Abstract
Background
Circulating insulin and insulin-like growth factor-1 (IGF-1) concentrations are positively correlated with adiposity. However, the causal effects of insulin and IGF-1 on adiposity are unclear.
Methods
We performed two-sample Mendelian randomization analyses to estimate the likely causal effects of fasting insulin and IGF-1 on relative childhood adiposity and adult body mass index (BMI). To improve accuracy and biological interpretation, we applied Steiger filtering (to avoid reverse causality) and ‘biological effect’ filtering of fasting insulin and IGF-1 associated variants.
Results
Fasting insulin-increasing alleles (35 variants also associated with higher fasting glucose, indicative of insulin resistance) were associated with lower relative childhood adiposity (P = 3.8 × 10−3) and lower adult BMI (P = 1.4 × 10−5). IGF-1-increasing alleles also associated with taller childhood height (351 variants indicative of greater IGF-1 bioaction) showed no association with relative childhood adiposity (P = 0.077) or adult BMI (P = 0.562). Conversely, IGF-1-increasing alleles also associated with shorter childhood height (306 variants indicative of IGF-1 resistance) were associated with lower relative childhood adiposity (P = 6.7 × 10−3), but effects on adult BMI were inconclusive.
Conclusions
Genetic causal modelling indicates negative effects of insulin resistance on childhood and adult adiposity, and negative effects of IGF-1 resistance on childhood adiposity. Our findings demonstrate the need to distinguish between bioaction and resistance when modelling variants associated with biomarker concentrations.
Similar content being viewed by others
Introduction
Obesity presents a major health challenge throughout the life-course, with childhood obesity persisting into adulthood and extending its implications beyond youth [1]. Both childhood and adult obesity are associated with changes in the levels of circulating insulin and insulin-like growth factor-1 (IGF-1) [2]. Investigating the potential causal effects of these biomarkers on obesity may help in designing effective strategies for preventing and managing obesity.
It is known that exogenous insulin therapy and medications such as sulfonylureas or glinides, which are used to enhance the body’s natural insulin production in patients with type 2 diabetes, confer increased weight gain [3]. An observational analysis of data from longitudinal studies and clinical trials found that increases in fasting insulin lead to subsequent increases in BMI [4]. However, it is unclear if this relationship reflects a positive effect of insulin on BMI as, conversely, hyperinsulinaemia is typically interpreted as a marker of insulin resistance [5].
The observed phenotypic association between IGF-1 and obesity is inconsistent. Studies have reported that the relationship between IGF-1 and BMI is negative [6, 7], positive [8] or non-linear [9, 10]. Additionally, the relationship between IGF-1 and adiposity may be influenced by its role as a positive marker of growth hormone axis activity, which potentially affects statural growth more than adipose tissue accumulation [11].
Due to the heterogeneous observations surrounding fasting insulin and IGF-1, the causal nature of these biomarkers on adiposity remains unclear. The identification of genetic variants by genome-wide association studies (GWAS) that are robustly associated with these hormones provides an opportunity to test for causal associations using Mendelian randomization (MR) [12]. Within the MR framework, the random allocation of alleles at conception helps eliminate possible confounding due to environmental factors. Additionally, methods such as Steiger filtering make this more robust by reducing the likelihood of reverse causality. In this study, we used ‘biological effect’ filtering to distinguish whether variants associated with higher biomarker concentrations are indicative of greater biomarker action or greater biomarker resistance. Using a two-sample MR approach, we aimed to examine the causal associations of fasting insulin and IGF-1 on relative childhood adiposity and adult body mass index (BMI).
Methods
Two-sample univariate MR analyses were performed to assess the effect between genetically predicted fasting insulin and IGF-1 on childhood and adult obesity. We conducted the analyses on 444,345 European individuals from the UK Biobank participants with self-reported relative childhood adiposity [13]. We verified any findings in a smaller sample of 39,725 participants with objectively measured childhood BMI from the Early Growth Genetics (EGG) consortium [14]. Additionally, we tested the associations with adult BMI using data from 456,426 European individuals from the UK Biobank [15].
MR utilizes GWAS summary statistics data to test the likely causal association between an exposure and outcome of interest. It relies on three key assumptions: (a) the instrumental variants are associated with the exposure; (b) the instrumental variants are not associated with confounders; and (c) the instrumental variants only influence the outcome via the exposure of interest. We selected as instrumental variants those single nucleotide polymorphisms (SNPs) that reached genome-wide significant threshold (P < 5.0 × 10−8) for circulating fasting insulin or IGF-1 concentrations. In all analyses, the variants were aligned to designate the exposure-increasing allele as the effect allele. For variants that were not present in the outcome data, a highly correlated proxy was selected (within 1 Mb and r2 > 0.7; Supplementary Table 1) using a linkage-disequilibrium panel from a random subsample of individuals in the UK Biobank study with self-identified and genetically confirmed white European ancestry.
MR assumes a consistent and linear relationship, but a higher biomarker level may indicate enhanced biomarker action or increased biomarker resistance. To distinguish between these contrasting effects, we applied ‘biological effect’ filtering in our MR models. This method involved filtering genetic variants based on their individual effect on a third phenotype, which serves as an established indicator of that biomarker’s action.
Instrumental variables
Fasting insulin
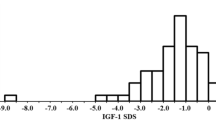
Genetic instruments for fasting insulin concentration (logn pmol/L, in models adjusted for BMI) were based on a European GWAS meta-analysis of approximately 150,000 individuals without diabetes, which reported 43 independent signals [16]. As higher insulin concentrations typically reflect greater insulin resistance [17], they can also reflect greater insulin secretion and bioaction [18]. Therefore, we stratified the insulin genetic instruments based on their effect on fasting glucose concentration, as the established role of insulin is to lower circulating glucose [16] (Fig. 1). Genetic data for fasting glucose (BMI adjusted) was based on a European GWAS meta-analysis of 200,621 individuals [16].
A Fasting insulin associated genetic instruments were filtered by their direction of effect on fasting glucose. B IGF-1 associated genetic instruments were filtered by their direction of effect on childhood height. IGF-1 insulin-like growth factor-1, SNPs single nucleotide polymorphisms.
Of the 43 insulin-increasing alleles, 35 were also directionally associated with higher fasting glucose (indicating insulin resistance), while 7 were directionally related to lower fasting glucose (indicating greater insulin secretion and bioaction). For the remaining one fasting insulin associated variants, there was no available proxy.
IGF-1
Genetic instruments for circulating IGF-1 concentration (nmol/L) were based on a European GWAS of 428,525 European participants from the UK Biobank, which we resolved to 828 independent signals [19]. Higher IGF-1 concentrations typically reflect higher IGF-1 secretion and bioaction; but can also reflect IGF-1 resistance (for example, due to damaging mutations in the IGF-1 receptor gene, IGF1R [20]). Therefore, we stratified the IGF-1 genetic instruments by their effect on childhood height, as the established role of IGF-1 is to increase height [11] (Fig. 1). GWAS data for childhood height at age 7 was based on 36,102 children from the Norwegian Mother, Father and Child Cohort Study (MoBa) [21].
Of the 828 IGF-1-increasing alleles, 351 were also directionally related to taller childhood height (indicating greater IGF-1 secretion and bioaction), while 306 were directionally related to shorter childhood height (indicating IGF-1 resistance). For the remaining 171 IGF-1 associated variants, there were no suitable proxies (r2 > 0.7) for childhood height in the MoBa data.
Outcome variables
Relative childhood adiposity
GWAS data for relative childhood adiposity in the UK Biobank study were modelled as a primary outcome. This was assessed in 444,345 European participants in response to the question “When you were 10 years old, compared to average would you describe yourself as thinner, plumper, or about average?” (UK Biobank field 1687). The GWAS model treated responses as a linear variable (thinner/average/plumper) [13].
To provide confirmation of the results obtained using relative childhood adiposity, we also used data on measured childhood BMI (in standardized units) from a much smaller GWAS meta-analysis. These data from the EGG consortium were based on a trans-ancestral meta-analysis GWAS of BMI in 39,620 children aged between 2 and 18 years [14].
Adult BMI
GWAS data for adult BMI was based on 456,426 European-ancestry participants from the UK Biobank using data from the first assessment centre visit (UK Biobank field 21001) [15].
Statistical analysis
MR models
Our primary analysis was the inverse-variance weighted (IVW) MR model with biological effect and Steiger filtering. Steiger filtering reduces reverse causality by removing instrumental variable SNPs if they have a stronger effect on the outcome than on the exposure (Supplementary Table 2). We performed sensitivity analyses that control for genetic pleiotropy, including MR-Egger, Weighted Median (WM), and Penalized Weighted Median (PWM) MR models. Additionally, we used the MR-Egger intercept P < 0.05, I2 statistic, Cochran’s Q-derived P value, funnel and dosage plots to assess evidence of balanced and unbalanced pleiotropy. When MR-Egger intercept is significant (P < 0.05), we considered the IVW model to be invalid. All analyses were performed using R (version 3.5.1) and a P < 0.0125 was considered statistically significant (calculated as 0.05 / (2 exposures * 2 outcomes)).
Results
Estimated effect of fasting insulin on adiposity
Fasting insulin (non-stratified SNPs)
Overall, genetically-predicted higher fasting insulin concentrations were associated with lower relative childhood adiposity (β = −0.053 categories per logn pmol/L; P = 1.2 × 10−2) and lower adult BMI (β = −0.669 Kg/m2 per logn pmol/L; P = 2.7 × 10−4) (Fig. 2 and Supplementary Table 3). Additional MR models using measured childhood BMI confirmed the relationship between higher fasting insulin and lower childhood BMI (β = −0.11 SD per logn pmol/L; P = 4.4 × 10−2) (Supplementary Fig. 1 and Supplementary Table 3).
Insulin resistance was represented by fasting insulin-increasing alleles also associated with higher fasting glucose. Insulin bioaction was represented by fasting insulin-increasing alleles also associated with lower fasting glucose. Estimates are from random-effects inverse-variance weighted analyses with Steiger filtering. BMI body mass index, IVW inverse-variance weighted, WM Weighted Median, PWM Penalized Weighted Median.
Insulin resistance
Fasting insulin-increasing alleles that were also associated with higher fasting glucose (indicative of insulin resistance) were associated with lower relative childhood adiposity (β = −0.066 category per logn pmol/L; P = 3.8 × 10−3) and lower adult BMI (β = −0.819 Kg/m2 per logn pmol/L; P = 1.4 × 10−5) (Fig. 2 and Supplementary Table 4). Additional MR models using measured childhood BMI confirmed the relationship between insulin resistance and lower childhood BMI (β = −0.126 SD per logn pmol/L; P = 3.2 × 10−2) (Supplementary Fig. 1 and Supplementary Table 4).
Insulin bioaction
Conversely, insulin-increasing alleles that were also associated with lower fasting glucose (indicators of insulin secretion and bioaction) showed no association with relative childhood adiposity nor with adult BMI (P = 0.521 and P = 0.478, respectively) (Fig. 2 and Supplementary Table 4). Additional MR models using measured childhood BMI showed no association between insulin bioaction and childhood BMI (P = 0.985) (Supplementary Fig. 1 and Supplementary Table 4).
Estimated effect of IGF-1 on adiposity
IGF-1 (non-stratified SNPs)
Overall, genetically-predicted higher IGF-1 concentrations were associated with lower relative childhood adiposity (β = −0.003 categories per nmol/L; P = 6.6 × 10−3) (Fig. 3 and Supplementary Table 3). Additional MR models using measured childhood BMI showed a directionally-consistent but non-significant association between higher IGF-1 and childhood BMI (P = 0.334) (Supplementary Fig. 2 and Supplementary Table 3). For adult BMI, IGF-1 showed directionally-inconsistent associations. The MR-Egger intercept indicated evidence of horizontal pleiotropy, and the MR-Egger estimate was in the opposite direction to estimates from the IVW, WM and PWM models (Fig. 3 and Supplementary Table 3).
IGF-1 bioaction was represented by IGF-1-increasing alleles also associated with taller childhood height. IGF-1 resistance was represented by IGF-1-increasing alleles also associated with shorter childhood height. Estimates are from random-effects inverse-variance weighted analyses with Steiger filtering. BMI body mass index, IGF-1 insulin-like growth factor-1, IVW inverse-variance weighted, WM Weighted Median, PWM Penalized Weighted Median. †Indicates significant MR-Egger intercept (P < 0.05).
IGF-1 bioaction
IGF-1-increasing alleles that were also associated with taller childhood height (indicative of greater IGF-1 secretion and bioaction) showed no association with relative childhood adiposity (β = −0.003 categories per nmol/L, P = 0.077) and adult BMI (β = −0.007 Kg/m2 per nmol/L, P = 0.562) (Fig. 3 and Supplementary Table 5). Additional MR models using measured childhood BMI showed no association with IGF-1 bioaction (P = 0.258) (Supplementary Fig. 2 and Supplementary Table 5).
IGF-1 resistance
Conversely, IGF-1-increasing alleles that were also associated with shorter childhood height (indicative of IGF-1 resistance) were associated with lower relative childhood adiposity (β = −0.005 categories per nmol/L; P = 6.7 × 10−3). Additional MR models using measured childhood BMI showed a similar negative association between IGF-1 resistance and childhood BMI (β = −0.016 SD per nmol/L; P = 9.5 × 10−4) (Supplementary Fig. 2 and Supplementary Table 5). For adult BMI, IGF-1 resistance showed directionally-inconsistent associations. The MR-Egger intercept indicated evidence of horizontal pleiotropy, and the MR-Egger estimate was in the opposite direction to estimates from the IVW, WM and PWM models (Fig. 3 and Supplementary Table 5).
Discussion
This study examined how genetically predicted fasting insulin and IGF-1 affect childhood and adult adiposity to infer the causal effects of these biomarkers. Using a biologically informed approach, our findings indicate negative effects of insulin resistance on childhood and adult adiposity, as well as negative effects of IGF-1 resistance on childhood adiposity. We show that using a naive MR approach, which ignores the biological effects of biomarker-associated variants, we might wrongly conclude that fasting insulin and IGF-1 secretion and bioaction have negative effects on childhood and adult adiposity. Instead, our results highlight the importance of distinguishing between bioaction and resistance when analysing variants associated with biomarker levels.
For fasting insulin, recent MR analyses have suggested the importance of biologically-informed MR approaches to select appropriate instruments. Gagnon et al. filtered fasting insulin SNPs using data on pancreatic islet expression to identify 3 SNPs that they considered to represent increased insulin secretion and found a positive effect of these on BMI, even though there was no overall effect seen with all fasting insulin SNPs [22]. However, they did not model an effect of insulin resistance. In our study, we stratified fasting insulin SNPs according to their association with blood glucose levels, and we found a negative effect of insulin resistance on BMI, whereas there was no effect of insulin secretion on BMI. Similarly, the lack of a causal effect of insulin secretion on BMI aligns with an earlier analysis that filtered T2D-associated SNPs by their reported biological effects [23]. Our findings support the importance of biological filtering of SNPs in MR studies of biomarkers.
We observed a negative effect of IGF-1 resistance on childhood adiposity, but no consistent effect on BMI among adults aged 40–69 years (where the significant MR-Egger intercept indicates potential bias). These results may be consistent with cross-sectional findings showing a negative IGF-1-BMI relationship in the youngest quartile of that study (age 24–58 years) and a positive relationship in the oldest quartile (87–110 years), while no significant effect was observed among adults aged 58–86 years [24]. While the term ‘IGF-1 resistance’ is used here in a broad sense and does not specify any particular condition or biological pathway, IGF-1 resistance has been associated with a potential compensatory mechanism involving increased secretion of growth hormone (GH). Elevated GH levels have been found to have lipolytic effects [11]. Thus, IGF-1 resistance may indirectly contribute to the regulation of adiposity through the upregulation of GH secretion.
There are several limitations in our study. Firstly, the studied phenotypic traits as measures of adiposity may have limitations in fully capturing the complexity of body composition. We relied on self-reported, ordered, measures of childhood adiposity. However, the recalled childhood adiposity trait in UK Biobank was recently reported to show very high genetic correlation (rg = 0.94) with objectively measured childhood BMI [13]. We also included as a sensitivity analysis data from a GWAS meta-analysis of childhood BMI, which supported most of our observed associations. Nevertheless, the use of ordered childhood obesity limits the interpretation of the effect estimates. Therefore, the effect sizes on childhood and adult adiposity could not be compared. Additionally, the biomarker measurements used in our study were from adults rather than children, and this MR is dependent on the assumption that the genetic determinants have consistent effects across the life-course. The reported genetic instruments for fasting insulin were from models adjusted for BMI, which could introduce collider bias [25]. However, this bias would affect all fasting insulin SNPs equally and does not explain the results of our biologically informed stratification. Our study was limited to European participants and the findings may not be applicable to other ethnicities. We also did not explore potential non-linear associations between IGF-1 and adiposity, as suggested by some phenotypic studies [6,7,8,9,10]. Finally, while we filtered biomarker-associated into two groups (representing bioaction or resistance), there is likely more complexity. Wang et al. used a statistical approach to separate IGF-1-associated SNPs into six clusters, although they were able to biologically annotate only two of these [26]. For example, it would be valuable in future to distinguish IGF-1 bioaction SNPs that act via increased GH secretion and signalling from those that act more directly.
In conclusion, this study provides new insights into the regulation of adiposity. Our findings indicate that insulin resistance has negative effects on childhood and adult adiposity, while IGF-1 resistance has negative effects on childhood adiposity. Furthermore, our study highlights the importance of employing “biological effect filtering” to differentiate between resistance and bioaction in MR studies, enhancing their interpretability and accuracy.
Data availability
Data from the Norwegian Mother, Father and Child Cohort Study and the Medical Birth Registry of Norway used in this study are managed by the national health register holders in Norway (Norwegian Institute of public health) and can be made available to researchers, provided approval from the Regional Committees for Medical and Health Research Ethics (REC), compliance with the EU General Data Protection Regulation (GDPR) and approval from the data owners. The consent given by the participants does not open for storage of data on an individual level in repositories or journals. Researchers who want access to data sets for replication should apply through helsedata.no. Access to data sets requires approval from The Regional Committee for Medical and Health Research Ethics in Norway and an agreement with MoBa.
References
Ogden CL, Carroll MD, Curtin LR, Lamb MM, Flegal KM. Prevalence of high body mass index in US children and adolescents, 2007–2008. JAMA 2010;303:242–9.
Kreitschmann-Andermahr I, Suarez P, Jennings R, Evers N, Brabant G. GH/IGF-I regulation in obesity – mechanisms and practical consequences in children and adults. Horm Res Paediatr. 2010;73:153–60.
Liu SC, Tu YK, Chien MN, Chien KL. Effect of antidiabetic agents added to metformin on glycaemic control, hypoglycaemia and weight change in patients with type 2 diabetes: a network meta-analysis. Diabetes Obes Metab. 2012;14:810–20.
Wiebe N, Ye F, Crumley ET, Bello A, Stenvinkel P, Tonelli M. Temporal associations among body mass index, fasting insulin, and systemic inflammation: a systematic review and meta-analysis. JAMA Netw Open. 2021;4:e211263.
Reaven GM. Banting lecture 1988. Role of insulin resistance in human disease. Diabetes. 1988;37:1595–607.
Faupel-Badger JM, Berrigan D, Ballard-Barbash R, Potischman N. Anthropometric correlates of insulin-like growth factor 1 (IGF-1) and IGF binding protein-3 (IGFBP-3) levels by race/ethnicity and gender. Ann Epidemiol. 2009;19:841–9.
Alderete TL, Byrd-Williams CE, Toledo-Corral CM, Conti DV, Weigensberg MJ, Goran MI. Relationships between IGF-1 and IGFBP-1 and adiposity in obese African-American and Latino adolescents. Obes Silver Spring Md 2011;19:933–8.
Laughlin GA, Barrett-Connor E, Criqui MH, Kritz-Silverstein D. The prospective association of serum insulin-like growth factor I (IGF-I) and IGF-binding protein-1 levels with all cause and cardiovascular disease mortality in older adults: the Rancho Bernardo Study. J Clin Endocrinol Metab. 2004;89:114–20.
Lukanova A, Söderberg S, Stattin P, Palmqvist R, Lundin E, Biessy C, et al. Nonlinear relationship of insulin-like growth factor (IGF)-I and IGF-I/IGF-binding protein-3 ratio with indices of adiposity and plasma insulin concentrations (Sweden). Cancer Causes Control. 2002;13:509–16.
Lukanova A, Lundin E, Toniolo P, Micheli A, Akhmedkhanov A, Rinaldi S, et al. Circulating levels of insulin-like growth factor-I and risk of ovarian cancer. Int J Cancer. 2002;101:549–54.
Aguiar-Oliveira MH, Bartke A. Growth hormone deficiency: health and longevity. Endocr Rev. 2019;40:575–601.
Lawlor DA, Harbord RM, Sterne JAC, Timpson N, Davey Smith G. Mendelian randomization: using genes as instruments for making causal inferences in epidemiology. Stat Med. 2008;27:1133–63.
Kaisinger LR, Kentistou KA, Stankovic S, Gardner EJ, Day FR, Zhao Y, et al. Large-scale exome sequence analysis identifies sex- and age-specific determinants of obesity. Cell Genomics. 2023;3:100362.
Vogelezang S, Bradfield JP, Ahluwalia TS, Curtin JA, Lakka TA, Grarup N, et al. Novel loci for childhood body mass index and shared heritability with adult cardiometabolic traits. PLOS Genet. 2020;16:e1008718.
Lotta LA, Mokrosiński J, Mendes de Oliveira E, Li C, Sharp SJ, Luan J, et al. Human gain-of-function MC4R variants show signaling bias and protect against obesity. Cell 2019;177:597–607.e9.
Chen J, Spracklen CN, Marenne G, Varshney A, Corbin LJ, Luan J, et al. The trans-ancestral genomic architecture of glycemic traits. Nat Genet. 2021;53:840–60.
Petersen MC, Shulman GI. Mechanisms of insulin action and insulin resistance. Physiol Rev. 2018;98:2133–223.
Saha S, Schwarz PEH. Impact of glycated hemoglobin (HbA1c) on identifying insulin resistance among apparently healthy individuals. J Public Health. 2017;25:505–12.
Elucidating the genetic architecture… | Wellcome Open Research [Internet]. [cited 2021 Apr 13]. Available from: https://wellcomeopenresearch.org/articles/6-20
Gardner EJ, Kentistou KA, Stankovic S, Lockhart S, Wheeler E, Day FR, et al. Damaging missense variants in IGF1R implicate a role for IGF-1 resistance in the etiology of type 2 diabetes. Cell. Genomics 2022;2:100208.
Helgeland Ø, Vaudel M, Sole-Navais P, Flatley C, Juodakis J, Bacelis J, et al. Characterization of the genetic architecture of infant and early childhood body mass index. Nat Metab. 2022;4:344–58.
Gagnon E, Mitchell PL, Arsenault BJ. Body fat distribution, fasting insulin levels, and insulin secretion: a bidirectional Mendelian randomization study. J Clin Endocrinol Metab. 2023;108:1308–17.
Scott RA, Fall T, Pasko D, Barker A, Sharp SJ, Arriola L, et al. Common genetic variants highlight the role of insulin resistance and body fat distribution in type 2 diabetes, independent of obesity. Diabetes 2014;63:4378–87.
Sherlala RA, Kammerer CM, Kuipers AL, Wojczynski MK, Ukraintseva SV, Feitosa MF, et al. Relationship between serum IGF-1 and BMI differs by age. J Gerontol Ser A 2021;76:1303–8.
Day FR, Loh PR, Scott RA, Ong KK, Perry JRB. A Robust example of collider bias in a genetic association study. Am J Hum Genet. 2016;98:392–3.
Wang W, Tesfay EB, van Klinken JB, Willems van Dijk K, Bartke A, van Heemst D, et al. Clustered Mendelian randomization analyses identify distinct and opposing pathways in the association between genetically influenced insulin-like growth factor-1 and type 2 diabetes mellitus. Int J Epidemiol. 2022;51:1874–85.
Acknowledgements
For the purpose of Open Access, the author has applied a Creative Commons Attribution (CC BY) licence to any Author Accepted Manuscript version arising. This work has been conducted using UK Biobank data, application number 9905. Data on glycaemic traits have been contributed by MAGIC investigators and have been downloaded from www.magicinvestigators.org. The authors were supported by the Medical Research Council (Unit programme MC_UU_00006/2). DO was also supported by a PhD studentship from King Abdullah International Medical Research Centre (KAIMRC), the Ministry of National Guard Health Affairs. The Norwegian Mother, Father and Child Cohort Study is supported by the Norwegian Ministry of Health and Care Services and the Ministry of Education and Research. We are grateful to all the participating families in Norway who take part in this ongoing cohort study. We thank the Norwegian Institute of Public Health (NIPH) for generating high-quality genomic data. This research is part of the HARVEST collaboration, supported by the Research Council of Norway (#229624). We also thank the NORMENT Centre for providing genotype data, funded by the Research Council of Norway (#223273), South East Norway Health Authorities and Stiftelsen Kristian Gerhard Jebsen. We further thank the Centre for Diabetes Research, the University of Bergen for providing genotype data funded by the ERC AdG project SELECTionPREDISPOSED, Stiftelsen Kristian Gerhard Jebsen, Trond Mohn Foundation, the Research Council of Norway, the Novo Nordisk Foundation, the University of Bergen, and the Western Norway Health Authorities. Supported by grants from the European Research Council (AdG #293574), the Bergen Research Foundation (“Utilizing the Mother and Child Cohort and the Medical Birth Registry for Better Health”), Stiftelsen Kristian Gerhard Jebsen (Translational Medical Centre), the University of Bergen, the Research Council of Norway (FRIPRO grant #240413), the Western Norway Regional Health Authority (Strategic Fund “Personalized Medicine for Children and Adults”), the Novo Nordisk Foundation (grant #54741), and the Norwegian Diabetes Association; and (to S.J.) Helse Vest’s Open Research Grant (grant #912250), the Research Council of Norway (FRIPRO grant #315599), and Novo Nordisk Foundation (grant #NNF21OC0070349). This work was partly supported by the Research Council of Norway through its Centres of Excellence funding scheme (#262700), Better Health by Harvesting Biobanks (#229624) and The Swedish Research Council, Stockholm, Sweden (2015-02559), The Research Council of Norway, Oslo, Norway (FRIMEDBIO #547711), March of Dimes (#21-FY16-121). The Norwegian Mother, Father, and Child Cohort Study is supported by the Norwegian Ministry of Health and Care Services and the Ministry of Education and Research, NIH/NIEHS (contract no N01-ES-75558), NIH/NINDS (grant no.1 UO1 NS 047537-01 and grant no.2 UO1 NS 047537-06A1). MV acknowledges the support of the Research Council of Norway (project #301178) and SJ the Novo Nordisk Foundation (project NNF20OC0063872). Analyses were performed using digital laboratories in HUNT Cloud at the Norwegian University of Science and Technology, Trondheim, Norway. We are grateful for outstanding support from the HUNT Cloud community.
Author information
Authors and Affiliations
Contributions
DIO, FRD, KKO, and JRBP designed the study. DIO, FRD, LRK, KAK, MV, SS, PRN, and SJ performed the analyses. DIO, FRD, and KKO wrote the manuscript and it was reviewed by all authors.
Corresponding author
Ethics declarations
Competing interests
JRBP is an employee of and holds shares in Insmed Inc.
Ethical approval
The establishment of MoBa and initial data collection was based on a licence from the Norwegian Data Protection Agency and approval from The Regional Committees for Medical and Health Research Ethics. The MoBa cohort is currently regulated by the Norwegian Health Registry Act. The current study was approved by The Regional Committees for Medical and Health Research Ethics (no. 2012/67). UK Biobank received ethical approval from the NHS National Research Ethics Service NorthWest (11/NW/0382).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Olwi, D.I., Kaisinger, L.R., Kentistou, K.A. et al. Likely causal effects of insulin resistance and IGF-1 bioaction on childhood and adult adiposity: a Mendelian randomization study. Int J Obes 48, 1650–1655 (2024). https://doi.org/10.1038/s41366-024-01605-4
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41366-024-01605-4