Abstract
Modular polyketide synthases (PKSs) can produce various secondary metabolites in a collinearity fashion. Although rational engineering of modular PKS can ultimately create a diverse array of compounds, de novo generation of defined structures usually results in the loss or remarkable decline of productivity due primarily to the incompatibility of different elements. Here, we present a modular PKS engineering strategy driven by an evolutionary event of gene conversion to accomplish successive engineering of the modular PKS in cinnamomycin biosynthetic gene cluster (cmm BGC). By simulating the gene conversion process, cmm BGC is consecutively reprogrammed to generate a macrolide with predicted structural features. Moreover, the intra-module KS domain is demonstrated to associate with the proofreading of extender units. Collectively, the gene conversion-associated approach may shed a light on modular PKS engineering.

Similar content being viewed by others
Introduction
Polyketides are biosynthesized by polyketide synthases (PKSs) to possess remarkable structural diversities and valuable medicinal utilities1. Among different types of PKS, type I PKS functions in a modular fashion2,3, in which the modules are linearly arranged to guide the unidirectional chain extension.
Since the early investigations on modular PKSs4,5, the concept of “Lego-ization” in polyketide biosynthesis has continuously been evolved6, allowing systematic reprogramming of PKSs to produce designed natural products with high stereo- and regio-specificity7,8,9. At the same time, various approaches, including site-directed mutagenesis, domain swapping and subunit insertions/deletions, have been implemented to the engineering of PKSs, aiming to side chain alteration or scaffold reconstruction. More recently, the strategies for PKS reprogramming have been moved forward from sequence-based design to structure-guided engineering10,11. However, the complex and dynamic conformations of PKSs12,13,14, along with sophisticated interactions and functional interdependencies of different PKS domains15,16, have posed considerable challenges for efficient engineering. Consequently, PKS assembly lines tend to become fragile following a single round of engineering, preventing successive reprogramming and applications17.
The findings on the evolutionary events in modular PKSs18, such as point mutation, gene duplication, gene loss, gene conversion, gene recombination, and horizontal gene transfer, have provided an opportunity for PKS engineering. In 2017, Abe lab proposed an evolutionary mechanism for structural diversification of polyketides by defining unique module organizations19. Keatinge-Clay lab demonstrated the improvement on success rate by using updated module boundaries for domain-fusion20. Based on the statistics of massive trans-AT PKS sequences, Piel lab applied the concept of evolutionary-guided engineering for trans-AT PKSs21. Collectively, the evolutionary information not only guides optimal recombination boundaries, but also facilitates the engineering efforts. For example, by simulating the evolutionary recombination of homologous modules, a ring-contracted mini-azalomycin was generated22. Regarding gene loss, the interconversions between polyene-pyrone structures, differing in the skeleton sizes, were accomplished23,24. Although engineering of PKS remains as a try-and-error, recent studies25,26 mutually demonstrate that evolutionary events-guided engineering is broadly applicable, extending the scenarios beyond isolated and special cases.
Gene conversion is a prevalent evolutionary phenomenon observed in PKSs, whereby genetic material is exchanged between adjacent and homologous modules particularly between regions with high sequence similarity27,28,29. Thus, the event of gene conversion can alter specific regions for fine-tuning the chemical diversity of polyketides. This evolutionary event is widely distributed in Streptomyces, frequently occurring in KS and AT domains of modular PKS assembly lines according to nucleotide homolog and phylogenetic tree analyses28,29,30. Consequently, the gene conversion process is regarded as an important feature for the alteration of macrolide structure28. Meanwhile, the intra-module KS-AT didomain is found to often engage in gene conversion as a complete entity25. Nevertheless, although current knowledge on gene conversion is limited, emulating gene conversion events may offer a possibility of achieving successive PKS engineering.
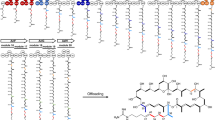
Previously, we unveiled an unusual type I/type III PKS hybrid cmm BGC and its products of cinnamomycin A-F (1,1b, 1c and 2–4), a series of 14-membered macrolides with significant and selective anti-proliferative activity31. When we performed further analysis of cmm BGC, we found the presence of gene conversion with high homology or even identical DNA fragments in module 2, 6 and 7 (Fig. 1a, Supplementary Table 2). These regions are specifically located in malonyl-CoA-specific AT domains, spanning from C-terminus of KS domain to post-AT linker (Fig. 1b). The 100% nucleotide sequence identity of gene conversion region between modules 2 and 6 strongly supporting the connection of gene conversion with the biosynthesis of cinnamomycin skeleton. Therefore, previous successes in utilizing evolutionary approach for PKS engineering promoted us to test whether gene conversion process could be applied to empowering PKS engineering.
a The locations of identical regions in cmm BGC. Rectangles colored in blue represent identical regions in CmmD1 and CmmD3. Numbers in rectangles are the abbreviation of module number; (b) The boundaries of gene conversion regions in module 2/6/7 and their encoding amino acid sequences are presented. Structural models of gene conversion regions in module 2/6/7 are created by AlphaFold 2.0. The regions colored in red represent the location neighboring KS domain; the portions colored in blue represent identical region; the part colored in brown represent post-AT region.
In this study, a homologous mgm BGC for possibly producing macrolides in S. mangrovisoli32 was discovered by gene conversion-oriented genome mining. This homologous mgm BGC could be combined with cmm BGC to serve as a pair of templates for mimicking the process of gene conversion. By proposing an approach for emulating gene conversion process, successive engineering of the modular PKSs was accomplished in cmm BGC for de novo production of mangromycin-like compounds. Moreover, the intra-module KS domain was revealed to act as a proofreading element for ensuring the fidelity of extender unit incorporation, expanding our knowledge on PKS synthetic logics.
Results
Discovery of a homologous mgm BGC by gene conversion-oriented genome mining
Bacterial natural products frequently exist as a large family of structural analogs, encoded by evolutionarily related homologous biosynthetic gene clusters. Similarly, the observation of gene conversion in cmm BGC could lead to the discovery of cinnamomycin-type compounds and their corresponding BGCs.
Since multiple ancestral cluster-specific fragments among homologous BGCs could be preserved by the process of gene conversion, these regions should be useful probes for genome mining. First, we used the gene conversion region in module2 of CmmD1 as the probe to mine BGC(s) with gene conversion characteristics in the NCBI genomic database. Unfortunately, the initial attempts did not yield any meaningful results, possibly due to a significant difference in sequence identity of AT domains caused by a large number of independent recombination events of AT domains in homologous BGCs. Since KS domains are usually accompanied with AT domains to possess gene conversion26,28, this association encouraged us to use KS fragments in cmm BGC for a BLAST search. As a result, an uncharacterized homologous mgm BGC in S. mangrovisoli (GCF_000974985.2) was revealed (Supplementary Fig. 1). The products from this BGC were designated as mangromycins.
While mgm BGC shows significant similarity to cmm BGC (Fig. 2a, Supplementary Table 3 and Supplementary Figs. 2–5), notable differences do present, particularly the genes involved in the biosynthesis of extender units and tailoring steps (Fig. 2a). Specifically, a unique mgmO gene, encoding a phenol-type FAD-dependent halogenase33, is located in the upstream of mgm BGC. After integration of mgmO gene in cmm BGC, chlorinated derivatives (5–8) were isolated and structurally elucidated. The structures of 5–8 revealed that the chlorination reaction took place at C4 position, whereas FAD-dependent halogenases typically undergo chlorination on the aromatic ring33 (Additional descriptions are presented in the section of Materials and Methods, supplementary Figs. 6-10, and Supplementary Tables 3, 6–9). By contrast, the homologous genes of a P450 monooxygenase encoding cmmA and a methyltransferase encoding cmmB are not present in mgm BGC, suggesting distinct tailoring processes. Since the substrate specificity of AT domains determines the structural diversity of side chains, the differences of AT domains in the modules 1, 4, and 5 of mgm BGC were compared based on the signature motifs2 (Supplementary Fig. 2). While MgmD1 is likely to incorporate methylmalonyl-CoA in module 1, MgmD2 might utilize ethylmalonyl-CoA in both modules 4 and 5, indicating acyl group variations between the skeletons of mangromycin and cinnamomycin (Fig. 2b). According to the biosynthetic logic and experimental evidence on the biosynthesis of cinnamomycin (Fig. 2c), we translated the genetic information of mgm BGC into possible structures of mangromycin A-C (Fig. 2d) along with a proposed biosynthetic pathway (Supplementary Fig. 11).
a Clinker analysis of mgm BGC and cmm BGC. Each ORF is labeled by a capital letter; (b) Domain organizations of cmm and mgm modular PKSs. Each circle represents a catalytic domain with its characteristic amino acid motif. The circles with the same color are originated from the same module. LM: 3,5-DHBA-specific loading module; DH0: non-functional dehydratase domain. The signature motif “LAF” of CmmD1-DH0 is present in MgmD1-DH0 (Supplementary Fig. 4). Signature motifs for acyltransferase specificity with the corresponding extender unit are labeled under each module. OMM-ACP: methoxymalonyl-ACP; MMCoA: methylmalonyl-CoA; HMCoA: hexylmalonyl-CoA; EMCoA: ethylmalonyl-CoA. c Structures of cinnamomycins (1, 1b, 1c); (d) Proposed structures of mangromycin A-C based on the biosynthetic logic and bioinformatics analysis.
Thus, the high degree of similarity between mgm and cmm provided a platform for us to validate the PKS reprogramming approach. By emulating the process of gene conversion and employing a rational selection of the embedded elements and their boundaries, it might be possible to expand the application scope of PKS engineering.
Gene conversion-associated successive engineering to create mangromycin skeleton
Mangromycin and cinnamomycin exhibit slight variations in their side chains, which could be attributed to the occurrence of gene conversion processes within the AT regions. Therefore, we speculated that such structural transformations could be achieved through artificial mimicking of the process of gene conversion. To reprogram cmm BGC for producing predicted mangromycin, various extender units incorporated by modules 1, 4, and 5 in cmm BGC were individually altered for matching those in the corresponding regions of mgm BGC (Fig. 2b). To mimic the gene conversion process, we proposed following guidelines for AT engineering: (i) the DNA fragments spanning the regions from “GTNAH” to “HHYWL” in each module, highly homologous to the reported replacement boundaries for AT domain replacement11,34, are designated as ATc region to locate the boundaries. (ii) the prioritization of the catalytic elements should be from the same BGC; (iii) if the elements originated from other sources are selected to replace, high sequence homology to host BGCs should be a major consideration on the selection.
Following the guidelines (i) and (ii), we utilized the ATc region from CmmD2-module 4, specific to methylmalonyl-CoA, to replace the corresponding region in CmmD1-module1, creating mutant S1 (Supplementary Table 4 and Supplementary Fig. 12), aiming to substitute the methoxy group by a methyl group at C14 (Fig. 3a).
a Diagram for domain replacements in wild-type S. cinnamoneus (left) and HPLC analysis of different strains (right). Abbreviations represent different catalytic domains. ATc colored in yellow represent CmmD2-AT4; ATc colored in red represent MgmD2-AT5; (b) Structures of compounds 9–15a; (c) HPLC analysis of strains of S3 and S3-CCR-HCD; d HPLC analysis of strains of S4 and S5 (left) and determined structures of the analogs (right).
In accordance with guideline iii), the MgmD2-AT5c region showed higher homology to CmmD2-AT4c than MgmD2-AT4c region (55.28% vs. 50.55%). Thus, this region was chosen for the exchange with the corresponding region in module 4 of CmmD2 to generate mutant S2 (Supplementary Table 4 and Supplementary Fig. 13) or module 5 to produce mutant S3 (Supplementary Table 4 and Supplementary Fig. 14).
After the fermentation of mutant strains S1, S2, and S3, HPLC analyses (Fig. 3a) indicated the production of a series of peaks (9–15a) to display identical UV-Vis spectra to cinnamomycin 1 (Supplementary Fig. 15), along with the disappearance of cinnamomycin 1 and 2 in all mutants. Then, fermentation was scaled up for isolation and structural characterization. LC/MS and 1D and 2D NMR analyses (Supplementary Figs. 16–23 and Supplementary Tables 10–17) confirmed the structures of compounds 9–15a (Fig. 3b). Compounds 9–12 from mutants S1 and S2 were in line with the alkyl group substitutions at C14 and C8 positions. Compared to 9 and 11, compounds 10 and 12 lack a C1’ hydroxyl group, due to lower catalytic activity of CmmA31. Notably, the yields of compounds 9–12 were not sacrificed, which was close to that of cinnamomycin 1 and 2 in wild-type strain (~50 mg/L).
Despite the utilization of Mgm-AT5 region in both S2 and S3 strains, they surprisingly behaved significantly different (Fig. 3a). In S2 strain, Mgm-AT5 region in module 4 resulted in ethyl substitution at C8 with high specificity. By contrast, S3 strain generated a group of structurally diverse cinnamomycin analogs 13a-15a, varying in acyl groups at C6 position, whereas the desired product of 14a was much less than expected. To enhance the biosynthesis of ethylmalonyl-CoA for improving the titer of 14a, the genes of crotonyl-CoA carboxylase (CCR) and 3-hydroxybutyryl-CoA dehydrogenase (HCD) from Streptomyces coelicolor A3(2) (GCA_008931305.1), critical in the ethylmalonyl-CoA biosynthesis pathway35, were inserted into pSET152, resulting in a construct of pSET152-CCR-HCD36 (Supplementary Fig. 24). Subsequently, CCR and HCD was co-expressed in mutant S3, yielding S3-CCR-HCD mutant strain. Unexpectedly, although 14a became the major product in the fermentation broth of S3-CCR-HCD strain (Fig. 3c), its titer remained comparable to that of S3 strain (~3 mg/L).
To further examine the feasibility for the creation of mangromycin skeleton under the direction of gene conversion, CmmD2-AT4 in S1 strain was replaced by MgmD2-AT5 to construct S4 strain (Supplementary Fig. 25). As expected, the S4 strain produced two compounds 16 and 17 at satisfactory titers (~15 mg/L). The structures of 16 and 17 were fully elucidated (Supplementary Figs. 26–27 and Supplementary Tables 18–19). Indeed, the substitutions at C8 and C14 took place to generate anticipated alkyl chains (Fig. 3d).
When CmmD2-AT5 in S4 strain was substituted by MgmD2-AT5, S5 strain was generated (Supplementary Fig. 25). However, different from S4 strain, HPLC-HRMS profiling of the fermentation broth of S5 strain showed six additional peaks (Fig. 3d). Following the scale-up fermentation of S5 strain and isolation, the structures of compounds 18b-c and 19a-c were characterized (Supplementary Figs. 28–29 and Supplementary Tables 20–24). Compounds 19a-c contain a butyl chain attached at C6 position, whereas compounds 18b-c feature desired ethyl substitution. Such a diversification of extender units in module 5 was similar to that in strain S3, indicating that AT domains are not the only elements for selective incorporation of extender units.
Identification of proofreading role of the intra-module KS domains for extender units
To increase the production of 18a-c, we sought to elucidate the underlying mechanism that discriminates different acyl groups. Base on previous observation on the substrate specificity of AT domain in module 5, we speculated that the situation in S5 strain could be caused by unnatural domain-domain interactions from exogenous MgmD2-AT5.
Theoretically, there could be three routes to decide extender unit specificity during chain extension (Fig. 4a–c). First, MgmD2-KS5, as an intra-module KS, might function as a proof-reading element during the Claisen-condensation step37. Second, CmmD3-KS6 might serve as a proofreader in the transfer of growing intermediates38,39,40. Third, the replacement of MgmD2-AT5 might result in unnatural AT5-ACP5 interactions41, affecting the fidelity of extender unit incorporation.
a–c Potential routes and domain organizations for different extender units by domain swapping in S5 strain. Red circles represent exogenous MgmD2-AT5, and purple and pink circles are natural domains in cmm BGC. ACP with red circle and KS with cyan circle indicate the counterpart domains from mgm BGC; (d) HPLC chromatograms of engineered strains at λ = 280 nm. The structures of 18a-c and 19a-c are shown in Fig. 3d along with their positions in HPLC analyses.
Next, to examine the actual route, we employed PKS domain counterparts from mgm BGC to replace internal elements for eliminating functional interferences from non-native domain-domain interactions. Additionally, through structural modeling by AlphaFold 2.0, the boundaries for the replacement of KS or ACP domains were precisely defined to ensure their compatibilities (Supplementary Figs. 30–32). Specifically, CmmD2-KS5 was replace by its counterpart MgmD2-KS5 from mgm BGC in S5 strain, resulting in S5-MgmKS5 strain (Supplementary Figs. 33). Furthermore, S5-MgmKS6 (Supplementary Fig. 34) and S5-MgmACP5 (Supplementary Fig. 35) strains were similarly generated (Fig. 4d). After fermentation of these mutant strains, HPLC analyses revealed that S5-MgmKS6 strain only produced trace amounts of 18a-c and 19a-c (Fig. 4), suggesting a low efficiency of the assembly line. On the other hand, S5-MgmACP5 strain produced products equivalent to S5 strain with a similar ratio of 18a-c and 19a-c, implying that the interaction between non-cognate MgmD2-AT5 and CmmD2-ACP5 did not affect extender unit incorporation (Fig. 4d). Intriguingly, unlike the cases in S5-MgmKS6 and S5-MgmACP5, the production of desired macrolides 18a-c was remarkably increased in the extracts of S5-MgmKS5 strain, along with complete abolishment of 19a-c (Fig. 4d), implying the involvement of MgmKS5 in the selection of extender units.
CmmD2-KS5 and MgmD2-KS5 shares 78.64% amino acid identity (Supplementary Fig. 36), but they possessed different specificity on extender units. This difference provided us an opportunity to identify the molecular basis in KS domains to determine the specificity. Multiple sequence alignments and structural modeling of these proteins indicated the presence of a specific region located near catalytically essential residues, which was previously named as “Active-Site Cap”42,43 (Fig. 5). Based on the structure and location, we then speculated that the “Active-Site Cap” might be closely related to the specificity of KS domain towards extender unit. To test this hypothesis, a S5-ASC mutant strain was constructed (Supplementary Fig. 37), in which the “Active-Site Cap” of CmmD2-KS5 was modified to match that of MgmD2-KS5. Similar to the findings in S3-mgmKS5 strain, the production of 19a-c was completely abolished along with the accumulation of 18a-c in S5-ASC strain (Fig. 5d), confirming that the specificity of KS domain on extender units is associated with the “Active Site Cap”.
a Structural models of CmmD2-KS5 and MgmD2-KS5 created by using AlphaFold 2.0. The catalytic essential residues are colored in yellow with the indication of residues 230; (b) Sequence comparison of “Active Site Cap” region of CmmD2-KS5 and MgmD2-KS5; (c) HPLC chromatograms of engineered strains with the same treatments and amounts of fermentation broth at λ = 280 nm with three independent repeats.
To pinpoint possible determinants for the specificity, amino acid residue 230 in KS domain, the position closest to catalytically essential residues within the “Active Site Cap”, was identified to show difference between CmmD2-KS5 and MgmD2-KS5 (Fig. 5b). To demonstrate functional role of residue 230, site-directed mutagenesis was undertaken to generate a mutant strain S3-A230T, where Ala was substituted by Thr in CmmD2-KS5 domain (Supplementary Fig. 38). Consequently, a significant decrease of 19a-c in the fermentation broth of S5-A230T strain (Fig. 5d) provided an evidence on the contribution of “Active Site Cap” of KS domain to the process of selecting a proper extender unit.
Biosynthesis of mangromycin C by systemic pathway reprogramming
With the establishment of macrolide skeleton (18a-c), complete biosynthesis of mangromycin required further engineering of tailoring steps. Since CmmA was unable to directly hydroxylate 18a-c, the first step was to disrupt cmmB in S5-MgmKS5 strain to yield S6 strain (Supplementary Fig. 39), thereby preventing the methylation at C19 (Fig. 6). Upon examining the fermentation broth of S6 strain, compound 20 was produced, but its yield was drastically decreased compared to 18a-c. Consequently, sufficient amount of compound 20 was unable to obtain from 50 L fermentation, and its structure therefore was proposed by detailed HRMS/MS analysis (Supplementary Figs. 40 and 41).
According to the characteristics of flavin-dependent halogenase MgmO, C4-chlorination would be the final step to complete the biosynthesis of mangromycin. Given the extremely low abundance of compound 20 and chemical inversion upon the absence of methyl group, CCR-HCD dual gene fragments, was then inserted, under the control of independent ermEp* promoter, into pSET152-mgmO plasmid to yield S7 strain (Supplementary Fig. 42). The fermentation of S7 strain led to the production of compound 21 (Supplementary Fig. 43). Following a 50 L fermentation, 1.5 mg of compound 21 was isolated for structural elucidation (Supplementary Table 25), confirming its structure as predicted mangromycin C (Fig. 2d and Fig. 6). Thus, through successive engineering efforts guided by the evolutionary event of gene conversion, a compound of mangromycin C was obtained by de novo construction of the biosynthetic pathway (Table 1).
In addition, a series of macrolides with diverse side chains were generated during the engineering processes (Table 1), which further exemplifies the functional compatibility and production efficiency in the present engineering.
Discussion
Polyketides biosynthesized by PKSs are a class important compounds with therapeutic values44. Currently, the emergence of drug-resistance and the growing demand for improved draggability have led to the need of structural diversity. The unique biosynthetic logics of modular PKSs provide an opportunity to utilize synthetic biology approach to reprogram PKSs. However, the production of structural analogs by conventional PKS reprogramming is still in the stage of trial-and-correct, due primarily to insufficient knowledge on dynamic conformations of different modules, domain-domain interactions, and speed-limiting bottlenecks in the complex biosynthetic processes.
Inspired by nature’s creation of diverse PKSs and their biosynthetic pathways, evolution-oriented strategies have shown the usefulness and promise in PKS engineering. For instance, mimicking and accelerating natural evolution in laboratory settings for modular PKSs have been practiced to yield various families of polyketides with high yields, such as ring-contracted or ring-expanded rapamycins27. Meanwhile, evolution-related PKS engineering approaches possess the capability of refining the assembly lines in well-organized and coordinated modular PKSs. Additionally, natural evolution contributes to the guidance of optimal selection of splice points during the designs for PKS engineering. Recent reports have extensively demonstrated that the modular unit for insertion or deletion can adhere to a “redefined module”19,20, spanning from upstream AT to downstream KS, rather than following canonical order from KS to ACP.
With the development of combinational biosynthesis, successive PKS engineering has emerged as a frontier to achieve double or even triple substitutions to produce “unnatural” natural products45. However, unpredictable functionality of chimeric assembly line and historically low yield by domain swapping have continuously hindered efficient PKS engineering because of the compatibility issues associated with sequence, structure and function. Similar to the other evolutionary process observed in PKSs, gene conversion has been observed and proposed to occur at a high frequency in regions containing AT domains or KS domains, and the gene conversion process is thought to enable structure diverse in polyketide skeletons. Despite the success on successive engineering in this study, the replacements of different regions of AT domains still exhibited noticeable differences, which reflects the complexity of PKS assembly lines influenced by various factors. Thus, further investigation on the rationale on compatibility and selection criteria will be required.
In this study, we confirmed a piece of evidence for gene conversion occurred in the AT domain of the cmm BGC, which may reflect how nature creates the diversity of acyl groups through certain evolutionary events like gene conversion. Subsequent genome mining resulted in the identification of a homologous mgm BGC containing gene conversion, which enabled us to test the usefulness of gene conversion-associated approach for PKS engineering. Specifically, our approach focused on selecting highly compatible fragments and the boundaries of AT and KS domains by imitating the process of gene conversion. After validating the approach, we performed a series of gene conversion-associated engineering to create a line of chimeric assemblies with four-fold substitution for the generation of a series of cinnamomycin derivatives.
Commonly, the quality control system of modular PKSs consists of a series of coordinated events to ensure irreversible assembly of substrate and the rate of product output. This system includes the dictation of the flow of inter-modular substrate transfer by docking domain pairing46, the gatekeeping roles of downstream KS domains and the hydrolysis of inaccurate intermediates by type II thioesterases47. On the other hand, it is also observed that some PKSs, such as the ones producing antimycin or epothilone, display a tolerance to incorporate different extender units, often attributing to substrate promiscuity of AT domain. Regardless of various outcomes, the precise mechanism on ensuring the fidelity of extender units during Claisen-condensation step is not fully elucidated. In this study, we identified the proofreading role of intra-module KS domains to control the fidelity of extender units, providing a possibility for the governance of polyketide synthase assembly line and the involvement of KS-AT didomain. Recently, the condensation domain in NRPS biosynthesis was also revealed to be proofreading for the control of substrate- and stereo-specificity48, suggesting a notable fashion of proofreading for well-coordinated stepwise biosynthesis of natural products. Therefore, because of the pivotal roles in determining substrate specificity and stereochemistry, organizational and functional characteristics of intra-module KS domains may largely decide the outcomes of PKS reprogramming.
In the past decades, the discovery of clinically useful molecules has been one of the major tasks in the field of natural product research49. However, some BGCs for producing natural products are virtually inaccessible regardless of the advancement of various tools for activating cryptic BGCs, thus limiting the access to the vast chemical space. Recent combination of genome mining with chemical synthesis resulted in a NRPS-like molecule, leading to the identification of an antibiotic capable of bypassing drug resistance50. The present work also showed that, the precise reprogramming of known biosynthetic pathways could be a feasible approach to obtain targeted molecules, which might be a valuable addition to the discovery of bioactive natural products.
Conclusions
In the present study, we have accomplished successive engineering of PKSs for targeted generation of polyketides based on the evolutionary event of gene conversion typically occurred in KS and AT domains. This approach has led to the creation of an artificial PKS gene cluster to produce a new-to-nature compound of mangromycin C. Moreover, we have demonstrated that the selectivity of extender units within the module is governed by intra-module KS domain, rather than previously proposed by AT domain alone. The present PKS engineering associated with gene conversion may facilitate the discovery of bacterial natural products and provide insights into the correlation between PKS biosynthetic rationale and evolutionary feature.
Materials and methods
Culture conditions and media
Streptomyces cinnamoneus ATCC 21532 and its mutant were cultured on solid ISP2 medium (4 g/L yeast extract, 4 g/L glucose, 10 g/L malt extract, 20 g/L agar) at 30 °C for 4 days for sporulation. Fresh spores of S. cinnamoneus were inoculated into 50 mL seed medium (15 g/L glucose, 2 g/L casamino acid, 1 g/L yeast extract, 1 g/L beef extract). After 3-day growth, SPC medium (40 g/L soybean meal, 30 g/L potato starch, 4 g/L CaCO3) was used for the fermentation of S. cinnamoneus at 30 °C, 220 rpm for 7 days.
Characterization of FAD-dependent halogenase mgmO to catalyze an unexpected olefin chlorination
To characterize the flavin-dependent halogenases (FDHs) are recognized for their capability to incorporate halogens into natural products, which are further categorized into subclasses based on their substrate-specificity and sequence homology. A phylogenetic comparison of MgmO in mgm BGC with known FDHs indicated that MgmO falls in the phenolic-type FDHs, known to chlorinate phenolic moieties (Supplementary Fig. 6). Accordingly, it was reasonable to speculate that MgmO catalyzes the chlorination at C19 position of 2,5-dihydroxy-p-benzoquinone moiety in cinnamomycins (Supplementary Fig. 8a), the site also to be methylated by CmmB. To examine the function of MgmO, the gene of mgmO was synthesized (Supplementary Table 2) and incorporated into pSET152 under the control of constitutive ermE*p promoter (Supplementary Table 4 and Supplementary Fig. 7). To eliminate potential interference from CmmB, pSET152-mgmO construct was introduced into wild-type S. cinnamoneus andΔcmmB mutant, resulting in the strains of WT-mgmO and ΔcmmB-mgmO for heterologous expression of mgmO. After fermentation of these strains, HPLC analyses showed the appearance of four peaks (Supplementary Fig. 8b), which were confirmed as chlorinated derivatives of cinnamomycin 1–4 based on their MS fragmentations (Supplementary Figs. 9 and 10). Surprisingly, the chlorination occurred at C4 position of the double bond in cinnamomycins after structural elucidation of compounds 5–8 from large-scale fermentation, purification and NMR spectroscopy (Supplementary Tables 6–9). The chlorination at the same position by the strains of WT-mgmO and ΔcmmB-mgmO further ruled out the possibility of competing for C19 position. Thus, the function of MgmO was characterized as a FAD-dependent halogenase to chlorinate the olefin group of cinnamomycin-type macrolides (Supplementary Fig. 8a).
Genetic manipulations
The plasmid pSET152-mgmO was used to construct the integrated mutant WT-mgmO and ΔcmmB-mgmO in vivo. Firstly, fragment of ermE*p-mgmO were synthesized by GenScript Biotechnology Co., Ltd (Nanjing, China). Then, the fragment was inserted into the EcoRI and EcoRV sites of Streptomyces-E. coli shuttle vector pSET152 to generate the plasmid pSET152-mgmO using T4 DNA ligase. Similar procedures are performed to obtain other plasmids pSET152-CCR-HCD and pSET152-CCR-HCD-mgmO, summarized in Supplementary Fig. 7 and Supplementary Table 4. All plasmids were sequenced and verified.
The plasmid pKC1139-S1 was used to construct the domain replacement mutant S1 in vivo. Firstly, two homologous fragments flanking gene conversion region in CmmD1-module 1 were amplified from S. cinnamoneus genome DNA by two pairs of primers S1-P1 and S1-P2, S1-P5 and S1-P6. Then, gene conversion region in CmmD2-module 4 were amplified from S. cinnamoneus genome DNA by primers S1-P3 and S1-P4. Then, these three fragments were inserted into the EcoRI and HindIII sites of Streptomyces-E. coli shuttle vector pKC1139 to generate the plasmid pKC1139-S1 using In-fusion cloning kit. Similar procedures are performed to obtain other plasmids for domain replacement mutants or in-frame deletion mutant, summarized in Supplementary Table 5. All plasmids were sequenced and verified.
Using mutant strain S1 as an example, the constructed plasmid pKC1139-S1 was transformed into the donor strain E. coli ET12567/pUZ8002. After E. coli-Streptomyces conjugation, and then apramycin-resistant ex-conjugants were incubated in ISP2 medium at 30 °C to generate double-crossover mutant. A pair of primers S1-C1 and S1-C2 was used to obtain the gene fragments, and PCR products were sequentially digested by NotI restriction enzyme and sequenced. Similar procedures were performed to obtain other mutant strains (Supplementary Figs. 12–14, 25, 33–35 and 37–39).
Isolation and purification of cinnamomycin analogues 5-21
For isolation of compounds 5 and 6 from strain WT+mgmO (Supplementary Table 4), a total of 5 L of fermentation media were extracted with an equal volume of ethyl acetate. The extracts were evaporated and dissolved in ethyl acetate. Then, the crude extracts were subjected to C18 silica gel column chromatography, and eluted stepwise using an acetonitrile/water gradient from 10% acetonitrile to 100% acetonitrile. The fractions containing the target compounds were confirmed by HPLC analyses, and the same fractions were combined. Finally, 80 mg of 5 and 60 mg of 6 was obtained.
Compound 5: yellow powder. NMR data, see Supplementary Table 6.
Compound 6: yellow powder. NMR data, see Supplementary Table 7.
For isolation of compounds 7 and 8 from strain ΔcmmB+mgmO (Supplementary Table 4), a total of 5 L of fermentation media were extracted with an equal volume of ethyl acetate. The extracts were evaporated and dissolved in ethyl acetate. Then, the crude extracts were subjected to C18 silica gel column chromatography, and eluted stepwise using an acetonitrile/water gradient from 10% acetonitrile to 100% acetonitrile. The fractions containing the target compounds were confirmed by HPLC analyses, and the same fractions were combined. Finally, 70 mg of 7 and 50 mg of 8 was obtained.
Compound 7: yellow powder. NMR data, see Supplementary Table 8.
Compound 8: yellow powder. NMR data, see Supplementary Table 9.
For isolation of compounds 9 and 10 from strain S1 (Supplementary Table 5), a total of 3 L of fermentation media were extracted with an equal volume of ethyl acetate. The extracts were evaporated and dissolved in ethyl acetate. Then, the crude extracts were subjected to C18 silica gel column chromatography, and eluted stepwise using an acetonitrile/water gradient from 10% acetonitrile to 100% acetonitrile. The fractions containing the target compounds were confirmed by HPLC analyses, and the same fractions were combined. Finally, 180 mg of 9 and 60 mg of 10 was obtained.
Compound 9: yellow powder. NMR data, see Supplementary Table 10.
Compound 10: yellow powder. NMR data, see Supplementary Table 11.
For isolation of compounds 11 and 12 from strain S2 (Supplementary Table 5), a total of 3 L of fermentation media were extracted with an equal volume of ethyl acetate. The extracts were evaporated and dissolved in ethyl acetate. Then, the crude extracts were subjected to C18 silica gel column chromatography, and eluted stepwise using an acetonitrile/water gradient from 10% acetonitrile to 100% acetonitrile. The fractions containing the target compounds were confirmed by HPLC analyses, and the same fractions were combined. Finally, 75 mg of 11 and 35 mg of 12 was obtained.
Compound 11: yellow powder. NMR data, see Supplementary Table 12.
Compound 12: yellow powder. NMR data, see Supplementary Table 13.
For isolation of compounds 13a, 13b, 14a and 15a from strain S3 (Supplementary Table 5), a total of 15 L of fermentation media were extracted with an equal volume of ethyl acetate. The extracts were evaporated and dissolved in ethyl acetate. Then, the crude extracts were subjected to C18 silica gel column chromatography, and eluted stepwise using an acetonitrile/water gradient from 10% acetonitrile to 100% acetonitrile. The fractions containing the target compounds were confirmed by HPLC analyses, and the same fractions were combined. Fractions containing the desired compounds were further purified using semi-preparative HPLC on a YMC-Pack ODS-A column with a water/acetonitrile gradient (35:65) over 25 min at a flow rate of 1.0 mL/min monitored at 280 nm. Finally, 75 mg of 13a, 7.5 mg of 13b, 30 mg of 14a and 20 mg of 15a was obtained.
Compound 13a: yellow powder. NMR data, see Supplementary Table 14.
Compound 13b: white powder. NMR data, see Supplementary Table 15.
Compound 14a: yellow powder. NMR data, see Supplementary Table 16.
Compound 15a: yellow powder. NMR data, see Supplementary Table 17.
For isolation of compounds 16 and 17 from strain S4 (Supplementary Table 5), a total of 3 L of fermentation media were extracted with an equal volume of ethyl acetate. The extracts were evaporated and dissolved in ethyl acetate. Then, the crude extracts were subjected to C18 silica gel column chromatography, and eluted stepwise using an acetonitrile/water gradient from 10% acetonitrile to 100% acetonitrile. The fractions containing the target compounds were confirmed by HPLC analyses, and the same fractions were combined. Finally, 75 mg of 16 and 30 mg of 17 was obtained.
Compound 16: yellow powder. NMR data, see Supplementary Table 18.
Compound 17: yellow powder. NMR data, see Supplementary Table 19.
For isolation of compounds 18b and 18c from strain S5-mgmKS5 (Supplementary Table 5), a total of 10 L of fermentation media were extracted with an equal volume of ethyl acetate. The extracts were evaporated and dissolved in ethyl acetate. Then, the crude extracts were subjected to C18 silica gel column chromatography, and eluted stepwise using an acetonitrile/water gradient from 10% acetonitrile to 100% acetonitrile. The fractions containing the target compounds were confirmed by HPLC analyses, and the same fractions were combined. Fractions containing the desired compounds were further purified using semi-preparative HPLC on a YMC-Pack ODS-A column with a water/acetonitrile gradient (35:65) over 25 min at a flow rate of 1.0 mL/min monitored at 280 nm. Compound 18a degrades during purification process. Finally, 35 mg of 18b and 30 mg of 15a was obtained.
Compound 18b: white powder. NMR data, see Supplementary Table 20.
Compound 18c: yellow powder. NMR data, see Supplementary Table 21.
For isolation of compounds 19a, 19b and 19c from strain S5 (Supplementary Table 5), a total of 10 L of fermentation media were extracted with an equal volume of ethyl acetate. The extracts were evaporated and dissolved in ethyl acetate. Then, the crude extracts were subjected to C18 silica gel column chromatography, and eluted stepwise using an acetonitrile/water gradient from 10% acetonitrile to 100% acetonitrile. The fractions containing the target compounds were confirmed by HPLC analyses, and the same fractions were combined. Fractions containing the desired compounds were further purified using semi-preparative HPLC on a YMC-Pack ODS-A column with a water/acetonitrile gradient (15:85) over 25 min at a flow rate of 1.0 mL/min monitored at 280 nm. Finally, 25 mg of 19a, 20 mg of 19b and 25 mg of 19c was obtained.
Compound 19a: yellow powder. NMR data, see Supplementary Table 22.
Compound 19b: white powder. NMR data, see Supplementary Table 23.
Compound 19c: yellow powder. NMR data, see Supplementary Table 24.
For isolation of compound 21 from strain S7 (Supplementary Table 5), a total of 20 L of fermentation media were extracted with an equal volume of ethyl acetate. The extracts were evaporated and dissolved in ethyl acetate. Then, the crude extracts were subjected to C18 silica gel column chromatography, and eluted stepwise using an acetonitrile/water gradient from 10% acetonitrile to 100% acetonitrile. Compound 21 easily dispersed on the column, so that we increased the column pressure during the purification process. The fractions containing the 21 were confirmed by HPLC analyses, and the same fractions were combined. Fractions containing the desired compounds were further purified using semi-preparative HPLC on a YMC-Pack ODS-A column with a water/acetonitrile gradient (35:65) over 25 min at a flow rate of 1.0 mL/min monitored at 280 nm. Finally, 3.5 mg of 21 was obtained.
Compound 21: yellow powder. NMR data, see Supplementary Table 25.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data supporting the findings of this work are available within the paper and Supplementary Information files. Experimental information, supporting tables and figures are available in Supplementary Information (PDF). Supplementary Data (NMR spectra) can be found in the separate file (PDF). Further information on research design is available in the Nature Research Reporting Summary linked to this article.
References
Hertweck, C. The biosynthetic logic of polyketide diversity. Angew. Chem. Int. Ed. 48, 4688–4716 (2009).
Keatinge-Clay, A. T. The structures of type I polyketide synthases. Nat. Prod. Rep. 29, 1050–1073 (2012).
Smith, S. & Tsai, S. C. The type I fatty acid and polyketide synthases: a tale of two megasynthases. Nat. Prod. Rep. 24, 1041–1072 (2007).
Cortes, J., Haydock, S. F., Roberts, G. A., Bevitt, D. J. & Leadlay, P. F. An unusually large multifunctional polypeptide in the erythromycin-producing polyketide synthase of Saccharopolyspora erythraea. Nature 348, 176–178 (1990).
Donadio, S. et al. Modular organization of genes required for complex polyketide biosynthesis. Science 252, 675–679 (1991).
Sherman, D. H. The Lego-ization of polyketide biosynthesis. Nat. Biotechnol. 23, 1083–1084 (2005).
Keatinge-Clay, A. T. Stereocontrol within polyketide assembly lines. Nat. Prod. Rep. 33, 141–149 (2016).
Weissman, K. J. & Leadlay, P. F. Combinatorial biosynthesis of reduced polyketides. Nat. Rev. Microbiol. 3, 925–936 (2005).
Weissman, K. J. Genetic engineering of modular PKSs: from combinatorial biosynthesis to synthetic biology. Nat. Prod. Rep. 33, 203–230 (2016).
Su, L. et al. Engineering the stambomycin modular polyketide synthase yields 37-membered mini-stambomycins. Nat. Commun. 13, 515 (2022).
Nava, A. A., Roberts, J., Haushalter, R. W., Wang, Z. & Keasling, J. D. Module-based polyketide synthase engineering for de novo polyketide biosynthesis. ACS Synth. Biol. 12, 3148–3155 (2023).
Whicher, J. R. et al. Structural rearrangements of a polyketide synthase module during its catalytic cycle. Nature 510, 560–564 (2014).
Dutta, S. et al. Structure of a modular polyketide synthase. Nature 510, 512–517 (2014).
Cogan, D. P. et al. Mapping the catalytic conformations of an assembly-line polyketide synthase module. Science 374, 729–734 (2021).
Xu, W., Qiao, K. & Tang, Y. Structural analysis of protein-protein interactions in type I polyketide synthases. Crit. Rev. Biochem. Mol. Biol. 48, 98–122 (2013).
Dodge, G. J., Maloney, F. P. & Smith, J. L. Protein-protein interactions in “cis-AT” polyketide synthases. Nat. Prod. Rep. 35, 1082–1096 (2018).
Klaus, M. & Grininger, M. Engineering strategies for rational polyketide synthase design. Nat. Prod. Rep. 35, 1070–1081 (2018).
Nivina, A., Yuet, K. P., Hsu, J. & Khosla, C. Evolution and diversity of assembly-line polyketide synthases. Chem. Rev. 119, 12524–12547 (2019).
Zhang, L. et al. Characterization of giant modular PKSs provides insight into genetic mechanism for structural diversification of aminopolyol polyketides. Angew. Chem. Int. Ed. 56, 1740–1745 (2017).
Miyazawa, T., Hirsch, M., Zhang, Z. & Keatinge-Clay, A. T. An in vitro platform for engineering and harnessing modular polyketide synthases. Nat. Commun. 11, 80 (2020).
Mabesoone, J. et al. Evolution-guided engineering of trans-acyltransferase polyketide synthases. Science 383, 1312–1317 (2024).
Zhai, G. et al. Insights into azalomycin F assembly-line contribute to evolution-guided polyketide synthase engineering and identification of intermodular recognition. Nat. Commun. 14, 612 (2023).
Sugimoto, Y., Ding, L., Ishida, K. & Hertweck, C. Rational design of modular polyketide synthases: morphing the aureothin pathway into a luteoreticulin assembly line. Angew. Chem. Int. Ed. 53, 1560–1564 (2014).
Peng, H., Ishida, K., Sugimoto, Y., Jenke-Kodama, H. & Hertweck, C. Emulating evolutionary processes to morph aureothin-type modular polyketide synthases and associated oxygenases. Nat. Commun. 10, 3918 (2019).
Ray, K. A. et al. Assessing and harnessing updated polyketide synthase modules through combinatorial engineering. Nat. Commun. 15, 6485 (2024).
Huang, Z. et al. Plug-and-play engineering of modular polyketide synthases. Nat. Chem. Bio. https://doi.org/10.1038/s41589-025-01878-4 (2025).
Wlodek, A. et al. Diversity oriented biosynthesis via accelerated evolution of modular gene clusters. Nat. Commun. 8, 1206 (2017). 2017.
Zucko, J., Long, P. F., Hranueli, D. & Cullum, J. Horizontal gene transfer and gene conversion drive evolution of modular polyketide synthases. J. Ind. Microbiol. Biotechnol. 39, 1541–1547 (2012).
Medema, M. H., Cimermancic, P., Sali, A., Takano, E. & Fischbach, M. A. A systematic computational analysis of biosynthetic gene cluster evolution: Lessons for engineering biosynthesis. PLoS Comput. Biol. 10, e1004016 (2014).
Nivina, A., Paredes, S. H., Fraser, H. B. & Khosla, C. GRINS: genetic elements that recode assembly-line polyketide synthases and accelerate their diversification. Proc. Natl. Acad. Sci. USA 118, e2100751118 (2021).
Zhang, B. et al. A type I/type III PKS hybrid generates cinnamomycin A-D. Org. Lett. 25, 2560–2564 (2023).
Ser, H.-L. et al. Presence of antioxidative agent, Pyrrolo[1,2-a]pyrazine-1,4-dione, hexahydro- in newly isolated Streptomyces mangrovisoli sp. nov. Front. Microbiol. 6, 854 (2015).
Latham, J. et al. Development of halogenase enzymes for use in synthesis. Chem. Rev. 118, 232–269 (2018).
Yuzawa, S. et al. Comprehensive in vitro analysis of acyltransferase domain exchanges in modular polyketide synthases and its application for short-chain ketone production. ACS Synth. Biol. 6, 139–147 (2017).
Wang, J. et al. Metabolic network model guided engineering ethylmalonyl-CoA pathway to improve ascomycin production in Streptomyces hygroscopicus var. ascomyceticus. Microb. Cell. Fact. 16, 169 (2017). 31.
Myronovskyi, M. & Luzhetskyy, A. Native and engineered promoters in natural product discovery. Nat. Prod. Rep. 33, 1006–1019 (2016).
Chen, A. Y. et al. Extender unit and acyl carrier protein specificity of ketosynthase domains of the 6-deoxyerythronolide B synthase. J. Am. Chem. Soc. 128, 3067–3074 (2006).
Klaus, M., Buyachuihan, L. & Grininger, M. Ketosynthase domain constrains the design of polyketide synthases. ACS Chem. Biol. 15, 2422–2432 (2020).
Yi, D. et al. Gatekeeping ketosynthases dictate initiation of assembly line biosynthesis of pyrrolic polyketides. J. Am. Chem. Soc. 143, 7617–7622 (2021).
Hirsch, M., Fitzgerald, B. J. & Keatinge-Clay, A. T. How cis-acyltransferase assembly-line ketosynthases gatekeep for processed polyketide intermediates. ACS Chem. Biol. 16, 2515–2526 (2021).
Wong, F. T. et al. Protein-protein recognition between acyltransferases and acyl carrier proteins in multimodular polyketide synthases. Biochemistry 49, 95–102 (2010).
Lohman, J. R. et al. Structural and evolutionary relationships of “AT-less” type I polyketide synthase ketosynthases. Proc. Natl. Acad. Sci. USA 112, 12693–12698 (2015).
Murphy, A. C. et al. Broadening substrate specificity of a chain-extending ketosynthase through a single active-site mutation. Chem. Commun. 52, 8373–8376 (2016).
Butler, M. S., Robertson, A. A. & Cooper, M. A. Natural product and natural product derived drugs in clinical trials. Nat. Prod. Rep. 31, 1612–1661 (2014).
McDaniel, R. et al. Multiple genetic modifications of the erythromycin polyketide synthase to produce a library of novel “unnatural” natural products. Proc. Natl. Acad. Sci. USA 96, 1846–1851 (1999).
Smith, H. G., Beech, M. J., Lewandowski, J. R., Challis, G. L. & Jenner, M. Docking domain-mediated subunit interactions in natural product megasynth(et)ases. J. Ind. Microbiol. Biotechnol. 48, kuab018 (2021).
Pourmasoumi, F. et al. Proof-reading thioesterase boosts activity of engineered nonribosomal peptide synthetase. ACS Chem. Biol. 17, 2382–2388 (2022).
Peng, H. et al. Controlling substrate- and stereospecificity of condensation domains in nonribosomal peptide synthetases. ACS Chem. Biol. 19, 599–606 (2024).
Scherlach, K. & Hertweck, C. Mining and unearthing hidden biosynthetic potential. Nat. Commun. 12, 3864 (2021).
Wang, Z. et al. A naturally inspired antibiotic to target multidrug-resistant pathogens. Nature 601, 606–611 (2022).
Acknowledgements
This work was supported by the grants of the Project Program of the State Key Laboratory of Natural Medicines, China Pharmaceutical University (SKLNMZZ2024JS06), National Key Research and Development Program of China (No.2018YFA0902000), Key Research and Development Project of Guangdong Province (2022B1111070004) and the “Double First-Class” University Project of China Pharmaceutical University (CPU2022QZ08).
Author information
Authors and Affiliations
Contributions
W.J.: design, data curation, formal analysis, methodology and writing–original draft. J.T.: data curation, formal analysis. B.Z.: data curation, methodology. X.W.: resources, supervision. Y.C.: conceptualization, funding acquisition, supervision and writing–review and editing.
Corresponding author
Ethics declarations
Competing interests
A Chinese patent application was filed with the number of 2024112777429. Y.C. and W.J. are the inventors of the patent, and China Pharmaceutical University owns the patent rights. All other authors declare no competing interests.
Peer review
Peer review information
Communications Chemistry thanks Yuhui Sun and the other, anonymous, review for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jin, W., Tu, J., Zhang, B. et al. Gene conversion-associated successive engineering of modular polyketide synthases. Commun Chem 8, 261 (2025). https://doi.org/10.1038/s42004-025-01671-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42004-025-01671-3