig. 7A, quercetin did not exhibit toxicity in PC12 cells, with cell viability remaining above 90% across the entire concentration range, and no significant differences were observed between the various concentrations and the control group (P > 0.05).
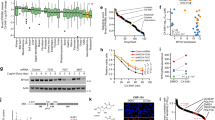
Cell viability of PC12 cells treated with quercetin (A), Cs@Se NPs (B), Tw-Cs@Se NPs (C), and Q-Tw-Cs@Se NPs (D), cisplatin (E) after 24 h. Each value is represented as mean ± SD, n = 3. A one-way ANOVA was used for statistical comparison among groups, followed by Dunnett’s’ post-test, and statistical significance is denoted as follows: *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001.
As shown in Fig. 7B and C, Cs@Se NPs and Tw-Cs@Se NPs exerted toxicity on PC12 cells at the concentration of 40 µg/mL, and cell viability was significantly less than that of the control group (P < 0.05 and P < 0.01, respectively). However, quercetin-loaded NPs at concentrations above 20 µg/mL demonstrated significant toxicity compared to the control group (P < 0.001) (Fig. 7D). Previous studies have shown that conjugating quercetin to other compounds through its functional groups reduces its antioxidant capacity. This reduction occurs because the number of available hydroxyl groups, which are responsible for its antiradical activity, is diminished71. Moreover, some studies have reported controversial results regarding the neuroprotective effects of quercetin. In fact, depending on the conditions under which cells are grown and their differentiation status, cells may exhibit different behaviors when exposed to quercetin72.
Additionally, the pro-oxidative activity of quercetin can be explained by its ortho-dihydroxy structure in the B ring and its vulnerability to autooxidation or enzymatic conversion into pro-oxidant intermediates under specific conditions73. Along with pro-oxidant activity, accumulating evidence has shown that flavonoids such as quercetin can induce cell death by changing the expression of heat shock proteins (HSP) and proteins associated with the MAPK pathway in undifferentiated PC12 cells72. Moreover, at the concentration of 20 µg/mL, the presence of Se NPs and their potential pro-oxidant effect might have triggered quercetin to activate the cell death pathways74.
This may explain why Q-Tw-Cs@Se NPs showed greater toxicity at a concentration of 20 µg/mL compared to the quercetin-free NPs, highlighting the importance of further research on the potential toxic effects of flavonoids and the appropriate concentration ranges for favorable effects in various conditions. Figure 7E shows the cytotoxicity of cisplatin on PC12 cells within a concentration range of 5 to 640 µg/mL. At 80 µg/mL, cisplatin induced approximately 50% toxicity, which was selected as the IC50 for subsequent experiments.
Overall, based on the MTT results, the selected concentration range was from 0.16 to 20 µg/mL to investigate the protective effects of the NPs on cisplatin-induced neurotoxicity.
Cytoprotective effect of Q-Tw-Cs@Se NPs
In this study, we investigated the protective effects of Q-Tw-Cs@Se NPs (0.16–20 µg/mL) and quercetin at an equivalent concentration, calculated based on the 4% loading capacity of the NPs, on PC12 cells 24 h before cisplatin (80 µg/mL) treatment (Fig. 8). According to the results, pretreatment with free quercetin and Q-Tw-Cs@Se NPs effectively inhibited cisplatin toxicity. A recent study demonstrated that, compared to cells treated with H2O2 alone, pretreatment with quercetin significantly enhanced cell viability and lactate dehydrogenase (LDH) release in a dose-dependent manner. Additionally, quercetin pretreatment significantly mitigated oxidative stress in PC12 cells exposed to H2O2, as evidenced by reduced intracellular ROS levels, malondialdehyde (MDA) production, and cell membrane lipoperoxidation. Ultimately, quercetin pretreatment reduced cell apoptosis in an H2O2-induced oxidative stress model75. In a similar study, the protective effects of quercetin on toxicity induced by H2O2 in a mouse colitis model were explored76. Due to their findings, superoxide dismutase (SOD) and glutathione peroxidase (GSH-Px) are among the endogenous antioxidant enzymes whose levels can be enhanced by quercetin. This increase in antioxidant enzymes facilitates the conversion of superoxide radicals into less harmful molecules, which reduces oxidative stress and enhances cellular antioxidant capacity77. As shown in Fig. 8, from the 0.16 to 0.62 µg/mL concentration, there is no significant difference between free quercetin and Q-Tw-Cs@Se NPs (P > 0.05). Unlike Yarjani et al.’s study, in which quercetin-loaded iron oxide NPs showed less protective effect than free quercetin, Q-Tw-Cs@Se NPs exhibited a protective effect comparable to free quercetin58. This result might have resulted from the synergistic antioxidant activity of Se NPs at low concentrations combined with quercetin. In agreement with the protective effect of Se NPs at low concentrations, Turovsky et al. revealed that Se NPs, only in concentrations below 0.5 µg/mL, are able to alleviate neurotoxicity induced by oxygen-glucose deprivation and reoxygenation on primary cortical neurons and astrocytes. While low doses of Se NPs activate the anti-apoptotic IRE1-XBP1 pathway and potentially the Nrf-2/HO-1 antioxidant signaling pathway, higher concentrations trigger necrosis and apoptosis70.
Cell viability of PC12 cells pre-exposed to Q-Tw-Cs@Se NPs and quercetin for 24 h before addition of cisplatin (80 µg/mL). Each value is represented as mean ± SD, n = 3. A two-way ANOVA was used for statistical comparison among groups, followed by Tukey’s post-test, and statistical significance compared to the model group (untreated + cisplatin) is denoted as follows: *P < 0.05, &P < 0.01, $P < 0.001, and #P < 0.0001. Statistical significance between two groups at the same concentration is indicated as follows: ^P < 0.05, ^^P < 0.01, ^^^P < 0.001, and ^^^^P < 0.0001.
Moreover, positively charged NPs can enhance the penetration of quercetin into cells. For instance, Ferreira et al. developed chitosan/sulfonyl-ether-β-cyclodextrin (SBE-β-CD) as a delivery system for quercetin. They demonstrated that positively charged NPs enhanced the antitumor activity of quercetin against ovarian cancer by improving the cellular uptake of the drug78. In our study, we used chitosan not only as a stabilizer but also to enhance the interaction of NPs with the negatively charged cell membrane through its natural positive charge.
At concentrations up to 1.25 µg/mL, a significant difference is observed between Q-Tw-Cs@Se NPs and the model group (cisplatin with no pretreatment). However, at higher concentrations, the protective effect of the NPs is significantly reduced compared to free quercetin. This might have resulted from the pro-oxidant effect of Se NPs at higher concentrations in the presence of cisplatin. As previously mentioned, Se NPs can exhibit both antioxidant and pro-oxidant activities, depending on their dose, duration, frequency of exposure, and oxidation state, which is supported by several studies9,74,79. In this case, although quercetin-free NPs at concentrations up to 40 µg/mL did not show toxicity, cisplatin might have stimulated the pro-oxidant activity of Se NPs even at lower concentrations, reducing the protective effects of quercetin released from Q-Tw-Cs@Se NPs. Overall, 24-h quercetin pretreatment at all concentrations provided protective effects and attenuated cisplatin toxicity; however, Q-Tw-Cs@Se NPs showed the greatest protection at 1.25 µg/mL.
Flow cytometry analysis of cytoprotection induced by Q-Tw-Cs@Se NPs
According to flow cytometry results, 24-h pre-exposure to Q-Tw-Cs@Se NPs before cisplatin treatment at its IC50 concentration inhibits both apoptosis and necrosis (Fig. 9A-D). As shown in Fig. 9B, cisplatin led to 39.11% necrosis and 5.62% apoptosis in PC12 cells. Meanwhile, in the sample in which only Q-Tw-Cs@Se NPs were added, 0.2% necrosis and 0.55% apoptosis were observed (Fig. 9C). According to Figs. 9D and 24-h pretreatment with Q-Tw-Cs@Se NPs reduced necrosis percentage from 39.11% to 8.45% and apoptosis percentage from 5.62% to 0.29%, confirming the inhibitory effect of Q-Tw-Cs@Se NPs pretreatment on cisplatin-induced toxicity in PC12 cells.
Several mechanistic insights can be proposed to explain how pretreatment with quercetin and Se NPs can effectively inhibit cell death. As mentioned, Se NPs can upregulate SOD/GSH-Px, creating a redox buffer against cisplatin-induced ROS80. Moreover, in another study, it has been revealed that Se NPs can modulate Bcl-2/Bax ratio and prevent cytochrome c release during cisplatin exposure81. It has also been reported that Se NPs can inhibit inflammatory cytokines like TNF-α and IL-6, thereby reducing secondary oxidative stress and mitigating both apoptosis and necrosis82. For quercetin, several studies have confirmed its anti-apoptotic and anti-inflammatory effects. For instance, Bournival et al. indicated that 3 h pretreatment with quercetin (0.1 µM) prior to exposure to 1-methyl 4-phenylpyridinium (MPP) suppressed the expression of inflammatory cytokines, including IL-6, TNF-α, and IL-1β83. Nichols et al. demonstrated that quercetin, epicatechin, or both mitigated apoptosis induced by glucose and oxygen deprivation in mouse cortical neurons and preserved mitochondrial function, while elevating mitochondrial gene expression and pro-survival signaling84.
Mechanistic insights predicted by network pharmacology
To identify potential targets responsible for the protective effects of Q-Tw-Cs@Se NPs, network pharmacology was employed. Firstly, genes targeted by quercetin and Se NPs were retrieved from the GeneCards database, CTD, and SwissTargetPrediction. After removing duplicates, 325 and 519 target genes were identified for quercetin and Se NPs, respectively. Then, genes associated with cisplatin toxicity and peripheral neuropathy, a common side effect of cisplatin, were collected from the GeneCards database and CTD. Duplicate removal resulted in a final set of 590 genes for cisplatin and 9407 genes for peripheral neuropathy.
Using a Venn diagram analysis, 73 overlapping genes were selected as intersection targets for further analysis (Fig. 10A). To better understand the interactions among these targets, a PPI network was constructed using the STRING database and visualized with Cytoscape. As illustrated in Fig. 10B, the resulting PPI network comprises 73 nodes and 481 edges. A topological network algorithm was employed to quantify each node by assigning a score and subsequently ranking the nodes based on their degree (k) using the cytoHubba plugin of Cytoscape. In the PPI network, darker colors represent higher scores, and red nodes indicate greater significance, suggesting their likely key roles within the network. Afterward, the top 10 hub nodes (targets) were identified with the highest degree of connectivity (Fig. 10C). The top 10 hub targets include B-cell lymphoma 2 (BCL2), Caspase 3 (CASP3), Serine/threonine kinase 1 (AKT1), Epidermal growth factor receptor (EGFR), Signal transducer and activator of transcription 3 (STAT3), Tumor necrosis factor (TNF), Heat Shock Protein 90 Alpha Family Class A Member 1 (HSP90AA1), Catenin beta 1 (CTNNB1), Interleukin 1 beta (IL-1β), and BCL2-like 1 (BCL2L1).
(A) Venn diagram showing overlapping genes. (B) Protein-protein interaction (PPI) network analysis of the 73 overlapping genes; darker colors indicate greater significance within network. (C) Identification of the 10 hub targets within the PPI network. (D) Four clusters identified from the PPI network via MCODE analysis.
The PPI network was clustered into four modules using the MCODE plugin of Cytoscape (Fig. 10D). Each cluster was analyzed individually to identify enriched biological processes using GO-BP terms. According to Table 2, Cluster 1 shows significant enrichment in the regulation of the apoptotic process; Cluster 2 is highly enriched in the glutathione metabolic process; Cluster 3 mainly participates in mitotic phase transition and the G2/M transition of mitotic cells; and Cluster 4 is mostly involved in the cellular response to unfolded proteins.
Notably, all top 10 hub targets are located within Cluster 1, suggesting a pivotal role for this cluster in the therapeutic effects of Q-Tw-Cs@Se NPs. GO and Reactome pathway analyses were then performed on Cluster 1 to further investigate the enrichment of these targets (Fig. 11). The results show that the 22 nodes in Cluster 1 are mainly related to the regulation of the apoptotic process (BP), intracellular membrane-bound organelles (CC), and kinase binding (MF). Pathway enrichment analysis indicates that the nodes in Cluster 1 are associated with intrinsic apoptosis pathways, programmed cell death, and signaling by interleukins.
According to network pharmacology, the major targets are BCL2, CASP3, and AKT1. In other words, these targets have a high potential to explain the protective effects of Q-Tw-Cs@Se NPs against cisplatin-induced neurotoxicity.
Apoptosis proceeds via two principal mechanisms: the extrinsic pathway, also known as the death receptor or caspase pathway, and the intrinsic pathway, commonly referred to as the mitochondrial or Bcl-2-regulated pathway, which is controlled by the interplay between proapoptotic and anti-apoptotic factors. Bcl-2 is one of the most important anti-apoptotic proteins in the Bcl-2 family, primarily responsible for maintaining mitochondrial integrity and preventing cell death. In contrast, CASP3 plays a central role in the extrinsic apoptotic pathway. Several studies have indicated that quercetin protects neuronal cells from toxic insults via the upregulation of Bcl-2 and downregulation of CASP3 expression85,86. For instance, it has been shown that quercetin alleviates neuronal death induced by aluminum by upregulating Bcl-2, downregulating Bax and p53, inactivating caspase-3, and reducing DNA fragmentation87. In addition to experimental studies, in silico studies have confirmed a strong binding affinity between quercetin and BCL-288,89. Besides quercetin, several studies have suggested a protective effect of Se NPs through the Bax/Bcl-2-mediated apoptotic pathway. For instance, in diabetic rats, chitosan-coated Se NPs combined with metformin significantly increased Bcl-2 expression90. Moreover, in neurotoxicity caused by oxygen-glucose deprivation, low doses of Se NPs exert anti-apoptotic effects through multiple mechanisms. These include upregulating Bcl-2 expression and suppressing mitochondrial and endoplasmic reticulum (ER) stress pathways, which ultimately lead to caspase-3 inactivation70. Therefore, the primary predicted mechanism for the neuroprotective effects of Q-Tw-Cs@Se NPs is the regulation of the apoptotic process, which is supported by the flow cytometric analysis of cell death. As mentioned, 24-h pretreatment with these NPs resulted in a considerable decrease in the percentage of apoptotic cells. These findings suggest that Q-Tw-Cs@Se NPs may offer a promising prophylactic strategy by modulating apoptotic pathways to protect neuronal cells from cisplatin-induced toxicity.
AKT1 plays a pivotal role in mediating survival signaling in a variety of neuronal cell types. The PI3K-AKT-mTOR pathway promotes neural repair by regulating cell survival and inhibiting oxidative stress and inflammation. Activation of AKT reduces free radical levels by upregulating antioxidant enzymes such as SOD, while concurrently suppressing the release of inflammatory cytokines, including TNF-α and IL-691. In this regard, an in vitro study has demonstrated that quercetin exerts its neuroprotective effects via the PI3K/Akt and ERK1/2 signaling pathways in copper-induced toxicity in P19 neuronal cells92. Consistent with these findings, the PI3K-AKT and EGFR/MAPK pathways have been proposed as major mechanisms underlying quercetin therapy in spinal cord injury93. However, researchers have emphasized the need for further investigations to elucidate the precise mechanisms by which quercetin contributes to neuroprotection through the PI3K and EGFR/MAPK pathways.
In addition to quercetin, a recent study has shown that Se NPs coated with lentinan attenuate neuronal apoptosis by upregulating the PI3K-AKT-mTOR and Ras-Raf-MEK-ERK signaling cascades, while modulating the expression of associated antioxidant and anti-inflammatory proteins94.
EGFR is one of the first discovered receptor tyrosine kinases, triggering signaling pathways that play crucial roles in neuronal proliferation, differentiation, and migration, including the Ras-Raf-MEK-ERK1/2, STAT3, and PI3K-Akt-mTOR pathways95. Although EGFR activates neurotrophic signaling pathways, its inhibition after injury can help reduce astrogliosis, suppress the release of proinflammatory factors, and protect neurons from cell death. The inhibitory effects of quercetin on EGFR have been well documented in various cancers; however, there are only a limited number of studies exploring the role of EGFR in quercetin’s neuroprotective effects96. Given the protective benefits observed from EGFR inhibition following quercetin treatment in nephropathy, it is plausible that quercetin alleviates neurotoxicity through EGFR inhibition, thereby reducing inflammatory responses, oxidative stress, and apoptosis97,98. Furthermore, molecular docking analysis has revealed that quercetin attaches to EGFR with a strong binding affinity via hydrogen bonds, suggesting that quercetin may influence EGFR through both direct and indirect mechanisms99.
HSPs are highly conserved chaperone proteins that play a vital role in cell survival. They protect cells under stress conditions, such as hypoxia, oxidative stress, and exposure to toxic agents, by activating various cytoprotective mechanisms. Among these, HSP90AA1, a member of the HSP90 family, is one of the most important HSPs100. Molecular docking analysis has indicated that quercetin can directly bind to HSP90AA1, and an in vitro study has confirmed the inhibitory effects of quercetin on HSP90 activity in breast cancer cells93,101. Nevertheless, there is insufficient evidence to elucidate how the interaction between quercetin and HSPs contributes to neuroprotection against toxic agents. In contrast, Se NPs have been shown to upregulate HSP90 expression, thereby protecting cells under stress conditions102.
STAT3 plays a multifaceted role in regulating cell proliferation, survival, death, and inflammatory responses. Because of its intricate function, STAT3 acts as a double-edged sword in neurotoxicity and brain injury. It can have both protective and harmful effects depending on the cell type and cellular environment. For example, activation of STAT3 has been demonstrated to exert a neuroprotective effect on dopaminergic neurons and axonal survival by reducing oxidative stress and cell death103. In contrast, STAT3 inhibition has been shown to significantly reduce apoptosis in SH-SY5Y cells exposed to hydrogen peroxide, thus protecting the cells from oxidative stress104. Several studies have reported the inhibitory effects of quercetin on STAT3 in both cancer and neurodegenerative models105,106,107. Due to the key role of STAT3 in inflammatory responses, STAT3 pathway inhibition may partially explain the anti-inflammatory mechanisms of quercetin108. Furthermore, suppressing the STAT3 pathway has been suggested as one of the anti-inflammatory mechanisms of Se NPs. For instance, a recent study demonstrated that the co-incubation of chitosan-coated Se NPs with bone marrow mesenchymal stem cells resulted in the downregulation of Th17 expression via the inhibition of the RORγt/STAT3 signaling pathway109.
TNF and IL-1β B are two of the major proinflammatory cytokines that contribute to neuroinflammation and oxidative stress. Several studies have confirmed quercetin’s suppressive effect on TNF and IL-1β expression by blocking the activation of the MAPK and NF-κB signaling pathways110,111. CTNNB1, which encodes β-catenin, is essential in the Wnt signaling pathway and the regulation of cellular homeostasis. The Wnt/β-catenin signaling pathway contributes to neuronal differentiation, axonal regeneration, local neuroinflammatory responses, and cell apoptosis. Consequently, inhibition of β-catenin can disrupt Wnt signaling, impair neuronal survival, and synaptic connections. Although strong evidence for a direct effect of quercetin on CTNNB1 and β-catenin is lacking, it is likely that quercetin indirectly stimulates the Wnt/β-catenin pathway112.
In conclusion, while these findings provide valuable insights, comprehensive in vitro and in vivo studies are essential to clarify the precise roles of quercetin and Se NPs in modulating these signaling pathways and genes/proteins, particularly in the context of cisplatin-induced neurotoxicity.
Conclusion
Cisplatin is one of the most widely used chemotherapy drugs for cancer treatment, while neurotoxicity remains a major limitation in cisplatin-based chemotherapy. Although quercetin has been reported as a promising neuroprotective polyphenol, its clinical application is hindered by poor bioavailability. To address this issue, we introduced innovative Se NPs containing quercetin, which demonstrated favorable physicochemical properties, biocompatibility, and protective effects against cisplatin-induced neurotoxicity in PC12 cells. Pretreatment with these NPs significantly reduced apoptosis and necrosis in this neuronal cell model.
Network pharmacology analysis identified 10 hub targets (BCL2, CASP3, AKT1, EGFR, STAT3, TNF, HSP90AA1, CTNNB1, IL-1β, and BCL2L1) that may underlie the neuroprotective mechanisms of the NPs. Functional enrichment analysis revealed that these targets are significantly involved in regulating apoptosis, a finding supported by the flow cytometric assessment of cell death.
Finally, by combining quercetin with Tw-Cs@Se NPs, we designed nanostructures that have not been previously reported for mitigating cisplatin neurotoxicity. While these findings suggest that Q-Tw-Cs@Se NPs may help alleviate cisplatin-induced neurotoxicity, further pharmacological research is undoubtedly required to elucidate the underlying mechanisms of neuroprotection and validate efficacy and safety in relevant animal models before considering clinical translation.
Data availability
The datasets used and analyzed during the current study are available from the corresponding author on reasonable request.
References
Santos, N. A. G., D., Ferreira, R. S. & Santos, A. C. D. Overview of cisplatin-induced neurotoxicity and ototoxicity, and the protective agents. Food Chem. Toxicol. 136, 111079. https://doi.org/10.1016/j.fct.2019.111079 (2020).
Dominiak, A., Wilkaniec, A., Wroczyński, P. & Adamczyk, A. Selenium in the therapy of neurological diseases. Where is it going? Curr. Neuropharmacol. 14, 282–299. https://doi.org/10.2174/1570159x14666151223100011 (2016).
Li, D. et al. Attenuation of cisplatin-induced neurotoxicity by cyanidin, a natural inhibitor of ROS-mediated apoptosis in PC12 cells. Cell. Mol. Neurobiol. 35, 995–1001. https://doi.org/10.1007/s10571-015-0194-6 (2015).
Elmorsy, E. A. et al. Advances in Understanding cisplatin-induced toxicity: molecular mechanisms and protective strategies. Eur. J. Pharm. Sci. 203, 106939. https://doi.org/10.1016/j.ejps.2024.106939 (2024).
Alhowail, A. H. Cisplatin induces hippocampal neurotoxicity and cognitive impairment in rats through neuroinflammation, oxidative stress, and overexpression of glutamatergic receptors mRNA. Front. Pharmacol. https://doi.org/10.3389/fphar.2025.1592511 (2025).
Tawfik, K. M., Moustafa, Y. M. & El-Azab, M. F. Neuroprotective mechanisms of sildenafil and selenium in PTZ-kindling model: Implications in epilepsy. Eur. J. Pharmacol. 833, 131–144. https://doi.org/10.1016/j.ejphar.2018.05.035 (2018).
Nazıroğlu, M., Öz, A. & Yıldızhan, K. Selenium and neurological diseases: Focus on peripheral pain and TRP channels. Curr. Neuropharmacol. 18, 501–517. https://doi.org/10.2174/1570159x18666200106152631 (2020).
Nazıroğlu, M. Role of selenium on calcium signaling and oxidative stress-induced molecular pathways in epilepsy. Neurochem. Res. 34, 2181–2191. https://doi.org/10.1007/s11064-009-0015-8 (2009).
Khurana, A., Tekula, S., Saifi, M. A., Venkatesh, P. & Godugu, C. Therapeutic applications of selenium nanoparticles. Biomed. Pharmacother. 111, 802–812. https://doi.org/10.1016/j.biopha.2018.12.146 (2019).
Savas, S. et al. A whole-genome SNP association study of NCI60 cell line panel indicates a role of Ca2+ signaling in selenium resistance. PLoS One. 5, e12601. https://doi.org/10.1371/journal.pone.0012601 (2010).
He, L., Zhang, L., Peng, Y. & He, Z. Selenium in cancer management: Exploring the therapeutic potential. Front. Oncol. https://doi.org/10.3389/fonc.2024.1490740 (2025).
MacFarquhar, J. K. et al. Acute selenium toxicity associated with a dietary supplement. Arch. Intern. Med. 170, 256–261. https://doi.org/10.1001/archinternmed.2009.495 (2010).
Sk, U. H., RoyMahapatra, D. & Bhattacharya, S. In Handbook of Oxidative Stress in Cancer: Therapeutic Aspects (ed Chakraborti, S.) 1–32 (Springer, 2021).
Varlamova, E. G., Turovsky, E. A. & Blinova, E. V. Therapeutic potential and main methods of obtaining selenium nanoparticles. Int. J. Mol. Sci. https://doi.org/10.3390/ijms221910808 (2021).
Hosnedlova, B. et al. Nano-selenium and its nanomedicine applications: A critical review. Int. J. Nanomed. 13, 2107–2128. https://doi.org/10.2147/ijn.S157541 (2018).
Chen, H., Shin, D. W., Nam, J. G., Kwon, K. W. & Yoo, J. B. Selenium nanowires and nanotubes synthesized via a facile template-free solution method. Mater. Res. Bull. 45, 699–704. https://doi.org/10.1016/j.materresbull.2010.02.016 (2010).
Zhang, C., Zhai, X., Zhao, G., Ren, F. & Leng, X. Synthesis, characterization, and controlled release of selenium nanoparticles stabilized by chitosan of different molecular weights. Carbohydr. Polym. 134, 158–166. https://doi.org/10.1016/j.carbpol.2015.07.065 (2015).
Zhang, S. Y., Zhang, J., Wang, H. Y. & Chen, H. Y. Synthesis of selenium nanoparticles in the presence of polysaccharides. Mater. Lett. 58, 2590–2594. https://doi.org/10.1016/j.matlet.2004.03.031 (2004).
Peng, S. et al. Preparation of polysaccharide-conjugated selenium nanoparticles from spent mushroom substrates and their growth-promoting effect on rice seedlings. Int. J. Biol. Macromol. 253, 126789. https://doi.org/10.1016/j.ijbiomac.2023.126789 (2023).
Dave, P. N. & Gor, A. In Handbook of Nanomaterials for Industrial Applications. 36–66 (eds Hussain, C. M.) (Elsevier, 2018).
Wang, B., Wang, L., Qu, Y., Lu, J. & Xia, W. Chitosan oligosaccharides exert neuroprotective effects via modulating the PI3K/Akt/Bcl-2 pathway in a parkinsonian model. Food Funct. 13, 5838–5853. https://doi.org/10.1039/D1FO04374A (2022).
Pangestuti, R. & Kim, S. K. Neuroprotective properties of chitosan and its derivatives. Mar. Drugs 8, 2117–2128 (2010).
Hao, C., Wang, W., Wang, S., Zhang, L. & Guo, Y. An overview of the protective effects of chitosan and acetylated chitosan oligosaccharides against neuronal disorders. Mar. Drugs 15, 89 (2017).
Magar, R. T. & Sohng, J. K. A review on structure, modifications and structure-activity relation of quercetin and its derivatives. J. Microbiol. Biotechnol. 30, 11–20. https://doi.org/10.4014/jmb.1907.07003 (2020).
Pawar, A. et al. A critical evaluation of risk to reward ratio of quercetin supplementation for COVID-19 and associated comorbid conditions. Phytother. Res. 36, 2394–2415. https://doi.org/10.1002/ptr.7461 (2022).
Islam, M. S. et al. Neuropharmacological effects of quercetin: A literature-based review. Front. Pharmacol. https://doi.org/10.3389/fphar.2021.665031 (2021).
Rangel-Ordóñez, L., Nöldner, M., Schubert-Zsilavecz, M. & Wurglics, M. Plasma levels and distribution of flavonoids in rat brain after single and repeated doses of standardized Ginkgo Biloba extract EGb 761®. Planta Med. 76, 1683–1690. https://doi.org/10.1055/s-0030-1249962 (2010).
Ader, P., Wessmann, A. & Wolffram, S. Bioavailability and metabolism of the flavonol quercetin in the pig. Free Radic. Biol. Med. 28, 1056–1067. https://doi.org/10.1016/S0891-5849(00)00195-7 (2000).
Ravichandran, V., Lee, M., Cao, N., Shim, M. S. & T. G. & Polysorbate-based drug formulations for brain-targeted drug delivery and anticancer therapy. Appl. Sci. 11, 9336. https://doi.org/10.3390/app11199336 (2021).
Kang, S. et al. Trichosanthes Kirilowii Maxim. And bioactive compound cucurbitacin D alleviate Cisplatin-Induced peripheral neuropathy in vitro and in vivo. Integr. Cancer Ther. 24, 15347354251339121. https://doi.org/10.1177/15347354251339121 (2025).
Ferreira, R. S. et al. Caffeic acid phenethyl ester (CAPE) protects PC12 cells against cisplatin-induced neurotoxicity by activating the AMPK/SIRT1, MAPK/Erk, and PI3k/Akt signaling pathways. Neurotox. Res. 36, 175–192. https://doi.org/10.1007/s12640-019-00042-w (2019).
Li, Y., Zheng, M., Sah, S. K., Mishra, A. & Singh, Y. Neuroprotective influence of sitagliptin against cisplatin-induced neurotoxicity, biochemical and behavioral alterations in Wistar rats. Mol. Cell. Biochem. 455, 91–97. https://doi.org/10.1007/s11010-018-3472-z (2019).
Westerink, R. H. S. & Ewing, A. G. The PC12 cell as model for neurosecretion. Acta Physiol. 192, 273–285. https://doi.org/10.1111/j.1748-1716.2007.01805.x (2008).
Wiatrak, B., Kubis-Kubiak, A., Piwowar, A. & Barg, E. PC12 cell line: Cell types, coating of culture Vessels, differentiation and other culture conditions. Cells 9, 958 (2020).
Abadi, B. et al. Chitosan-sialic acid nanoparticles of selenium: statistical optimization of production, characterization, and assessment of cytotoxic effects against two human glioblastoma cell lines. Int. J. Pharm. 637, 122884. https://doi.org/10.1016/j.ijpharm.2023.122884 (2023).
Shakibaie, M. et al. Microwave assisted biosynthesis of cadmium nanoparticles: Characterization, antioxidant and cytotoxicity studies. J. Cluster Sci. 33, 1877–1887. https://doi.org/10.1007/s10876-021-02107-3 (2022).
Roacho-Pérez, J. A. et al. A full set of in vitro assays in chitosan/tween 80 microspheres loaded with magnetite nanoparticles. Polymers 13, 400. https://doi.org/10.3390/polym13030400 (2021).
Wang, H. et al. Utilizing network Pharmacological analysis to investigate the key targets and mechanisms of kaempferol against oxaliplatin-induced neurotoxicity. Toxicol. Mech. Methods. 33, 38–46. https://doi.org/10.1080/15376516.2022.2069531 (2023).
Zhang, L., Cai, J., Xiao, J. & Ye, Z. Identification of core genes and pathways between geriatric multimorbidity and renal insufficiency: Potential therapeutic agents discovered using bioinformatics analysis. BMC Med. Genom. 15, 212. https://doi.org/10.1186/s12920-022-01370-1 (2022).
Kuleshov, M. V. et al. Enrichr: A comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97. https://doi.org/10.1093/nar/gkw377 (2016).
Hernández-Díaz, J. A. et al. Antibacterial activity of biosynthesized selenium nanoparticles using extracts of calendula officinalis against potentially clinical bacterial strains. Molecules 26, 5929. https://doi.org/10.3390/molecules26195929 (2021).
Maiyo, F. & Singh, M. Folate-targeted mRNA delivery using chitosan-functionalized selenium nanoparticles: Potential in cancer immunotherapy. Pharmaceuticals 12, 164. https://doi.org/10.3390/ph12040164 (2019).
Lin, Z. H. & Chris Wang, C. R. Evidence on the size-dependent absorption spectral evolution of selenium nanoparticles. Mater. Chem. Phys. 92, 591–594. https://doi.org/10.1016/j.matchemphys.2005.02.023 (2005).
Luesakul, U., Komenek, S., Puthong, S. & Muangsin, N. Shape-controlled synthesis of cubic-like selenium nanoparticles via the self-assembly method. Carbohydr. Polym. 153, 435–444. https://doi.org/10.1016/j.carbpol.2016.08.004 (2016).
Johnson, J. A., Saboungi, M. L., Thiyagarajan, P., Csencsits, R. & Meisel, D. Selenium nanoparticles: A small-angle neutron scattering study. J. Phys. Chem. B 103, 59–63. https://doi.org/10.1021/jp983229y (1999).
Song, X., Chen, Y., Sun, H., Liu, X. & Leng, X. Physicochemical and functional properties of chitosan-stabilized selenium nanoparticles under different processing treatments. Food Chem. 331, 127378. https://doi.org/10.1016/j.foodchem.2020.127378 (2020).
Karthikeyan, C., Varaprasad, K., Akbari-Fakhrabadi, A., Hameed, A. S. H. & Sadiku, R. Biomolecule chitosan, curcumin and ZnO-based antibacterial nanomaterial, via a one-pot process. Carbohydr. Polym. 249, 116825. https://doi.org/10.1016/j.carbpol.2020.116825 (2020).
Raul, P. K. et al. CuO nanorods: A potential and efficient adsorbent in water purification. RSC Adv. 4, 40580–40587. https://doi.org/10.1039/C4RA04619F (2014).
Bide, Y., Fashapoyeh, M. A. & Shokrollahzadeh, S. Structural investigation and application of tween 80-choline chloride self-assemblies as osmotic agent for water desalination. Sci. Rep. 11, 17068. https://doi.org/10.1038/s41598-021-96199-6 (2021).
Catauro, M. et al. Silica/quercetin sol–gel hybrids as antioxidant dental implant materials. Sci. Technol. Adv. Mater. 16, 035001. https://doi.org/10.1088/1468-6996/16/3/035001 (2015).
Tran, T. T. & Hadinoto, K. A. Potential quorum-sensing inhibitor for bronchiectasis therapy: Quercetin–chitosan nanoparticle complex exhibiting superior inhibition of biofilm formation and swimming motility of pseudomonas aeruginosa to the native quercetin. Int. J. Mol. Sci. 22, 1541. https://doi.org/10.3390/ijms22041541 (2021).
Prestianni, L. et al. Synthesis and characterization of Quercetin–Iron complex nanoparticles for overcoming drug resistance. Pharmaceutics 15, 1041. https://doi.org/10.3390/pharmaceutics15041041 (2023).
Hansen, P. E. & Spanget-Larsen, J. NMR and IR investigations of strong intramolecular hydrogen bonds. Molecules 22, 552. https://doi.org/10.3390/molecules22040552 (2017).
Elsayed, A. M. et al. Novel quercetin encapsulated chitosan functionalized copper oxide nanoparticles as anti-breast cancer agent via regulating p53 in rat model. Int. J. Biol. Macromol. 185, 134–152. https://doi.org/10.1016/j.ijbiomac.2021.06.085 (2021).
Horky, P. et al. Selenium nanoparticles as potential antioxidants to improve semen quality in boars. Animals 13, 2460. https://doi.org/10.3390/molecules2619592910.3390/ani13152460 (2023).
Zhai, X. et al. Antioxidant capacities of the selenium nanoparticles stabilized by chitosan. J. Nanobiotechnol. 15, 4. https://doi.org/10.1186/s12951-016-0243-4 (2017).
Zhou, F., Peterson, T., Fan, Z. & Wang, S. The commonly used stabilizers for phytochemical-based nanoparticles: Stabilization effects, mechanisms, and applications. Nutrients 15, 3881. https://doi.org/10.3390/nu15183881 (2023).
Yarjanli, Z., Ghaedi, K., Esmaeili, A., Zarrabi, A. & Rahgozar, S. The antitoxic effects of quercetin and quercetin-conjugated iron oxide nanoparticles (QNPs) against H(2)O(2)-induced toxicity in PC12 cells. Int. J. Nanomed. 14, 6813–6830. https://doi.org/10.2147/ijn.S212582 (2019).
de Menezes, B. B., Frescura, L. M., Duarte, R. & Villetti, M. A. & Da Rosa, M. B. A critical examination of the DPPH method: Mistakes and inconsistencies in stoichiometry and IC50 determination by UV–Vis spectroscopy. Anal. Chim. Acta. 1157, 338398. https://doi.org/10.1016/j.aca.2021.338398 (2021).
Arriagada, F., Günther, G. & Morales, J. Nanoantioxidant-based silica particles as flavonoid carrier for drug delivery applications. Pharmaceutics 12, 302 (2020).
Wu, Y. et al. Pectin-decorated selenium nanoparticles as a nanocarrier of curcumin to achieve enhanced physicochemical and biological properties. IET Nanobiotechnol. 13, 880–886. https://doi.org/10.1049/iet-nbt.2019.0144 (2019).
Guo, S. et al. Relationship and improvement strategies between drug nanocarrier characteristics and hemocompatibility: What can we learn from the literature. Asian J. Pharm. Sci. 16, 551–576. https://doi.org/10.1016/j.ajps.2020.12.002 (2021).
ASTMF-756. Standard practice for assessment of hemolytic properties of materials, Annual Book of ASTM Standards. Committee F04 Medical and Surgical Materials and Devices, Subcommittee F04.16 Biocompatibility Test Methods. Annual Book of ASTM Standards. (2009).
Macías-Martínez, B. I., Cortés-Hernández, D. A., Zugasti-Cruz, A., Cruz-Ortíz, B. R. & Múzquiz-Ramos, E. M. Heating ability and hemolysis test of magnetite nanoparticles obtained by a simple co-precipitation method. J. Appl. Res. Technol. 14, 239–244. https://doi.org/10.1016/j.jart.2016.05.007 (2016).
Sampathi, S., Tiriya, P. K., Dodoala, S., Junnuthula, V. & Dyawanapelly, S. Development of biocompatible ciprofloxacin–gold nanoparticle coated sutures for surgical site infections. Pharmaceutics 14, 2130 (2022).
Sukhanova, A. et al. Dependence of nanoparticle toxicity on their physical and chemical properties. Nanoscale Res. Lett. 13, 44. https://doi.org/10.1186/s11671-018-2457-x (2018).
Yang, L., Wang, N. & Zheng, G. Enhanced effect of combining chlorogenic acid on selenium nanoparticles in inhibiting amyloid β aggregation and reactive oxygen species formation in vitro. Nanoscale Res. Lett. 13, 303. https://doi.org/10.1186/s11671-018-2720-1 (2018).
Qiao, L., Chen, Y., Dou, X., Song, X. & Xu, C. Biogenic selenium nanoparticles attenuate Aβ25–35-Induced toxicity in PC12 cells via Akt/CREB/BDNF signaling pathway. Neurotox. Res. 40, 1869–1881. https://doi.org/10.1007/s12640-022-00590-8 (2022).
Wang, J. et al. Blood brain barrier-targeted delivery of double selenium nanospheres ameliorates neural ferroptosis in Alzheimer’s disease. Biomaterials 302, 122359. https://doi.org/10.1016/j.biomaterials.2023.122359 (2023).
Turovsky, E. A. et al. Features of the cytoprotective effect of selenium nanoparticles on primary cortical neurons and astrocytes during oxygen–glucose deprivation and reoxygenation. Sci. Rep. 12, 1710. https://doi.org/10.1038/s41598-022-05674-1 (2022).
Ravichandran, R., Rajendran, M. & Devapiriam, D. Antioxidant study of quercetin and their metal complex and determination of stability constant by spectrophotometry method. Food Chem. 146, 472–478. https://doi.org/10.1016/j.foodchem.2013.09.080 (2014).
Sasaki, M. et al. Quercetin-induced PC12 cell death accompanied by caspase-mediated DNA fragmentation. Biol. Pharm. Bull. 30, 682–686. https://doi.org/10.1248/bpb.30.682 (2007).
Dajas, F. et al. Quercetin in brain diseases: Potential and limits. Neurochem. Int. 89, 140–148. https://doi.org/10.1016/j.neuint.2015.07.002 (2015).
Menon, S. & Shanmugam, V. K. Chemopreventive mechanism of action by oxidative stress and toxicity induced surface decorated selenium nanoparticles. J. Trace Elem. Med Biol. 62, 126549. https://doi.org/10.1016/j.jtemb.2020.126549 (2020).
Bao, D., Wang, J., Pang, X. & Liu, H. Protective effect of Quercetin against oxidative Stress-Induced cytotoxicity in rat pheochromocytoma (PC-12) cells. Molecules 22, 1122. https://doi.org/10.3390/molecules22071122 (2017).
Dong, Y. et al. Quercetin alleviates intestinal oxidative damage induced by H2O2 via modulation of GSH: In vitro screening and in vivo evaluation in a colitis model of mice. ACS Omega. 5, 8334–8346. https://doi.org/10.1021/acsomega.0c00804 (2020).
Magalingam, K. B., Radhakrishnan, A. & Haleagrahara, N. Protective effects of Quercetin glycosides, rutin, and Isoquercetrin against 6-hydroxydopamine (6-OHDA)-induced neurotoxicity in rat pheochromocytoma (PC-12) cells. Int. J. ImmunoPathol Pharmacol. 29, 30–39. https://doi.org/10.1177/0394632015613039 (2016).
Ferreira, M. et al. Development and characterization of quercetin-loaded delivery systems for increasing its bioavailability in cervical cancer cells. Pharmaceutics 15, 936. https://doi.org/10.3390/pharmaceutics15030936 (2023).
Zhao, G. et al. Selenium nanoparticles are more efficient than sodium selenite in producing reactive oxygen species and hyper-accumulation of selenium nanoparticles in cancer cells generates potent therapeutic effects. Free Radic. Biol. Med. 126, 55–66. https://doi.org/10.1016/j.freeradbiomed.2018.07.017 (2018).
Li, Y. et al. The reversal of cisplatin-induced nephrotoxicity by selenium nanoparticles functionalized with 11-mercapto-1-undecanol by Inhibition of ROS-mediated apoptosis. Biomaterials 32, 9068–9076. https://doi.org/10.1016/j.biomaterials.2011.08.001 (2011).
Karavelioglu, E. et al. Selenium protects cerebral cells by cisplatin induced neurotoxicity. Acta Cirúrgica Brasileira https://doi.org/10.1590/S0102-865020150060000004 (2015).
ALRashdi, B. M. et al. Therapeutic activity of green synthesized selenium nanoparticles from turmeric against cisplatin-induced oxido-inflammatory stress and cell death in mice kidney. Biosci. Rep. https://doi.org/10.1042/bsr20231130 (2023).
Bournival, J., Plouffe, M., Renaud, J., Provencher, C. & Martinoli, M. G. Quercetin and sesamin protect dopaminergic cells from MPP+-induced neuroinflammation in a microglial (N9)-neuronal (PC12) coculture system. Oxidative Med. Cell. Longev. 2012(921941), https://doi.org/10.1155/2012/921941 (2012).
Nichols, M. et al. Synergistic neuroprotection by epicatechin and quercetin: Activation of convergent mitochondrial signaling pathways. Neuroscience 308, 75–94. https://doi.org/10.1016/j.neuroscience.2015.09.012 (2015).
Yardim, A. et al. Quercetin provides protection against the peripheral nerve damage caused by vincristine in rats by suppressing caspase 3, NF-κB, ATF-6 pathways and activating Nrf2, Akt pathways. NeuroToxicology 81, 137–146. https://doi.org/10.1016/j.neuro.2020.10.001 (2020).
Rahman, M. Neuroprotective effect of rutin on iron induced oxidative stress and cytotoxicity in PC12 neuronal cells via Bcl-2/Bax/caspase-3 signaling pathways. Parkinsonism Relat. Disord. 46, e38–e39. https://doi.org/10.1016/j.parkreldis.2017.11.125 (2018).
Sharma, D. R. et al. Quercetin attenuates neuronal death against aluminum-induced neurodegeneration in the rat hippocampus. Neuroscience 324, 163–176. https://doi.org/10.1016/j.neuroscience.2016.02.055 (2016).
Alam, S., Mohammad, T., Padder, R. A., Hassan, M. I. & Husain, M. Thymoquinone and Quercetin induce enhanced apoptosis in non-small cell lung cancer in combination through the Bax/Bcl2 cascade. J. Cell. Biochem. 123, 259–274. https://doi.org/10.1002/jcb.30162 (2022).
Primikyri, A. et al. Direct binding of Bcl-2 family proteins by quercetin triggers its pro-apoptotic activity. ACS Chem. Biol. 9, 2737–2741. https://doi.org/10.1021/cb500259e (2014).
Mohamed, A. A. R. et al. Chitosan-stabilized selenium nanoparticles alleviate cardio-hepatic damage in type 2 diabetes mellitus model via regulation of caspase, Bax/Bcl-2, and Fas/FasL-pathway. Gene 768, 145288. https://doi.org/10.1016/j.gene.2020.145288 (2021).
He, X. et al. The PI3K/AKT signalling pathway in inflammation, cell death and glial Scar formation after traumatic spinal cord injury: mechanisms and therapeutic opportunities. Cell Prolif. 55, e13275. https://doi.org/10.1111/cpr.13275 (2022).
Zubčić, K. et al. PI3K/Akt and ERK1/2 signalling are involved in quercetin-mediated neuroprotection against copper-induced injury. Oxid. Med. Cell. Long. https://doi.org/10.1155/2020/9834742(2020) (2020).
Shen, W. et al. Exploration of the effect and potential mechanism of Quercetin in repairing spinal cord injury based on network pharmacology and in vivo experimental verification. Heliyon 9, e20024. https://doi.org/10.1016/j.heliyon.2023.e20024 (2023).
Liu, P. et al. Size effect-based improved antioxidant activity of selenium nanoparticles regulating Anti-PI3K-mTOR and Ras-MEK pathways for treating spinal cord injury to avoid hormone shock-induced immunosuppression. J. Nanobiotechnol. 23, 17. https://doi.org/10.1186/s12951-024-03054-7 (2025).
Romano, R. & Bucci, C. Role of EGFR in the nervous system. Cells 9, 1887. https://doi.org/10.3390/cells9081887 (2020).
Asgharian, P. et al. Potential mechanisms of quercetin in cancer prevention: Focus on cellular and molecular targets. Cancer Cell Int. 22, 257. https://doi.org/10.1186/s12935-022-02677-w (2022).
Liu, Y. et al. Quercetin attenuates podocyte apoptosis of diabetic nephropathy through targeting EGFR signaling. Front. Pharmacol. https://doi.org/10.3389/fphar.2021.792777(2022) (2021).
Wang, Q., Wang, F., Li, X., Ma, Z. & Jiang, D. Quercetin inhibits the amphiregulin/EGFR signaling-mediated renal tubular epithelial-mesenchymal transition and renal fibrosis in obstructive nephropathy. Phytother. Res. 37, 111–123. https://doi.org/10.1002/ptr.7599 (2023).
Xie, Y., Xu, S., Chen, Z., Song, C. & Yan, W. Unveiling the therapeutic potential of airpotato yam rhizome against colorectal cancer: A network pharmacology approach. Front. Oncol. 2024. https://doi.org/10.3389/fonc.2024.1414766 (2024).
Xiao, X. et al. HSP90AA1-mediated autophagy promotes drug resistance in osteosarcoma. J. Experimental Clin. Cancer Res. 37, 201. https://doi.org/10.1186/s13046-018-0880-6 (2018).
Kıyga, E., Şengelen, A. & Adıgüzel, Z. Önay Uçar, E. Investigation of the role of quercetin as a heat shock protein inhibitor on apoptosis in human breast cancer cells. Mol. Biol. Rep. 47, 4957–4967. https://doi.org/10.1007/s11033-020-05641-x (2020).
Kojouri, G. A., Faramarzi, P., Ahadi, A. M. & Parchami, A. Effect of selenium nanoparticles on expression of HSP90 gene in myocytes after an intense exercise. J. Equine Vet. Sci. 33, 1054–1056. https://doi.org/10.1016/j.jevs.2013.04.001 (2013).
Smit, R. D. et al. STAT3 protects dopaminergic neurons against degeneration in animal model of Parkinson’s disease. Brain Res. 1824, 148691. https://doi.org/10.1016/j.brainres.2023.148691 (2024).
Kim, S. K. et al. Neuroprotective effects of STAT3 inhibitor on hydrogen peroxide-induced neuronal cell death via the ERK/CREB signaling pathway. Neurochem. Res. 50, 52. https://doi.org/10.1007/s11064-024-04252-3 (2024).
Michaud-Levesque, J., Bousquet-Gagnon, N. & Béliveau, R. Quercetin abrogates IL-6/STAT3 signaling and inhibits glioblastoma cell line growth and migration. Exp. Cell Res. 318, 925–935. https://doi.org/10.1016/j.yexcr.2012.02.017 (2012).
Wu, L. et al. Quercetin shows anti-tumor effect in hepatocellular carcinoma LM3 cells by abrogating JAK2/STAT3 signaling pathway. Cancer Med. 8, 4806–4820. https://doi.org/10.1002/cam4.2388 (2019).
Zalpoor, H. et al. Quercetin as a JAK–STAT inhibitor: A potential role in solid tumors and neurodegenerative diseases. Cell. Mol. Biol. Lett. 27, 60. https://doi.org/10.1186/s11658-022-00355-3 (2022).
Zou, Y. et al. Quercetin regulates microglia M1/M2 polarization and alleviates retinal inflammation via ERK/STAT3 pathway. Inflammation 47, 1616–1633. https://doi.org/10.1007/s10753-024-01997-5 (2024).
Gu, W. J. et al. Selenium nanoparticles activate selenoproteins to mitigate septic lung injury through miR-20b-mediated RORγt/STAT3/Th17 axis inhibition and enhanced mitochondrial transfer in BMSCs. J. Nanobiotechnol. 23, 226. https://doi.org/10.1186/s12951-025-03312-2 (2025).
Chen, T. et al. Quercetin inhibits TNF-α induced HUVECs apoptosis and inflammation via downregulating NF-kB and AP-1 signaling pathway in vitro. Medicine 99, e22241. https://doi.org/10.1097/md.0000000000022241 (2020).
Cheng, S. C., Huang, W. C., Pang, S., Wu, J. H., Cheng, C. Y. & Y.-H. & Quercetin inhibits the production of IL-1β-Induced inflammatory cytokines and chemokines in ARPE-19 cells via the MAPK and NF-κB signaling pathways. Int. J. Mol. Sci. 20, 2957 (2019).
Predes, D. et al. The flavonol quercitrin hinders GSK3 activity and potentiates the Wnt/β-catenin signaling pathway. Int. J. Mol. Sci. 23, 12078. https://doi.org/10.3390/ijms232012078 (2022).
Acknowledgements
We thank the Pharmaceutical Sciences and Cosmetic Products Research Center, Institute of Pharmaceutical Sciences, Kerman University of Medical Sciences (Kerman, Iran) for its admirable laboratory and equipment participation in this study.
Funding
This work was supported by Elite Researcher Grant Committee under award number from the National Institutes for Medical Research Development, Tehran, Iran (NIMAD, 4010032).
Author information
Authors and Affiliations
Contributions
Banafshe Abadi: Investigation, Methodology, Formal analysis, Data curation, Writing–original draft, Visualization, Conceptualization. Mahboubeh Adeli-Sardou: Investigation. Hamid Forootanfar: Conceptualization, Writing–review & editing. Mojtaba Shakibaie: Supervision, Methodology, Validation, Funding acquisition, Conceptualization.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This study was approved by the Biomedical Research Ethics Committee of National Institutes for Medical Research Development, Tehran, Iran (Ethical approval ID: IR.NIMAD.REC.1402.035).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Abadi, B., Adeli-Sardou, M., Forootanfar, H. et al. Protective potential of tween 80-functionalized chitosan-selenium nanoparticles containing quercetin against cisplatin neurotoxicity in PC12 cells: an in vitro and network pharmacology study. Sci Rep 16, 1940 (2026). https://doi.org/10.1038/s41598-025-31744-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-31744-1