Abstract
Antigenic variation of the Plasmodium falciparum multicopy var gene family enables parasite evasion of immune destruction by host antibodies1,2. Expression of a particular var subgroup, termed upsA, is linked to the obstruction of blood vessels in the brain and to the pathogenesis of human cerebral malaria3,4,5,6. The mechanism determining upsA activation remains unknown. Here we show that an entirely new type of gene silencing mechanism involving an exonuclease-mediated degradation of nascent RNA controls the silencing of genes linked to severe malaria. We identify a novel chromatin-associated exoribonuclease, termed PfRNase II, that controls the silencing of upsA var genes by marking their transcription start site and intron-promoter regions leading to short-lived cryptic RNA. Parasites carrying a deficient PfRNase II gene produce full-length upsA var transcripts and intron-derived antisense long non-coding RNA. The presence of stable upsA var transcripts overcomes monoallelic expression, resulting in the simultaneous expression of both upsA and upsC type PfEMP1 proteins on the surface of individual infected red blood cells. In addition, we observe an inverse relationship between transcript levels of PfRNase II and upsA-type var genes in parasites from severe malaria patients, implying a crucial role of PfRNase II in severe malaria. Our results uncover a previously unknown type of post-transcriptional gene silencing mechanism in malaria parasites with repercussions for other organisms. Additionally, the identification of RNase II as a parasite protein controlling the expression of virulence genes involved in pathogenesis in patients with severe malaria may provide new strategies for reducing malaria mortality.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
European Nucleotide Archive
Data deposits
The RNA-seq data generated for this study have been deposited in the European Nucleotide Archive under accession number PRJEB4511, and RNA-seq data from wild-type 3D7 cells25 used for this study are available under accession no. ERP001849.
References
Miller, L. H., Baruch, D. I., Marsh, K. & Doumbo, O. K. The pathogenic basis of malaria. Nature 415, 673–679 (2002)
Scherf, A., Riviere, L. & Lopez-Rubio, J. J. SnapShot: var gene expression in the malaria parasite. Cell 134, 190–190.e1 (2008)
Deitsch, K. W. & Chitnis, C. E. Molecular basis of severe malaria. Proc. Natl Acad. Sci. USA 109, 10130–10131 (2012)
Avril, M. et al. A restricted subset of var genes mediates adherence of Plasmodium falciparum-infected erythrocytes to brain endothelial cells. Proc. Natl Acad. Sci. USA 109, E1782–E1790 (2012)
Claessens, A. et al. A subset of group A-like var genes encodes the malaria parasite ligands for binding to human brain endothelial cells. Proc. Natl Acad. Sci. USA 109, E1772–E1781 (2012)
Turner, L. et al. Severe malaria is associated with parasite binding to endothelial protein C receptor. Nature 498, 502–505 (2013)
Jiang, L. et al. PfSETvs methylation of histone H3K36 represses virulence genes in Plasmodium falciparum. Nature 499, 223–227 (2013)
Lopez-Rubio, J. J., Mancio-Silva, L. & Scherf, A. Genome-wide analysis of heterochromatin associates clonally variant gene regulation with perinuclear repressive centers in malaria parasites. Cell Host Microbe 5, 179–190 (2009)
Freitas-Junior, L. H. et al. Telomeric heterochromatin propagation and histone acetylation control mutually exclusive expression of antigenic variation genes in malaria parasites. Cell 121, 25–36 (2005)
Duraisingh, M. T. et al. Heterochromatin silencing and locus repositioning linked to regulation of virulence genes in Plasmodium falciparum. Cell 121, 13–24 (2005)
Jacquier, A. The complex eukaryotic transcriptome: unexpected pervasive transcription and novel small RNAs. Nature Rev. Genet. 10, 833–844 (2009)
Kiss, D. L. & Andrulis, E. D. The exozyme model: a continuum of functionally distinct complexes. RNA 17, 1–13 (2011)
Armstrong, C. M. & Goldberg, D. E. An FKBP destabilization domain modulates protein levels in Plasmodium falciparum. Nature Methods 4, 1007–1009 (2007)
Mourier, T. et al. Genome-wide discovery and verification of novel structured RNAs in Plasmodium falciparum. Genome Res. 18, 281–292 (2008)
Frank, M., Dzikowski, R., Amulic, B. & Deitsch, K. Variable switching rates of malaria virulence genes are associated with chromosomal position. Mol. Microbiol. 64, 1486–1498 (2007)
Horrocks, P., Pinches, R., Christodoulou, Z., Kyes, S. A. & Newbold, C. I. Variable var transition rates underlie antigenic variation in malaria. Proc. Natl Acad. Sci. USA 101, 11129–11134 (2004)
Ralph, S. A. et al. Transcriptome analysis of antigenic variation in Plasmodium falciparum-var silencing is not dependent on antisense RNA. Genome Biol. 6, R93 (2005)
Epp, C., Li, F., Howitt, C. A., Chookajorn, T. & Deitsch, K. W. Chromatin associated sense and antisense noncoding RNAs are transcribed from the var gene family of virulence genes of the malaria parasite Plasmodium falciparum. RNA 15, 116–127 (2009)
Jensen, A. T. et al. Plasmodium falciparum associated with severe childhood malaria preferentially expresses PfEMP1 encoded by group A var genes. J. Exp. Med. 199, 1179–1190 (2004)
Mair, G. R. et al. Regulation of sexual development of Plasmodium by translational repression. Science 313, 667–669 (2006)
Chen, Q. et al. Developmental selection of var gene expression in Plasmodium falciparum. Nature 394, 392–395 (1998)
Gudipati, R. K., Neil, H., Feuerbach, F., Malabat, C. & Jacquier, A. The yeast RPL9B gene is regulated by modulation between two modes of transcription termination. EMBO J. 31, 2427–2437 (2012)
Chess, A., Simon, I., Cedar, H. & Axel, R. Allelic inactivation regulates olfactory receptor gene expression. Cell 78, 823–834 (1994)
Deitsch, K. W., Lukehart, S. A. & Stringer, J. R. Common strategies for antigenic variation by bacterial, fungal and protozoan pathogens. Nature Rev. Microbiol. 7, 493–503 (2009)
Siegel, T. N. et al. Strand-specific RNA-Seq reveals widespread and developmentally regulated transcription of natural antisense transcripts in Plasmodium falciparum. BMC Genomics 15, 150 (2014)
Nkrumah, L. J. et al. Efficient site-specific integration in Plasmodium falciparum chromosomes mediated by mycobacteriophage Bxb1 integrase. Nature Methods 3, 615–621 (2006)
Armstrong, C. M. & Goldberg, D. E. An FKBP destabilization domain modulates protein levels in Plasmodium falciparum. Nature Methods 4, 1007–1009 (2007)
Zhang, Q. et al. A critical role of perinuclear filamentous actin in spatial repositioning and mutually exclusive expression of virulence genes in malaria parasites. Cell Host Microbe 10, 451–463 (2011)
Fidock, D. A. & Wellems, T. E. Transformation with human dihydrofolate reductase renders malaria parasites insensitive to WR99210 but does not affect the intrinsic activity of proguanil. Proc. Natl Acad. Sci. USA 94, 10931–10936 (1997)
Baker J. R., ed. Severe falciparum malaria. Trans. R. Soc. Trop. Med. Hyg. 94 (Suppl. 1). S1–S90 (2000)
Rottmann, M. et al. Differential expression of var gene groups is associated with morbidity caused by Plasmodium falciparum infection in Tanzanian children. Infect. Immun. 74, 3904–3911 (2006)
Kyes, S., Pinches, R. & Newbold, C. A simple RNA analysis method shows var and rif multigene family expression patterns in Plasmodium falciparum. Mol. Biochem. Parasitol. 105, 311–315 (2000)
Amblar, M., Barbas, A., Fialho, A. M. & Arraiano, C. M. Characterization of the functional domains of Escherichia coli RNase II. J. Mol. Biol. 360, 921–933 (2006)
Joergensen, L. et al. Surface co-expression of two different PfEMP1 antigens on single Plasmodium falciparum-infected erythrocytes facilitates binding to ICAM1 and PECAM1. PLoS Pathog. 6, e1001083 (2010)
Staals, R. H. et al. Dis3-like 1: a novel exoribonuclease associated with the human exosome. EMBO J. 29, 2358–2367 (2010)
Lopez-Rubio, J. J., Mancio-Silva, L. & Scherf, A. Genome-wide analysis of heterochromatin associates clonally variant gene regulation with perinuclear repressive centers in malaria parasites. Cell Host Microbe 5, 179–190 (2009)
Mancio-Silva, L. & Scherf, A. In situ fluorescence visualization of transcription sites and genomic loci in blood stages of Plasmodium falciparum. Methods Mol. Biol. 923, 335–351 (2013)
Kyes, S. A. et al. A well-conserved Plasmodium falciparum var gene shows an unusual stage-specific transcript pattern. Mol. Microbiol. 48, 1339–1348 (2003)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nature Methods 5, 621–628 (2008)
Xue, Z. et al. Genetic programs in human and mouse early embryos revealed by single-cell RNA sequencing. Nature 500, 593–597 (2013)
Schaeffer, D. et al. The exosome contains domains with specific endoribonuclease, exoribonuclease and cytoplasmic mRNA decay activities. Nature Struct. Mol. Biol. 16, 56–62 (2009)
Acknowledgements
We thank T. Wandless for providing Shld1 compound. This work was supported by a European Research Council Advanced Grant (PlasmoEscape 250320), the French Parasitology consortium ParaFrap (ANR-11-LABX0024), the National Natural Science Foundation of China (NSFC; no. 31271388), the French National Research Agency (13-ISV3-0003-01)–NSFC (no. 81361130411) International Collaboration Project, and the Fundamental Research Funds for the Central Universities of China (20123283). T.N.S. was supported by the Human Frontier Science Program and a European Molecular Biology Organization long-term fellowship. J.C. was supported by the NSFC (no. 81271870). J.G. was supported by the Human Frontier Science Program.
Author information
Authors and Affiliations
Contributions
Q.Z. and A.S. conceived and designed experiments. Q.Z., T.N.S. and R.M.M. performed most of the experiments. Q.Z., T.N.S. and C.H. performed RNA sequencing and data analysis. R.M.M. produced recombinant proteins and performed the exoribonuclease assay in vitro. L.J., X.C., F.W. and H.S. generated constructions, transfectants and parasite material. J.C. and Q.G. collected the field isolates and performed gene transcription analysis. C.S. performed the northern blot assay. L.T. and A.T.R.J. generated the antibodies against individual PfEMP1. J.G. and N.A.M. performed IFA and FACS. Q.Z. and A.S. analysed all the data and wrote the manuscript. All authors discussed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Modelling of crystal structure of conserved catalytic RNase II domains.
a, PfDis3 (putative P. falciparum Dis3, PF3D7_1359300 (PlasmoDB)); b, PfRNase II (PF3D7_0906000 (PlasmoDB)); c, hDis3 (Homo sapiens Dis3, BAF92610.1 (Genbank)); d, yRrp44 (Saccharomyces cerevisiae Dis3, NP_014621.1 (Genbank)). The highly conserved and critical residues in the active centre of the catalytic domain of RNase II are highlighted by pink dots. The amino and carboxy termini are shown in blue and red, respectively.
Extended Data Figure 2 Sequence alignment of RNase II domain of exoribonucleases from P. falciparum, Saccharomyces cerevisiae and Homo sapiens.
The predicted second structures of yRrp44 (top) and PfRNase II (bottom) are shown with arrows (β sheets) and spirals (α helices), respectively. The four critical aspartic residues (D) in the catalytic site that have been shown to be essential for RNase activity are highlighted with red arrows and were replaced by glycine (G) in the PfRNase II-dead-mutant recombinant protein (see Fig. 1d).
Extended Data Figure 3 Biochemical characterization of PfRNase II protein in P. falciparum.
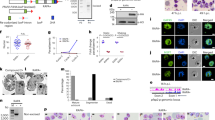
a–c, Schematic representation of the construct of PfRrp4–HA/pLN (a), PfRrp4–HA fusion protein (b), and the recognized site of anti-PfRrp45 antibody (c). d, Western blot of PfRrp4–HA and wild-type 3D7 with anti-HA antibody (left) or anti-PfRrp45 (right). Anti-aldolase antibody was used as a loading control. The full-length proteins are indicated by arrows. e, f, Immunoelectron microscropy assay of PfRNase II in early-stage (e) and mature-stage (f) wild-type 3D7 parasites. Rabbit antibody against PfRNase II was used in this assay (10-nm gold particles). The nuclear membrane is indicated by an arrow. N, nucleus; C, cytoplasm. Scale bar, 500 nm. g, Coomassie staining of three recombinant RNase II domains. h, Sequences of single-stranded (ss) and double-stranded (ds) RNA probes used in exoribonuclease activity analysis in vitro. i, Exoribonuclease activity analysis of recombinant RNase II domains and RNase I (positive control) in vitro with dsRNA probe. j, Exoribonuclease activity analysis of recombinant GST protein (negative control) in vitro with ssRNA probe. Data in d–g, i and j are representative of at least two independent experiments.
Extended Data Figure 4 Strategies of knockout and knockdown approaches of PfRNase II in the 3D7 line.
a, Top: schematic representation of the full-length PfRNase II gene, and the fragments chosen as box 1 or box 2 in pCC1 vector-based knockout constructs. Bottom: various attempts including different combination of box 1 and box 2, parasite lines, and drug on/off selective cyclings are listed in the table. b, Conditional knockout (knockdown) approach of PfRNase II by FKBP destabilization system. c, Three positive clones (G1, H3 and C3), shown in red. d, Western blot assay of PfRNase II protein in the absence of Shld1 in culture. The total extracts of PfRNase II–FKBP-Shld1+ or Shld1− parasites (H3 clone) collected at various time points are indicated on the top of each well. The Shld1 drug was removed in the culture from the ring stage of the first cycle. Rabbit antibody against PfRNase II was used in this assay with anti-aldolase antibody as a loading control. The full-length band of PfRNase II–FKBP is indicated by an arrow. R, ring; S, schizont. Data in c and d are representative of three independent experiments.
Extended Data Figure 5 Comparative analysis of transcriptome changes in PfRNase II–FKBP (H3) and wild-type 3D7-G7 clones by RNA sequencing.
a, Samples and strategies of the analysis. b, Fold change of transcript levels of mitochondrial genes in PfRNase II–FKBP-Shld1+ versus Shld1−. c, List of gene families with a more than fivefold change in transcript levels in the comparison of PfRNase II–FKBP-Shld1+ versus 3D7-G7. Plus sign indicates upregulation of transcript levels. d, Transcription levels of non-coding GC elements and subtelomeric repetitive region (TARE3) in PfRNase II–FKBP-H3 clone with 3D7-G7 as control, determined by qPCR. Samples were harvested at the ring stage. Expression levels were normalized to expression of the seryl-tRNA synthetase gene. Data are represented as means ± s.e.m. for three independent experiments.
Extended Data Figure 6 qPCR analysis of individual var genes in various parasite clones.
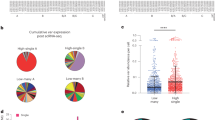
a, Transcription profile of 3D7-G7, PfRNase II–FKBP-H3, G1, C3 clones and two subclones of C3 (scB7 and scB1). The predominantly transcribed var genes with distinct ups type are indicated on each column. Expression levels are normalized to expression of the seryl-tRNA synthetase gene. b, Control parasite clones only predominantly transcribe a single var gene. The five subtype var genes and four control genes are shown on the top of the graph. Data are shown as relative copy numbers related to the seryl-tRNA synthetase gene. Samples were harvested at the ring stage.
Extended Data Figure 7 In situ degradation of nascent RNA from promoter regions is linked to the transcriptional regulation of upsA var genes.
a, Switching pattern of var gene family in PfRNase II–FKBP clones during cultivation in vitro for 70 days. The wild-type 3D7-G7 clone was used as control. The predominantly transcribed var genes are indicated beside the columns. Samples were harvested at the ring stage. The five subtype var genes are shown on the top of the graph. Data are shown as relative copy numbers related to the seryl-tRNA synthetase gene. b, Flow cytometry assay of various PfEMP1 expression in mature trophozoite-stage 3D7-G7 and PfRNase II–FKBP-G1 clones. c, Comparative qPCR var transcription analysis from nascent and steady-state mRNA in ring-stage 3D7-G7 wild-type parasites. Primers are designed against different regions of upsA-type var genes. The seryl-tRNA synthetase gene was used as an internal control for the calculation of relative copy numbers. Error bars represent s.e.m. for three independent experiments. d, Transcription analysis of upsA var gene in field isolates from patients with severe malaria or uncomplicated malaria. Data are shown as relative copy numbers related to the seryl-tRNA synthetase gene. **, P < 0.01 (two-tailed Student’s t-test).
Extended Data Figure 8 RNA-seq coverage of var genes in ring-stage wild-type 3D7-G7 clones (green line) and PfRNase II–FKBP H3 clones (red line).
Each plot contains the data of one var locus; that is, from 200 nt upstream of the start codon to 200 nt downstream of the stop codon. Accession numbers are shown above each plot. RNA-seq coverage on sense and antisense strands are shown in the upper and lower panels for each var gene, respectively. The y axis shows the RNA-seq read pileup coverage on a log10 scale, normalized between samples by scaling with the inter-sample mean number of reads mapped to mRNA. The x axis shows the relative position of the var locus (left and right as the 5′ and 3′ ends of the locus, respectively), with the actual genomic range indicated in the axis label in the form [chromosome: strand: start position–end position]. The grey boxes on the x axis refer to the var exons. The repetitiveness of each relative position on the x axis is shown in an extra panel at the bottom, according to the colour scale at the right (from 1 to 20). a–d, Four representative var genes with different ups types and transcription statuses, as indicated on the top of each plot.
Extended Data Figure 9 Chromatin-associated exoribonuclease PfRNase II and plasmodial exosome have distinct functions in P. falciparum.
a, Schematic representation of the PfRNase II–HA fusion protein. b, Examination of the integration event of PfRNase II–HA transfectants by PCR with genomic DNA. c, Western blot of PfRNase II–HA line with anti-HA and anti-aldolase antibodies. d, qPCR analysis of individual var genes in two independent PfRNase II–HA clones. Samples were harvested at the ring stage. The data are shown as relative copy numbers related to the seryl-tRNA synthetase gene. e, ChIP–qPCR of PfRNase II–HA transfectant. The enrichment of distinct var-subtype var genes with anti-HA antibody is shown. Error bars represent s.e.m. for three independent experiments. f, The plasmodial exosome is expected to exert its functions as described in other eukaryotic organisms (i), whereas the additional non-exosome exoribonuclease PfRNase II has evolved as a regulator of expression of upsA-type var genes in P. falciparum (ii). The 3′–5′ exoribonuclease activity may need other helper molecules to access the 3′ end of nascent RNA. Alternatively, a potential N-terminal PIN-like domain of PfRNase II may help to degrade the nascent RNA by its endonuclease activity as described in yeast42. For b and c, data are representative of three independent experiments.
Supplementary information
Supplementary Table 1
This table shows changes of transcripts level in PfRNase II-FKBP H3 clone versus wild-type 3D7-G7 clone. The upregulation and downregulation of 5-fold are shown in two sheets, respectively. (XLS 77 kb)
Supplementary Table 2
This table contains the sequences of the primers used in this study. (XLS 20 kb)
Rights and permissions
About this article
Cite this article
Zhang, Q., Siegel, T., Martins, R. et al. Exonuclease-mediated degradation of nascent RNA silences genes linked to severe malaria. Nature 513, 431–435 (2014). https://doi.org/10.1038/nature13468
Received:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/nature13468
This article is cited by
-
Extracellular vesicles could be a putative posttranscriptional regulatory mechanism that shapes intracellular RNA levels in Plasmodium falciparum
Nature Communications (2023)
-
Erythrocyte miRNA-92a-3p interactions with PfEMP1 as determinants of clinical malaria
Functional & Integrative Genomics (2023)
-
A single point mutation in the Plasmodium falciparum 3′–5′ exonuclease does not alter piperaquine susceptibility
Malaria Journal (2022)
-
Role of chromatin modulation in the establishment of protozoan parasite infection for developing targeted chemotherapeutics
The Nucleus (2021)
-
Actin-related protein Arp4 regulates euchromatic gene expression and development through H2A.Z deposition in blood-stage Plasmodium falciparum
Parasites & Vectors (2020)