Abstract
A carbon nanotube (CNT) can affect biological systems, ranging from toxicity to changes in functionality. Here, a series of long-scale (1–2 µs) molecular dynamics simulations were conducted to investigate the adsorption and interaction of lysozyme with the CNT, a possible mechanism for altering protein flexibility and function. Four systems were examined: native lysozyme/CNT, denatured lysozyme/CNT, and both systems post-docking. Our results indicate that native lysozyme does not undergo conformational changes when initially captured by a CNT. However, after docking, the native lysozyme/CNT complex exhibits conformational changes. In contrast, the denatured lysozyme binds more effectively to the CNT in both pre- and post-docking scenarios. Key amino acid residues, arginine and tryptophan, have been identified as crucial for lysozyme/CNT interactions. The surface of the CNT adsorbs lysozyme through π-π stacking and van der Waals interactions, with these multimodal interactions serving as the main driving force for protein anchoring to the nanotube. These results also underscore the significance of docking in the simulation of protein/nanoparticle interactions, which can lead to entirely different conclusions regarding, for example, the toxicity or functionality of a given nanoparticle life.
Similar content being viewed by others
Introduction
Carbon nanotubes (CNTs), especially with complex proteins, have a wide range of applications, from nanotechnology to medicine1,2,3,4,5,6. The practical viewpoint of the CNT/protein complex system is the ability to tune the physical and chemical properties of CNT, especially the negative attributes of CNT, such as toxicity and the absence of biodegradability. Due to their unique needle-like shape, the tunable CNT/protein complexes have been mainly functionalized to enable and solve cell penetration issues as a delivery agent3. Depending on the protein, the mentioned complex can be endowed with more biocompatibility options7, as well as can change toxicity to be non-toxic or less toxic8,9, i.e., “protein corona” modulates the toxicity of CNTs.
The studies of protein-functionalized CNT systems have been intensively reported during the last decade using experimental10,11,12,13,14,15 and computational8,16,17,18,19,20,21 methods. Beyond the importance of main parameters, like pH14,15,22, temperature23, and protein concentration24, aspects such as protein denaturation should also become important. The denatured protein with a CNT effective covering is displayed as the best coating agent18. However, the dynamic behavior of denatured protein/nanotube complex and the covering issues at the atomistic level must be better understood. In this context, the molecular dynamics method is an excellent tool to help us understand the insight of covering effectiveness issues and will answer some critical questions.
In this paper, we study the effect of lysozyme adsorption (native and denatured) on a single-walled CNT at the atomistic level. As a water-soluble protein, the chicken egg white lysozyme is selected. The globular protein—lysozyme is found in many tissues endowed with key functions and has attracted much attention due to its antimicrobial properties25. We have discussed some important aspects concerning the coverage of nanotubes and their effectiveness in binding. In our study, we have calculated several key parameters to understand the interaction between lysozyme and single-walled carbon nanotubes (SWCNTs). These parameters include Root Mean Square Deviation (RMSD), Root Mean Square Fluctuation (RMSF), Solvent Accessible Surface Area (SASA), time evolution of secondary structure, radius of gyration (\({R}_{g}\)), binding lifetime, and the distance between the residues and SWCNT carbons. These calculations provide a comprehensive view of how the interaction with CNTs can evolve the structure of lysozyme.
Modeling setup
All MD simulations were carried out using the GROMACS software package26 (2020 version) with GPU support27.
The crystal structure of hen egg-white lysozyme was taken from the protein data bank (ID: 1LYZ)28. The already denatured lysozyme PDB was taken from our previous simulation29. The protein consists of 129 residues with two structural domains—α and β. The structure is stabilized via four disulfide bridges. The five helices and 310-helix (81–85) are inside of α-domain: Helix 1 (5–14), Helix 2 (25–35), Helix 3 (90–100) and Helix 4 (110–115). The β-domain covers two β-sheets: Beta-strand 1 (43–46) and Beta-strand 2 (51–54).
An armchair single-walled carbon nanotube (SWNT) with chirality (n = 10, m = 10) was constructed using the CHARMM GUI Nanomaterial modeler30 plugin. The SWNT was used in two simulations, with a diameter of 13.6 Å and a tube length of 61.5 Å.
Two independent systems were created: System 1—Native Lysozyme/SWCNT in water and System 2—Denatured Lysozyme/SWCNT in water. Two systems were constructed using the CHARMM GUI Multicomponent Assembler31 plugin, which allows the creation of complex models with many components (merging .PSF and CHARMM coordinate .CRD files). We set an initial distance of nanotube-protein about 1 nm to ensure no interaction between protein atoms and the nanotube carbons. Two systems were solvated in water with a volume of 10 × 10 × 10 nm3. To neutralize the system, additional chloride and 150 mM NaCl were added. The minimization/equilibrium procedures were done for two systems according to CHARMM GUI standards, and the production runs (1000 ns for each) were carried out at NPT ensemble.
Additionally, we utilized the AutoDock Vina package32 to generate docked versions of the two systems: System 3—Native Lysozyme/SWCNT in water, and System 4—Denatured Lysozyme/SWCNT in water. This approach enabled us to predict more effective binding sites and affinities of lysozyme on the SWCNTs. The initial configurations obtained from the docking simulations were further refined through molecular dynamics simulations, providing a more accurate representation of the interaction dynamics between lysozyme and SWCNTs. The docking parameters are: the volume with X, Y, and Z dimensions set as 50 Å × 50 Å × 50 Å(size_x, size_y, size_z), the center of grid box was set to 5.033 Å, 26.243 Å, and 36.061 Å for center_x, center_y, and center_z, the maximum number of binding modes, which implemented via num_modes option, was fixed to 6, the exhaustiveness of search was set to 32 and the rest of the docking parameters were set at default values. The outcome of docking was the binding affinity values (in kcal/mol) and root-mean-square deviation (RMSD) lower and upper bounds. Further, the most significant negative value of affinities was selected for all series of runs.
The interactions between the protein, SWCNT, and the ions were modeled with CHARMM36m force-field33, and the TIP3P34 was used to model water molecules. The room temperature (300 K) and standard pressure were maintained via Nose–Hoover thermostat35 and Parrinello-Rahman barostat36, correspondingly, and the temperatures of each component were independently controlled. The bonds were constrained using the LINCS37 algorithm. For electrostatic interactions, the Particle-mesh Ewald (PME)38 algorithm was used with a cut-off of 1.2 nm. The same cut-off of 1.2 nm was used for Van der Waals interactions. The decay of the interaction potential was approximated using the switch method. Analysis was performed on the last 100 ns of the trajectory where the systems are in equilibrium.
All simulations were performed on Armenian e-infrastructure computational resources39. The snapshots and visualizations were generated using the VMD graphical package40.
Results and discussion
Secondary structures
First, the change in secondary structure of the protein throughout the trajectory was calculated. In Fig. 1, the secondary structure for each run is shown. The secondary structure analysis was done using the Kabsch and Sander method implemented in the do_dssp module provided by GROMACS.
Time evolution of secondary structure on residue sequence number (a-top)-System-1—Native Lysozyme/SWCNTand (a-bottom) System-2 Denatured Lysozyme/SWCNT without docking process. In (b) cases, the same is shown for both cases after the docking procedure, i.e. (b-top) System 3 Native Lysozyme/SWCNT (2 µs), and (b) System-4 Denatured Lysozyme/SWCNT (1 µs). The GROMACS do_dssp module assigns the secondary structure.
Examining the secondary structure of native lysozyme and visual inspection of trajectories in the undocked case (System 1) shows that SWCNT does not lead to a more significant disruption of the protein’s secondary structure elements (Fig. 1—a-top). Just minor fluctuations are observed in the helical region (Helix 4—(110–115)), while other areas remain stable throughout the production run, i.e., the structural content does not undergo significant changes (~ 1–5% loss of structural content). Thus, it can be argued that denaturation does not occur, meaning that the secondary structure remains unchanged when the protein is adsorbed onto SWCNTs. Several groups have experimentally verified this hypothesis14,15,17,18,24.
In the denatured lysozyme/SWCNT case, we track that the protein remains denatured throughout the simulation run. We used the protein from our previous simulation as a starting configuration, where the protein undergoes complete denaturation via surface active agents. However, small helical parts (5–14) can be seen in the α-domain, which is almost stable until the end of the simulation run. In addition to some helical parts in the structure, the random coils bend and turn dominantly.
The same calculations for native lysozyme/SWCNT in System 3 (docked) revealed some similarities compared to System 1 but a few notable changes. Similar to System 1, multiple well-defined horizontal bands indicate stable structural elements, indicating that the pattern remains relatively consistent throughout the 2000 ns simulation—a higher density of blue and yellow bands suggesting maintained alpha-helices and beta-sheets. The docking simulations indicated that the interaction with SWCNTs led to more significant disruptions, but it was impossible to observe in a one µs time scale. Still, it was revealed by extending the simulation to 2 µs. Visual inspection of the trajectories showed increased fluctuations, a subtle but noticeable shift in the secondary structure pattern, and a change in the density and continuity of some bands (see below). These changes suggest that the docking process induces partial denaturation, altering the stability and conformation of the lysozyme when adsorbed onto SWCNTs.
By comparing the DSSP calculation results in Fig. 1 a-bottom and b-bottom, we found that the overall patterns of secondary structure disruption and unfolding are similar in the docked and non-docked denatured cases. However, the non-docked denatured state shows slightly more persistent secondary structure elements, with fewer gaps and discontinuities in the bands. The transitions between secondary structure types seem slightly less abrupt in this case.
Overall, the longer simulation time (2000 ns) for the native state shows that the protein maintains its structure better, while the denatured state (1000 ns) shows significant structural changes even in the shorter time frame. This indicates the protein’s unfolding process and loss of native conformation.
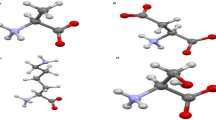
Figures 2 and 3 show snapshots of non-docked and docked systems extracted from the starting and ending time points to visualize our ongoing processes. Comparing these two sets reveals that the initial configuration in docked cases interacts more with SWCN than their corresponding non-docked cases. The final configurations of both denatured cases (Systems 2 and 4) are almost the same, but the final snapshots of the docked ones (Systems 1 and 3) are different.
Non-docked systems; snapshots extracted from the first (a) and last (b,c) configuration System-1—Native Lysozyme/SWCNT and from the first (d) and last (e,f) configuration System-2—Denatured Lysozyme/SWCNT. For clarity, the different views (top and side views) for last configuration are shown. The snapshots were designed with VMD40.
The docked systems; snapshots extracted from the first (a) and last (b,c) configuration System-3—Native Lysozyme/SWCNT and from the first (d) and last (e,f) configuration System-4—Denatured Lysozyme/SWCNT. For clarity, the different views (top and side views) for last configuration are shown. The snapshots were designed with VMD40.
To create a clearer picture of the evolution of each system, the radius of gyration of proteins in each case with respect to time is shown in Fig. 4. To understand lysozyme evolution in native and denatured cases and to find the effect of SWCN, we also calculated this parameter for the system in water without the CNT. The top figure shows a more fluctuating nature of denatured lysozyme (red) rather than the native one (blue) in water, indicating more fluctuating features of denatured rather than the native one, with the average \({R}_{g}\)~ 1.5 and 1.4 nm, respectively. The data for System 1 support the stability of lysozyme in the presence of SWCNT; for System 1, nearly 1.4 of already denatured protein displays large fluctuations until ~ 300 ns and remains more stable until the end of the production run. The stabilization is because the nanotube completely captures the protein; the value is estimated at ~ 1.8 nm. The same trend can be observed in docked systems (Systems 3 and 4), at least up to 1000 ns. However, we had some extra fluctuation in the native lysozyme case near 1000 ns, which led us to extend System 3’s simulation run to 2000 ns (see below).
Next, the adsorption of protein on SWCNT was examined. We have calculated the number of residues in close contact with SWCNT atoms to characterize the binding affinity. Note that the mentioned parameter was calculated via GROMACS g_rdf and g_mindist modules. As a close contact, we assumed that the residues/atoms are within 0.5 nm cut-off from SWCNT atoms20,21. The distance of protein residues from SWCNT and the number of residues in close contact with SWCNT is monitored in Fig. 5 for System 1.
The lysozyme is adsorbed onto SWCNT, and the amount of protein residues attached to SWCNT is nearly 14, as seen in Fig. 5 (the residues in the region < 0.5 nm). Note that the adsorbed residues are in the α structural domain, including the turns, bends, and partial Helix 4. We found [as demonstrated in Fig. 5 (below plot)] four arginines (ARG61, ARG68, ARG73, and ARG112), three tryptophans (TRP62, TRP63, and TRP111), three asparagines (ASN103, ASN106 and ASN113), a proline (PRO70), a threonine (THR69), an alanine (ALA107) and a lysine (LYS116) are strongly binds to the SWCNT. We divide the adsorbed amino acids into four binding sites (BSs) for protein onto SWCNT: BS1 includes ARG61, TRP62, and TRP63, BS2 consists of ARG68, THR69, PRO70, and ARG73, BS3—ASN103, ASN106, and ALA107. The residues denoted as TRP111, ARG112, ASN113 and LYS116 cover BS4. The mentioned amino acids and binding sites are the same as those reported by other authors17,19,41,42.Based on distance calculations, we argue that in native protein mode, three amino acids (ARG, TRP, and ASN) are identified as key units in the binding process, and particularly, one should argue that the mentioned residues are located on the surface of theprotein (see Fig. 6). In overall, we track that 50% of all tryptophans, ~ 37% of all arginines, and 22% of all asparagines are involved in the binding process.
To understand more about the binding lifetime and distance between the residues and SWCNT carbons, we have calculated the minimum distances of some residues of the protein and the nanotube. The latter was done using GROMACS g_mindist module. The curves are provided in Fig. 6, where the distances of three key residues were identified, i.e. ARG112, ASN106 and TRP62. As we see, the adsorption process occurs very fast and after a couple of nanoseconds, the residues are trapped and stay bound till the end of production run. Note that the average distances are estimated to be at the range of ~ 0.25–0.32 nm.
It is known that the Helix-Loop-Helix (HLH) or α-helical hairpin (residues 89–116) located at the upper lip of the active site of molecule, in terms of motif positioning, plays a crucial role in biological functions of protein and serves as a key element in the mechanism of bactericidal action42,43,44. We argue that the binding aminoblocks are in the helical hairpin, i.e., the HLH domain serves as an anchor to make a “protein corona.” Examining the dynamical properties of lysozyme, Artymiuk and coauthors45 show the high displacement inherent with the active site of lysozyme. They also show that the six-membered part of the indole ring of TRP63 and TRP111 has a larger displacement than other atoms45. A carbon nanotube and lysozyme’s tryptophan residue exhibit favorable π–π interaction, facilitating effortless non-covalent functionalization46. The extensive MD simulations performed by Calvaresi and coauthors17 shows the adsorption favorite sites and determining the binding energy, they argue that ARG21, ARG112 residues have strong binding values, as well as other residues (denoted as Asn27, Asn103, Asn106, Lys116, and Trp111) have a major contribution to the binding. Note that the high-propensity close contacts with the arginine residues (denoted as ARG112, ARG21) were reported when considering the lysozyme with sulfonate-based surfactants29.The role of arginine in protein-nanotube interaction is recently reviewed by Wu and coworkers19. They show that arginine residues, which are evenly distributed on the protein surface, play an important role in binding to the surface of single-walled CNT. Four arginines, denoted as ARG21, ARG68, ARG112, and ARG128, established close contact of the lysozyme with CNT. Besides arginine, the authors show that the prolines (PRO70) also quickly make fleeting contact with CNT42. The guanidinium group of arginine predominantly contributed to the binding process by frequently engaging the favorable interaction with the sidewall of single-walled carbon nanotubes47.
When discussing the height of the binding protein, one can mention that it is about ~ 3 nm (see Fig. 5), which is almost the same as the diameterofthe lysozyme. This is obvious as the protein undergoes no structural changes and remains stable.
Examining the binding affinity and contacts in the case of System 2, we have calculated the same parameter as in the case of System 1. First, we have provided all amino acid distances from the carbon nanotube and the number of residues in close contact with SWCNT. The criterion was the atoms within a 0.5 nm cutoff from SWCNT carbons. As demonstrated in Fig. 7, we found seven arginines (ARG5, ARG61, ARG68, ARG112, ARG114, ARG125, and ARG128), six alanines (ALA31, ALA42, ALA82, ALA90, ALA95, and ALA110), five threonines (THR40, THR43, THR47, THR51, and THR69), five serines (SER36, SER50, SER85, SER86, and SER91), four asparagines (ASN37, ASN44, ASN59, and ASN113), four glycines (GLY49, GLY71, GLY117, and GLY126), four cysteines (CYS30, CYS94, CYS115, and CYS127), four tryptophans (TRP28, TRP62, TPR111, and TRP123), three aspartic acids (ASP48, ASP52, and ASP57), three leucines (LEU56, LEU83, and LEU84), two valines (VAL29 and VAL109), two lysines (LYS33 and LYS116), a tyrosine (TYR53), an isoleucine (ILE88), a phenylalanine (PHE34), and a proline (PRO70) firmly attached to the SWCNT. We see that in the case of System 2, where the protein was already denatured, the lysozyme effectively coats a CNT. In the binding process, the amino acids participate in the following ratio when we take into account all the amino acids: THR-72%, TRP–67%, ARG-64%, ALA-50%, SER-50%, ASP-44%, LEU-37%, VAL-34%, LYS-34%, CLY-32%, ASN-29%.
Unlike the native form, approximately 46 residues are involved in the adsorption process in the denatured case (System 2). In contrast, the number of amino acids in the native case is nearly three times smaller, suggesting that denatured lysozyme binds more efficiently than its native counterpart18 (see Fig. 8). As mentioned, the thickness of the protein layer in System 1 is the same as the lysozyme diameter (~ 3 nm). Meanwhile, in System 2, we observe that the thickness of the protein layer is less than 2 nm, as shown in Fig. 8 case).
To support these interpretations, we calculated the RMSD and RMSF for all systems, as shown in Figs. 9 and 10. The RMSD analysis reveals structural instability increases with system complexity, moving from pure water to CNT environments. In pure water (Fig. 9—top), the native lysozyme maintains relatively stable RMSD values around 0.2 nm, while the denatured form exhibits higher but consistent fluctuations around 0.9 nm. When CNT is introduced without docking (Fig. 9—middle), both forms show increased RMSD values and more fluctuations, with the denatured form demonstrating exceptionally high values of 1.2–1.6 nm, suggesting that the mere presence of CNT affects protein stability. The most dramatic changes occur after docking to CNT (Fig. 9—bottom), where both forms display significant structural perturbations. The native form shows a gradual increase in RMSD over time, reaching approximately 0.4 nm, while the denatured form exhibits large fluctuations between 0.8 and 1.2 nm. This indicates that direct interaction with CNT substantially impacts the protein’s structural integrity, particularly affecting the stability of the native state. In summary, the RMSD plots demonstrate that the docking of lysozyme to CNT significantly increases structural flexibility and destabilization in both forms, with a more pronounced effect on the native structure.
In pure water, the RMSF values show (Fig. 10—top) a clear distinction between native and denatured states, with the native form exhibiting consistently low fluctuations (~ 0.1–0.2 nm) for most residues, while the denatured form shows higher flexibility (0.2–0.4 nm) with several peaks indicating locally increased mobility. When CNT is included without docking (Fig. 10—middle), both forms maintain similar patterns but with increased fluctuations, particularly in the denatured state. However, after docking (Fig. 10—bottom), the RMSF profiles change dramatically: the native form develops significant peaks around residues 50, 75, and 125 (reaching up to 0.3 nm), while the denatured form shows even more pronounced fluctuations (up to 0.8 nm), especially in the C-terminal region. This progressive increase in local flexibility, particularly in the specific areas, suggests that CNT interaction enhances overall protein mobility and induces region-specific structural perturbations, with the effect being most pronounced in the docked state.
To show how this drastic change has been undergone for System-4, we calculate the SASA, as shown in Fig. 11. Looking at the SASA plot in conjunction with the previous RMSD and RMSF data (Figs. 9 and 10) provides an interesting correlation in the structural changes of native lysozyme interacting with CNT: The SASA plot shows a gradual increase over time, starting from around 70–75 nm2 and eventually reaching higher values of around 80–85 nm2 (with a notable peak near 1500 ns reaching ~ 94 nm2). This increment in SASA aligns well with the RMSD data, which showed progressive structural deviation in the native state after CNT docking (reaching ~ 0.4 nm). The RMSF results revealed increased local flexibility in specific regions. The parallel increase in SASA suggests that the interaction with CNT is causing the protein20 to partially unfold or adopt more extended conformations, exposing more surface area to the solvent. This is particularly evident after 1250 ns, where we see a significant jump in SASA values, indicating a substantial conformational change that correlates with the higher RMSF values observed in specific residues (especially around residues 50, 75, and the C-terminal region) and the gradually increasing RMSD trend. Together, these measurements paint a consistent picture of CNT-induced structural perturbation in the native lysozyme, leading to a more loosened and solvent-exposed structure over time.
A more precise visualization of System 4 is shown in Fig. 12. We capture a critical conformational transition of native lysozyme interacting with a carbon nanotube (CNT) surface over a brief period (1440–1467 ns). Initially, at 1440 ns, the protein maintains its native structure while making localized contact with the CNT surface. At 1463 ns, a significant conformational change begins as the protein reorients itself, leading to a dramatic structural reorganization by 1467 ns. During this transition, the protein’s interaction with the CNT surface evolves from a localized contact point to a more extensive surface interaction, as indicated by the dashed lines in both the top and bottom views. While the protein undergoes this spatial reorientation and structural change, it maintains some key secondary structural elements (α-helices shown in purple and β-sheets in yellow), though in a reorganized configuration. This visualization provides clear structural evidence for the changes previously observed in the DSSP plots (Fig. 2) around the 1500 ns time point, demonstrating how the protein-CNT surface interaction drives this conformational adaptation.
Conclusion
In this study, we conducted an extensive set of long-timescale simulations on lysozyme/carbon nanotube (CNT) systems under four different conditions: native and denatured lysozyme; each examined both with and without the use of docking algorithms to identify favorable initial configurations.
Our results show that native lysozyme, when initially placed without docking, generally preserves its secondary structure and maintains a relatively stable conformation throughout the simulation. These findings are consistent with prior experimental work14,15,17,18,24, which suggests that although proteins adjust their conformation upon adsorption to CNTs, complete denaturation rarely occurs. In contrast, the docked native lysozyme undergoes a pronounced conformational reorientation after approximately 1463 ns (Fig. 12). This rearrangement increases the lysozyme’s surface contact with the CNT while retaining core structural elements. The interaction with the CNT surface primarily involves three key amino acid residues: ARG, TRP, and ASN.
For the denatured lysozyme systems, both docked and non-docked configurations exhibit more extensive and efficient adsorption than the native cases. The denatured lysozyme interacts with roughly three times as many amino acids contacting the CNT surface, resulting in a more compact, lower-profile protein layer—less than 2 nm in thickness (height)—compared to the native maintained thickness of approximately 3 nm. Among these, the docked denatured system shows especially enhanced surface interactions, underscoring the importance of initial configuration in modulating the extent and nature of protein-CNT binding.
Furthermore, these findings have important biological implications. The conformational changes, resulting from the protein-CNT interactions, suggest potential functional alterations. Such changes could influence the protein’s antimicrobial activity, enzymatic behavior, and interaction with other biomolecules. Additionally, the ability of denatured lysozyme to form a compact and stable layer on CNT surfaces may open avenues for designing functionalized nanomaterials with enhanced biocompatibility and controlled bioactivity. This highlights the significance of understanding protein-CNT interactions for applications in nanomedicine, biosensing48, and drug delivery49.
In summary, these simulations also provide molecular-level insights into how initial configuration and protein conformation influence lysozyme adsorption on CNT surfaces. The results highlight that while proteins can achieve stable binding and even undergo significant reorientation to optimize surface contact, the underlying secondary structure remains largely intact in the native state. In contrast, denatured proteins readily form more densely adsorbed layers.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Rutherglen, C., Jain, D. & Burke, P. Nanotube electronics for radiofrequency applications. Nat. Nanotechnol. 4, 811–819 (2009).
Petrov, A. & Audette, G. F. Peptide and protein-based nanotubes for nanobiotechnology. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 4, 575–585 (2012).
Zare, H. et al. Carbon nanotubes: Smart drug/gene delivery carriers. Int. J. Nanomed. 16, 1681–1706 (2021).
Karimi, M. et al. Nanotechnology in diagnosis and treatment of coronary artery disease. Nanomedicine 11, 513–530 (2016).
Son, K. H., Hong, J. H. & Lee, J. W. Carbon nanotubes as cancer therapeutic carriers and mediators. Int. J. Nanomed. 11, 5163–5185 (2016).
Ganazzoli, F. & Raffaini, G. Classical atomistic simulations of protein adsorption on carbon nanomaterials. Curr. Opin. Colloid Interface Sci. 41, 11–26. https://doi.org/10.1016/j.cocis.2018.11.008 (2019).
Nel, A. E. et al. Understanding biophysicochemical interactions at the nano-bio interface. Nat. Mater. 8(7), 543–557 (2009).
Ge, C. et al. Binding of blood proteins to carbon nanotubes reduces cytotoxicity. Proc. Natl. Acad. Sci. U.S.A. 108(41), 16968–16973 (2011).
Shim, M., Kam, N. W. S., Chen, R. J., Li, Y. & Dai, H. Functionalization of carbon nanotubes for biocompatibility and biomolecular recognition. Nano Lett. 2, 285–288 (2002).
Ramanathan, T., Fisher, F. T., Ruoff, R. S. & Brinson, L. C. Amino-functionalized carbon nanotubes for binding to polymers and biological systems. Chem. Mater. 17(6), 1290–1295 (2005).
Stevens, J. L. et al. Side-wall functionalization of single walled carbon nanotubes through fluorination and subsequent reactions with terminal domains. Nano Lett. 3(3), 331–336 (2003).
Calvaresi, M. & Zerbetto, F. The devil and holy water: protein and carbon nanotube hybrids. Acc. Chem. Res. 46(11), 2454–2463 (2013).
Matsuura, K. et al. Selectivity of water-soluble proteins in single-walled carbon nanotube dispersions. Chem. Phys. Lett. 429, 497–502 (2006).
Nepal, D. & Geckeler, K. pH-sensitive dispersion and debundling of single-walled carbon nanotubes: Lysozyme as a tool. Small 2, 406–412 (2006).
Bomboi, F., Bonincontro, A., La Mesa, C. & Tardani, F. Interactions between single-walled carbon nanotubes and lysozyme. J. Colloid Interface Sci. 355(2), 342–347 (2011).
Noordadi, M., Mehrnejad, F., Sajedi, R. H., Jafari, M. & Ranjbar, B. The potential impact of carboxylic-functionalized multi-walled carbon nanotubes on trypsin: A comprehensive spectroscopic and molecular dynamics simulation study. PLoS ONE 13(6), e0198519 (2018).
Calvaresi, M., Hoefinger, S. & Zerbetto, F. Probing the structure of lysozyme–carbon-nanotube hybrids with molecular dynamics. Chem. Eur. J. 18, 4308–4313 (2012).
Siepi, M. et al. Denatured lysozyme-coated carbon nanotubes: A versatile biohybrid material. Sci. Rep. 9(1), 16643 (2019).
Wu, E., Coppens, M. O. & Garde, S. Role of arginine in mediating protein-carbon nanotube interactions. Langmuir 31(5), 1683–1692 (2015).
Raffaini, G. & Ganazzoli, F. Simulation study of the interaction of some albumin subdomains with a flat graphite surface. Langmuir 19, 3403–3412 (2003).
Raffaini, G. & Ganazzoli, F. Protein adsorption on a hydrophobic surface: A molecular dynamics study of lysozyme on graphite. Langmuir 26, 5679–5689 (2010).
Borzooeian, Z., Taslim, M. E., Borzooeian, G., Ghasemi, O. & Aminlari, M. Activity and stability analysis of covalent conjugated lysozyme-single walled carbon nanotubes: Potential biomedical and industrial applications. RSC Adv. 7, 48692–48701 (2017).
Kumar, S., Mehta, D., Negi, S. Role of temperature variation on the dynamics of carbon nanotube and protein interactions. AIP Conf. Proc. 2752 (2023).
Horn, D. W. & Davis, V. A. Concentration of lysozyme/single-walled carbon nanotube dispersions. Colloids Surf. B 139, 237–243 (2016).
Ferraboschi, P., Ciceri, S. & Grisenti, P. Applications of lysozyme, an innate immune defense factor, as an alternative antibiotic. Antibiotics (Basel, Switzerland) 10(12), 1534 (2021).
Berendsen, H. J. C., van der Spoel, D. & van Drunen, R. GROMACS: A message-passing parallel molecular dynamics implementation. Comput. Phys. Commun. 91, 43–56 (1995).
Abraham, M. J., Murtola, T., Schulz, R., Páll, S., Smith, J. C., Hess, B., Lindahl, E. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX. 1, 19–25 (2015).
Diamond, R. Real-space refinement of the structure of hen egg-white lysozyme. J. Mol. Biol. 82(3), 371–391 (1974).
Poghosyan, A. H. et al. A molecular dynamics study of protein denaturation induced by sulfonate-based surfactants. J. Mol. Model. 27(9), 1–15 (2021).
Choi, Y. K., Kern, N. R., Kim, S., Kanhaiya, K., Jeon, S. H., Afshar, Y., Jo, S., Brooks, B. R., Lee, J., Tadmor, E. B., Heinz, H., Im, W. CHARMM-GUI nanomaterial modeler for modeling and simulation of nanomaterial systems (2022).
Kern, N. R. CHARMM-GUI multicomponent assembler for modeling and simulation of complex heterogeneous biomolecular systems. Biophys. J. 116(3), 290a (2019).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J. Comput. Chem. 31, 455–546 (2010).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14(1), 71–73 (2017).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Nosé, S. A unified formulation of the constant temperature molecular-dynamics methods. J. Chem. Phys. 81(1), 511–519 (1984).
Rahman, A. & Parrinello, N. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182–7189 (1981).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1987).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: anN·log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089 (1993).
Shahinyan, A. A., Poghosyan, A. H. & Astsatryan, H. V. Parallel peculiarities and performance of GROMACS package on HPC platforms. Int. J. Sci. Eng. Res. 4(12), 1755–1761 (2013).
Humphrey, W., Dalke, A. & Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Ibrahim, H. R., Ursula, T. & Pellegrini, A. A helix-loop-helix peptide at the upper lip of the active site cleft of lysozyme confers potent antimicrobial activity with membrane permeabilization action. J. Biol. Chem. 276(47), 43767–43774 (2001).
Yanamala, N., Kagan, V. E. & Shvedova, A. A. Molecular modeling in structural nano-toxicology: Interactions of nanoparticles with nano-machinery of cells. Adv. Drug. Deliv. Rev. 65, 2070–2077 (2013).
Engelman, D. M. & Steitz, T. A. The spontaneous insertion of proteins into and across membranes: The helical hairpin hypothesis. Cell 23(2), 411–422 (1981).
Nawaz, N. et al. Lysozyme and its application as antibacterial agent in food industry. Molecules (Basel, Switzerland) 27(19), 6305 (2022).
Artymiuk, P. J. et al. Crystallographic studies of the dynamic properties of lysozyme. Nature 280(5723), 563–568 (1979).
Noor, M. M., Goswami, J. & Davis, V. A. Comparison of attachment and antibacterial activity of covalent and noncovalent lysozyme-functionalized single-walled carbon nanotubes. ACS Omega 5(5), 2254–2259 (2020).
Hirano, A., Tanak, T., Kataura, H. & Kameda, T. Arginine side chains as a dispersant for individual single-walled carbon nanotubes. Chem. Eur. J. 20, 4922–4930 (2014).
Baldacchini, C., Chamorro, M. A. H., Prato, M. & Cannistraro, S. Highly conductive redox protein-carbon nanotube complex for biosensing applications. Adv. Funct. Mater. 21, 153–157. https://doi.org/10.1002/adfm.201001650 (2011).
Wakekar, S., Tiwari, A., Chaskar, J. & Chaskar, A. Protein nanotubes as drug delivery systems: an overview. J. Nanopart. Res. 25, 132. https://doi.org/10.1007/s11051-023-05786-3 (2023).
Acknowledgements
We acknowledge computational resources (“Aznavour” supercomputer) provided at the Institute of Informatics and Automation Problems of NAS RA. H.I. acknowledges the Higher Education and Science Committee of the Republic of Armenia for supporting this research (Grant No. N 24RL-1C009). The work of S. M. V. A. and M. A. were partially supported by the research council of the University of Tehran.
Author information
Authors and Affiliations
Contributions
S.M.V.A. and A.H.P. conceived the original idea, planned the project. H.I. and A.H.P. carried out all simulations. All authors worked on postprocessing and analyzing the data. All authors provided critical feedback and helped shape the research, figures, analysis and manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Vaez Allaei, S.M., Amininasab, M., Ishkhanyan, H. et al. Potential functional changes in native lysozyme induced by carbon nanotubes studied by molecular dynamics simulations. Sci Rep 15, 11593 (2025). https://doi.org/10.1038/s41598-025-96435-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-96435-3