Abstract
Active elimination of apoptotic cells is very important for maintaining homeostasis of early embryos. Recent observations on mouse blastocysts freshly isolated from healthy dams have shown that the majority of incidentally occurring apoptotic cells is eliminated by neighbouring embryonic cells. Some apoptotic cells escape phagocytosis, but the frequency of such processes usually does not exceed 10%. The aim of the current study was to evaluate whether the non-professional embryonic phagocytes can respond to experimentally induced increase in apoptosis by increasing the frequency of efferocytosis and whether their activity can be decreased by selective inhibition of specific component of efferoctosis machinery. Experiments were performed in vitro on cultured mouse blastocysts with a differentiated trophectoderm and inner cell mass and on the human trophoblast cell line Ac-1M88. Samples were assessed using fluorescence immunostaining: Apoptotic cells (TUNEL) internalised within the cytoplasm of non-professional embryonic phagocytes (phalloidin T membrane staining) were considered ingested; apoptotic cells co-localised with acidified phagosomes (LysoTracker) were considered digested. First, we tested the ability of embryonic phagocytes to respond to elevated incidence of apoptosis induced by actinomycin D (4 nM). The results showed that the increase in apoptosis was accompanied by a significant elevation of the phagocytosis and digestion of dead cells in both mouse blastocysts and human trophoblast cells. We then assessed the effect of selective inhibition of lysosomal acidification in embryonic phagocytes using a specific V-ATPase inhibitor bafilomycin A1. The results showed that the inhibitor at 0.1 and 0.2 nM was able to negatively affect the execution of both initiative and terminal phases of efferocytosis in mouse blastocysts, although the decrease was not as profound as expected. When compared to mouse trophectoderm cells, human hybrid cells displayed a very low sensitivity to bafilomycin A1. Higher concentrations of bafilomycin A1 had a more harmful impact on overall cell viability than on digestive activity. The results show that the ability of non-professional embryonic phagocytes to successfully execute all stages of efferocytosis is not limited by the frequency of apoptosis and is preserved even at elevated rates of the apoptotic process. The effectiveness of embryonic phagocytes can be partially decreased by selective inhibition of lysosomal acidification conducted via V-ATPase.
Similar content being viewed by others
Introduction
Apoptosis is a physiological process occurring spontaneously in the majority of cell populations. The main function of programmed cell death during preimplantation development is the self-destruction of the minority of cells with abnormal or detrimental potential1,2.
Apoptotic cells in blastocysts are eliminated by neighbouring blastomeres. Our previous study provided the first insight into the machinery and physiological occurrence of embryonic efferocytosis3. We have shown that intact embryonic cells in mouse blastocysts can act as non-professional phagocytes and possess all the mechanisms necessary for the recognition, engulfment and digestion of damaged blastomeres. The process of embryonic efferocytosis begins with the recognition of the apoptotic cell via the binding of various phagocytic receptors to externalised phosphatidyl serine or modified lipoproteins. Signalling leads to the recruitment of Rho family GTPase RAC1, the actin-dependent formation of the phagocytic cup and targeted internalisation, i.e. taking the apoptotic cell into a vacuole (the phagosome), which then undergoes maturation (progressive acidification) and gradual degradation. Intact early embryonic cells undertake the clearance of the majority of dying cells in blastocysts. Some apoptotic cells escape phagocytosis, but the frequency of such processes usually does not exceed 10%3.
The main role of efferocytosis is to prevent the release of potentially noxious or immunogenic intracellular materials from decaying apoptotic cells. Thus, their active elimination is very important for maintaining embryonic homeostasis. During early mammalian embryogenesis, the cells subjected to elimination can be categorized into three distinct groups: cells demonstrating disparities in differentiation status, karyotypically aberrant cells, and cells exhibiting mitochondrial dysfunction4,5.
The aim of the current study was to evaluate whether the exposure of blastocyst to specific non-physiological conditions can affect efficiency of embryonic efferocytosis. We tested following hypotheses: 1. whether the non-professional embryonic phagocytes can respond to experimentally induced increase in apoptosis by increasing the frequency of engulfment and digestion of dead cells, and 2. whether the activity of non-professional embryonic phagocytes can be decreased by selective inhibition of specific component of efferoctosis machinery – an enzyme responsible for acidification of phagosomes. Analyzing the fundamental processes that regulate cell will provide a comprehensive understanding of early embryonic development and offer novel approaches for therapeutic interventions in cases of early pregnancy loss6. We first tested the ability of embryonic phagocytes to respond to an elevation in the frequency of apoptosis induced by a well-established inductor. For such purpose, actinomycin D was used. Actinomycin D is a cyclic polypeptide (usually used as an antibiotic) that binds to DNA, inhibits RNA synthesis7,8,9 and results in increased apoptosis and decreased proliferation10,11,12.
Several previous studies have shown that actinomycin D can negatively affect development of mouse embryos13,14,15 and significantly increase the incidence of apoptotic cells in blastocysts16,17,18,19. Secondly, we assessed the impact of selective inhibition of lysosomal acidification in embryonic phagocytes on efferocytosis efficiency. For this purpose, a specific V-ATPase inhibitor bafilomycin A1 was used. Bafilomycin A1 specifically targets the vacuolar-type H + ATPase (V-ATPase) enzyme, a membrane-spanning proton pump that acidifies either the extracellular environment or the intracellular organelles, such as the lysosome20. Reduced lysosomal acidification subsequently inhibits the activity of lysosomal proteases21.
At higher (micromolar) concentrations, bafilomycin A1 can also act on P-type ATPases20,22,23,24,25. Previous study has shown that bafilomycin A1 at 160.6 nM can disrupt the frequency of autophagy in aneuploid mouse blastocysts26. However, since no other information on the effect of bafilomycin A1 on embryonic cells is available, in the current study, we used a wider range of concentrations of the inhibitor (from 0.01 nM to 2.00 nM). In addition, to assess the general validity of the findings obtained on a mouse blastocyst model, all experiments were performed on the human hybrid trophoblast cell line Ac-1M88 in parallel.
Results
Response of embryonic phagocytes to increased incidence of apoptosis
Mouse blastocysts exposed to actinomycin D showed a significantly decreased mean number of cells when compared to untreated controls (Tables 1 and 4). The presence of the apoptotic inductor in culture media significantly increased the mean number of apoptotic cells per blastocyst (Tables 1 and 4). Such increase was accompanied by increased extrusion and engulfment of apoptotic cells (Table 1). However, the proportion of extruded, internalised and non-internalised apoptotic cells did not differ from their proportion in control blastocysts (Table 1). In both groups, approx. 80% of evaluated apoptotic cells were localised in the cytoplasm of neighbouring embryonic cells. This suggests the ability of non-professional embryonic phagocytes to respond to an increase in apoptosis by elevating the phagocytosis of dead cells. What’s more, in both treated and untreated embryos, the majority of evaluated apoptotic cells displayed co-localisation with acidified phagosomes (Table 4). In mouse blastocysts exposed to actinomycin D, the average % and overall proportion of LysoTracker-positive apoptotic cells were even higher than in the controls. This suggests that the increase in engulfment of apoptotic cells is followed by their successful digestion.
Similar results were observed when the trophectoderm cell line in mouse blastocyst was evaluated separately (Tables 2 and 5). There was only one exception: Although the trend was the same, the increase in the proportion of LysoTracker-positive apoptotic cells in the group exposed to actinomycin D did not reach statistical significance.
Actinomycin D showed an analogous effect on cell number and apoptosis incidence in the human trophoblast cell line, although the overall incidence of spontaneous apoptosis in the control samples and incidence of induced apoptosis in the treated samples appeared to be lower than in mouse trophectoderm cell line (Tables 3 and 6). Hybrid cells were able to respond to the increase in apoptosis by elevating phagocytic and digestive activity, too (Tables 3 and 6).
The results show that the ability of non-professional embryonic phagocytes to successfully execute all stages of efferocytosis is not limited by physiological conditions, i.e. frequency of apoptosis (Tables 4, 5 and 6).
The effect of inhibition of lysosomal acidification on embryonic efferocytosis
Mouse blastocysts exposed to bafilomycin A1 showed a significantly decreased mean number of cells when compared to untreated controls (Tables 7 and 10). The presence of a lysosomal acidification inhibitor in the culture media did not affect the mean number of apoptotic cells per blastocyst (Tables 8 and 9).
Bafilomycin A1 at 0.1 and 0.2 nM significantly increased average % (P < 0.01 and P < 0.001, respectively) and the overall proportion (P < 0.001 for both concentrations) of LysoTracker-negative apoptotic cells (Table 10). This suggests its ability to negatively affect the execution of the terminal phases of efferocytosis in non-professional embryonic phagocytes. Surprisingly, mouse blastocysts exposed to bafilomycin A1 at 0.1 and 0.2 nM also showed a significant decrease in the proportion of internalised apoptotic cells (Table 7, P < 0.01 for both concentrations) and a slight increase in the proportion of non-internalised apoptotic cells (Table 7, P < 0.05 for 0.1 nM, and P = 0.09 for 0.2 nM). Furthermore, embryos treated with inhibitor at 0.2 nM showed an increase in the % of extruded apoptotic cells, i.e. in the % of cells that escaped phagocytosis. This suggests the ability of bafilomycin A1 to negatively affect even the early phase of embryonic efferocytosis – the engulfment process.
Again, similar results were observed when the trophectoderm cell line in mouse blastocyst was evaluated separately (Tables 8 and 11). There were only two exceptions: Although the trend was the same, the increase in the proportion of LysoTracker-negative apoptotic cells in the group exposed to bafilomycin A1 at 0.2 nM did not reach statistical significance (Table 11); statistical analysis proved negative effect on apoptotic cell internalisation just when trophectoderm cells were exposed to bafilomycin A1 at a concentration of 0.2 nM (Table 8). This suggests a more prominent effect of bafilomycin A1 on processes of efferocytosis in the inner cell mass than in the trophectoderm cell line.
The response of the human trophoblast cell line to the lysosomal acidification inhibitor differed from that of the mouse trophectoderm cell line. Bafilomycin A1 at equivalent concentrations (from 0.01 to 0.2 nM) did not affect the growth of human trophoblast cells (Tables 9 and 12). The tendency to decrease the mean number of cells per optical field was recorded when the concentration of 2 nM was used; however, even in this case the difference from the control group did not reach statistical significance (Table 12, P = 0.0061). On the other hand, bafilomycin A1 at concentrations of 0.2 (Table 9), 0.5 and 2 nM (Table 12) significantly elevated the incidence of apoptotic cells in the hybrid cell line.
The presence of the lysosomal acidification inhibitor at concentrations up to 0.2 nM did not affect either the percentage of trophoblast apoptotic cells co-localised with acidified phagosomes (Table 12) or the percentage of internalised trophoblast apoptotic cells (Table 9, P > 0.05 for all evaluated parameters). This suggests a lower sensitivity of human hybrid cells to bafilomycin A1 when compared to mouse normal trophectoderm cells. When bafilomycin A1 at 2 nM was used, massive elevation in the mean number of apoptotic cells per optical field was accompanied with a significant increase in the average % of both LysoTracker-negative and LysoTracker-positive apoptotic cells when compared to the control group (P < 0.0001 for both cases). The proportion of LysoTracker-negative apoptotic cells tended to be higher; however, the difference from the control group did not reach statistical significance (Table 12, 25.31% vs 17.02%, P = 0.33). Apparently, higher concentrations of bafilomycin A1 had a more harmful impact on overall cell viability than on the phagocytic or digestive activity of hybrid cells.
Effect of low concentrations of actinomycin D on transcription in early embryonic cells
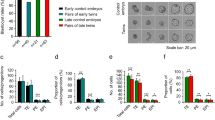
To evaluate the effect of low concentrations of actinomycin D on transcription in early embryonic cells, we compared the abundance of 10 transcripts in mouse blastocysts treated with 4 nM actinomycin D and in control blastocysts. Transcripts of five genes involved in the regulation of apoptosis (BCL2-associated X protein, Bax; BCL2-antagonist/killer 1, Bak1; BCL2 binding component 3, Bbc3 / PUMA; B cell leukemia/lymphoma 2, Bcl2; BCL2-like 1, Bcl2l1) and five transcripts of housekeeping genes (actin beta, Actb; succinate dehydrogenase complex subunit A, Sdha; peptidylprolyl isomerase A, Ppia; H2A histone family member Z, H2afz; beta-2 microglobulin, B2m) were analyzed using real-time RT-PCR. Our results showed that actinomycin D significantly reduced the abundance of four housekeeping gene transcripts (P = 0.0002 for ActB, P = 0.0022 for Sdha, P = 0.0021 for H2afz and P = 0.023 for Ppia; Fig. 1A). The abundance of the two antiapoptotic gene transcripts examined was slightly (Bcl2) or significantly (Bcl2l1, P = 0.024) reduced (Fig. 1B). Conversely, the abundances of proapoptotic gene transcripts were slightly (Bak1) or significantly (Bax, P = 0.0073; Bbc3/PUMA, P = 0.0126) increased (Fig. 1B).
Effect of actinomycin D on the expression of selected genes. (A) Relative quantity of Actb, Sdha, Ppia, H2afz and B2m transcripts in blastocysts. (B) Relative quantity of Bbc3/PUMA, Bax, Bak1, Bcl2, and Bcl2l1 transcripts in blastocysts. The values are arithmetic means + SEMs; Cont, control blastocysts; ActD, actinomycin D-treated blastocysts; n = 5. Statistical differences between the groups were assessed via the unpaired T-test (*P < 0.05), (**P < 0.01), (***P < 0.001).
Discussion
Response of embryonic phagocytes to increased incidence of apoptosis
The removal of dead cells by both professional and non-professional phagocytes is a crucial process that maintains the balance of most cellular systems10. It has been suggested that if the rate of cell death by apoptosis exceeds the capacity of macrophages to clear away dead cells, these apoptotic cells may become necrotic, leading to the release of harmful cellular contents and tissue damage11. Since non-professional phagocytes have a more restricted set of targets and engulf them more slowly, their clearance capacity is considered to be even lower12,27.
It is generally accepted that an increase in apoptosis is accompanied by the recruitment of phagocytes and an increase in their activity. However, to our knowledge, no experimental model in vivo or in vitro has previously been executed to evaluate the clearance capacity of non-professional phagocytes.
Our results obtained on a mouse blastocyst model show that the ability of non-professional embryonic phagocytes to successfully execute efferocytosis is not limited by the frequency of apoptosis: the increase in the frequency of apoptosis induced by actinomycin D was accompanied by an increase in the engulfment of apoptotic cells.
The ability of actinomycin D to induce apoptosis in preimplantation embryos was shown in several studies17,18,28,29, and results of our previous study showed that, except of general lowering transcript amounts, actinomycinD affected the ratio of regulatory pro-apoptotic and anti-apoptotic proteins (Bad and Bax vs. Bcl2l2) in favour of pro-apoptotic ones18. Actinomycin D have also been shown to induce apoptosis in various cell lines and tissues and several mechanisms involved in this process have been proposed30,31.
Mouse blastocysts exposed to actinomycin D for 24 h showed a significantly decreased mean number of cells when compared to controls. A similar cleavage-inhibiting effect of actinomycin D was documented in previous studies on in vitro cultured mouse, rabbit and bovine oocytes32,33,34, developing preimplantation embryos35, and blastocysts17,28.
In the current study, the inhibition of cell cycle progression and cell division by actinomycin D was substantiated through immunohistochemical analysis of Ki-67, a well-established marker of cell proliferation. Results revealed a statistically significant reduction in the mean number of Ki67-positive embryonic cells in mouse blastocysts treated with actinomycin D compared to controls (Supplementary Table S1). Ki-67 is a nuclear protein that is ubiquitously expressed in proliferating cells, whereas it is absent in quiescent or differentiated cells36,37.
Actinomycin D inhibits transcription through all three types of RNA polymerase, with RNA polymerase I (which transcribes ribosomal RNA encoding genes) being the most sensitive (it can be inhibited with nanomolar ranges of actinomycin D concentrations)38. Several studies, using various cell types, have demonstrated that low concentrations of actinomycin D (1–10 nM) can inhibit cell proliferation and induce apoptosis; the p53 pathway has been shown to be involved in this effect39,40. Due to feedback loops, decreased rRNA synthesis (and impaired ribosome biogenesis) induced by low doses of actinomycin D can result in elevated transcription of p53 target genes, which trigger apoptosis30,41,42. Transcriptome analysis of two neural cell lines (EP1NS and SJ-BT57) treated with high doses (100 nM) or low doses (5 nM) of actinomycin D revealed significant differences in the abundances of many transcripts.
Some genes were downregulated after high-dose and upregulated after low-dose actinomycin D treatment; among the upregulated pathways after low-dose treatment were p53 pathway genes, such as the pro-apoptotic gene PUMA43. Our results indicate that even low (4 nM) concentrations of actinomycin D can inhibit transcription of non-ribosomal genes in early embryonic cells. On the other hand, consistent with the above results, transcription (or mRNA stability, or both) of some proapoptotic genes was stimulated by low concentrations of actinomycin D in blastocyst cells.
In the current study, the sensitivity of the human hybrid trophoblast cell line to apoptosis induction with actinomycin D was documented for the first time. However, the frequency of apoptosis in treated trophoblast cells did not reach the level recorded in the mouse trophectoderm cell line (130 apoptotic cells per 1957 cells vs 217 apoptotic cells per 4664 cells). This fact, together with the extremely low incidence of spontaneous apoptosis, suggests differences in the apoptosis machinery between human hybrid trophoblast cells and normal embryonic cells.
Still, regardless of the model used, in both the control and the actinomycin D groups, similar percentages of apoptotic cells were localised in the cytoplasm of other embryonic cells (approx. 80% in mouse blastocysts and 90% in human hybrid cells) and the majority of them displayed co-localisation with acidified organelles. The first hypothesis was confirmed: The increase in apoptosis was apparently accompanied by increased engulfment and digestion of apoptotic cells. Clearance capacity of non-professional embryonic phagocytes remained at the same level despite the fact that actinomycin D reduced overall transcription in blastocysts. However, using transcription inhibitor to induce apoptosis is also the key limitation of the study. Efficiency of embryonic efferocytosis in response to apoptosis elevation triggered by other inductors or environmental factors is remains to be evaluated.
The effect of inhibition of lysosomal acidification on embryonic efferocytosis
V-ATPase functions as a proton pump primarily located in cellular organelles, such as lysosomes, secretory vesicles, endosomes, and the plasma membrane. Responsible for regulating intracellular and extracellular pH levels, V-ATPases play crucial roles in various physiological processes, from membrane trafficking and autophagy to apoptosis, bone resorption, and sperm maturation20,21,22,23,24,25. Mutations in genes encoding V-ATPase subunits in yeast, Drosophila, and mice often result in lethality due to its widespread distribution and essential functions in cells44.
Studies have revealed the presence of acidic compartments generated by V-ATPase as early as the one-cell stage in mouse preimplantation embryos. As trophoblasts and the inner cell mass undergo differentiation at the blastocyst stage, these compartments demonstrate a distinct perinuclear distribution pattern45. Analysis of V-ATPase isoforms in mouse embryos collected at ED6.5 highlighted the expression of all four isoforms46. Similarly, investigations in bovine embryos at the 17th day of pregnancy, during the initial interaction with the uterine epithelium, showcased the expression of all three subunits of V-ATPase in the trophoblast47. Noteworthy findings from murine studies indicate the critical role of V-ATPase during the post-implantation period in establishing and maintaining apico-basolateral cell polarity within the embryonic epithelium46.
Bafilomycin A1 has been shown to be a potent inhibitor of cellular autophagy in previous studies. The indirect proof of the ability of bafilomycin A1 to disrupt autophagy in mouse embryos was also given: Aneuploid embryos from the early to late blastocyst stage were exposed to 160.6 nM bafilomycin A1 for 24 h, and such treatment reduced the number of dying cells (evaluated by SYTOX-viability assay) in the inner cell mass. However, the effect was not shown in diploid embryos21,22.
In the current study, we investigated the impact of V-ATPase inhibition by bafilomycin A1 on embryonic efferocytosis. Our results proved the ability of bafilomycin A to negatively affect the execution of terminal phases of efferocytosis in non-professional embryonic phagocytes: Bafilomycin A1 at 0.1 and 0.2 nM significantly increased the average percentage and overall proportion of LysoTracker-negative apoptotic cells. Similar to the study on aneuploid embryos26, the results of the current study suggest a more prominent effect of bafilomycin A1 on processes in the inner cell mass than in the trophectoderm cell line.
Interestingly, our results suggest that bafilomycin A1 is able to negatively affect even the early phase of embryonic efferocytosis – the engulfment process (as shown by a significant decrease in the proportion of internalised apoptotic cells and an increase in the proportion of extruded apoptotic cells). Since V-ATPase is widely distributed within the cell, bafilomycin A1 is probably not only specific as a lysosomal acidification inhibitor. Indeed, other effects were previously documented, including the blocking of the fusion of autophagosomes with lysosome48. Based on the findings in yeast, which suggest a direct role of the integral V domain of the ATPase in the process of membrane fusion of vacuoles that is independent of acidification, we might hypothesise that the drug blocked the fusion of membranes involved in the process of engulfing embryonic apoptotic cells by non-professional phagocytes24,49.
Mouse blastocysts exposed to bafilomycin A1 for 24 h showed a significantly decreased mean number of cells when compared to controls. Since the presence of a lysosomal acidification inhibitor did not cause a significant increase in apoptosis (Tables 7 and 10) or a significant decrease in cell proliferation (Supplementary Table S1), we can assume that the decrease in cell number was probably a consequence of the slowing down of blastocyst growth. Similarly, bafilomycin A1 inhibited the growth of a variety of cultured cells, including embryonic cells (golden hamster embryo), NIH-3T3 fibroblasts, PC12 cells and HeLa cells50. Consistent with our data, the growth-depriving effect was usually connected with nanomolar concentrations, e.g.: Bafilomycin A1 at 1 nM effectively suppressed cell growth of human paediatric B-cell acute lymphoblastic leukaemia cells cultured for 74h51; bafilomycin A1 at 5 nM inhibited growth of hepatocellular carcinoma cell lines cultured for 48h52, and bafilomycin A1 at 0.5 nM effectively inhibited human diffuse large B cell lymphoma cells cultured for 24h53.
Surprisingly, in the current study, bafilomycin A1 at concentrations equivalent to those used in blastocysts (from 0.01 to 0.2 nM) did not affect the proliferation or viability of human trophoblast cells. This suggests a lower sensitivity of the human hybrid cell line to the inhibitor. However, higher concentrations of bafilomycin A1 (2 nM) had a more harmful impact on both cell growth and cell viability of hybrid cells. This suggests that the wider number of V-ATPase functions might be disrupted in exposed cells at such concentrations.
To summarize, results obtained on blastocysts confirmed the second tested hypothesis: Selective inhibition of V-ATPase partially reduced activity of embryonic phagocytes. However, the decrease was not as profound as expected. The ability of embryonic cells to engulf and digest neighbouring dead cells appears to be fundamental process of early development which remains functional even at unfavourable conditions (disbalanced membrane transport of protons by bafilomycin A1, reduced transcription by actinomycin D).
Conclusions
Our results show that the ability of non-professional embryonic phagocytes to successfully execute all stages of efferocytosis is not limited by physiological conditions. The increase in the frequency of apoptosis induced by actinomycin D was accompanied by increased engulfment of apoptotic cells. V-ATPase is important for the execution of embryonic efferocytosis and its inhibition negatively affects both the engulfment and digestion of apoptotic cells. However, the specificity of bafilomycin A1 to inhibit V-ATPase seems to be relatively low.
Methods
Unless otherwise indicated, all chemicals were purchased from Merck, Darmstadt, Germany.
Animals and embryo recovery
Female mice (30–35 days old) underwent hormonal synchronisation using pregnant mare’s serum gonadotropin (eCG, 5 IU intraperitoneally) (Folligon, Intervet International, Boxmeer, Holland), followed 47 h later by human chorionic gonadotropin (hCG, 4 IU intrape classifi ritoneally) (Pregynal, Organon, Oss, The Netherlands). They were then mated with males of the same strain overnight. Successful mating was confirmed by identification of a vaginal plug on the following morning (Day 1 of pregnancy). On Day 4 of pregnancy (at 97 h post hCG administration), the fertilised mice were killed by cervical dislocation and their embryos isolated by flushing the uterus and oviducts using an in-house flushing-holding medium54 supplemented with 1% bovine serum albumin (BSA). The collected embryos underwent immediate cation using a Nikon SMZ 745 T stereoscope (Nikon, Tokyo, Japan); only those at the blastocyst stage were selected for further processing. Mouse blastocyst from blood-unrelated females were randomly divided into several subgroups and cultured in vitro under standard conditions (humidified atmosphere with 5% CO2 and 37 °C) in 400 μl of synthetic oviduct medium (EmbryoMax KSOM [potassium simplex optimised medium] with amino acids and D-glucose [Millipore, Darmstadt, Germany]) supplemented with 0.1% embryo culture tested BSA with/without the presence of actinomycin D (Sigma-Aldrich) at 4 nM concentration or bafilomycin A1 (Abcam) at 0.01 nM, 0.1 nM and 0.2 nM concentrations for 24 h.
Human trophoblast cell line Ac-1M88 culture
Ac-1M88 cells were purchased from a commercial cell bank (Leibniz-Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH, Braunschweig, Germany). The cell line was produced by the fusion of human normal invasive extravillous cytotrophoblasts from Chorion laeve with AC1-1, an HGPRT-negative mutant of the choriocarcinoma cell line JEG-355. The Ac-1M88 cells were cultured in vitro under standard conditions (humidified atmosphere with 5% CO2 and 37 °C) in DMEM (DMEM Glutamax F12, Gibco) with 10% FBS to 70% confluency56. Then they were cultured in fresh media with/without the presence of actinomycin D at 4 nM concentration or bafilomycin A1 at 0.01 nM, 0.1 nM, 0.2 nM, 0.5 nM and 2.0 nM concentrations for 24 h.
Analysis of internalisation of apoptotic cells in mouse blastocysts and the human Ac-1M88 cell line
The microscopic analysis of apoptotic cells internalisation was based on the combination of a TUNEL assay (terminal deoxynucleotidyl transferase dUTP nick end labelling) using a DeadEnd Fluorometric TUNEL System (Promega Corporation, Madison, USA; green labelling) with a method of visualising the plasma membranes via fluorescence staining of F-actin with phalloidin-TRITC conjugate (Santa Cruz Biotechnology, Inc., Santa Cruz, CA, USA; red labelling) (Figs. 2, 3).
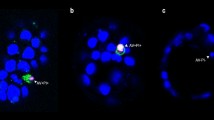
Analysis of apoptotic cell internalisation in mouse blastocysts. Illustrative images were obtained by confocal laser scanning microscopy (magnifcation × 400). Single optical sections of one treated blastocyst with 0.2 nM bafilomycin A (A) and two untreated blastocysts (B, C) are shown in the pictures. To visualise cell internalisation, fluorescence staining of F-actin in plasma membrane (red) was combined with TUNEL labelling (T+ , green) and Hoechst 33342 DNA staining (blue) (A1–C1): merge of all 3 channels, (A2–C2): merge of red and green channel). Abbreviations: i, internalised apoptotic nuclei (T+ nuclei in cytoplasm of embryonic cells with a normal nucleus and intact plasma membrane); e, extruded apoptotic nuclei (in blastocoele).
Analysis of apoptotic cell internalisation in human trophoblast cells. Illustrative images were obtained by confocal laser scanning microscopy (magnifcation × 400). Single optical sections one untreated monolayer of cell line AC-1M88 (A) and one monolayer of cell line AC-1M88 treated with 2 nM bafilomycin A (B) are shown in the pictures. To visualise cell internalisation, fluorescence staining of F-actin in plasma membrane (red) was combined with TUNEL labelling (T+, green) and Hoechst 33342 DNA staining (blue) (A1–B1): merge of all 3 channels, (A2–B2): merge of red and green channel). Abbreviations: i, internalised apoptotic nuclei (T+ nuclei in cytoplasm of embryonic cells with a normal nucleus and intact plasma membrane); e, extruded apoptotic nuclei (in blastocoele).
The in vitro cultured blastocysts and Ac-1M88 cells were fixed in 4% w/v paraformaldehyde in phosphate buffered saline (PBS) (Invitrogen Life Technologies) at room temperature for 10 min and stored in 1% w/v paraformaldehyde in PBS at 4 °C for up to one week, as needed. Fixed blastocysts and Ac-1M88 cells were washed in PBS containing 0.1% BSA or PBS, respectively, and transferred into PBS with 0.5% v/v Triton X-100. After 1 h permeabilisation, the blastocysts and the Ac-1M88 cells were washed and incubated in TUNEL assay reagents for 1 h at 37 °C in the dark57. After further washing in PBS with BSA or PBS, respectively, the embryos and the Ac-1M88 cells were transferred into a 3 μM solution of fluorescent phalloidin-TRITC (phalloidin tetramethylrhodamine B isothiocyanate) (Santa Cruz Biotechnology) prepared according to the manufacturer’s instructions. Labelling took 30 min and was performed at room temperature. After a final washing in PBS with BSA or PBS, respectively, the blastocysts and the Ac-1M88 cells were counterstained with Hoechst 33342 DNA (20 μl/ml in PBS with BSA, blue labelling) for 5 min at room temperature and mounted on glass slides3. As a positive staining control for each TUNEL assay, two to four randomly selected blastocysts were pre-incubated in 50 U/ml DNase I (Invitrogen Life Technologies, Karlsruhe, Germany) for 30 min at 37 °C, an enzyme inducing DNA nicks (Supplementary Fig. S1).
Analysis of co-localisation of apoptotic cells and acidified phagosomes in mouse blastocysts and human Ac-1M88 cell line
The microscopic analysis of apoptotic cells digestion was based on a combination of a TUNEL assay (green labelling) with fluorescence staining of phagosomes using a LysoTracker kit (pinkish labelling, Fig. 4). The LysoTracker Red DND-99 staining kit (Cell Signaling Technology, Inc., Danvers, MA, USA) was used to label acid organelles according to the manufacturer’s instructions. Briefly, in vitro cultured blastocysts and Ac-1M88 cells were washed in flushing-holding medium supplemented with BSA or in PBS, respectively, and incubated in 50 nM LysoTracker Red DND-99 diluted in flushing-holding medium or in PBS for 1 h at 37 °C3. Subsequently, the blastocysts and the Ac-1M88 cells were fixed in 4% w/v paraformaldehyde (Merck) in PBS for 10 min at room temperature and stored in 1% w/v paraformaldehyde in PBS at 4 °C until the next day. Fixed blastocysts and Ac-1M88 cells were washed in PBS with BSA or PBS, respectively, permeabilised for 1 h in 0.5% Triton X-100, incubated in TUNEL assay reagents for 1 h at 37 °C in the dark and subjected to further washing in PBS with BSA or PBS, respectively57. Finally, the blastocysts and the Ac-1M88 cells were counterstained with Hoechst 33342 (20 μl/ml in PBS) for 5 min at room temperature and mounted on glass slides. A positive staining control for each TUNEL assay was obtained as stated above.
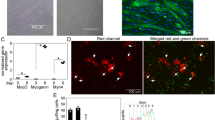
Analysis of co-localisation of apoptotic cells and acid organelles in mouse blastocysts and human trophoblast cells. Illustrative images were obtained by confocal laser scanning microscopy (magnifcation × 400). Single optical sections of blastocyst untreated blastocyst (A), treated with 0.2 nM bafilomycin A (B), untreated monolayer of cell line AC-1M88 (C) and monolayer of cell line AC-1M88 treated with 2 nM bafilomycin A (D) and are shown in the pictures. To visualise the acid organelles (lysosomes/phagosomes) in embryonic cells, LysoTracker labelling was used (lt+, red, A3–D3). To visualise apoptotic cells, staining was combined with TUNEL labelling (T+, green, A2–D2) and Hoechst 33342 DNA staining (blue) (A1–D1: merge). Abbreviations: T+lt+, TUNEL positive apoptotic nuclei (or their condensed fragments) showing co-localisation with acid organelles; T-lt+, TUNEL negative nucleus showing co-localisation with acid organelle; T+lt−, TUNEL positive apoptotic nuclei (or their condensed fragments) without the presence of acid organelle staining.
Analysis of cell proliferation in mouse blastocyst
Proliferation marker protein Ki-67 was immunohistochemically localized in blastocysts using a rabbit recombinant monoclonal antibody Anti-Ki67. Fixed blastocysts were washed in PBS with BSA, permeabilized 1 h in 0.1% Triton X-100. Non-specific immunoreactions were blocked by incubating the embryos in a solution containing 10% v/v normal goat serum (Santa Cruz Biotechnology) and 2% w/v BSA fraction V in PBS for 2 h at room temperature. After blocking, the blastocysts were incubated with primary antibody (Rabbit recombinant monoclonal antibody [SP6], 1:100 dilution) (Abcam, Cambridge, UK) diluted in blocking solution at 4 °C overnight. The next day, the blastocysts were extensively washed in the blocking solution and incubated with Cy™3 AffiniPure™ Goat Anti-Rabbit IgG secondary antibody (1:100 dilution) (Jackson ImmunoResearch Laboratories, West Grove, PA, USA) for 1 h at room temperature. Finally, the blastocysts were counterstained with Hoechst 33342 (20 μl/ml in PBS) for 5 min at room temperature and mounted on glass slides. Immunochemical negative control staining was performed by omission of either the primary or secondary antibody (Supplementary Fig. S2).
Microscopic analysis
All stained blastocysts and Ac-1M88 cells were mounted on glass slides with Vectashield (Vector Laboratories, Burlingame, CA, USA) and examined using a Leica TCS SPE confocal microscope (magnification × 400). Identification of apoptotic cells was based on an evaluation of the TUNEL-labelling and the nuclear morphology: a TUNEL-assay was used to track specific DNA fragmentation in the nucleoplasm of apoptotic cells, and Hoechst 33342 DNA staining was used to distinguish between normal chromatin organisation, chromatin condensation and karyorrhexis. Clusters of nuclear fragments taking up space comparable to the size of a normal nucleus were counted as one apoptotic nucleus. Nuclear fragments standing alone were considered remnants of apoptotic nuclei only when they were TUNEL-positive. TUNEL-positive nuclei localised within embryonic cells with a normal nucleus and intact plasma membrane were classified as internalised apoptotic cells. TUNEL-positive nuclei surrounded with intact plasma membranes from at least three neighbouring cells were classified as non-internalised apoptotic cells. TUNEL-positive nuclei (or nuclei showing chromatin condensation) in the blastocoele cavity or perivitelline space were classified as extruded apoptotic cells. TUNEL-positive nuclei co-localised with acid organelle were classified as digested apoptotic cells.
RT-PCR and transcript relative quantification
Total RNA was extracted from batches of 50 mouse blastocysts (the number of blastocysts in each pool was exactly determined), and from mouse brain or kidney (positive tissue control). TRIzol Reagent (Invitrogen Life technologies, Karlsruhe, Germany) was used for the extraction (according to the manufacturer’s instructions). Contaminating DNA in RNA preparations was digested with amplification-grade DNase I (Invitrogen Life Technologies). To correct differences in RNA recovery and loading of RT-PCR reactions 0.08 pg of luciferase (Luc) mRNA (Promega, Madison, WI) per blastocyst was added to the TRIzol lysis reagent before the RNA extraction.
The RNA was reverse transcribed with Superscript™ III RNase H− Reverse Transcriptase (Invitrogen Life technologies) using 4 μM anchored oligo dT primers and 1.5 μM random hexamer primers (Thermo Fisher Scientific, Epsom, UK). To check for the presence of genomic DNA contamination in the RNA preparations, reverse transcriptase negative controls (no reverse transcriptase in the reaction) were carried out in parallel using a portion of each RNA sample. The cDNA preparations were then cleaned by ethanol precipitation, and the cDNA pellets from blastocysts were diluted in an appropriate amount of 10 mM Tris (pH 8.3) so that 1 μl of the cDNA corresponded theoretically to 2.5 embryo/equivalents. Five independent cDNA preparations from control blastocysts and blastocyst treated with 4 nM Actinomycin D were used. To check for the presence of cross contamination, the reaction with water instead of cDNA was performed concurrently (blank reaction).
PCR amplifications were performed in a Light Cycler 480 real-time PCR system (Roche Diagnostics, Rotkreuz, Switzerland). The reactions were carried out in 25 μl volumes containing 1ul of the cDNA, SYBR Green qPCR mastermix (Qiagen) and commercial primer sets from Qiagen (product numbers: PPM02917E, PPM03410F, PPM03118E, PPM03562A, PPM31938A, PPM04997A, PPM02918F, PPM02920F, PPM02945B, PPM03717B). An initial step at 95 °C for 10 min was followed by 45 cycles at 95 °C for 15 s, 60 °C for 30 s, 72 °C for 20 s, and 77–87 °C for 20 s (acquisition of fluorescence). For amplification of luciferase, we used primers designed by Offenberg et. al (2005; primer sequences: 5′- GACGATGACGCCGGTGAACTT-3′, 5′-ACACAACTCCTCCGCGCAACT-3′). An initial step at 95 °C for 10 min was followed by 35 cycles at 95 °C for 15 s, 60 °C for 30 s, 72 °C for 20 s, and 81 °C for 20 s (acquisition of fluorescence). Amplification specificity was assessed with a melting curve analysis and agarose gel electrophoresis. The fluorescence data obtained from amplifications were transformed to the relative mRNA quantity values using the relative standard curve (created with serial dilutions of mouse brain or kidney cDNA)58. The expression of mouse genes was normalized with the external control (luciferase mRNA). The relative amounts of transcripts were divided by the average number of cells in control and actinomycin D-treated blastocysts to account for the significantly different cell numbers in the two groups of blastocysts.
Statistical analysis
Statistical analysis was performed using PRISM v.5.01@2007 (GraphPad Software Inc., La Jolla, CA, USA). To determine if data sets are well-modelled by a normal distribution, data were analysed using the D’Agostino-Pearson normality test. The differences between data showing normal Gaussian distribution were assessed using the unpaired Student t-test or ANOVA followed by Tukey’s post-hoc test. The differences between data which did not pass the normality tests were assessed using the Mann–Whitney test or the Kruskal–Wallis test, followed by Dunn’s post hoc test. The results are expressed as means ± standard error (S.E.M.), or as medians and interquartile range. To assess the differences between score-type data, standard chi-square tests with one degree of freedom were used, or the Fischer test. All details of the statistical analysis are given in the footnotes to the individual tables.
Data availability
No datasets were generated or analysed during the current study.
References
Pampfer, S. & Donnay, I. Apoptosis at the time of embryo implantation in mouse and rat. Cell Death Differ. 6, 533–545 (1999).
Huppertz, B. & Herrler, A. Regulation of proliferation and apoptosis during development of the preimplantation embryo and the placenta. Birth Defects Res. C Embryo Today 75, 249–261 (2005).
Pisko, J. et al. Apoptotic cells in mouse blastocysts are eliminated by neighbouring blastomeres. Sci. Rep. 11, 9228 (2021).
Nichols, J., Lima, A. & Rodríguez, A. T. Cell competition and the regulative nature of early mammalian development. Cell Stem Cell 29, 1018–1030 (2022).
Regin, M., Spits, C. & Sermon, K. On the origins and fate of chromosomal abnormalities in human preimplantation embryos: an unsolved riddle. Mol. Hum. Reprod. 28, 1–13 (2022).
Shahbazi, N. M. Mechanisms of human embryo development: From cell fate to tissue shape and back. Development 147, 1–14 (2020).
Kirk, J. M. The mode of action of actinomycin D. Biochim. Biophys. Acta 42, 167–169 (1960).
Goldberg, I. H., Rabinowitz, M. & Reich, E. Basis of actinomycin action. I. DNA binding and inhibition of RNA-polymerase synthetic reactions by actinomycin. Proc. Natl. Acad. Sci. U. S. A. 48, 2094–2101 (1962).
Reich, E. & Goldberg, I. H. Actinomycin and nucleic acid function. Prog. Nucleic Acid Res. Mol. Biol. 3, 183–234 (1964).
Arandjelovic, S. & Ravichandran, K. S. Phagocytosis of apoptotic cells in homeostasis. Nat. Immunol. 16, 907–917 (2015).
Hart, S. P., Dransfield, I. & Rossi, A. G. Phagocytosis of apoptotic cells. Methods 44, 280–285 (2008).
Lim, J. J., Grinstein, S. & Roth, Z. Diversity and versatility of phagocytosis: Roles in innate immunity, tissue remodeling, and homeostasis. Front. Cell Infect. Microbiol. 7, 191 (2017).
Skalko, R. G. & Morse, J. M. The differential response of the early mouse embryo to actinomycin D treatment in vitro. Teratology 2, 47–54 (1969).
Bell, P. S. & Glass, R. H. Development of the mouse blastocyst after actinomycin D treatment. Fertil. Steril. 5, 449–454 (1975).
Glass, R. H., Spindle, A. I. & Pedersen, R. A. Differential inhibition of trophoblast outgrowth and inner cell mass growth by actinomycin D in cultured mouse embryos. J. Reprod. Fertil. 48, 443–445 (1976).
Fabian, D. et al. Inhibitory effect of IGF-I on induced apoptosis in mouse preimplantation embryos cultured in vitro. Theriogenology 61, 745–755 (2004).
Fabian, D., Juhás, S., Il’ková, G. & Koppel, J. Dose- and time-dependent effects of TNFalpha and actinomycin D on cell death incidence and embryo growth in mouse blastocysts. Zygote 15, 241–249 (2007).
Fabian, D., Cikos, S. & Koppel, J. Gene expression in mouse preimplantation embryos affected by apoptotic inductor actinomycin D. J. Reprod. Dev. 55, 576–582 (2009).
Riley, J. K., Heeley, J. M., Wyman, A. H., Schlichting, E. L. & Moley, K. H. TRAIL and KILLER are expressed and induce apoptosis in the murine preimplantation embryo. Biol. Reprod. 71, 871–877 (2004).
Stransky, L., Cotter, K. & Forgac, M. The function of V-ATPases in cancer. Physiol. Rev. 96, 1071–1091 (2016).
Mauvezin, C. & Neufeld, T. P. Bafilomycin A1 disrupts autophagic flux by inhibiting both V-ATPase-dependent acidification and Ca-P60A/SERCA-dependent autophagosome-lysosome fusion. Autophagy 11, 1437–1438 (2015).
Wang, R. et al. Molecular basis of V-ATPase inhibition by bafilomycin A1. Nat. Commun. 12, 1782 (2021).
Cotter, K. et al. Activity of plasma membrane V-ATPases is critical for the invasion of MDA-MB231 breast cancer cells. J. Biol. Chem. 290, 3680–3692 (2015).
Forgac, M. Vacuolar ATPases: Rotary proton pumps in physiology and pathophysiology. Nat. Rev. Mol. Cell Biol. 8, 917–929 (2007).
Oot, R. A., Couoh-Cardel, S., Sharma, S., Stam, N. J. & Wilkens, S. Breaking up and making up: The secret life of the vacuolar H+ -ATPase. Protein Sci. 26, 896–909 (2017).
Singla, S., Iwamoto-Stohl, L. K., Zhu, M. & Zernicka-Goetz, M. Autophagy-mediated apoptosis eliminates aneuploid cells in a mouse model of chromosome mosaicism. Nat. Commun. 11, 2958 (2020).
Popov, Y. et al. Macrophage-mediated phagocytosis of apoptotic cholangiocytes contributes to reversal of experimental biliary fibrosis. Am. J. Physiol. Gastrointest. Liver Physiol. 298, 323–334 (2010).
Niknafs, B., Mehdipour, A. & Mohammadi Roushandeh, A. Melatonin improves development of early mouse embryos impaired by actinomycin-D and TNF-α. Iran. J. Reprod. Med. 12, 799–804 (2014).
Adeluola, A. A., Bosomtwe, N., Long, T. E. & Amin, A. R. M. R. Context-dependent activation of p53 target genes and induction of apoptosis by actinomycin D in aerodigestive tract cancers. Apoptosis 27, 342–353 (2022).
Donati, G., Peddigari, S., Mercer, C. A. & Thomas, G. 5S ribosomal RNA is an essential component of a nascent ribosomal precursor complex that regulates the Hdm2-p53 checkpoint. Cell Rep. 4, 87–89 (2013).
Liu, X. F. et al. Actinomycin D enhances killing of cancer cells by immunotoxin RG7787 through activation of the extrinsic pathway of apoptosis. Proc. Natl. Acad. Sci. U. S. A. 113, 10666–10671 (2016).
Morovic, M. et al. Maternally inherited rRNA triggers de novo nucleolus formation in porcine embryos. Zygote 26, 395–402 (2018).
Moura, M. T., de Sousa, R. V., de Oliveira Leme, L. & Rumpf, R. Analysis of actinomycin D treated cattle oocytes and their use for somatic cell nuclear transfer. Anim. Reprod. Sci. 109, 40–49 (2008).
Li, T., Liu, C., Zhen, X., Yu, Y. & Qiao, J. Actinomycin D causes oocyte maturation failure by inhibiting chromosome separation and spindle assembly. Biol. Reprod. 104, 94–105 (2021).
Fabian, D., Makarevich, A. V., Chrenek, P., Bukovská, A. & Koppel, J. Chronological appearance of spontaneous and induced apoptosis during preimplantation development of rabbit and mouse embryos. Theriogenology 68, 1271–1281 (2007).
Goodlad, A. R. Quantification of epithelial cell proliferation, cell dynamics, and cell kinetics in vivo. WIREs Dev. Biol. 6, 1–18 (2017).
Winking, H., Gerdes, J. & Traut, W. Expression of the proliferation marker Ki-67 during early mouse development. Cytogenet. Genome Res. 105, 251–256 (2004).
Perry, R. P. & Kelley, D. E. Inhibition of RNA synthesis by actinomycin D: Characteristic dose-response of different RNA species. J. Cell. Physiol. 76, 127–139 (1970).
Cortes, C. L. et al. Effect of low doses of actinomycin D on neuroblastoma cell lines. Mol. Cancer 15, 1–13 (2016).
Brodská, B., Holoubek, A., Otevřelová, P. & Kuželová, K. Low-dose actinomycin-D induces redistribution of wild-type and mutated nucleophosmin followed by cell death in leukemic cells. J. Cell. Biochem. 117, 1273–1481 (2016).
Choong, L. M., Yang, H., Lee, A. M. & Lane, D. L. Specific activation of the p53 pathway by low dose actinomycin D: A new route to p53 based cyclotherapy. Cell Cycle 8, 2810–2818 (2009).
Hölzel, M. et al. Defects in 18 S or 28 S rRNA Processing Activate the p53 Pathway. J. Biol. Chem. 285, 6364–6370 (2010).
Tzaridis, T. et al. Low-dose Actinomycin-D treatment re-establishes the tumoursuppressive function of P53 in RELA-positive ependymoma. Oncotarget 7, 61860–61873 (2016).
Horng, J. L. et al. Knockdown of V-ATPase subunit A (atp6v1a) impairs acid secretion and ion balance in zebrafish (Danio rerio). Am. J. Physiol. Regul. Integr. Comp. Physiol. 292, 2068–2076 (2007).
Sun-Wada, G. et al. Acidic endomembrane organelles are required for mouse postimplantation development. Dev. Biol. 228, 315–325 (2000).
Sun-Wada, G. H., Tabata, H. & Wada, Y. Vacuolar-type proton ATPase is required for maintenance of apicobasal polarity of embryonic visceral endoderm. Sci. Rep. 11, 19355 (2021).
Skinner, M. A., MacLaren, L. A. & Wildeman, A. G. Stage-dependent redistribution of the V-ATPase during bovine implantation. J. Histochem. Cytochem. 47, 1247–1254 (1999).
Yoshimori, T., Yamamoto, A., Moriyama, Y., Futai, M. & Tashiro, Y. Bafilomycin A1, a specific inhibitor of vacuolar-type H(+)-ATPase, inhibits acidification and protein degradation in lysosomes of cultured cells. J. Biol. Chem. 266, 17707–17712 (1991).
Peters, C. et al. Trans-complex formation by proteolipid channels in the terminal phase of membrane fusion. Nature 409, 581–588 (2001).
Ohkuma, S. et al. Inhibition of cell growth by bafilomycin A1, a selective inhibitor of vacuolar H(+)-ATPase. In Vitro Cell Dev. Biol. Anim. 29A, 862–866 (1993).
Yuan, N. et al. Bafilomycin A1 targets both autophagy and apoptosis pathways in pediatric B-cell acute lymphoblastic leukemia. Haematologica 100, 345–356 (2015).
Yan, Y. et al. Bafilomycin A1 induces caspase-independent cell death in hepatocellular carcinoma cells via targeting of autophagy and MAPK pathways. Sci. Rep. 6, 37052 (2016).
Li, F., Hu, Y., Hu, Y., Zhou, R. & Mao, Z. Bafilomycin A1 induces caspase-dependent apoptosis and inhibits autophagy flux in diffuse large B cell lymphoma (2021).
Lawitts, J. A. & Biggers, J. D. Culture of preimplantation embryos. Methods Enzymol. 225, 153–164 (1993).
Frank, H. G. et al. Cytogenetic and DNA-fingerprint characterization of choriocarcinoma cell lines and a trophoblast/choriocarcinoma cell hybrid. Cancer Genet. Cytogenet. 116, 16–22 (2000).
Seeling, T. et al. Glyoxalase 1 expression is downregulated in preimplantation blastocysts of diabetic rabbits. Reprod. Domest. Anim. 54, 4–11 (2019).
Babeľová, J. et al. In vitro exposure to pyrethroid-based products disrupts development of mouse preimplantation embryos. Toxicol. In Vitro 57, 184–193 (2019).
Čikoš, Š & Koppel, J. Transformation of real-time PCR fluorescence data to target gene quantity. Anal. Biochem. 384, 1–10 (2009).
Acknowledgements
The study was funded by the Slovak Research and Development Agency project APVV-22-0071 and by the Slovak Academy of Sciences project VEGA 2/0041/23. The authors thank Anne Navarrete Santos and Maria Schindler (Institute of Anatomy and Cell Biology Faculty of Medicine, Martin Luther University Halle-Wittenberg) for helpful advice with the cultivation of the Ac-1M88 cells, Dana Čigašová and Anna Olšavská for technical assistance, and David Lee McLean for English editing.
Author information
Authors and Affiliations
Contributions
J.P. performed the experiments, obtained images using CLSM, and wrote the manuscript, prepared figures. Z.Š., V.K. performed the experiments. A.Š. performed RT-PCR analysis. Š.Č. performed RT-PCR analysis, assisted with data interpretation, prepared figures. D.F. designed the work and critically revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
All experiments were performed in mice of the outbred CD-1 strain (Velaz, Prague, Czech Republic). Mice were maintained in plexiglass cages under standard conditions (temperature 22 ± 2 °C, humidity 55 ± 5%, 12:12 h light–dark cycle with lights on 6:00 a.m.), with free access to food and water. All animal experiments were approved by the Ethics Committee for Animal Experimentation of the Institute of Animal Physiology, and by the State Veterinary and Food Administration of the Slovak Republic, and were performed in accordance with Slovak legislation based on EU Directive 2010/63/EU on the protection of animals used for experimental and other scientific purposes. The study was carried out in compliance with the Animal Research: Reporting of In Vivo Experiments (ARRIVE) guidelines.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pisko, J., Čikoš, Š., Špirková, A. et al. Elimination of apoptotic cells by non-professional embryonic phagocytes can be stimulated or inhibited by external stimuli. Sci Rep 15, 5494 (2025). https://doi.org/10.1038/s41598-025-88800-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-88800-z