Summary
Anonymized biomedical data sharing faces several challenges. This systematic review analyzes 1084 PubMed-indexed studies (2018–2022) using anonymized biomedical data to quantify usage trends across geographic, regulatory, and cultural regions to identify effective approaches and inform implementation agendas. We identified a significant yearly increase in such studies with a slope of 2.16 articles per 100,000 when normalized against the total number of PubMed-indexed articles (p = 0.021). Most studies used data from the US, UK, and Australia (78.2%). This trend remained when normalized by country-specific research output. Cross-border sharing was rare (10.5% of studies). We identified twelve common data sources, primarily in the US (seven) and UK (three), including commercial (seven) and public entities (five). The prevalence of anonymization in the US, UK, and Australia suggests their practices could guide broader adoption. Rare cross-border anonymized data sharing and differences between countries with comparable regulations underscore the need for global standards.
Similar content being viewed by others
Introduction
Digital technology is transforming healthcare delivery, improving patient outcomes, and facilitating more efficient public health strategies1. Data plays a central role in these advancements and is the foundation for personalized medicine2,3, epidemiological research4,5, and the development of artificial intelligence (AI) and machine learning models in healthcare6. Realizing the full potential of these developments requires the ability to re-use and share health data across systems, disciplines, and borders7. Such data flows face significant ethical, legal, and societal challenges which remain a major roadblock, as highlighted also during the COVID-19 pandemic8. While obtaining informed consent, i.e., having patients opt-in to their data being shared, is a common solution, this process is faced with several challenges, including impracticality9 and varying levels of comprehension10. As an alternative, and generally as part of a privacy-by-design approach to medical research, technology can be employed to ensure that patients remain anonymous. In this context, the scientific community is working on a wide range of technologies, including federated learning11,12, distributed data analytics platforms13, differential privacy14, and synthetic data generation15. However, these techniques have not yet been adopted on a broad scale in biomedical research due to their high infrastructure requirements, complexity of use, and potential impacts on the reliability of results16,17. The traditional solution is data anonymization.
Anonymization, in essence, denotes a technical process of altering data in such a manner that the risk of it being traced back to individuals is significantly reduced18. Common methods include the removal of data, the masking of values, the addition of noise or the reduction of the fidelity of data points. The feasibility of achieving privacy protection with such methods while ensuring that the data remains useful for scientific research is a subject of intense debate within scholarly discourse19. Some experts argue that privacy risks are not sufficiently mitigated by this approach, suggesting a potential inadequacy in legal frameworks20,21. Studies that focus on the uniqueness of data as a proxy for privacy risks support this view22,23. Conversely, others argue that the chances of privacy breaches to actually happen are low, especially when anonymization adheres to best practices and established policies24,25,26. A range of such best practices and case studies have been published27,28,29,30.
Diverse perspectives on anonymization in medical research also appear across various legal frameworks. For example, in the United States (US) biomedical data is frequently shared in a “de-identified” form (see Section “Search strategy and selection criteria”) for research purposes based on the concrete requirements laid out in the Safe Harbor method of the Privacy Rule of the Health Insurance Portability and Accountability Act (HIPAA)31. As another example, the National Health Services (NHS) in the United Kingdom (UK) has provided specific guidance regarding the use of statistical anonymization methods32. In contrast, in the European Union (EU) anonymization is more challenging to apply in practice, due to ambiguities in the legal definition33 and a lack of a common understanding and standards34. To address these obstacles and facilitate data sharing, the European Union (EU) is currently introducing legislation on the so-called European Health Data Space (EHDS), which will provide additional legal bases for sharing biomedical data on an opt-out basis35.
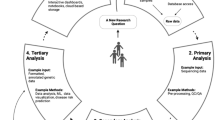
The objective of this paper is to investigate, on a quantitative level, how diverse perceptions and legal ambiguities actually impact the use of anonymization in biomedical research. To this end, we analyzed a corpus of medical studies explicitly stating in their title or abstract that they used anonymized data. With our results, we aim to support policymakers, healthcare organizations, and researchers with data for improving existing policies and structures as well as developing future research agendas. Specifically, we aim to provide insights along three dimensions:
-
(1)
Temporal trends: Is the number of studies based on anonymized data increasing or decreasing? Did recent developments, such as the COVID-19 pandemic, have measurable impacts on the use of anonymization in biomedical research?
-
(2)
Geographical differences: In which countries is the sharing of anonymized data more prevalent than in others? Do frequently discussed regulatory and cultural differences across regions influence the use of anonymization in biomedical research, including in cross-border settings?
-
(3)
Common data sources: Are there specific organizations that frequently share data, and do these organizations target research on particular disease categories?
To the best of our knowledge, this is the first review that adopts a bottom-up approach by analyzing scientific studies that utilized anonymized data, rather than analyzing policies or anonymization practices.
Results
Through the PubMed search, 1641 articles were identified. After filtering for language, peer-review status, and publication date, 1551 articles were initially screened based on their title and abstract. Of the remaining 1150 articles, 50 were inaccessible, and 16 were further excluded during full-text screening. Ultimately, 1084 articles were included in the review. The detailed selection process is depicted in the flowchart in Fig. 1.
Temporal trends
Figure 2 shows the number of included articles per year of publication, categorically separated into non-COVID-19 and COVID-19 related research. We divided the number by the total number of articles published on PubMed for each respective year to account for the general growth in research output. Additionally, labels indicate the absolute number of included articles for each year.
The number of articles was normalized per 100,000 articles registered on PubMed in each year. Articles related to COVID-19 research are highlighted. Two regression lines with a 95% confidence interval, shown as two shaded areas, depict trends for the total number of articles (gray) and the number of articles without COVID-19 focus (green). Labels above the bars indicate the absolute number of included articles.
We identified a clear upwards trend in papers published that utilize anonymized data over the years, which further intensifies with the onset of COVID-19. The regression analysis, conducted after normalizing against the annual total research registered on PubMed, reveals a statistically significant yearly increase of 2.16 included articles per 100,000 articles (p = 0.021). Even when excluding articles with COVID-19 focus, the upward trend amounts to an increase of 1.14 articles per year (p = 0.030).
Geographical differences
Figure 3 shows the distribution of the location of the first authors and the origin of the data sorted by frequency of occurrence for the papers where the data originated from exactly one country (1027 [94.7%] of 1084). In the great majority of these cases (970 [94.4%] of 1027), the location of the first author and the data origin was the same. The first authors came from a total of 53 different countries, while the data originated from 55 different countries.
Among the 1027 articles with data originating from a single country, the most frequent country of first authors was the US (545 [53.1%]), followed by the UK (187 [18.2%]), and Australia (54 [5.3%]). Similarly, for these articles, most data originated from the US (563 [54.8%]), followed by the UK (186 [18.1%]), and Australia (54 [5.3%]). Notably, these are all countries from the so-called Core Anglosphere, i.e. a group of English-speaking countries with historical, cultural, and political links, which includes the US, UK, Canada, Australia, and New Zealand. In contrast, only 10.1% (104 of 1027) of first authors and 8.7% (89 of 1027) of the data came from continental EU countries, Norway, or Switzerland, which all operate under the EU General Data Protection Regulation (GDPR) or a highly convergent law, and to which we henceforth refer to as Continental Europe. In the studies that sourced data from multiple countries (57 [5.2%] of 1084), the most common countries of first authors were the US (16 studies), the UK (11 studies), and Australia (5 studies), whereas the top countries from which the data originated were the US and UK (used in 13 studies), Germany (11 studies), Spain, France, and Italy (9 studies each). It should be noted that for 27 of the articles with multiple data origins it was not possible to break down the individual countries.
Figure 4 displays the number of articles included per country, normalized by the number of citable documents of the country in the subject area of medicine (as a proxy for overall research output) in the years 2018–2022, according to the SCImago Journal and Country Rank (SJR)36. Countries to which we assigned articles and which are among the top-20 according to their medical research output in the SJR are shown individually (16 in total). All other countries listed in SJR are summarized in the category “other”, including 34 countries to which we also assigned papers. For an included article to be assigned to a country, both the first author and the data must be from that country, and from that country only.
The number of articles with same author and data origin was normalized by the overall medical research output for each country, approximated by the number of citable documents in the field of medicine from 2018 to 2022, as indexed in SJR. Countries among the top-20 in research output assigned to at least one article are shown individually. All remaining countries were categorized as “other”.
In addition to the grouping of countries belonging to the Core Anglosphere, which was already identified as particularly relevant in the previous analysis, and Continental Europe, the categorization was extended with groups for countries within Asia, Eurasia, and South America. Eurasia was added to accommodate Russia’s unique geographic and geopolitical positioning across two continents.
For each group we calculated an average over all countries assigned to this group in the previous step. We did this by first summing up the number of articles assigned to those countries, and then dividing this total by the number of citable documents in the SJR for those countries. Countries from the Core Anglosphere, i.e. the UK, US, Australia, and Canada have by far the highest average ratio of articles included to the total number of publications (average of 0.345 articles per 1000 citable documents), exceeding the average of the top-20 countries shown in Fig. 4 (0.198) as well as the global average (0.157). Conversely, in Continental Europe, anonymized data sharing is comparatively uncommon, with such studies being underrepresented relative to the overall research output of these countries (average of 0.061). A similar picture can be observed for countries in Asia (average of 0.044). Notably, Germany and Japan rank fourth and fifth overall, with 0.129 and 0.081 articles per 1000 citable documents, respectively.
Next, we examined individual data flows by looking at the origin of the data and the first authors to understand how data has been transferred. We found that 10.5% (114 of 1084) of first authors have used data from one or multiple foreign countries, resulting in a total of 202 cross-border data flows. The great majority of authors (991 [91.4%] of 1084) used data from their own country either exclusively or in addition to foreign data, which we have counted as a total of 991 domestic data flows. Figure 5a shows the relative frequency of data origins in cross-border data flows as well as domestic data usages. Countries that are not listed at least twenty times as the origin of a data flow were categorized as “other”. Figure 5b illustrates specific cross-border data flows identified in at least three studies.
a Proportion of data origins in cross-border and domestic data flows. Labels above the bars indicate the absolute number of data flows. Countries not listed at least twenty times as the origin of a data flow were categorized as “other”. b Sankey diagram of cross-border data flows found at least three times (39 data flows).
As can be seen in Fig. 5a, the US, which we found to be the country sharing data most often, was also the country sharing data most frequently in cross-border scenarios. Nevertheless, for both the US and the UK, cross-border data sharing was much less common than in domestic scenarios, both in absolute and proportional terms. This also explains why countries categorized under “other” have a comparatively large share of cross-border flows. Among these, there are 26 countries where we found only studies that used data in cross-border scenarios and none with domestic usage. The most frequent patterns shown in Fig. 5b reveal a notable predominance of cross-border data flows within the Core Anglosphere, as well as between the Core Anglosphere and Continental Europe.
Common data sources
Of the 1084 studies included, 85.8% (930 of 1084) were identified through the use of healthcare related terms to describe their data, 8.4% (91 of 1084) used research data related terms, and 5.8% (63 of 1084) used both (see Table 3 for a list of corresponding terms). To gain a better understanding of the entities providing the data, we examined common sources, which we define as data sources utilized in at least ten studies. Table 1 summarizes these sources. More than 40% of the studies (460 [42.4%] of 1084) used data from one or multiple common sources. The share of articles utilizing common data sources has steadily increased over time, from 31.1% (44 of 140) in 2018 to 50.1% (167 of 330) in 2022. We also assessed whether repeated citations by author groups influenced a source’s classification as common by calculating the share of distinct authors among all citing authors. This was not the case: half (6 of 12) of the common sources exceeded 80%, and 83.3% (10 of 12) reached at least 70%. The only exceptions were SLaM NHS (59.3%) and VUMC (66.1%), which, however, had high citation counts (25 and 52), well above the 10-reference threshold. The distribution of the use of healthcare and research data terms in studies utilizing common data sources is comparable to that across the corpus.
Consistent with our previous findings, 58.3% (7 of 12) of the identified common sources are from the US. The vast majority of US common sources (6 [85.7%] of 7), as well as the German InGef are commercial enterprises. The only non-commercial US provider is the Vanderbilt University Medical Center, which predominantly supports intra-mural research. Common data sources located in the UK and Australia, are all publicly funded. All common sources offer at least some data that was routinely collected during healthcare processes. Except for TriNetX and IQVIA, for which some of the included articles indicate that data was gathered from multiple countries, the origin of the data used in the studies corresponded to the origin of the respective data source. We note that MIMIC, another well-known source of anonymized data37, was also used by some studies, but did not classify as a common source per our definition.
Figure 6 provides an overview of how many articles used data from each common source. Due to some articles using multiple common data sources, the total number of usages amounts to 472. Notably, two healthcare and information technology companies, Optum and Flatiron, together account for 43.9% (207 of 472) of all uses of common data sources. VUMC stands out as the only common source operated by a single academic institution, ranking third in frequency of use and accounting for 11.0% (52 of 472) of all uses. It should be noted that 86.5% (45 of 52) of these uses involved intra-mural provisioning, where the first author was also affiliated with VUMC. Publicly funded entities in the UK cumulatively represent 24.6% (116 of 472) of the used common sources.
Figure 7 illustrates the data flow from common sources to studies targeting specific disease categories. A substantial proportion of the studies on neoplasms use data from Flatiron (88 [51.8%] of 170), reflecting its specialization in cancer. Optum enabled a significant proportion of the studies on diseases of the circulatory system (14 [16.5%] of 85), while data from SLaM NHS was used often in studies on mental and behavioral disorders (24 [20.7%] of 116). Most of the common sources are not specialized but provide data for studies across a broad spectrum of diseases.
Discussion
Our results show that the number of studies relying on anonymized data has steadily increased in recent years. The COVID-19 pandemic response, necessitating enhanced information flow, has further amplified this trend and the advancement of anonymization technologies is also called for in agendas for pandemic preparedness, such as the GLoPID-R Roadmap for Data Sharing in Public Health Emergencies38.
Regarding geographic, regulatory, and cultural differences, the sharing of anonymized data is particularly common in countries belonging to the Core Anglosphere. The US, UK, and Australia share anonymized data more frequently than other countries, even when put in relation to their overall research output. In contrast, countries in Asia, Continental Europe, Eurasia, and South America, seem to provide less anonymized data to biomedical studies. One reason for this are differences in legal and regulatory frameworks. In the US, anonymization is often performed based on the requirements laid out in the HIPAA Privacy Rule, for example in Vanderbilt’s Synthetic Derivative39 or MIMIC37, illustrating that the legal requirements are interpretable and can be applied consistently as a standard of practice. The GDPR, however, is interpreted differently across EU member states (and countries with similar laws, such as Switzerland, Japan, and Korea40) with ambiguous perceptions of anonymity41. It can be understood as an absolute state achieved solely by transforming the data41,42, or as a state that exists within a certain context33. As a result, despite a legal framework comparable to the GDPR, the sharing of anonymized data is much more common in the UK. In a report from 2006, the UK Academy of Medical Sciences noted that anonymity should be assessed within the context in which data is being shared43. This interpretation remained unchanged with the introduction of the GDPR in 201844. Also in Australia, a context-sensitive approach is being pursued45. One common way to control the context is the sharing of data through secure platforms, called Safe Havens or Secure Processing Environments, which are for example used by SAIL46, SLaM47, and MIMIC37. This approach also forms a core concept of the upcoming European Health Data Space, which provides an example of a case where mechanisms and protocols successfully used in some countries have informed implementation strategies in others.
We acknowledge that further factors, such as technical, motivational, and economic barriers, influence data sharing practices, which may also impact anonymized data sharing48. However, there is ample evidence that data protection is a fundamental roadblock and that only once basic legal preconditions are sufficient, further aspects become critical factors. For example, a frequently cited barrier to data sharing is the lack of incentives, even within countries where anonymized data sharing is common, such as in the US49,50,51. At the same time, there are numerous large-scale data sharing initiatives in regions where we found anonymization to be less commonly adopted52,53. They are usually based on obtaining informed consent or on dedicated laws. TEHDAS, a collaborative initiative aimed at preparing and shaping the EHDS, also identified the lack of clear guidance on anonymization as a key barrier to data sharing54.
Our findings also showed that cross-border flows of anonymized data are relatively uncommon, even for countries which have a robust culture of domestic or intra-mural anonymized data sharing. This aligns with previous results. One explanation is that there are significant differences in data protection frameworks across jurisdictions, including the US, UK, and Australia, resulting in a lack of a uniform understanding55 and varying interpretations56. Moreover, it has been found that requirements for anonymization are often vague and that more comprehensive guidelines like the UK Anonymisation Decision-Making Framework30, which has also been adopted in Australia, are needed55. However, among recommendations on how to anonymize clinical trial dataset, a growing, yet incomplete, consensus on anonymization methods has been observed57. Recent publications from the International Standardization Organization (ISO) can be seen as a further step in the right direction18,58.
Finally, we observed that a significant portion of studies rely on anonymized data from common sources. These are usually focused on anonymized routine healthcare data, which is a natural application area for anonymization mechanisms. Many of the sources are located in the US and managed by commercial entities, alongside publicly funded entities in the UK and Australia. The only source identified outside the Core Anglosphere was InGef from Germany, a commercial provider with ties to statutory health insurances. The Vanderbilt University Medical Center was the only identified single academic institution that established a frequently used source of anonymized biomedical data. Some common sources as well as their data have also been described in the scientific literature39,46,47,59,60,61. Most common sources are not specialized but provide data to studies on a wide range of different diseases. Nevertheless, some identified sources focus on specific disease areas and contribute significantly to respective research using anonymized data.
We conclude that sharing and accessing anonymized data is an important mechanism enabling biomedical studies. Moreover, differences in regulatory frameworks and policies in different geographies have a measurable impact on how anonymization is adopted in practice. Our analysis focused on data protection-related differences and found that consistent policies and a context-sensitive interpretation of the concept of anonymity may help to implement anonymization on a broader scale. Moreover, a significant proportion of anonymized data sharing occurs through publicly funded as well as commercially operated common sources and their methodologies could serve as blueprints for developing approaches that also work in other regions. Finally, we found that cross-border data sharing is rare in practice, highlighting the need to establish uniform anonymization standards. Our overall recommendations for improving existing data protection policies and structures as well as developing future research agendas are summarized in Table 2.
Limitations include a review scope restricted to studies indexed in PubMed, written in English, and published between 2018 and 2022, which could have introduced bias. Additionally, the interpretation and jurisdictional variations of the terms anonymization, de-identification, and pseudonymization might impact the generalizability of our findings. However, PubMed is one of the most comprehensive databases for biomedical literature and the selected timeframe is characterized by increasing privacy awareness and evolving regulatory landscapes (e.g., the introduction of the GDPR in the EU). We also carefully selected a terminology for our search criteria, and our findings confirm results of previous research conducted on the topic at a conceptual level. Our geographical analysis, based on the first author’s institution and data origin, may not fully capture global collaboration networks, though the rarity of cross-border data flows suggests limited impact on our conclusions. Moreover, while our study aims to provide a global overview, our findings, based on a subset of literature explicitly mentioning the use of anonymized data, might not be fully comprehensive. This can be exemplified by the absence of some widely recognized sources of anonymized data as common data sources, such as MIMIC37 and the UK Biobank62, possibly due to their prominence and policies affecting how studies reference anonymization. However, as our analysis mainly relies on the comparison of relative frequencies and trends, we believe that our results are representative. Ultimately, while we identified regional differences in the use of anonymized data that correspond with differences in data protection laws and their interpretation, there are further factors that also need to be considered when measures are planned to make anonymized data sharing more common.
Methods
Search strategy and selection criteria
We identified biomedical studies that produced findings based on the analysis of anonymized datasets published between 2018 and 2022 through searches of PubMed. This period includes the introduction of new data protection regulations in several countries and regions, such as in Brazil, China, the EU, Japan, and Thailand, the peak of the COVID-19 pandemic (2020-2021) and the time thereafter. Where applicable, we reported our review process in accordance with the PRISMA checklist for systematic reviews63, and employed a step-wise procedure for fine-tuning our search query.
Selecting studies that utilized anonymized biomedical data published by authors across the globe is complicated by the varying terminologies used in different regions and contexts56. The terms anonymization and pseudonymization are, for example, used in countries within the EU, the UK, and China. We deliberately focused on anonymization and not on pseudonymization (sometimes also referred to as pseudo-anonymization18), as it is generally understood that pseudonymized data retains a protected but explicit link back to the individual. Despite debates on what exactly distinguishes pseudonymized from anonymized data, the latter is commonly understood as data that cannot be traced back to individuals. This aligns with the common understanding of the term de-identified data56, for example in the US, Canada, or Australia. For instance, HIPAA defines de-identified health information as data for which there is no reasonable basis to assume it can be linked back to an individual64. In contrast, pseudonymization is usually used during data collection or when patients have explicitly consented to the use or sharing of their data65.
Our initial search utilized terms related to anonymization and de-identification along with their spelling variants, combined with keywords pertaining to various types of healthcare or research data. We refined our search by excluding terms associated with privacy, surveys, and legal issues to focus specifically on studies that use anonymized health data rather than addressing the topic on a meta-level. All search terms were matched against titles and abstracts. The individual groups of terms as well as the final search string is shown in Table 3. The final search was performed on 06.02.2025.
During the screening process, we excluded studies (1) focusing on anonymization technology, (2) which did not present findings that were produced using anonymized data, (3) which did not focus on a health-related outcome, (4) which were solely based on hard-to-anonymize unstructured data, (5) were based on anonymous surveys, or (6) focused on the implementation of data sharing. The screening process involved an initial screening of titles and abstracts by two reviewers assigned at random. Discrepancies in their decisions were resolved through discussion with a third reviewer. Subsequently, full-text articles were evaluated for inclusion using the same procedure.
Data collection and analysis
Table 4 lists the data items we collected to provide insights into the three dimensions of interest. To investigate temporal trends, we charted the year of publication as well as whether a study produced COVID-19-related findings, to see whether the pandemic response had an effect. To provide insights into geographical differences, we collected the country in which the first author’s first affiliation is located as well as the country of origin of the dataset used in the study. We defined the dataset’s origin as the country in which the data was initially collected, typically corresponding to the location of the data source (e.g., hospitals, insurance companies, or research institutions) providing the data. In cases where data was obtained from multiple countries, all contributing countries were recorded, provided they were reported separately. We note that we also collected information on the senior author’s locations, but those were largely consistent with the origin of the first author (same in 1014 [93.5%] of 1084 cases). Finally, to study common data sources, we charted whether a paper used terms related to healthcare or research data or both (see Table 3), the name of the source providing the data, as well as the ICD-1066 chapter of the diseases of interest. Analogously to the data origin, several entities could be recorded as data sources for a single article. When a study spanned morbidities from multiple ICD-10 chapters, the journal’s subject area was utilized to identify a primary chapter. With the exception of the items “Year” and “Data type”, which were automatically inferred by querying the database, the charting was performed manually. Each item was charted by two independent reviewers and discrepancies were resolved by discussion with a third reviewer.
According to our dimensions of interest, we prepare the results using primarily descriptive statistics, complemented by data flow mappings and clustering of data sources. Additionally, we supplement our data with external information on the general research output of countries using the SCImago Journal and Country Rank36.
Data availability
All data collected for this review is publicly available on GitHub (https://github.com/bih-mi/anonymization-review/).
Code availability
The data analysis was performed using Python 3.11. The code is publicly available on GitHub (https://github.com/bih-mi/anonymization-review/).
References
Topol, E. J. Deep Medicine: How Artificial Intelligence Can Make Healthcare Human Again. Basic Books: New York, (2019).
The “All of Us” Research Program. N. Engl. J. Med. 381, 668–676 (2019).
Denny, J. C. et al. PheWAS: demonstrating the feasibility of a phenome-wide scan to discover gene-disease associations. Bioinformatics 26, 1205–1210 (2010).
Mahmood, S. S., Levy, D., Vasan, R. S. & Wang, T. J. The Framingham Heart Study and the epidemiology of cardiovascular disease: a historical perspective. Lancet 383, 999–1008 (2014).
Murray, C. J. L. The Global Burden of Disease Study at 30 years. Nat. Med. 28, 2019–2026 (2022).
Ngiam, K. Y. & Khor, I. W. Big data and machine learning algorithms for health-care delivery. Lancet Oncol. 20, e262–e273 (2019).
Näher, A.-F. et al. Secondary data for global health digitalisation. Lancet Digit Health 5, e93–e101 (2023).
Dron, L. et al. Data capture and sharing in the COVID-19 pandemic: a cause for concern. Lancet Digit Health 4, e748–e756 (2022).
Laurijssen, S. J. et al. When is it impractical to ask informed consent? A systematic review. Clin. Trials 19, 545–560 (2022).
Pietrzykowski, T. & Smilowska, K. The reality of informed consent: empirical studies on patient comprehension—systematic review. Trials 22, 57 (2021).
Sadilek, A. et al. Privacy-first health research with federated learning. npj Digit. Med. 4, 132 (2021).
Crowson, M. G. et al. A systematic review of federated learning applications for biomedical data. PLOS Digit Health 1, e0000033 (2022).
Wirth, F. N., Meurers, T., Johns, M. & Prasser, F. Privacy-preserving data sharing infrastructures for medical research: systematization and comparison. BMC Med. Inf. Decis. Mak. 21, 242 (2021).
Dwork, C. Differential Privacy. In Automata, Languages and Programming (eds. Bugliesi, M., Preneel, B., Sassone, V. & Wegener, I.) vol. 4052 1–12 (Springer, 2006).
Giuffrè, M. & Shung, D. L. Harnessing the power of synthetic data in healthcare: innovation, application, and privacy. npj Digit. Med. 6, 186 (2023).
Jordan, S., Fontaine, C. & Hendricks-Sturrup, R. Selecting Privacy-Enhancing Technologies for Managing Health Data Use. Front. Public Health 10, 814163 (2022).
Scheibner, J. et al. Revolutionizing Medical Data Sharing Using Advanced Privacy-Enhancing Technologies: Technical, Legal, and Ethical Synthesis. J. Med Internet Res 23, e25120 (2021).
ISO/IEC 20889:2018, Privacy enhancing data de-identification terminology and classification of techniques.
Schwartz, P. M. & Solove, D. J. The PII Problem: Privacy and a New Concept of Personally Identifiable Information. SSRN Scholarly Paper at https://papers.ssrn.com/abstract=1909366 (2011).
Ohm, P. Broken promises of privacy: Responding to the surprising failure of anonymization. UCLA l. Rev. 57, 1701 (2009).
Narayanan, A. & Felten, E. W. No silver bullet: De-identification still doesn’t work. https://randomwalker.info/publications/no-silver-bullet-de-identification.pdf (2014).
Rocher, L., Hendrickx, J. M. & De Montjoye, Y.-A. Estimating the success of re-identifications in incomplete datasets using generative models. Nat. Commun. 10, 3069 (2019).
Narayanan, A. & Shmatikov, V. Robust De-anonymization of Large Sparse Datasets. in 2008 IEEE Symposium on Security and Privacy (sp 2008) 111–125 https://doi.org/10.1109/SP.2008.33 (IEEE, 2008).
El Emam, K., Jonker, E., Arbuckle, L. & Malin, B. A Systematic Review of Re-Identification Attacks on Health Data. PLoS ONE 6, e28071 (2011).
Information, O. O. of the, Commissioner, P., Cavoukian, A. & Castro, D. Big Data and Innovation, Setting the Record Straight: De-Identification Does Work (Information and Privacy Commissioner, 2014).
Seastedt, K. P. et al. Global healthcare fairness: We should be sharing more, not less, data. PLOS Digit Health 1, e0000102 (2022).
El Emam, K., Rodgers, S. & Malin, B. Anonymising and sharing individual patient data. BMJ 350, h1139–h1139 (2015).
Wagner, I. & Eckhoff, D. Technical Privacy Metrics: A Systematic Survey. ACM Comput. Surv. 51, 1–38 (2019).
Jakob, C. E. M., Kohlmayer, F., Meurers, T., Vehreschild, J. J. & Prasser, F. Design and evaluation of a data anonymization pipeline to promote Open Science on COVID-19. Sci. Data 7, 435 (2020).
Elliot, M., Mackey, E. & O’Hara, K. The anonymisation decision-making framework, 2nd. ed. (European practitioners’ guide, 2020).
Office for Civil Rights, H. Standards for privacy of individually identifiable health information. Final rule. Fed. Register 67, 53181–53273 (2002).
NHS Digital. ISB1523: Anonymisation standard for publishing health and social care data (NHS Digital, 2020).
Mourby, M. Anonymity in EU Health Law: Not an Alternative to Information Governance. Med Law Rev. 28, 478–501 (2020).
Peloquin, D., DiMaio, M., Bierer, B. & Barnes, M. Disruptive and avoidable: GDPR challenges to secondary research uses of data. Eur. J. Hum. Genet 28, 697–705 (2020).
European Commission. Proposal for a regulation of the European Parliament and of the Council on the European Health Data Space (European Commission, 2022).
SCImago. SJR — SCImago Journal & Country Rank [Portal] (SCImago, 2023).
Johnson, A. E. W. et al. MIMIC-IV, a freely accessible electronic health record dataset. Sci. Data 10, 1 (2023).
Norton, A., Pardinaz-Solis, R. & Carson, G. GloPID-R Roadmap for data sharing in public health emergencies (GloPID-R, 2019).
Danciu, I. et al. Secondary use of clinical data: the Vanderbilt approach. J. Biomed. Inf. 52, 28–35 (2014).
Joo, M.-H. & Kwon, H.-Y. Comparison of personal information de-identification policies and laws within the EU, the US, Japan, and South Korea. Gov. Inf. Q. 40, 101805 (2023).
Molnár-Gábor, F. et al. Harmonization after the GDPR? Divergences in the rules for genetic and health data sharing in four member states and ways to overcome them by EU measures: Insights from Germany, Greece, Latvia and Sweden. Semin Cancer Bio 84, 271–283 (2022).
Article 29 Data Protection Working Party. Opinion 05/2014 on Anonymization Techniques, WP216, (0829/14/ EN) (2014).
Academy of medical sciences. Personal Data for Public Good: Using Health Information in Medical Research (Academy of Medical Sciences, 2006).
Mourby, M. et al. Are ‘pseudonymised’ data always personal data? Implications of the GDPR for administrative data research in the UK. Comput. Law Secur. Rev. 34, 222–233 (2018).
Office of the Australian Information Commissioner. De-Identification and the Privacy Act. (Office of the Australian Information Commissioner, 2018).
Jones, K. H., Ford, D. V., Thompson, S. & Lyons, R. A Profile of the SAIL Databank on the UK Secure Research Platform. IJPDS 4, (1134, 2020).
Fernandes, A. C. et al. Development and evaluation of a de-identification procedure for a case register sourced from mental health electronic records. BMC Med. Inf. Decis. Mak. 13, 71 (2013).
Van Panhuis, W. G. et al. A systematic review of barriers to data sharing in public health. BMC Public Health 14, 1144 (2014).
Tenopir, C. et al. Data Sharing by Scientists: Practices and Perceptions. PLoS ONE 6, e21101 (2011).
Lewin, J. H. et al. Determining barriers to effective data sharing in cancer genomic sequencing initiatives: A Global Alliance for Genomics and Health (GA4GH) survey. JCO 34, 11502–11502 (2016).
Gaba, J. F., Siebert, M., Dupuy, A., Moher, D. & Naudet, F. Funders’ data-sharing policies in therapeutic research: A survey of commercial and non-commercial funders. PLoS ONE 15, e0237464 (2020).
Cuggia, M. & Combes, S. The French Health Data Hub and the German Medical Informatics Initiatives: Two National Projects to Promote Data Sharing in Healthcare. Yearb. Med. Inf. 28, 195–202 (2019).
Lawrence Adrien K., Selter Liselotte, & Frey Urs. SPHN – The Swiss Personalized Health Network Initiative. In Studies in Health Technology and Informatics https://doi.org/10.3233/SHTI200344 (IOS Press, 2020).
Abboud, L. et al. Summary of results: Case studies on barriers to cross-border sharing of health data for secondary use (TEHDAS consortium partners, 2021).
Scheibner, J. et al. Data protection and ethics requirements for multisite research with health data: a comparative examination of legislative governance frameworks and the role of data protection technologies. J. Law Biosci. 7, lsaa010 (2020).
Chevrier, R., Foufi, V., Gaudet-Blavignac, C., Robert, A. & Lovis, C. Use and Understanding of Anonymization and De-Identification in the Biomedical Literature: Scoping Review. J. Med. Internet Res. 21, e13484 (2019).
Rodriguez, A. et al. Current recommendations/practices for anonymising data from clinical trials in order to make it available for sharing: A scoping review. Clin. Trials 19, 452–463 (2022).
ISO/IEC 27559:2022, Information security, cybersecurity and privacy protection – Privacy enhancing data de-identification framework.
Ehwerhemuepha, L. et al. Cerner real-world data (CRWD) - A de-identified multicenter electronic health records database. Data Brief. 42, 108120 (2022).
Herrett, E. et al. Data Resource Profile: Clinical Practice Research Datalink (CPRD). Int. J. Epidemiol. 44, 827–836 (2015).
Busingye, D. et al. Data Resource Profile: MedicineInsight, an Australian national primary health care database. Int J. Epidemiol. 48, 1741–1741h (2019).
Sudlow, C. et al. UK Biobank: An Open Access Resource for Identifying the Causes of a Wide Range of Complex Diseases of Middle and Old Age. PLoS Med. 12, e1001779 (2015).
Page, M. J. et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ 372, n71 (2021).
U.S. Department of Health and Human Services - Office for Civil Rights. HIPAA Administrative Simplification. (U.S. Department of Health and Human Services - Office for Civil Rights, 2013).
Kohlmayer, F., Lautenschläger, R. & Prasser, F. Pseudonymization for research data collection: is the juice worth the squeeze?. BMC Med. Inf. Decis. Mak. 19, 178 (2019).
World Health Organization. International statistical classification of diseases and related health problems, 10th ed. (WHO, 2019).
Davis, S. et al. Standardized Health data and Research Exchange (SHaRE): promoting a learning health system. JAMIA Open 5, ooab120 (2022).
Company. Flatiron Health, https://flatiron.com/about-us. (2025)
Evidence Solutions. Real-World Evidence. Flatiron Health. https://flatiron.com/real-world-evidence (2025).
IBM Explorys Cohort Discovery, IBM Explorys Therapeutic Datasets, and IBM Explorys Virtual Workbench provide life sciences insights into realworld care delivery. https://www.ibm.com/docs/en/announcement_archive/ENUSZP16-0471/ENUSZP16-0471.PDF (2016).
Francisco Partners Completes Acquisition of IBM’s Healthcare Data and Analytics Assets; Launches Healthcare Data Company Merative. Media. Francisco Partners https://www.franciscopartners.com/media/Merative (2022).
Andersohn, F. & Walker, J. Institute for Applied Health Research Berlin (InGef) Database. in Databases for Pharmacoepidemiological Research (eds. Sturkenboom, M. & Schink, T.) 125–129 https://doi.org/10.1007/978-3-030-51455-6_9 (Springer International Publishing, 2021).
Real World & Healt Data Sets. IQVIA. https://www.iqvia.com/solutions/real-world-evidence/real-world-data-and-insights (2025).
Optum. Real world health care experiences. https://www.optum.com/content/dam/optum/resources/productSheets/5302_Data_Assets_Chart_Sheet_ISPOR.pdf (2015).
Palchuk, M. B. et al. A global federated real-world data and analytics platform for research. JAMIA Open 6, ooad035 (2023).
Roden, D. et al. Development of a Large-Scale De-Identified DNA Biobank to Enable Personalized Medicine. Clin. Pharm. Ther. 84, 362–369 (2008).
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
F.P., T.M., J.D., and J.R. conceptualized the study. T.M. and F.P. designed the method. T.M., K.O., H.A., F.B., M.H., B.K., V.M., A.M., G.P., and, F.W. performed the screening and data charting process. T.M., F.P., J.D., and J.R. analyzed and interpreted the data. T.M. and F.P. drafted the manuscript. J.D., J.R., K.O., H.A., F.B., M.H., B.K., V.M., A.M., G.P., and F.W. edited and reviewed the final manuscript. All authors have had access to the data presented in this study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Meurers, T., Otte, K., Abu Attieh, H. et al. A quantitative analysis of the use of anonymization in biomedical research. npj Digit. Med. 8, 279 (2025). https://doi.org/10.1038/s41746-025-01644-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-01644-9