Abstract
N6-methyladenosine (m6A) is involved in the development of non-alcoholic fatty liver disease (NAFLD). Here, we aimed to investigate the effect of m6A methyltransferase METTL3 on liver damage in high-fat diet (HFD)-induced mouse model and hepatocyte damage treated with free fatty acid (FFA). Plasma lipid, lipogenesis, viability, and apoptosis were measured to assess injury. m6A methylation was evaluated using m6A dot blot, methylated RNA immunoprecipitation, dual-luciferase reporter assay, and RNA decay assay. The results indicated that METTL3 was highly expressed in the liver of HFD mice, which knockdown improved plasma lipid and reduced liver lipids. Additionally, silencing of METTL3 promoted cell viability, inhibited apoptosis, reduced lipid concentrations, and downregulated lipogenesis-related marker levels. Moreover, METTL3 promoted the m6A methylation of FAS and enhanced its stability. In conclusion, silencing of METTL3 attenuates the progression of NAFLD by FAS m6A methylation, suggesting that METTL3 may be a promising target for treating NAFLD.
Similar content being viewed by others
Introduction
Non-alcoholic fatty liver disease (NAFLD) is a metabolic syndrome arising from steatosis alone, steatosis with mild inflammation, to non-alcoholic steatohepatitis with rapid fibrosis1. Currently, NAFLD has a global prevalence of up to 25% and is the leading cause of liver fibrosis, cirrhosis, and primary liver cancer2. The diabetes epidemic is an important reason for the increased morbidity of NAFLD, which is present in more than 70% of patients with type 2 diabetes3. In addition, obesity is an important risk factor for the progression of NAFLD and its fibrosis4. Of note, the development of NAFLD can further promote the progression of these metabolic diseases. Lifestyle modification is the primary treatment for NAFLD. Besides, it is also considered feasible to regulate lipid metabolism, protect liver, and improve inflammation through drug therapy5,6. However, the guidelines7 recommend few treatments and do not apply to all patients. Therefore, it is still urgent to further elucidate the complex pathogenesis of NAFLD in order to provide a new strategy for the treatment.
N6-methyl adenosine (m6A) methylation is a common RNA modification in mammalian cells that affects RNA fate8. M6A methylation is involved in the development of diseases. In NAFLD, m6A methylation is dysregulated and regulates lipogenesis and plasma lipids9,10. Growing evidence has reported that methyltransferases (“writers”), including METTL3, METTL14, and WTAP, catalyzed m6A methylation. Inversely, demethylases (“erasers”) reversibly regulate m6A modification to remove m6A. Besides, the binding proteins (“readers”) decode m6A methylation11,12. METTL3 is the first discovered member of m6A “writer” that plays a critical role in diseases including NAFLD13. Thus, to understand the role of m6A methylation in the progression of NAFLD, there is an urgent need to establish a bridge between METTL3 and hepatic steatosis.
Fatty acids are key components of cell membranes and are essential for cell survival14. Excess glucose can be converted into fatty acids, preventing sugar toxicity, and providing energy reserves. Aberrant upregulation of de novo lipogenesis (DNL) is associated with multiple pathologies, involving the regulation of fatty acid synthase (FAS), ATP-citrate lyase (ACLY), and acetyl-CoA carboxylase (ACC)15. FAS, also known as FASN, is a homodimeric multi-enzyme in mammals with two multifunctional polypeptides, which is essential for fatty acid biosynthesis16. Previous studies have revealed that DNL is increased in patients with NAFLD, leading to lipid deposition in the liver, indicating that it may be a target to treat NAFLD17,18. Consistently, FAS expression is upregulated in the liver tissues of patients19. However, whether FAS can be modified by m6A is not clear.
In this study, we investigated the role of m6A methylation in NAFLD. Previous studies have reported that METTL3 expression is increased in the liver tissues of NAFLD mouse model, affecting lipid accumulation and macrophage functions10,13,20,21. Here, we also found that METTL3 expression was upregulated in HFD mice. Thus, we explored whether METTL3 affected hepatic steatosis and its m6A modification of FAS. This study provides a new idea for the pathogenesis of NAFLD.
Materials and methods
Establishment of NAFLD mouse model
The animal experiments were carried out according to the protocol approved by the ethics committee of Affiliated Hospital of Chengdu University. C57BL/6J mice (male, 7 weeks old, 20 ± 2 g) were purchased from Vital River (Beijing, China). All mice were kept in a 12 h light/dark cycle condition at 23–25 °C with free access to food and water. After one week of acclimation, all mice were randomly divided into six groups: control, high-fat diet (HFD), HFD + Ad-sh-NC, HFD + Ad-sh-METTL3, HFD + Ad-sh-METTL3 + Ad-NC, and HFD + Ad-sh-METTL3 + Ad-FAS, with six mice each group.
To establish the mouse model, the mice were fed HFD (60% energy from fat, HFK Bio-technology, Beijing, China) for 12 weeks as previously described22. The mice in the control group were fed normal diet (10% energy from fat, HFK Bio-technology) for 12 weeks. Then, mice were euthanized by intraperitoneally injection of 160 mg/kg pentobarbital sodium. The blood and liver were collected.
To explore the role of METTL3 and FAS in mice, adenovirus containing METTL3 short hairpin RNA (ad-sh-METTL3), adenovirus containing FAS (Ad-FAS), and their negative controls (ad-sh-NC, Ad-NC) were acquired from Genomeditech (Shanghai, China). The adenovirus (109 PFU) was injected into the mice through the tail vein every 2 weeks from the beginning of model establishment.
Plasma lipid measurement
Plasma was isolated from the blood by centrifuging at 3500 rpm for 10 min. Triglyceride (TG) and total cholesterol (TC) contents in the plasma of mice were measured using the TG and TCH assay kits (ERKN, Wenzhou, China) according to the manufacturer’s protocol, respectively.
Clinical samples
A total of 33 patients with NAFLD and 23 nonalcoholic steatohepatitis (NASH). They were diagnosed based on the pathological staging of liver biopsies. Additionally, 28 age and sex matched healthy individuals were enrolled as the heathyl control group. The blood samples were acquired from each participant. This study was approved by the Ethics committee of Affiliated Hospital of Chengdu University. Writtern informed consent was approved by all participants.
Cell culture
Primary hepatocytes were isolated from the liver tissues of mice (8–10 weeks old) as previously described23. The cells were maintained in Dulbecco’s modified eagle medium (DMEM) supplemented with 10% fetal bovine serum (FBS), 1% penicillin/streptomycin, and 2 mM glutamine at 37 °C with 5% CO2.
To induce lipid accumulation, the cells were incubated with 1 mM free fatty acid (FFA) mixture containing oleic acid and palmitic acid (2:1 ratio) for 48 h.
Cell transfection
Sh-METTL3 (target sequences: CCTCAGTGGATCTGTTGTGAT), sh-NC (CAACAAGATGAAGAGCACCAA), METTL3 overexpression vector (pcDNA3.1-METTL3), FAS overexpression vector (pcDNA3.1-FAS), and pcDNA3.1 were purchased from Genepharma (Shanghai, China). The vectors were transfected into hepatocytes in six-well plates using Lipofectamine 2000 reagent (Invitrogen, Carlsbad, CA, USA) in line with the manufacturer’s instructions for 48 h.
Quantitative real-time PCR (qPCR)
Total RNA was isolated from the liver of mice, serum of participants, and primary hepatocytes using a total RNA extraction kit (Solarbio, Beijing, China). Reverse transcription was carried out using the RT Master Mix for qPCR II (MedChemExpress, Monmouth Junction, NJ, USA). Then, the mRNA expression was quantified using SYBR Green qPCR Master Mix (MedChemExpress) on a CFX384 real-time PCR system (Bio-Rad, Hercules, CA, USA). The relative mRNA expression was calculated using the 2−ΔΔCt method by normalizing to GAPDH. The sequences of primers are listed in Table 1.
Hematoxylin and Eosin (HE) and oil red O staining assays
For HE staining, the liver tissues were fixed with 10% formalin and embedded in paraffin. Then, the paraffin Sect. (5 μm) were deparaffined and rehydrated. After staining with hematoxylin and eosin (Sigma-Aldrich, St. Louis, MO, USA) in turn, the results were viewed using a light microplate.
For Oil Red O staining, the liver tissues were embedded in optimum cutting temperature (OCT) to make frozen Sect. (10 μm). The sections were stained with 0.5% Oil Red O (Sigma-Aldrich) solution for 10 min. After decolorizing with 60% isopropanol and counterstaining with hematoxylin for 5 min, the results were captured under a light microplate.
For hepatocyte Oil Red O staining, primary hepatocytes cells were washed with PBS and fixed in 10% neutral formalin for 30 min. Then, the cells were stained with Oil Red O solution for 10 min. After decolorizing with 60% isopropanol, the cells were counterstained with hematoxylin for 5 min. The results were captured under a light microplate.
M6A dot blot
The RNA samples were heated at 95℃ for 3 min to destroy the secondary structure and then immediately frozen. The RNA was added to the nitrocellulose membrane and crosslinked using UV for 5 min. The membranes were washed with TBST and blocked with 5% skim milk. Afterwards, the membranes were incubated with anti-m6A at 4 °C for one night and probed with secondary antibody for 1 h. The dots were visualized using an excellent chemiluminescent (ECL) substrate detection kit (Elabscience).
Cell counting kit-8 (CCK-8)
Cell viability was measured using a CCK-8 kit (Dojindo, Kumamoto, Japan). The cells were seeded at the density of 2000 cells/well in 96-well plates. After incubation for 24 h, the cells were incubated with 10 µL of CCK-8 solution for 4 h. The absorbance was detected using a microplate reader (Bio-Rad).
Flow cytometry
Cell apoptosis was determined using flow cytometry using an Annexin V-FITC/PI cell apoptosis assay kit (YEASEN, Shanghai, China). Briefly, the cells were suspended in 100 µL binding buffer and incubated with 5 µL Annexin V-FITC and 5 µL PI solution for 15 min. After adding 400 µL binding buffer, cell apoptosis was analyzed using a FACSymphony flow cytometer (BD Bioscience, San Jose, CA, USA).
Western blotting
Cell samples were lysed using the RIPA lysis buffer containing protease inhibitors. Then, protein concentration was measured using a BCA protein colorimetric assay kit (Elabscience, Wuhan, China). Equal amounts of proteins were added in each lane to perform SDS-PAGE, followed by transfer onto PVDF membranes. Primary antibodies were incubated with the membranes at 4 °C for 12 h and probed with secondary antibody. The signals were viewed using an ECL substrate detection kit (Elabscience).
Methylated RNA immunoprecipitation (MeRIP)
The m6A expression of FAS was measured using a MeRIP m6A transcriptome profiling kit (Ribobio, Guangzhou, China). The isolated total RNA was fragmented. Additionally, anti-m6A antibody was incubated with magnetic beads A/G for 30 min. The fragmented RNA was incubated with anti-m6A beads at 4 °C for 2 h. After eluting, RNA was purified and qPCR was performed to measure the expression of mRNAs.
RNA immunoprecipitation (RIP)
The interaction between METTL3 and FAS was analyzed using an Imprint RIP kit (Sigma-Aldrich). The cells were lysed using RIP lysis buffer for 15 min. After centrifuging at 16,000 g at 4 °C, the supernatant was collected. Antibodies (anti-METTL3 and IgG) and magnetic beads were added to the lysate at 4 °C overnight. After washing the beads five times using lysis buffer, RNA was purified and FAS expression was measured using qPCR.
Dual-luciferase reporter assay
The m6A sites in FAS were predicted using the SRAMP database. The top 2 confidence sites, sites 1 and 2, were chosen. The sequences containing sites 1 and 2 were cloned into pGL3 basic vectors (Promega, Madison, WI, USA) to construct wild-type (WT) plasmids. To construct corresponding mutant (MUT) plasmids, adenosine (A) in the m6A motif was replaced by cytosine (C), and the sequences were cloned into pGL3 basic vectors. Hepatocytes were transfected with WT or MUT plasmids, pcDNA3.1-NC or pcDNA3.1-METTL3, and pRL-TK plasmids (Promega) using Lipofectamine 2000. After 48 h, the luciferase activity (firefly/Renilla ratio) was measured using a dual-luciferase reporter assay system (Promega).
RNA decay assay
The cells were cultured in six-well plates and incubated with 5 µg/mL Actinomycin D for 1, 4, 8, and 12 h. Total RNA was isolated from cells at each point time, and FAS expression was measured using qPCR.
Statistical analysis
Statistical analyses were conducted with GraphPad Prism 8.0 software (GraphPad, La Jolla, CA, USA). Data were represented as the mean ± SD. Quantitative data were compared using two-tail Student t-test or one-way analysis of variance (ANOVA). Statistical significance was defined as P < 0.05.
Results
METTL3 is highly expressed in HFD mice
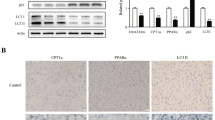
M6A methylation is a kind of epigenetic modification for gene expression that is commonly dysregulated in a variety of diseases, including NAFLD. A previous study revealed that aberrant m6A methylation promotes the development of NAFLD10. To confirm the role of m6A modification in NAFLD, we established the HFD mouse model. M6A levels were higher in the liver of HFD mice than that in the control mice (Fig. 1A), suggesting that m6A methylation may be involved in NAFLD progression. ALKBH5 and FTO are m6A “erasers”, and METTL3, METTL14, and WTAP are m6A “writers”. To explore which “eraser” or “writer” acts in NAFLD, their expression was measured using qPCR. The results of qPCR showed that ALKBH5 and METTL3 levels were increased in the liver of mice, compared with the control group. However, FTO, METTL14, and WTAP levels showed no significant difference between the control and HFD groups (Fig. 1B-F). Additionally, we also measured the protein levels of these m6A-related enzymes using western blotting. As shown in Fig. 1G, METTL3 levels were highr in the HFD group than that in the control group, while ALKBH5, FTO, METTL14, and WTAP levels had no obvious difference between these two groups. Therefore, METTL3 with the most significant difference was chosen for subsequent study. The expression of METTL3 was measured in patients with NAFLD, NASH, and healthy controls using qPCR. The results indicated that compared with the healthy group, METTL3 expression was increased in the serum of patients with NAFLD and NASH (Fig. 1H). These results suggest that METTL3 may be involved in the progression of NAFLD.
METTL3 is highly expressed in HFD mice. (A) m6A levels in the liver of mice were measured using m6A dot blot. (B) ALKBH5, (C) FTO, (D) METTL3, (E) METTL14, and (F) WTAP expression in the liver of mice were detected in qPCR. (G) Western blotting was performed to determine the protein levels of ALKBH5, FTO, METTL3, METTL14, and WTAP in the liver of mice. (H) METTL3 expression in the serum of healthy individuals, patients with NAFLD, and NASH was measured using qPCR. n = 6. NS no significance.
Knockdown of METTL3 alleviates HFD-induced hepatic damage
To explore the role of METTL3 in vivo, Ad-sh-METTL3 was injected into HFD mice, and Ad-sh-NC was injected as the negative control. After Ad-sh-METTL3 injection, METTL3 expression was downregulated in the liver of mice (Fig. 2A). Then, TC and TC were detected to evaluate plasma lipid. The results showed that TC and TG were increased in HFD mice, whereas METTL3 knockdown reversed this increase (Fig. 2B and C). Then, hepatic steatosis was evaluated using HE and Oil Red O staining assays. The results showed that HFD increased hepatic lipid droplets, which was partly abolished by METTL3 knockdown (Fig. 2D and E). Taken together, silencing of METTL3 improves elevated plasma lipid and hepatic steatosis in HFD mice.
Knockdown of METTL3 alleviates HFD-induced hepatic damage. (A) METTL3 expression was measured in the liver of mice after injecting adenovirus using qPCR. (B) TC and (C) TG levels in mice were measured using kits to assess plasma lipid. Representative images of lipid droplets in the liver that assessed using (D) HE staining and (E) Oil Red O staining. n = 6.
Knockdown of METTL3 inhibits FFA-induced hepatocyte damage
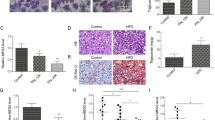
To further investigate the benefits of METTL3 knockdown in NAFLD, primary hepatocytes cells were treated with FFA to generate the cell model in vitro to mimic NAFLD. After sh-METTL3 transfection, the expression of METTL3 was decreased (Fig. 3A). Subsequently, cell viability was measured using CCK-8. The results indicated that FFA suppressed hepatocyte viability, whereas METTL3 knockdown reversed this suppression (Fig. 3B). Cell apoptosis was determined using flow cytometry. Inversely, cell apoptosis was facilitated by FFA, which was counteracted by interference with METTL3 (Fig. 3C and D). Later, the results of Oil Red O staining illustrated that the lipid concentration was elevated in FFA-treated cells, and METTL3 knockdown partly abrogated this elevation (Fig. 3E). SCD1, SREBP1, and FAS are well-known lipogenesis-related factors24. Therefore, their levels in cells were measured using qPCR and western blotting. As expected, the mRNA and protein levels of SCD1, SREBP1, and FAS were upregulated in FFA-treated cells, whereas sh-METTL3 reversed the upregulation caused by FFA (Fig. 3F and G). ACAT2 and HMGCR are associated with cholesterol absorption and biosynthesis, respectively25, and their levels were also measured using western blotting. FFA treatment increased ACAT2 and HMGCR levels in hepatocytes, while METTL3 knockdown abrogated this increase (Fig. 3G). In summary, interfering with the expression of METTL3 improves hepatocyte damage induced by FFA.
Knockdown of METTL3 inhibits FAA-induced hepatocyte damage. (A) The expression of METTL3 was measured using qPCR in sh-NC and sh-METTL3 transfected cells. Primary hepatocytes were exposed to FFA and transfected with shRNAs, and (B) cell viability was measured using CCK-8; (C) apoptosis was assessed using flow cytometry and (D) apoptosis rate was quantified; (E) lipid was observed using Oil Red O staining assay; (F) mRNA expression of SCD1, SREBP1, and FAS were detected using qPCR; (G) protein levels of ACAT2, HMGCR, SCD1, SREBP1, and FAS were measured using western blotting. n = 3.
METTL3 promotes m6A methylation of FAS
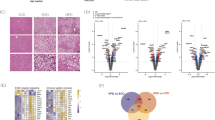
As an m6A “writer”, METTL3 promotes the m6A modification of RNAs. Hence, we first examined the effects of METTL3 overexpression on total m6A levels. METTL3 expression was elevated after pcDNA3.1-METTL3 transfection, compared with pcDNA3.1-NC (Fig. 4A). Then, m6A levels were increased after METTL3 overexpression (Fig. 4B). We performed MeRIP to measure the m6A levels of mRNAs related to lipid metabolism that were measured in our study, and found that overexpression of METTL3 increased m6A expression of FAS, but did not change the m6A levels of ACAT2, HMGCR, SCD1, and SREBP1 (Fig. 4C). Thus, FAS was chose as the target of METTL3 for the following experiments. Then, METTL3 was found to interact with FAS using RIP analysis (Fig. 4D). Later, the methylation sites in FAS were identified. Four sites in FAS could be modified by m6A (Fig. 4E). We chose the two sites with the very high confidence, and their sequences are shown in Fig. 4F. To confirm which sites can be methylated, we constructed WT and MUT reporter plasmids according to their sequences to perform dual-luciferase reporter analysis. The data showed that METTL3 increased the luciferase activity in the WT site 2 group, but failed to change the luciferase activity in the WT site 1 group (Fig. 4G and H), suggesting METTL3 promotes m6A methylation of FAS in site 2. Finally, the stability of FAS was measured. METTL3 enhanced the FAS mRNA stability (Fig. 4I). Based on these results, METTL3 promotes m6A methylation of FAS in site 2 and enhances FAS stability.
METTL3 promotes m6A methylation of FAS. (A) METTL3 expression was detected using qPCR following pcDNA3.1-NC and pcDNA3.1-METTL3 transfection. (B) Effect of METTL3 on m6A levels was assessed using m6A dot blot. (C) The m6A expression of ACAT2, HMGCR, SCD1, SREBP1, and FAS mediated by METTL3 was measured using MeRIP. (D) RIP was used to determine the interaction between METTL3 and FAS. (E) Potential m6A methylation sites in FAS were predicted. (F) The sequences of all potential modifing sites. Dual-luciferase reporter assay revealed the combined relationship between FAS containing (G) site 1 or (H) site 2 and METTL3. (I) Effect of METTL3 on the stability of FAS. n = 3. NS no significance.
FAS rescues the effects of METTL3 knockdown on hepatocyte damage
Afterwards, we performed rescue experiments to investigate the effects of METTL3 and FAS on hepatocyte injury. pcDNA3.1-FAS was transfected into hepatocytes, and FAS expression was upregulated (Fig. 5A). Cell viability, apoptosis, and lipid were analyzed using CCK-8, flow cytometry, and Oil Red O staining, respectively. The results indicated that silencing of METTL3 promoted viability, inhibited apoptosis, and reduced lipid concentration of hepatocytes, whereas overexpression of FAS abrogated these effects (Fig. 5B-E). The levels of ACAT2, HMGCR, SCD1, SREBP1, and FAS downregulated by METTL3 knockdown were reversed by FAS in FFA-stimulated cells (Fig. 5F and G). The results demonstrated that interfering with METTL3 inhibits injury of FFA-induced cells by decreasing FAS expression.
FAS rescues the effects of METTL3 knockdown on hepatocyte damage. (A) The expression of FAS was measured using qPCR in pcDNA3.1-NC and pcDNA3.1-FAS transfected cells. Primary hepatocytes were exposed to FFA and transfected with shRNAs and pcDNA3.1 plasmids, and (B) cell viability was measured using CCK-8; (C) apoptosis was assessed using flow cytometry and (D) apoptosis rate was quantified; (E) lipid was observed using Oil Red O staining assay; (F) mRNA expression of SCD1, SREBP1, and FAS were detected using qPCR; (G) Protein levels of ACAT2, HMGCR, SCD1, SREBP1, and FAS were measured using western blotting. n = 3.
FAS rescues the hepatic damage caused by METTL3 knockdown in HFD mice
Finally, the effects of METTL3 and FAS in vivo were explored. Following Ad-FAS injection into mice, its expression in the liver of mice was upregulated (Fig. 6A). The results of plasma lipid detection showed that METTL3 knockdown decreased TC and TG levels in HFD mice, whereas FAS overexpression abrogated this reduction (Fig. 6B and C). Lipid droplets in liver tissues were reduced by silencing of METTL3, which was rescued by FAS overexpression (Fig. 6D and E). The results demonstrated that silenced METTL3 inhibits hepatic steatosis in HFD mice by downregulating FAS expression.
FAS rescues the hepatic damage caused by METTL3 knockdown in HFD mice. (A) FAS expression was measured using qPCR in the liver of mice after injecting adenovirus. (B) TC and (C) TG levels in mice were measured using kits to assess plasma lipid. Representative images of lipid droplets in liver assessed using (D) HE staining and (E) Oil Red O staining. n = 6.
Discussion
Herein, we provided evidence to demonstrate that METTL3 mediated m6A methylation of FAS to promote NAFLD progress by regulating plasma lipid and lipid droplets. This study serves a potential target for NAFLD treatment.
M6A methylation is involved in the physiological and pathological processes of human diseases, such as neurodegenerative disease, cardiovascular diseases, and metabolic syndrome26. Interestingly, m6A methylation regulates mRNA metabolism to affect adipocyte differentiation and adipogenesis in adipose tissue expansion27. In NAFLD, several m6A-modified mRNAs are differentially expressed and participate in metabolism, transcription, and translation9. In HFD mice, differential m6A methylation is found to be related to lipid metabolism28. In the present study, m6A levels were increased in HFD mice, compared with control mice, suggesting m6A methylation is associated with NAFLD, which is consistent with previous studies10,20. M6A “writers” and “erasers” reversibly control m6A methylation. Thus, we measured their levels in HFD mice. METTL3 was upregulated by HFD in mice, consistent with previous research13. Then, METTL3 was chosen for further study. We also found that METTL3 expression was increased in patients with NAFLD and its serious form NASH. However, a previous study indicates that METTL3 expression is decreased in patients with NAFLD29, which seems to be at odds with our findings. This difference may be due to limitations in clinical sample collection or mouse sample size. Therefore, it is of great significance to explore the role of METTL3 in NAFLD. We further identified that knockdown of METTL3 inhibited lipid droplets and improved plasmid lipid in HFD mice, suggesting that METTL3 knockdown alleviates NAFLD.
METTL3 regulates NAFLD progression through multiple mechanisms. For example, Peng et al.13 report that downregulation of METTL3 facilitates autophagic flux in liver tissues, thereby regulating lipid metabolism to eliminate lipid droplets. Qin et al.30 found that METTL3 prevents the development of NAFLD by modulating macrophage metabolic reprogramming. Xu et al. study31 shows that FFA reduces METTL3 expression in AML12 cells, and silenced METTL3 aggravates steatosis in this cell line. These studies suggest that the role of METTL3 in NAFLD is contradictory. In the present study, we found that METTL3 knockdown promoted hepatocyte viability, inhibited apoptosis and lipid levels, and downregulated the levels of lipogenesis-related factors. The findings demonstrated that METTL3 contributes to promoting the progress of NAFLD. A previous study has indicated that overexpression of METTL3 promoted lipid production and thus facilitating the development of NAFLD10, consistent with our fundings.
METTL3 knockdown reduced FAS levels in FFA-treated hepatocytes. As an m6A “writer”, we subsequently investigated the m6A methylation of FAS mediated by METTL3. FAS has been confirmed to play an important role in lipogenic tissues, such as liver tissues. The expression of FAS in different tissues and developmental stages is differentiated and subject to transcriptional control and post-translational modification15. It regulates energy uptake and consumption, lipid deposition, and insulin sensitivity32. The expression of FAS in NAFLD is undoubtedly elevated in HFD mice and FFA-induced cells33,34. However, whether FAS can be modified by m6A is unknown. In this study, we found that METTL3 interacted with FAS and promoted m6A methylation of FAS. Several possible methylation sites in FAS were predicted. We verified that METTL3 promoted the methylation at site 2 rather than site 1. However, are there other sites on FAS that can be confirmed for m6A methylation? This will be further confirmed in our future studies. Importantly, m6A methylation regulates RNA splicing, translation, and stability35. Therefore, we analyzed the effects of METTL3 on the stability of FAS. The results showed that METTL3 enhanced FAS mRNA stability in hepatocytes. In summary, METTL3 promotes FAS m6A methylation, thereby enhancing its stability to increase FAS expression. Different from this study, a previous study has found that METTL3/METTL14 promotes m6A methylation of ACLY and SCD1 and up-regulates their expression levels to participate in lipid production10, indicating that the regulatory mechanism of METTL3 in NAFLD involves multiple target genes. Although our study and this previous study revealed different target genes, they all revealed an important role of METTL3 regulation in m6A modification of lipid metabolism in NAFLD. Furthermore, rescue experiments verified that METTL3 regulates hepatic steatosis and hepatocyte functions by positively regulating FAS, demonstrating that METTL3 promotes NAFLD by m6A methylation of FAS.
In conclusion, the present study demonstrated that silencing of METTL3 inhibits apoptosis and lipid of hepatocytes, and then improves steatosis and lipids by inhibiting m6A methylation of FAS, suggesting that downregulation of METTL3 expression decelerates the progression of NAFLD. The findings provide a strong theoretical basis for METTL3 to be a therapeutic target of NAFLD.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Pouwels, S. et al. Non-alcoholic fatty liver disease (NAFLD): a review of pathophysiology, clinical management and effects of weight loss. BMC Endocr. Disord. 22 (1), 63 (2022).
Powell, E. E., Wong, V. W. & Rinella, M. Non-alcoholic fatty liver disease. Lancet 397 (10290), 2212–2224 (2021).
Lee, C. H., Lui, D. T. & Lam, K. S. Non-alcoholic fatty liver disease and type 2 diabetes: an update. J. Diabetes Investig. 13 (6), 930–940 (2022).
Man, S. et al. Association between metabolically healthy obesity and non-alcoholic fatty liver disease. Hepatol. Int. 16 (6), 1412–1423 (2022).
Rong, L. et al. Advancements in the treatment of non-alcoholic fatty liver disease (NAFLD). Front. Endocrinol. (Lausanne). 13, 1087260 (2022).
Paternostro, R. & Trauner, M. Current treatment of non-alcoholic fatty liver disease. J. Intern. Med. 292 (2), 190–204 (2022).
EASL-EASD-EASO Clinical practice guidelines for the management of non-alcoholic fatty liver disease. J. Hepatol. 64 (6), 1388–1402 (2016).
Sendinc, E. & Shi, Y. RNA m6A methylation across the transcriptome. Mol. Cell. 83 (3), 428–441 (2023).
Li, Z. et al. Abnormal m6A modification in non-alcoholic fatty liver disease. Zhong Nan Da Xue Xue Bao Yi Xue Ban. 46 (8), 785–792 (2021).
Yang, Y. et al. Dysregulated m6A modification promotes lipogenesis and development of non-alcoholic fatty liver disease and hepatocellular carcinoma. Mol. Ther. 30 (6), 2342–2353 (2022).
Yang, B. et al. RNA methylation and cancer treatment. Pharmacol. Res. 174, 105937 (2021).
Garbo, S., Zwergel, C. & Battistelli, C. m6A RNA methylation and beyond - the epigenetic machinery and potential treatment options. Drug Discov. Today 26 (11), 2559–2574 (2021).
Peng, Z. et al. METTL3-m(6)A-Rubicon axis inhibits autophagy in nonalcoholic fatty liver disease. Mol. Ther. 30 (2), 932–946 (2022).
Paiva, P. et al. Animal fatty acid synthase: a chemical nanofactory. Chem. Rev. 121 (15), 9502–9553 (2021).
Batchuluun, B., Pinkosky, S. L. & Steinberg, G. R. Lipogenesis inhibitors: therapeutic opportunities and challenges. Nat. Rev. Drug Discov. 21 (4), 283–305 (2022).
Maier, T., Leibundgut, M. & Ban, N. The crystal structure of a mammalian fatty acid synthase. Science 321 (5894), 1315–1322 (2008).
Smith, G. I. et al. Insulin resistance drives hepatic de novo lipogenesis in nonalcoholic fatty liver disease. J. Clin. Invest. 130 (3), 1453–1460 (2020).
Heeren, J. & Scheja, L. Metabolic-associated fatty liver disease and lipoprotein metabolism. Mol. Metab. 50, 101238 (2021).
Higuchi, N. et al. Liver X receptor in cooperation with SREBP-1c is a major lipid synthesis regulator in nonalcoholic fatty liver disease. Hepatol. Res. 38 (11), 1122–1129 (2008).
Zhu, X. et al. N6-methyladenosine in macrophage function: a novel target for metabolic diseases. Trends Endocrinol. Metab. 34 (2), 66–84 (2023).
Li, Y. et al. M(6)a regulates Liver Metabolic disorders and Hepatogenous Diabetes. Genomics Proteom. Bioinf. 18 (4), 371–383 (2020).
Wu, Y. K. et al. Sulforaphane ameliorates non-alcoholic fatty liver disease in mice by promoting FGF21/FGFR1 signaling pathway. Acta Pharmacol. Sin. 43 (6), 1473–1483 (2022).
Charni-Natan, M. & Goldstein, I. Protocol for primary mouse hepatocyte isolation. STAR. Protoc. 1 (2), 100086 (2020).
Chen, X. et al. Long non-coding RNA AC012668 suppresses non-alcoholic fatty liver disease by competing for microRNA mir-380-5p with lipoprotein-related protein LRP2. Bioengineered 12 (1), 6738–6747 (2021).
Zheng, M. et al. Korean red ginseng formula attenuates non-alcoholic fatty liver disease in oleic acid-induced HepG2 cells and high-fat diet-induced rats. Heliyon 9 (11), e21846 (2023).
Jiang, X. et al. The role of m6A modification in the biological functions and diseases. Signal. Transduct. Target. Ther. 6 (1), 74 (2021).
Wang, X. & Wang, Y. From histones to RNA: role of methylation in Signal Proteins Involved in Adipogenesis. Curr. Protein Pept. Sci. 18 (6), 589–598 (2017).
Luo, Z. et al. Comprehensive analysis of differences of N(6)-methyladenosine RNA methylomes between high-fat-fed and normal mouse livers. Epigenomics 11 (11), 1267–1282 (2019).
Cheng, W. et al. New roles of N6-methyladenosine methylation system regulating the occurrence of non-alcoholic fatty liver disease with N6-methyladenosine-modified MYC. Front. Pharmacol. 13, 973116 (2022).
Qin, Y. et al. M(6)a mRNA methylation-directed myeloid cell activation controls progression of NAFLD and obesity. Cell. Rep. 37 (6), 109968 (2021).
Xu, R. et al. The methyltransferase METTL3-mediated fatty acid metabolism revealed the mechanism of cinnamaldehyde on alleviating steatosis. Biomed. Pharmacother. 153, 113367 (2022).
Paul, B., Lewinska, M. & Andersen, J. B. Lipid alterations in chronic liver disease and liver cancer. JHEP Rep. 4 (6), 100479 (2022).
Dong, R. et al. Yangonin protects against non-alcoholic fatty liver disease through farnesoid X receptor. Phytomedicine 53, 134–142 (2019).
Wang, X. Down-regulation of lncRNA-NEAT1 alleviated the non-alcoholic fatty liver disease via mTOR/S6K1 signaling pathway. J. Cell. Biochem. 119 (2), 1567–1574 (2018).
An, Y. & Duan, H. The role of m6A RNA methylation in cancer metabolism. Mol. Cancer 21 (1), 14 (2022).
Author information
Authors and Affiliations
Contributions
JX conceived the study; QL conducted the experiments; JX analyzed the data; QL was a major contributor in writing the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The study was approved by the Ethics Committee of Affiliated Hospital of Chengdu University. This study was performed in line with the principles of the Declaration of Helsinki. Informed consent was obtained from all individual participants included in the study. All experiments were performed in accordance with ARRIVE guidelines.
Consent for publication
All authors approved the final manuscript and the submission to this journal.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Q., Xiang, J. METTL3 promotes the progression of non-alcoholic fatty liver disease by mediating m6A methylation of FAS. Sci Rep 15, 6162 (2025). https://doi.org/10.1038/s41598-025-90419-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-90419-z
Keywords
This article is cited by
-
Epitranscriptomic modifications in programmed cell death: mechanistic insights and implications for liver diseases
Cellular & Molecular Biology Letters (2025)