Abstract
Purpose
Comorbid familial nonobstructive azoospermia (NOA) and congenital cataract (CC) have not been reported previously, and no single human gene has been associated with both diseases in humans. Our purpose was to uncover novel human mutations and genes causing familial NOA and CC.
Methods
We performed whole-exome sequencing for two brothers with both NOA and CC from a consanguineous family. Mutation screening of TDRD7 was performed in another similar consanguineous family and 176 patients with azoospermia or CC alone and 520 healthy controls. Histological analysis was performed for the biopsied testicle sample in one patient, and knockout mice were constructed to verify the phenotype of the mutation in TDRD7.
Results
Two novel loss-of-function mutations (c.324_325insA (T110Nfs*30) and c.688_689insA (p.Y230X), respectively) of TDRD7 were found in the affected patients from the two unrelated consanguineous families. Histological analysis demonstrated a lack of mature sperm in the male patient’s seminiferous tubules. The mutations were not detected in patients with CC or NOA alone. Mice with Tdrd7 gene disrupted at a similar position precisely replicated the human syndrome.
Conclusion
We identified TDRD7 causing CC as a new pathogenic gene for male azoospermia in human, with an autosomal recessive mode of inheritance.
Similar content being viewed by others
Introduction
Azoospermia, defined as the absence of spermatozoa in the seminal fluid, is the main reason for male infertility, and it consists of two main types, obstructive azoospermia and nonobstructive azoospermia (NOA).1 The latter affects ~0.6% of men in the general population and ~10% of infertile men,2 with its etiology largely unknown. Spermatogenesis involves over 1,000 genes with mouse models identifying over 400 genes that are specifically linked to azoospermia,3 including Rfx2, Brd7, and Tdrd7.4,5,6 Despite substantial effort over the last few decades, outside the candidate genes in the Y-chromosome AZF region,7 only a small number of the genes associated with azoospermia proposed by mouse models have been verified in humans (MEI0B, SYCP3, NR5A1, and TEX11).8,9,10,11 It is estimated that the genetic factors associated with more than 80% of the azoospermia cases remain unknown in humans.
Congenital cataract (CC), caused by lens opacity resulting from metabolic abnormalities during early development, is the principal cause of permanent blindness in children, with an estimated frequency of 1–6 per 10,000 live births worldwide.12 Approximately one-third of all CCs are caused by genetic defects13,14 and over 100 pathogenic genes, including TDRD7,15 have been identified. Although azoospermia and CCs are clinically distinct diseases affecting different organs, they occasionally present together as manifestations of a syndrome, such as Lowe syndrome,16 Kallmann syndrome in humans,17 and Asherman syndrome in mice.18 However, to our knowledge, familial NOA associated with CC has not been reported as a single disease entity in humans, and the genetic etiology of such a condition remains elusive. In this study, we report our findings from the genetic analyses of two unrelated consanguineous Chinese families, which include three male patients with NOA and CC and two female patients with CC alone.
Materials and methods
Families and subjects
Family 1
The proband (family member IV-3) was 32 years old and the third brother in the family of Han Chinese origin. He and his wife visited the Reproductive & Genetic Hospital of CITIC-Xiangya for infertility. His wife had a prior history of pregnancy before this marriage, and her infertility-related examinations (endocrine examinations, hysterosalpingogram, menorrhagia and B ultrasonography, etc.) did not show any known abnormalities. Routine semen analysis of the proband revealed a complete azoospermia with normal volume. The testes were palpable but in a smaller size, left-side testicular volume of 6 ml (31 × 13 × 20 mm), right-side testicular volume of 5.8 ml (29 × 14 × 20 mm) measured by B ultrasonography (Supplementary Figure S1a,b online). Internal sexual organs, including bilateral vas deferens, scrotal, epididymis, and bilateral spermatic varices, were shown to be normal. Detection of sex hormones (testosterone, follicle-stimulating hormone, luteinizing hormone, prolactin, estradiol and inhibin B) showed a high follicle-stimulating hormone and prolactin level, and low inhibin B level (Supplementary Table S1). Testis size, semen analysis, and hormone detection all pointed to NOA. Additional examinations included standard karyotyping and analysis for deletions in the Y chromosome showed to be normal. His physical examination measurements, including height, weight, hair distribution, mentality, and external genital organs, were normal, but he was diagnosed as having bilateral congenital cataracts after birth, and received intraocular lens replacement surgery at 3 years. The ophthalmic medical records were lost.
The oldest brother of the proband (IV-1) was 40 years old with similar clinical characteristics as the proband. He had a history of infertility from his two marriages and was diagnosed as NOA, with normal semen volume (Supplementary Table S1). The left-side testicular volume of 6.2 ml (31 × 15 × 20 mm) was smaller than the right side with a testicular volume of 13.2 ml (39 × 19 × 25 mm) measured by B ultrasonography. The examination results for internal sexual organs, hormone, karyotype, deletions in the Y chromosome, and physical examination were also determined to be normal. He was diagnosed as bilateral congenital lamellar cataract in Xiangya Hospital and received bilateral small incision and intraocular lens replacement cataract surgery at 8 years old. His bilateral vision was 0.1 prior to the cataract surgery and he had eyeball horizontal vibration without proptosis. Eye examination in January 2015 showed his left eye vision was 0.2, 0.02 in right; with vitreous opacity and bilateral intraocular lens (Supplementary Figure S2).
The sister of the proband (family member IV-2), 35 years old, born with poor vision, silver-white binocular lens opacity, and eyeball horizontal vibration, was diagnosed as bilateral congenital cataract in Xiangya Hospital and received bilateral small incision and intraocular lens replacement cataract surgery at the age of 4. Other examinations including hormone detection, karyotype analysis, and physical examination were shown to be normal. She had been married and had two children without known abnormalities.
Their parents (III-1 and III-2) were first-degree cousins who were born and lived in the same village in Hunan province. They were healthy with normal vision.
Family 2
Two affected individuals (IV-2 and IV-6) from another unrelated consanguineous marriage between first-degree cousins from Yunnan province, were presented with congenital cataract. IV-6, 24 years old, the youngest sister in the family, received surgical management at 13 years old. She presented eyeball horizontal vibration, and no pregnancy after 5 years of marriage. Related gynecological examinations showed a fallopian tube obstruction. Other examinations were normal, including endocrine and karyotype analysis of the couple and semen analysis of her husband. IV-2 was 46 years old, had no child after 11 years of marriage. Hormone evaluation showed high follicle-stimulating hormone and prolactin level, and low inhibin B level, similar to IV-1 and IV-3 in family 1 (Supplementary Table S1). The semen analysis showed complete azoospermia. Two siblings (family members IV-3 and IV-5) were heterozygous in TDRD7. Some members in the family declined genetic analysis.
Whole-exome sequencing and variant analysis
Genomic DNA samples from IV-1 and IV-3 in family 1 were extracted from peripheral blood using a QIAamp DNA blood midi kit (Qiagen, Hilden, Germany). Whole-exome sequencing was performed by Beijing Genome Institute at Shenzhen on the HiSeq2000 sequencing platform (Illumina, San Diego, CA, USA) as described previously.19 The analysis of whole-exome sequencing data was performed following the Genome Analysis Toolkit best practices (https://software.broadinstitute.org/gatk/best-practices/). Briefly, the whole-exome sequencing raw reads, after removing adaptors, were aligned to National Center for Biotechnology Information GRCh37 using the Burrows–Wheeler Aligner,20 followed by removal of polymerase chain reaction (PCR) duplicates and sorting using Picard (http://broadinstitute.github.io/picard/). The variant identification was performed using the Genome Analysis Toolkit package21 following its recommended best practices including base recalibration variant calling with Haplotype Caller, variant quality score recalibration, and variant annotation using snpEFF. A candidate gene was considered a variant that matched the following criteria: homozygous for alternate allele, predicted to be deleterious variants, associated with azoospermia or cataract, and either coexisted in IV-1 and IV-3, or had a frequency below 0.1% in three public databases: the 1000 Genomes variant database; Human Gene Mutation Database (HGMD Professional 2016.3); National Heart, Lung, and Blood Institute Exome Sequencing Project; and Exome Aggregation Consortium. The mutation (NM_014290: c.324_325insA (exon 3)) of TDRD7 was validated by Sanger sequencing using specific primers, TDRD7-F: 5’- GAACTGGCCTCTAGGCAACA-3’ and TDRD7-R: 5’- TCAGGATCTTTCCCCTGACATA-3’, which flanked the third exon. The mutation site in TDRD7 was amplified by PCR using Ex Taq DNA polymerase (Bio-Rad, Hercules, CA, USA) in all family 1 members. All PCR products were sequenced on a 3730XL sequencer (Applied Biosystems, Foster City, CA, USA) according to the manufacturer’s instructions.
Single-nucleotide polymorphism array analysis
Genomic DNA sample from IV-1 in family 1 was first subjected to whole-genome amplification using a WGA4 GenomePlex Single Cell Whole Genome Amplification Kit (Sigma-Aldrich, St. Louis, MO, USA) and purified as previously described.22 The whole-genome amplification products were used for single-nucleotide polymorphism array to screen the microchromosomal abnormalities except for the Y chromosome. The patient’s sample was hybridized to a GeneChip Mapping Nsp I 262-K microarray (Affymetrix, Santa Clara, CA, USA). Copy number variation (CNV) and loss of heterozygosity were analyzed using the Gene Chip Genotyping Analysis Software. The National Center for Biotechnology Information’s Map Viewer database was used to map the genomic coordinates and search for genes within the regions of CNVs. All CNVs were checked with the University of California–Santa Cruz Genome Browser on Human Feb. 2009 (GRCh37/hg19) Assembly and the DECIPHER database. Alerted regions of CNVs without OMIM genes were excluded.
Bioinformatics prediction
MutationTaster (http://www.mutationtaster.org/) was applied to predict the possible impact of frameshift mutation on protein function. The mutated TDRD7 complementary DNA sequence was used to compare homology with the wild-type amino acid sequence, which was performed by Ensembl (http://useast.ensembl.org) and BLASTP (http://blast.ncbi.nlm.nih.gov). The domain organization of TDRD7 protein was analyzed by Transeq (http://www.ebi.ac.uk/services).
Generation of Tdrd7-knockout mice
Single guide RNA (TAGCCTGCACAGAAACTGCAAGG) plasmids against the exon 3 of Tdrd7 were designed and constructed. The Cas9 messenger RNA (mRNA) and single guide RNA transcribed by T7 RNA polymerase in vitro were mixed and co-microinjected into fertilized eggs of C57BL/6 mice. Homozygous targeted mice were obtained by intercrossing heterozygous mutant mice. Offspring were genotyped by PCR of tail genomic DNA with the following primers: mTdrd7-gRNA-F: AGTTGTGTACCCCGGGCTCTGG and mTdrd7-gRNA-R: CAATGGAAATCCCAGTTGCAGG. All animals were treated in accordance with the protocols established by the Institutional Animal Care and Use Committee of Central South University (Changsha, China). To determine whether Tdrd7 loss had any impact on fertility, we mated +/+, +/−, and −/− female mice with +/+, +/−, and −/− male mice, and the pups from the combination with different genotypes of males and females were recorded as a measure of their fertility.
Histological and immunochemistry analysis
For histology, testicular tissue from IV-1 in family 1 and from a prostate cancer patient with normal fertility (as normal control testis), and from wild-type and knockout mice were fixed in Bouin’s solution (Sigma-Aldrich), embedded in paraffin, sectioned, processed, and stained with hematoxylin and eosin. For immunostaining, sections of testicular tissue were stained with a primary anti-TDRD7 antibody (1:50 dilution of goat polyclonal antibody) (Abcam, Cambridge, UK), then incubated with a secondary antibody (1:200 dilution of horseradish peroxidase conjugated chicken anti-goat secondary antibody (Santa Cruz Biotechnology, Santa Cruz County, CA, USA). Staining was visualized with the use of 3,3’-diaminobenzidine (Sigma-Aldrich) and hematoxylin as the counter stain. Periodic acid–Schiff staining was performed as previously described;23 briefly, the testes were fixed overnight in Bouin’s solution and then transferred to 70% ethanol. Finally, 6-μm sections were stained with hematoxylin and periodic acid–Schiff reagent to visualize the acrosome.
Fluorescence in situ hybridization
Three types of probes, for chromosome 18, X chromosome, and Y chromosome, which were labeled with white, green, and red fluorescent, respectively, were purchased from Abbott-Vysis (Downers Grove, IL). The fluorescence in situ hybridization (FISH) procedure was performed as we have previously described.24 Briefly, the semen from IV-1 (family 1) and normal control were exposed for 5 min to hypotonic solution (1% sodium citrate in 6 mg/ml bovine serum albumin) and transferred into a small drop of Tween 20 fixative (0.01 N HCl, 0.1% Tween 20) on a clean slide. Fixed cells were analyzed using three types of probes. The FISH signals were observed using an Olympus BX-51 fluorescence microscope (Olympus, Tokyo, Japan). Images were captured using the VideoTesT-FISH 2.0 software (version number 5.0.74.4803, VideoTesT, Petersburg, Russia).
Transmission electron microscopy
Normal control and patient testis biopsy tissues were fixed overnight with 2.5% glutaraldehyde (Sigma-Aldrich) in 0.1 M phosphate buffer (pH 7.4) and subsequently for 2 h in 1.0% osmium tetroxide. They were then subjected to postfixation with 1% OsO4 and 0.1 M sucrose in 0.1 M phosphate buffer, dehydrated with graded concentrations of ethanol, and then embedded in Epon812, dodecenylsuccinic anhydride, methylnadic anhydride, and dimethylaminomethyl phenol at 60 °C for 24 h. Semithin 1-μm-thick sections were stained with toluidine blue for light microscopy. Ultrathin 70–90-nm-thick sections were contrasted with uranyl acetate and lead citrate and examined using a H7700 Hitachi electron microscope (Hitachi, Japan). Digital images were captured using a MegaView III digital camera (Munster, Germany).
Real-time quantitative PCR
Total RNA was extracted from skin fibroblast tissues from IV-1 in family 1 and normal control using Trizol extraction kit (Invitrogen, Carlsbad, CA). Real-time quantitative PCR was performed using SYBR Green PCR Master Mix (Promega, Madison, WI, USA) and a CFX96 Real-Time PCR Detection System (Bio-Rad, Berkeley, CA, USA) according to manufacturer’s instructions. The mRNA expression level of TDRD7 was normalized to the endogenous expression of GAPDH using the following primers: mTDRD7 PCR-F 5′-TGCAGGTTGACGCCATGTA-3′, R 5′-AGAGGCAGATTTTCCCACAGA-3′ and mGAPDH PCR-F 5′-CGAGATCCCTCCAAAATCAA-3′, R 5′-TTCACACCCATGACGAACAT-3′. The assays were done in triplicate.
Western blots
Skin fibroblast tissues from IV-1 in family 1 and normal control were prepared by homogenization of tissues in SDS–PAGE buffer. Proteins were blotted to a polyvinylidene difluoride membrane and incubated overnight at 4 °C with anti-TDRD7 antibody (1:200 dilution, Abcam), then incubated with goat anti-rabbit IgG conjugated to horseradish peroxidase conjugated second antibodies and visualized using enhanced chemiluminescence (Pierced, Grand Island, NY, USA). The relative levels of test protein to control β-actin were analyzed by ImagJ2 software (Madison, WI, USA).
Population screening of TDRD7 variations
For mutation screening, genomic DNA samples for members in family 2 and 140 sporadic subjects with NOA alone, 36 subjects with CC alone (27 males and 9 females), and 520 healthy controls of unrelated Han Chinese population were obtained and subjected to Sanger sequencing. The target regions were amplified by PCR primers listed in Supplementary Table S8, followed by Sanger sequencing.
Statistical analysis
Statistical analysis was determined by Student’s t-tests and one-way analysis of variance using SPSS software, version 19.0 (SPSS, Chicago, IL). Differences were considered significant when P < 0.05.
Results
Clinical characterization of patients with familial NOA and CC
Two Chinese consanguineous families were included in the study. In family 1, both the proband (family member IV-3) and his brother (family member IV-1) suffered from bilateral CC and male infertility due to NOA (Figure 1a). Their sister (IV-2) was diagnosed with bilateral CC but with normal fertility. All three siblings underwent cataract surgery with intraocular lens implantation at the ages of 8 (IV-1), 3 (IV-2), and 4 (IV-3), respectively (Supplementary Figure S2). In family 2, the proband (IV-6) is a women affected with CC. One of her brothers (IV-2) from the same consanguineous marriage of the first-degree cousins also had bilateral CC and infertility due to azoospermia (Figure 1c).
(a) Family 1 involving a consanguineous marriage between two first cousins (III-1 and III-2) in generation 3 with three children (IV-1, IV-2, and IV-3). All siblings (IV-1, IV-2, and IV-3) were affected with CC (horizontal stripes), while both of the two males (IV-1 and IV-3) were also affected with male infertility (solid black). (b) Sanger sequencing chromatograms for subjects from family 1. (c) Family 2 is another consanguineous marriage between first-degree cousins in generation 3 with two affected among six siblings. IV-6 only affected by CC and IV-2 affected by NOA and CC. (d) Sanger sequencing chromatograms for subjects from family 2. Probands in the families are indicated as “p” plus a red arrow. The genotype for the two TRDR7 variants for available family members is indicated with “+” and “−”, indicating normal allele and the mutant alleles, respectively.
To characterize the nature of azoospermia observed in our patients, we performed standard histology analysis of testis biopsy samples from patient IV-1 in family 1. Primary spermatocytes and round spermatids were observed in the patient’s seminiferous tubules of the testis (Figure 2a) as in the normal control (Figure 2b), but no elongated sperms were seen in the patient sample in contrast with the normal control (Figure 2a). The presence of haploid spermatids was further confirmed using FISH with probes for chromosomes 18, X, and Y (Figure 2c). The data indicated that the lack of mature sperms (i.e., NOA) in the patient is due to the arrest of spermatogenesis at the spermiogenesis stage, during which a round haploid spermatid develops into an elongated spermatozoon.
(a,b) Hematoxylin and eosin staining of cross-sections of a single seminiferous tubule at 400 × using testicular biopsy sample for (a) IV-1 from family 1 and (b) a normal control (NC). (c,d) Detection of haploid spermatids using fluorescence in situ hybridization (FISH) with probes for chromosome 18 (white), chromosome X (red), and chromosome Y (green), showing microscopic pictures of cross-sections of a single seminiferous tubule from (c) the patient and (d) a normal control. Haploid spermatids were those with a white dot and either a green or red dot but not both. Red arrow, haploid spermatid.
Identification of TDRD7 mutations
To identify the genetic defects of the patients with CC and NOA phenotypes, we performed whole-exome sequencing and single-nucleotide polymorphism array analysis using blood genomic DNA samples from IV-1 and IV-3 in family 1 using Illumina HiSeq2000 and GeneChip Mapping Nsp I 262 K microarray (Affymetrix) as described previously, respectively.19,25 A large number of variations and no deleterious CNVs except loss of heterozygosity were detected in both samples (Supplementary Tables S2–5). To identify candidate pathogenic variants, we focused on homozygous variants, which are predicted to be deleterious and shared between the two affected individuals (IV-1 and IV-3). We identified a novel frameshift insertion in TDRD7 (NM_014290.2:c.324_325insA (T110Nfs*30)) as the only variant fulfilling the criteria (Supplementary Table S6). We then performed Sanger sequencing and cosegregation analysis using the available DNA samples from other members of this family. Our genotype–phenotype correlation data were consistent with the notion that the homozygous TDRD7 p.T110Nfs*30 mutation is linked to NOA and CC in men and to CC alone in women (Figure 1a and Supplementary Figure S3a).
To validate our observations, we then sequenced the coding regions of TDRD7 using the DNA samples from family 2, which included a brother with NOA and CC (IV-2) and a sister with CC alone (IV-6) from a consanguineous marriage. We discovered another novel nucleotide insertion mutation (c.688_689insA (p.Y230X)), which is predicted also to result in a premature stop codon (Figure 1c and Supplementary Figure S3b). The unaffected parents (III-1 and III-2) and the two siblings (IV-3 and IV-5) were shown to be heterozygous carriers for the same TDRD7 mutation (Figure 1c).
To test whether these two mutations in TDRD7 are independently linked to CC alone or NOA alone, we screened for TDRD7 mutations in 140 cases with NOA alone and 36 cases with CC alone (27 males and 9 females). A control cohort of 520 healthy Han Chinese individuals was also screened. We did not find any apparently deleterious variants, such as those leading to premature stop codons, suggesting that TDRD7 mutations are not a common cause for NOA alone or CC alone. These data also provide additional evidence that TDRD7 defect causes a unique disease, which is associated with NOA and CC in men, and with CC only in women.
The impact of TDRD7 mutations
In silico analysis predicted that both TRDR7 mutations described above are frameshift insertions, which generate premature stop codons (T110Nfs*30 and Y230X) resulting in severely truncated proteins missing most of the known functional domains (Figure 3a–c). In conjunction with the mechanism of nonsense-mediated mRNA decay,26 these mutations in homozygous genotype are predicted to lead to a complete loss of the gene function. To verify the impact of these deleterious mutations in vivo, we analyzed the level of TDRD7 transcripts and protein in the patent’s skin fibroblast tissue. As shown in Figure 3d, real-time quantitative PCR revealed a markedly reduced level of TDRD7 transcripts in the affected subject compared with the normal control. Western blot analysis demonstrated a complete absence of the TDRD7 protein in the patient sample as expected (Figure 3e). Immunohistochemistry using the testis biopsy sample from an affected male patient did not show TDRD7 staining (Figure 3f). Taken together, these data suggest that loss of TDRD7 underlies the pathogenesis of NOA and CC in men.
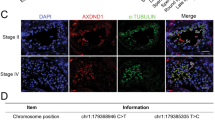
(a) The locations of the two TDRD7 mutations in context of the gene structure and protein coding. Top horizontal bars with the numbers indicate the chromosome locations in chromosome 9. The green horizontal line with vertical bars indicates the locations of exons in scale to gene length, and the arrows indicate the orientation of the gene from 5′ to 3′. Colored text boxes and vertical arrows indicate the positions of the mutations in context of the gene structure. (b) Complementary DNA (cDNA) sequence for the wild-type and mutant alleles with corresponding protein translations. The numbers at the two sides of the sequences indicate the locations in the coding region (CDS) and protein sequences. (c) Location of the insertions in context of the known protein domains from the National Center for Biotechnology Information Conserved Domain Database drawn to the scale of relative sizes and positions with the domains labeled in text boxes. The red and purple vertical bars indicate the locations of the two mutations. (d) Analysis of TDRD7 expression in the skin fibroblast samples from IV-1 and control using real-time quantitative polymerase chain reaction. (e) Western blot analysis showing the absence of TDRD7 protein in the skin fibroblasts of patient IV-1 from family 1 (left) compared with the sample from a normal control (NC, right). (f,g) Immunostaining of testis biopsy samples using an anti-TDRD7 antibody (Abcam ab10767, brown color) for samples from (f) IV-1 in family 1 and (g) normal control (NC). (h,i) Scanning electronic microscopy of spermatid in testicular biopsy samples showing an abnormal chromatid body in (h) IV-1 compared with (i) normal control (NC). Red arrows, chromatid body.
A previous study has suggested that TDRD7, a component of chromatoid bodies (CBs), participates in the assembly of CBs in male germ cells and affects spermatogenesis in mice.6 We therefore examined the integrity of CBs in the male germ cells using scanning electron microscopy. Abnormal CB structures that appeared to be loose and fragmented were observed (Figure 3h) compared with round and tight CBs observed in the control (Figure 3i). These data suggest that azoospermia in the patients was caused by the block of spermatogenesis in the round spermatid cell phase (i.e., arrest of spermiogenesis) in association with the function of CBs.
Validation of TDRD7 mutations’ impact using a mouse model
To further validate our findings from patients, we generated a Tdrd7-knockout mouse model by using clustered regularly interspaced short palindromic repeats (CRISPR)/Cas9 systems by targeting exon 3 (Supplementary Figure S4a,b). In the mouse model, we followed the eye development of mice in different Tdrd7 genotypes (+/+, +/−, and −/−) for 1 month after birth by using a slit lamp. Homozygous Tdrd7-knockout mice showed lens opacity and were diagnosed with CCs, while the wild-type and heterozygous mutant mice had normal lens (Figure 4a). Further, we surveyed the number of pups obtained from crosses between females and males in different Tdrd7 genotypes. The homozygous Tdrd7 mutant males failed to produce any pups (Supplementary Table S7), while the heterozygous males generated comparable numbers of pups with the wild-type males. Moreover, homozygous Tdrd7 mutant males had testes with significantly smaller size and lower weight than their wild-type and heterozygous mutant litter mates (P < 0.001, Figure 4b). Histological analysis of the testes showed that the homozygous Tdrd7 mutant mice had spermatocytes and round spermatids but no elongated sperm in the seminiferous tubules (Figure 4c) and no mature sperm in the epididymis (Figure 4d), indicating that spermatogenesis was arrested at the spermiogenesis stage. This is consistent with the phenotypes of human males carrying the homozygous TDRD7 mutations. Furthermore, abnormal acrosomes were observed in round sperms during steps VII and VIII of spermatogenesis in Tdrd7 −/− mice using periodic acid–Schiff staining, suggesting that spermatogenesis was blocked between steps VII and VIII, during which round spermatids deform to become elongated spermatids (Figure 4e).
(a) Absence of cataract in 2-month-old Tdrd7 +/− mouse lens (control, left) and presence of cataract in age-matched Tdrd7 −/− mouse lens (right). (b) The size, weight, and volume of testes from male mice in three Tdrd7 phenotypes (+/+, +/−, and −/− testes, n = 3 for all groups, left) at 10 weeks, showing Tdrd7 −/− testes were significantly smaller/lower than those from the other two genotypes (**P<0.01 and ***P<0.001, right). (c) Hematoxylin and eosin stained cross-sections of testes from mice in the three Tdrd7 genotypes at 2 months, showing the presence of mature sperms in the Tdrd7 +/+ and −/− mice, but the absence of mature sperm in Tdrd7 −/− mice. (d) Eosin staining of sperm smears from epididymis of male mice in the three Tdrd7 genotypes showing the absence of sperms in the semen of Tdrd7 −/− mice. (e) Periodic acid–Schiff staining of testicular biopsy samples from male mice in the Tdrd7 +/+ and −/− genotypes. The arrowheads indicate abnormal acrosome, and arrows indicate pachytene spermatocytes (P) or round spermatids (RS), respectively.
Discussion
A homozygous mutation with a single amino acid deletion (V618del) in TDRD7 has been previously reported in a family with four affected children with CC alone, including two boys and two girls.15 In this study, NOA was not reported, likely due to the young ages of the male patients. In the same study, characterization of the mice with an N-ethyl-N-nitrosourea–induced Q723X mutation of Tdrd7 suggested male infertility in homozygous condition in mice.15 However, it remains unknown whether loss of TDRD7 causes male infertility in humans. In this report, we show that two loss-of-function mutations of TDRD7 cause NOA and CC in men and cause CC alone in women in two unrelated families. Our additional studies of a new Tdrd7-knockout mouse model generated using the CRISPR/Cas9 approach fully validated our findings in humans. These data provide robust evidence that loss of TDRD7 causes a rare syndrome that involves NOA and CC in men and CC alone in women. For this reason, it would be interesting for Lackhe et al.15 to follow up and see whether NOA can be seen in their male patients.
TDRD7 encodes a member of a large family of Tudor domain–containing proteins and contains LOTUS/OST-HTH and Tudor domains.15,27 In eukaryotic cells, proteins encoding these domains play an important role in the recognition and localization of ribonucleoprotein complexes,27 and constitute an essential class of gametogenesis genes.28 In mice, each member of the Tdrd family has a crucial function at different differentiation stages, and knockout mice for Tdrd1, Tdrd3, Tdrd5, Tdrd6, Tdrd7, and Tdrd12 have been shown to have either disruptions in the dynamic ribonucleoprotein complex remodeling in CBs or metabolism of piwi-interacting RNAs, which protects against retrotransposition of LINE1 that results in male infertility.6,29,30,31,32
We observed that, in the patients with the homozygous TDRD7 mutations, lens formation seemed to be completed, but presented with opacity, while the process of spermatogenesis seemed to be completely blocked in the spermiogenesis stage, and confirmed the arrest of spermiogenesis was blocked between steps VII and VIII in the knockout mice model, leading to the failure of spermatid elongation. This may suggest that TDRD7-associated RNA granules (or the protected mRNA species) are not required for lens formation but for maintaining the function, specifically lens transparency, while it is required for spermatogenesis. Although the Tudor-containing proteins have been reported to be essential for oogenesis in Drosophila,33 both women carrying the homozygous TDRD7 loss-of-function mutations from the two families included in our study were fertile, with similar observation also seen in the Tdrd7-knockout mouse model (Supplementary Table S7). These data imply that TDRD7 is not required in oogenesis in mammals.
It is interesting to observe that function loss of TDRD7 leads to a syndrome that involves two developmentally and anatomically very different organs, while other organs appeared to be unaffected. This suggests that a unique cellular process is shared between the development of lens and spermiogenesis. Although no data are available to directly link these two developmental processes, it is possible that the development of lens cells and spermatozoa requires a similarly sophisticated and lengthy post–cell division maturing process to develop into highly specialized cells with unique cellular morphologies and functions.34,35 A notable important similarity between the two developmental processes is that they both proceed without a nucleus (mature lens fibers) or without an active nucleus (mature sperm). It seems likely that the unique TDRD7-containing RNA granules play a critical role in regulating the posttranscriptional processes of some specific genes in these two cell types, and loss of TDRD7 leads to the impairment of these processes and cellular dysfunction. Indeed, the unique TDRD7-containing RNA granules have been shown to be molecularly distinct from other known RNA granules, and the TDRD7-containing RNA granules are enriched in lens and testis.15,29 Although some RNA transcripts regulated by TDRD7 have been reported, the pathogenic mechanism of TDRD7-mediated NOA and CC has not been well understood.6,29,36 Further studies of the TDRD7-mediated NOA and CC may provide molecular insights into understanding the tissue- or cell-specific role of the unique TDRD7-containing RNA granules in the lens and testis, and therefore, provide a mechanistic basis for development of therapeutic approaches.
References
Lee JY, Dada R, Sabanegh E et al. Role of genetics in azoospermia. Urology 2011;77:598–601.
Poongothai J, Gopenath TS, Manonayaki S. Genetics of human male infertility. Singapore Med J 2009;50:336–347.
Matzuk MM, Lamb DJ. The biology of infertility: research advances and clinical challenges. Nat Med 2008;14:1197–1213.
Wu Y, Hu X, Li Z et al,. Transcription factor RFX2 is a key regulator of mouse spermiogenesis. Sci Rep 2016;6:20435.
Wang H, Zhao R, Guo C et al. Knockout of BRD7 results in impaired spermatogenesis and male infertility. Sci Rep 2016;6:21776.
Tanaka T, Hosokawa M, Vagin VV et al. Tudor domain containing 7 (Tdrd7) is essential for dynamic ribonucleoprotein (RNP) remodeling of chromatoid bodies during spermatogenesis. Proc Natl Acad Sci U S A 2011;108:10579–10584.
Singh K, Raman R. Male infertility: Y-chromosome deletion and testicular aetiology in cases of azoo-/oligospermia. Indian J Exp Biol 2005;43:1088–1092.
Gershoni M, Hauser R, Yogev L et al. A familial study of azoospermic men identifies three novel causative mutations in three new human azoospermia genes. Genet Med; e-pub ahead of print 16 February 2017.
Miyamoto T, Hasuike S, Yogev L et al. Azoospermia in patients heterozygous for a mutation in SYCP3. Lancet 2003;362:1714–1719.
Ropke A, Tewes AC, Gromoll J et al. Comprehensive sequence analysis of the NR5A1 gene encoding steroidogenic factor 1 in a large group of infertile males. Eur J Hum Genet 2013;21:1012–1015.
Yatsenko AN, Georgiadis AP, Ropke A et al. X-linked TEX11 mutations, meiotic arrest, and azoospermia in infertile men. N Engl J Med 2015;372:2097–2107.
Sheeladevi S, Lawrenson JG, Fielder AR et al. Global prevalence of childhood cataract: a systematic review. Eye (Lond) 2016;30:1160–1169.
Pichi F, Lembo A, Serafino M et al. Genetics of congenital cataract. Dev Ophthalmol 2016;57:1–14.
Zhang DD, Du JZ, Topolewski J et al. Review recent progress in identification and characterization of loci associated with sex-linked congenital cataract. Genet Mol Res 2016;15:mr.15038600.
Lachke SA, Alkuraya FS, Kneeland SC et al. Mutations in the RNA granule component TDRD7 cause cataract and glaucoma. Science 2011;331:1571–1576.
Munns CF, Fahiminiya S, Poudel N et al. Homozygosity for frameshift mutations in XYLT2 result in a spondylo-ocular syndrome with bone fragility, cataracts, and hearing defects. Am J Hum Genet 2015;96:971–978.
Legouis R, Hardelin JP, Levilliers J et al. The candidate gene for the X-linked Kallmann syndrome encodes a protein related to adhesion molecules. Cell 1991;67:423–435.
Alawadhi F, Du H, Cakmak H et al. Bone marrow-derived stem cell (BMDSC) transplantation improves fertility in a murine model of Asherman’s syndrome. PLoS One 2014;9:e96662.
Zhu F, Wang F, Yang X et al. Biallelic SUN5 mutations cause autosomal-recessive acephalic spermatozoa syndrome. Am J Hum Genet 2016;99:942–949.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009;25:1754–1760.
McKenna A, Hanna M, Banks E et al. The genome analysis toolkit: a map reduce framework for analyzing next-generation DNA sequencing data. Genome Res 2010;20:1297–1303.
Tan YQ, Tan K, Zhang SP et al. Single-nucleotide polymorphism microarray-based preimplantation genetic diagnosis is likely to improve the clinical outcome for translocation carriers. Hum Reprod 2013;28:2581–2592.
Meistrich ML, Hess RA. Assessment of spermatogenesis through staging of seminiferous tubules. Methods Mol Biol 2013;927:299–307.
Cheng DH, Gong F, Lu CF et al. Risk evaluation and preimplantation genetic diagnosis in an infertile man with an unbalanced translocation t(10;15) resulting in a healthy baby. J Assist Reprod Genet 2012;29:1299–1304.
Gonsalves J, Sun F, Schlegel PN et al. Defective recombination in infertile men. Hum Mol Genet 2004;13:2875–2883.
Kotaja N, Sassone-Corsi P. The chromatoid body: a germ-cell-specific RNA-processing centre. Nat Rev Mol Cell Biol 2007;8:85–90.
Cui G, Botuyan MV, Mer G. (1)H, (15)N and (13)C resonance assignments for the three LOTUS RNA binding domains of Tudor domain-containing protein TDRD7. Biomol NMR Assign 2013;7:79–83.
Pek JW, Anand A, Kai T. Tudor domain proteins in development. Development 2012;139:2255–2266.
Hosokawa M, Shoji M, Kitamura K et al. Tudor-related proteins TDRD1/MTR-1, TDRD6 and TDRD7/TRAP: domain composition, intracellular localization, and function in male germ cells in mice. Dev Biol 2007;301:38–52.
Yabuta Y, Ohta H, Abe T et al. TDRD5 is required for retrotransposon silencing, chromatoid body assembly, and spermiogenesis in mice. J Cell Biol 2011;192:781–795.
Pandey RR, Tokuzawa Y, Yang Z et al. Tudor domain containing 12 (TDRD12) is essential for secondary PIWI interacting RNA biogenesis in mice. Proc Natl Acad Sci USA 2013;110:16492–16497.
Vasileva A, Tiedau D, Firooznia A et al. Tdrd6 is required for spermiogenesis, chromatoid body architecture, and regulation of miRNA expression. Curr Biol 2009;19:630–639.
Boswell RE, Mahowald AP. Tudor, a gene required for assembly of the germ plasm in Drosophila melanogaster. Cell 1985;43:97–104.
Forrester JV, Dick AD, McMenamin PG et al. The Eye: Basic Sciences in Practice, 4th edn. Elsevier Health Sciences: Edinburgh, UK, 2015: 102.
Clermont Y. Kinetics of spermatogenesis in mammals: seminiferous epithelium cycle and spermatogonial renewal. Physiol Rev 1972;52:198–236.
Lachke SA, Maas RL. RNA granules and cataract. Expert Rev Ophthalmol 2011;6:497–500.
The research team acknowledges the support of the National Natural Science Foundation of China (81471432 to Y.T. and 81471510 to G.L.) and the National Key Research and Development Program of China (2016YFC1000206 to G.L.). We acknowledge the excellent technical support provided by Weina Li and Ruiling Tang, as well as support from the clinical and nursing staff at the Reproductive and Genetic Hospital of CITIC-Xiangya. We also thank Xiaobo Xia and Yanxiu Li in the Department of Ophthalmology, Xiangya Hospital for their technical support in cataract analysis and Songqing Fan at the Department of Pathology, the Second Hospital of Xiangya for his generous help with the pathological analysis. The authors also thank all patient families and individuals who participated in this study.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Disclosure
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Tan, YQ., Tu, C., Meng, L. et al. Loss-of-function mutations in TDRD7 lead to a rare novel syndrome combining congenital cataract and nonobstructive azoospermia in humans. Genet Med 21, 1209–1217 (2019). https://doi.org/10.1038/gim.2017.130
Received:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/gim.2017.130
Keywords
This article is cited by
-
Novel INSL3 variants cause male infertility with cryptorchidism
Journal of Assisted Reproduction and Genetics (2026)
-
Novel mutations in ZMYND15 are associated with male infertility with oligozoospermia/azoospermia
Journal of Assisted Reproduction and Genetics (2025)
-
Exploiting autophagy and related pathways: pioneering new horizons in cataract therapy
Apoptosis (2025)
-
CEP112 coordinates translational regulation of essential fertility genes during spermiogenesis through phase separation in humans and mice
Nature Communications (2024)
-
Novel SPEF2 variants cause male infertility and likely primary ciliary dyskinesia
Journal of Assisted Reproduction and Genetics (2024)